Abstract
Colorectal cancer is the third most common cancer worldwide, and its incidence continues to grow. Approximately one-third of patients with colorectal cancer develop liver metastases during the natural course of disease. Complete surgical resection is associated with very low mortality in colorectal liver metastasis patients, but only a small fraction of colorectal liver metastasis patients fulfill the selection criteria for surgical treatment. We herein describe a high-risk stage-IV rectal carcinoma patient who was initially unresectable according to the National Comprehensive Cancer Network guidelines with a clinical risk score of 4 but received conversion surgery combined with systemic chemotherapy and achieved a favorable long-term clinical outcome (pathologic complete response) of approximately 28 months. Furthermore, serial circulating tumor DNA monitoring using next-generation sequencing provided a comprehensive view of the patient’s clinical and pathologic status for better clinical decision support over the course of the disease. The absence of circulating tumor DNA/cells after conversion surgery was correlated with pathologic complete response. This case study not only demonstrated that a curative oncosurgical approach could be considered for high-risk colorectal liver metastasis patients under specific circumstances but also highlighted the role of circulating tumor DNA monitoring to gain further insight into the evolution of a patient’s response over time.
Introduction
Colorectal cancer is the third most common cancer worldwide, and its incidence continues to grow. Approximately one-third of patients with colorectal cancer develop liver metastases (CRLM) during the natural course of disease. 1 Complete surgical resection is associated with very low mortality in CRLM patients and can provide long-term survival approaching 40%, 2 compared to a median survival rarely exceeding 9 months without such treatment. 3 Only a small fraction of CRLM patients (10%–20%) fulfill the selection criteria for surgical resection, including resection margin, size, number, and location of liver metastasis (LM), synchronous LM, stage of the primary tumor, and peritoneal dissemination. 4 However, recent advances in multimodality therapy, including portal vein embolization, ablative techniques, and chemotherapy, known as conversion therapy, 5 has increased the resectability of CRLM patients. Furthermore, the analysis of circulating tumor DNA (ctDNA), also known as “liquid biopsy,” is a noninvasive way to detect and measure cancer-specific molecular alterations in blood. 6 Recent data suggest that serial assessment of ctDNA may represent a promising approach for monitoring treatment response, with early decreases in ctDNA serving as a predictor of response.7,8
Case presentation
A 34-year-old male patient with right upper quadrant discomfort and hematochezia was admitted to the Hospital on 10 January 2018. The levels of carcinoembryonic antigen (CEA) were 73 ng/mL, CA19-9 > 12,000 U/mL, CA125 338 U/mL, and AFP (alpha-fetoprotein) 3 ng/mL. The colonoscopy identified a tumor located in the rectum about 5–7 cm above the anal verge (Figure 1(a) and (b)). Colonic stenosis was present, but no obstruction. The computed tomography (CT) and magnetic resonance imaging (MRI) scans confirmed that the tumor was a synchronous CRLM consisting of over 100 LMs (Figure 1(c)). The patient was identified with a KRAS G12D mutation, while NRAS, BRAF, and PIK3CA were wild type based on next-generation sequencing (NGS), HER2 amplification (−) determined by fluorescence in situ hybridization (FISH), and programmed death-1 (PD-1)/ programmed death ligand-1 (PD-L1) (−) probed by immunohistochemistry (IHC) staining. According to the National Comprehensive Cancer Network (NCCN) guidelines (2018), 9 this was an unresectable cT3N+M1 stage-IV rectal adenocarcinoma patient with a clinical risk score (CRS) 10 of 4. The patient did not have any family medical history.
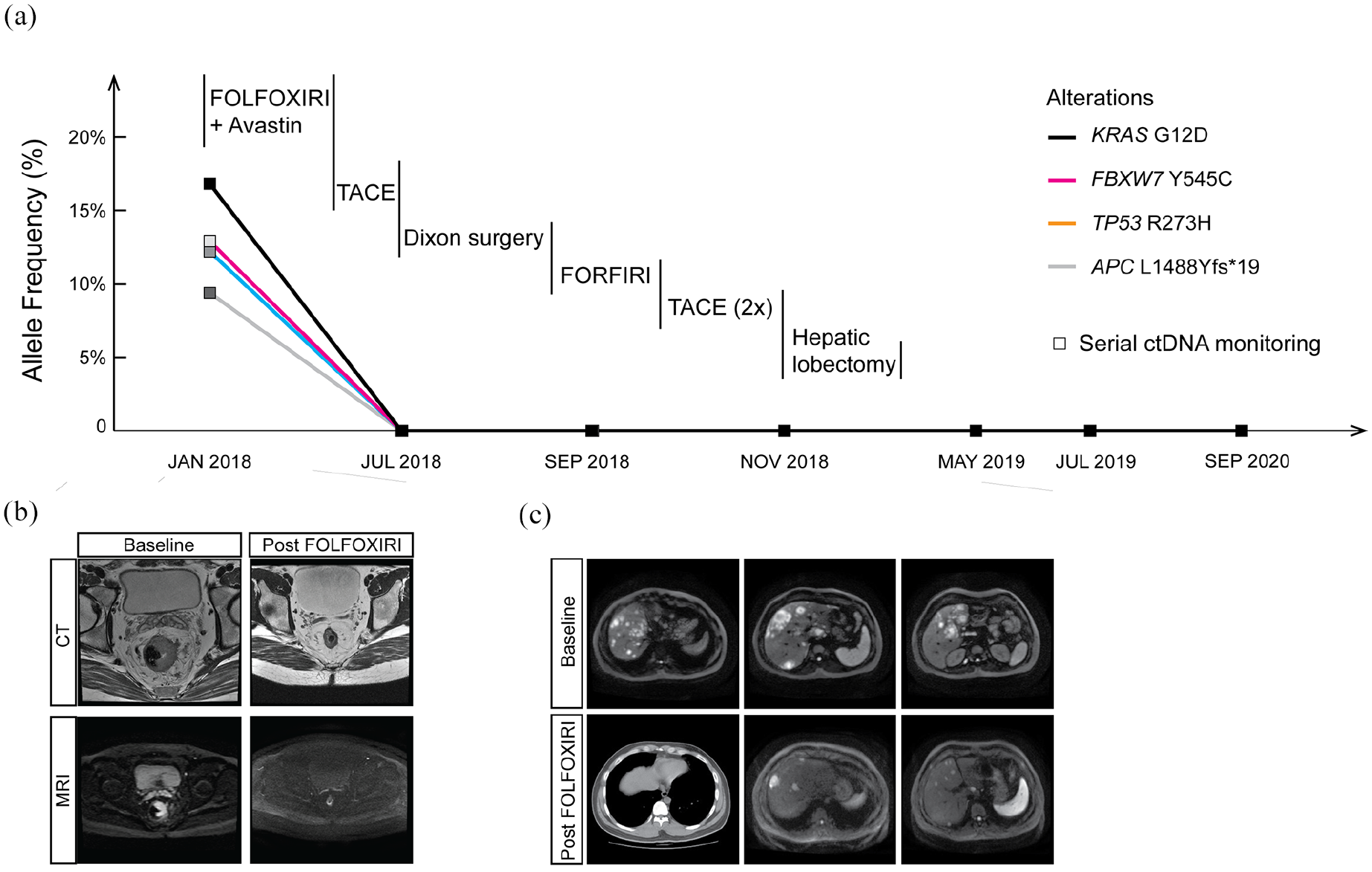
Circulating tumor DNA (ctDNA) monitoring in plasma during the clinical disease course: (a) serial ctDNA monitoring in response to conversion therapy in a high-risk stage-IV rectal carcinoma patient with synchronous liver metastases. (b) The magnetic resonance imaging (MRI) scans of the baseline and the post-chemotherapy phase of the patient who was diagnosed with advanced colon carcinoma with numerous synchronous liver metastases. (c) Computed tomography (CT) scans of the baseline and the post-chemotherapy phase of the patient’s liver metastases.
The patient received a combination therapy of FOLFOXIRI with bevacizumab (Avastin) for six cycles. Tumor biomarkers dramatically decreased, and MRI scans showed a partial response (PR) in the primary tumor and LMs according to RECIST v1.1 11 (Figure 1(b) and (c)). Following three additional cycles of FOLFIRI, the patient underwent a transarterial chemoembolization (TACE) 12 treatment along with 100 mg of CPT-11. Radiography showed no apparent activity in tumors, as well as undetectable ctDNA or circulating tumor cells (CTCs) (Figure 1(a) and Methods). The patient was declared no evidence of disease (NED) with only a small ulcer detected in the rectum by colonoscopy.
On 18 July 2018, the patient received laparoscopic Dixon surgery after multidisciplinary consultation and experienced a pathologic complete response (pCR). The surgical margins had no sign of malignancy, but many foamy macrophages and lymphoplasmacytic cells. Plasma ctDNA and CTC remained undetectable (Figure 1(a)). MRI revealed that there were at least 20 nodules in the liver. The patient was thereby given another round of FORFIRI but experienced severe fatigue. The patient subsequently received two TACE treatments and according to radiography, the tumors remained inactive. This led us to consider if the patient could benefit further from liver surgery, given the likelihood of pCR of the liver lesion. The patient was informed of all possible risks of surgery considering the presence of numerous nodules and the lack of precedent cases. On 19 December 2018, the patient underwent surgical liver treatment, including middle hepatic lobectomy, left partial hepatic lobectomy, right posterior hepatic lobectomy, partial hepatic caudate lobectomy, and intraoperative radiofrequency ablation (RFA; Figure 2). A total of 17 nodules were surgically removed, two were radiofrequency ablated, while another two nodules in the liver S3 and S6 were not located by MRI. The patient achieved pCR (CRS 0) and was relapse free at the last follow-up on 9 March 2021 (Figure 1(a)).
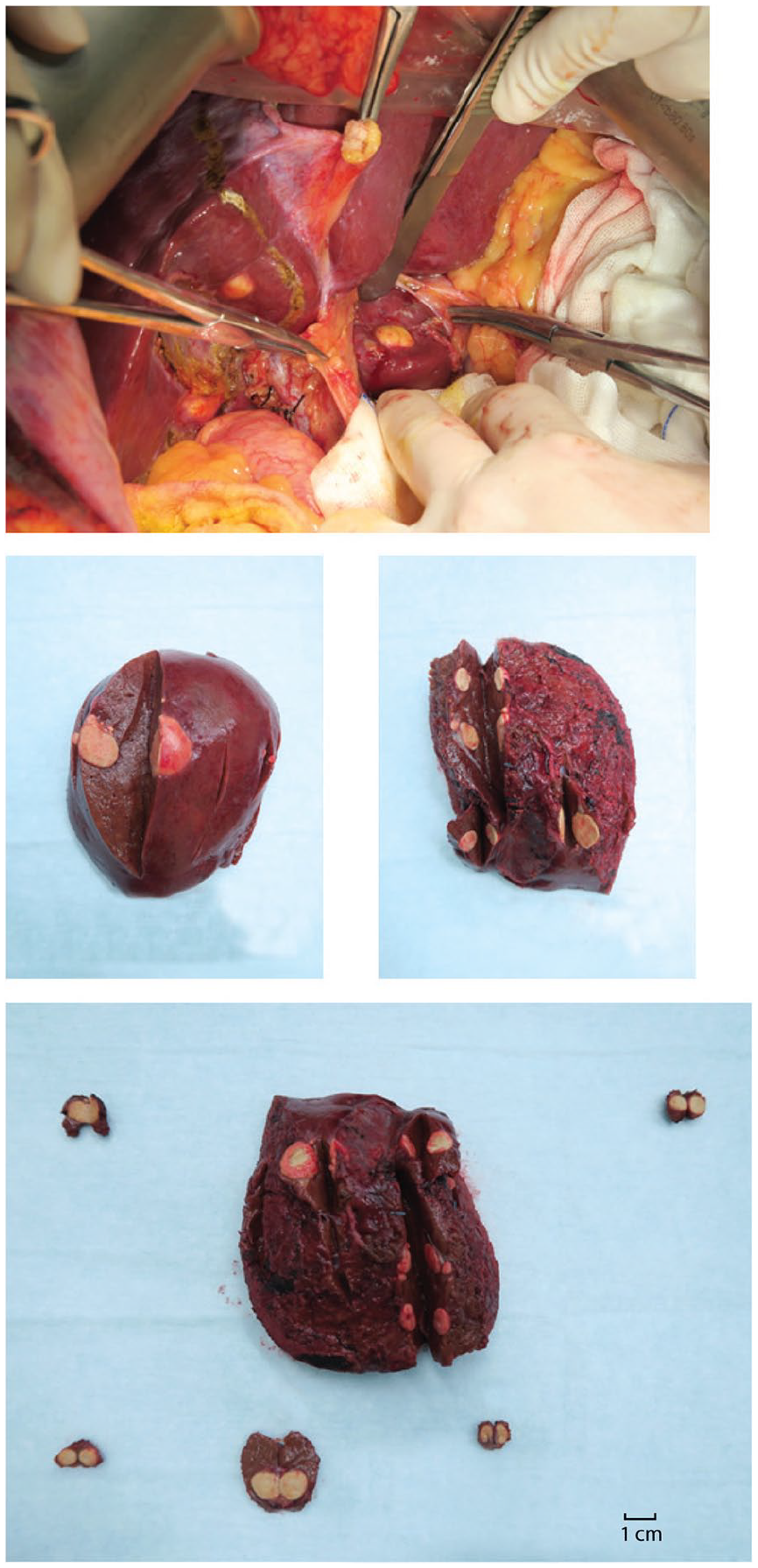
A curative surgical procedure of the patient’s hepatic metastases.
Discussion
Stage-IV colorectal cancer patients present with various metastatic sites, and the liver is one of the most common sites of synchronous and metachronous colorectal metastases. Conversion surgery therapies, including liver resection, RFA, or microwave coagulation therapy (MCT) combined with systemic chemotherapy, have been reported for unresectable advanced gastric cancer patients with LMs. 13 However, the clinical significance of conversion surgery remains controversial for late-stage colorectal cancer patients. Herein, we described a high-risk stage-IV rectal carcinoma patient who was initially unresectable according to the NCCN guidelines, with a high CRS score of 4, but received conversion surgery combined with systemic chemotherapy and achieved a favorable long-term clinical outcome (pCR) of approximately 28 months. Furthermore, serial ctDNA monitoring using NGS provided a comprehensive view of the patient’s clinical and pathologic status, thus, providing clinical decision-making support over the course of the disease. The absence of ctDNA/CTC after conversion surgery was correlated with pCR. This case study not only demonstrated that a curative oncosurgical approach could be considered for high-risk CRLM patients under specific circumstances but also highlighted the value of ctDNA monitoring to gain further insight into the evolution of a patient’s response over time.
Methods
Patient approval
This study was approved by the Institutional Review Board/Ethics Committee of Beijing Hospital, China. Written informed consent was obtained from the patient for publication of the clinical data, sequencing data, and accompanying images. The FISH assay for HER2 testing and IHC staining for PD1/PD-L1 proteins were performed according to the protocols approved by the Department of Pathology of Beijing Hospital.
ctDNA extraction, targeted enrichment, and NGS
Genomic profiling of the patient’s ctDNA was performed by a Clinical Laboratory Improvement Amendments–certified, College of American Pathologists–accredited laboratory (Nanjing Geneseeq Technology, Jiangsu, China). ctDNA from plasma was purified using the Circulating Nucleic Acid Kit (Qiagen, Germany) following the manufacturer’s protocol. DNA was quantified using the dsDNA HS Assay Kit on a Qubit Fluorometer (Life Technologies, CA, USA). Sequencing libraries were prepared using the KAPA Hyper Prep Kit (Roche, Switzerland), as described previously. 14 Indexed DNA libraries were pooled for probe-based hybridization capture of the targeted gene regions covering 425 cancer-related genes (GeneseeqPrimeTM).
Sequencing was performed on the Illumina HiSeq4000 platform followed by data analysis, as previously described. 15 In brief, sequencing data were analyzed by Trimmomatic 16 to remove low-quality (quality < 15) or N bases and then mapped to the human reference genome hg19 using the Burrows-Wheeler Aligner (https://github.com/lh3/bwa/tree/master/bwakit). Polymerase chain reaction (PCR) duplicates were removed by Picard (available at: https://broadinstitute.github.io/picard/). The Genome Analysis Toolkit (GATK) (https://software.broadinstitute.org/gatk/) was used to perform local realignments around indels and base quality reassurance. Single-nucleotide polymorphisms (SNPs) and indels were analyzed by VarScan2 17 and HaplotypeCaller/UnifiedGenotyper in GATK, with a mutant allele frequency (MAF) cut-off value of 0.1% for ctDNA samples and a minimum of three unique mutant reads. Common SNPs were excluded if they were present in >1% population frequency in the 1000 Genomes Project or the Exome Aggregation Consortium (ExAC) 65,000 exomes database. The resulting mutation list was further filtered by an in-house list of recurrent artifacts based on a normal pool of whole blood samples.
CTC detection
CTCs in the peripheral blood were efficiently captured using the CTCBIOPSY® device (Wuhan YZY Medical Science and Technology Co., Ltd., Wuhan, China) by size of epithelial tumor cells through a polymer membrane made by biocompatible parylene and Wright’s staining as previously described. 18 The specific procedure was carried out by the laboratory professionals of Wuhan YZY Medical Science and Technology company following the manufacturer’s protocol19,20 approved by the Department of Pathology of Beijing Hospital. The cut-off value of CTC detection was CTC count ⩾3.
Footnotes
Acknowledgements
The authors would like to thank the patient and his family for providing consent to participate in this study, as well as all of the research staff and co-investigators involved.
Author contributions
Y.H. contributed to the study’s conception and design, data acquisition, manuscript writing, and study supervision; W.J. contributed to the acquisition of data; L.W. contributed to the acquisition of data; Q.O. contributed to the data analysis and interpretation, and manuscript writing; X.W. contributed to data analysis and interpretation; B.X. provided administrative and material support.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Key Research Program for Health Care of China (grant no. W2016ZD01).
Conflict of interest statement
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Q.O. and X.W. are employees of Geneseeq Technology Inc., Canada. The remaining authors have nothing to declare.
Data availability statement
Data sharing is not applicable to this article as no new data were created or analyzed in this study.
