Abstract
A single event can cause a life-long memory. Memories physically reside in neurons, and changes in neuronal gene expression are considered to be central to memory. Early models proposed that specific DNA methylations of cytosines in neuronal DNA encode memories in a stable biochemical form. This review describes recent research that elucidates the molecular mechanisms used by the mammalian brain to form DNA methylcytosine encoded memories. For example, neuron activation initiates cytosine demethylation by stimulating DNA topoisomerase II beta (TOP2B) protein to make a temporary DNA double-strand break (repaired within about 2 hours) at a promoter of an immediate early gene,
Keywords
Introduction
Griffith and Mahler 1 and Holliday 2 were among the first to propose that memories reside in a code of DNA cytosine methylations (Figure 1) in brain neurons.
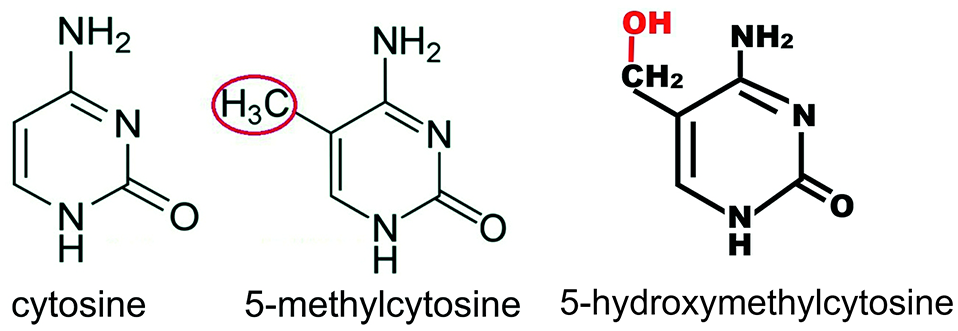
Cytosine, when methylated, has a methyl group added on the 5 position as shown in red oval. The methyl group can be oxidized to form 5-hydroxymethylcytosine.
Recently, several thousand alterations were found to occur in neuronal methylated cytosines during formation of a long-term memory.3,4
Blocking the occurrence of these alterations blocks memory formation. This review describes the neuronal factors that catalyze the formation or removal of cytosine methylation in the DNA of neurons. It also describes how cytosine methylation functions to control gene transcription. In addition, the review indicates how alterations in neuronal nuclear messenger RNA expression affect protein expression within the neuronal dendrites that form synaptic interconnections.
Early Approaches
The model for the location and formation of memories proposed by Griffith and Mahler in 1969 was based on the chemical nature of DNA. 1 They noted that, in man, memories may survive for almost the entire lifetime. Further, DNA is the one molecule which, apart from possible minor alterations due to genome damage and repair, is surely present in neurons for the entire lifetime of the organism. They noted that nerve cells do not divide and suggested that this may have been selected for, in order to avoid disturbing the learned information which is somehow stored in their DNA. In particular, Griffith and Mahler 1 suggested that the physical basis of memory could lie in the enzymatic modification of the DNA of nerve cells. They suggested that the DNA of a nerve cell may be thought of as a long tape, most of which contains in coded form a representation of the amino-acid sequences of proteins, but with short regions concerned only with control of the transcription or translation of this information. Their model of memory involved the chemical alteration of these control regions, likely by the methylation or demethylation of DNA cytosine. They pointed out that enzymatic modification (methylation of cytosine) at the control regions may affect the level of recognition of the DNA by enzymes, or that the modifications of the control regions may affect other control elements such as inducers or repressors.
Holliday 2 in 1999 also noted that long-term human memory can be retained for many decades. He suggested that the exceptional stability required implies that essential memory components are based on chemical changes. He proposed that the enzymatic modification of cytosine in DNA to 5-methylcytosine may provide this necessary stability. His general model was that specific sites in the DNA of neurons required for memory can exist in methylated or non-methylated states. The initial signal, that is to be memorized, switches the DNA from a modified to an unmodified state, or vice versa. He noted that the presence or absence of DNA methylation at a particular sequence of DNA can be thought of as a 0, 1 binary code. Thus, as noted by Holliday, “10 such sites have 210 (1024) epigenotypes and potential phenotypes, and 30 such sites could have up to 230, or 1.07 × 109 epigenotypes.” He indicated that such a set of control mechanisms has enormous potential for neuronal specificity.
When the Giffith and Mahler 1 and Holliday 2 proposals for the location of memory were presented, the mechanisms by which DNA cytosine methylation and demethylation occur were unknown.
The elements of the models of Griffiths and Mahler and of Holliday were impressively predictive of what has recently been found to be a major feature of the basis of memory. By 2002, Bird 5 reviewed the evidence that methylcytosine in the promoter (control) region of genes participates in repression of gene expression. In 2011, Yu et al 6 published an article titled “DNA methylation mediated control of learning and memory” that identified cytosine methylation changes at specific gene loci in response to memorable events. They also noted the presence of methylating enzymes in neurons. In 2015, Heyward and Sweatt 7 reviewed the evidence that there is short-lived DNA methylation within the hippocampus for memory consolidation, which is a temporary precursor to subsequent cortex-mediated long-lasting memory storage in methylated DNA. By 2016, Oliveira 8 reviewed a large body of evidence indicating that methylations and demethylations of cytosines in promoter regions of genes in neurons have a central role in memory formation and maintenance. In 2020, Poon et al 9 gave a detailed description of the actions of the major enzymes (DNMT3a, MeCP2, and DNA demethylases) central to carrying out the DNA methylations/demethylations that are the basis of memory formation.
Control of Expression by Cytosine Methylation
Methylation of cytosine is now recognized as one of the main epigenetic mechanisms that regulate gene expression. 10 Methylated cytosine is frequently present in the linear DNA sequence where a cytosine is followed by a guanine in a 5′ to 3′ direction (a CpG site). In mammals, DNA methyltransferases (which add methyl groups to DNA bases) exhibit a sequence preference for adding methyl groups to cytosines at CpG sites. 11 About 28 million CpG dinucleotides occur in the human genome. 12 In most tissues of mammals, on average, 70% to 80% of CpG cytosines are methylated (forming 5-mCpG). 13 Methylated cytosines within 5′cytosine-guanine 3′ sequences often occur in groups, called CpG islands, within the promoter regions of genes. Promoter regions are DNA sequences upstream of the genes they control. Methylation of cytosines in CpG sites in promoter regions can reduce or silence gene expression. 5 Methylated cytosines in the gene body, however, are positively correlated with expression. 14
The mechanism by which 5-mCpG sites in promoters repress gene transcription appears to depend on the family of proteins with a methyl-CpG binding domain (MBD). These proteins include MBD1, MBD2, MBD4, and MeCP2. As described by Baubec et al, 15 when these MBD proteins are bound to 5-mCpGs at promoters, they recruit chromatin remodelers, histone deacetylases and methylases, leading to transcriptional repression. There is a strong negative correlation between MBD binding and promoter activity.
Alteration of expression by 5-hydroxymethylcytosine in neurons
The methyl group on cytosine in DNA can be converted into a hydroxymethyl group, forming 5-hydroxymethylcytosine (5-hmC) (Figure 1). This conversion is carried out by the enzyme tet methylcytosine dioxygenase 1 (TET1). TET1 is present at high levels in neurons in brain regions where memories are first formed. 16 Conversions of 5-mC to 5-hmC is the initial step in a sequence of oxidative steps, followed by DNA base excision repair, resulting in un-modified cytosine. However, 5-hmC itself is fairly stable and is likely the main product of TET1 enzyme activity in neurons. 17
As reviewed by Clemens and Gabel,
18
DNA cytosines in brain neurons are methylated and hydroxmethylated at levels that are very different than in most tissues of the body. Neurons have about 11 million 5-mCpG sites and about 6 million 5-hmCpG sites in their DNA. By comparison, other tissues have about 17 million 5-mCpG sites in their DNA and about ½ million 5-hmCpG sites. In addition, neurons have about 23 million 5-mCp
Oxidation of 5-mCpG to 5-hmCpG converts high-affinity MeCP2 binding sites to low-affinity sites, resulting in diminished MeCP2 binding, release of its repressive effect, and increased transcription of the genes affected. 19 In contrast, oxidation of 5-mCpH to 5-hmCpH does not affect MeCP2 binding and does not appear to affect transcription. 19
Initial Events in Neural Activity
A memorable event in the environment can initiate neural activation. Memorable events often include things that are seen. An example of a very simple event is a sudden alteration of light at a spot in the environment. Such an event can activate an optic neuron. Light-altered neuron activation occurs within about 5-20 ms in a rabbit retinal neuron. 20 The initial activation can be detected by an action potential spike, a sudden spike in neuron membrane electric voltage. A more complex event may consist of the presentation to a person of an anomalous word. If the person is shown a sentence, presented as a sequence of single words on a computer screen with a puzzling word out of place in the sequence, the sight of the puzzling word can register on an electroencephalogram (EEG). In one experiment, human readers wore an elastic cap with 64 embedded electrodes distributed over their scalp surface. 21 Within 230 ms of encountering the anomalous word, the human readers generated an event-related electrical potential alteration of their EEG at the left occipital-temporal channel, over the left occipital lobe and temporal lobe (Figure 2 top).

Panel A: Outer parts of the brain including cerebrum lobes. Panel B: Deep structures including brain regions involved in memory formation .
Potentially memorable events are thought to initially register in the brain by activating neurons. After the initial electrical spike in a rabbit activated optical neuron, there may be several further spikes, spaced about 10 ms apart followed by return of the neuron to its basal state. Similarly, with the more complex event in humans (an aberrant word in a sentence) the electrical state in the brain is altered for a brief period but then returns to the basal level.
The electrical spikes measured on the neural membrane, or the cerebral surface charges measured in an EEG reflect what can be seen from the outside of the neuron or the cerebrum. However, electron configurations altered by nerve cells not only affect the outer electrical fields but also affect the electron configurations at the cell nucleus. In cultured neurons from the hippocampus (Figure 2 bottom), spontaneous electrical activity is sufficient to cause nuclear export of histone deacetylase 4 protein. 22
Although these findings indicate that external events can cause transient electrical changes in neurons, they do not explain long term effects.
Changes in Gene Expression are Central to Memory
Long-term plastic changes in the brain, including memory formation, depend on permanent functional alterations in neurons, rather than temporary voltage spikes. Memory formation requires neuronal reprogramming of gene expression.
23
In the brain, neuronal gene expression is changed in response to neuronal activity. Immediate early genes (IEGs) are defined as genes that are expressed rapidly and transiently in response to a stimulus. In particular, the expression of neuronal IEGs, such as
A Major Pathway From Neuronal Activation to Neuronal DNA Cytosine Demethylations
Brief overview of the pathway
This sub-section is a very brief overview of the actions of the neuronal cellular elements that control gene expression leading to memory formation. Enhancer regions of the genome are key elements of this process. Enhancers are DNA sequences that are often tens of thousands of nucleotides distant along the DNA sequence from their target promoters but never-the-less cooperate with their target promoter DNA sequences to control target gene transcription. Chromosome loops bring enhancers to promoters (Figure 3).

Enhancer promoter interactions. An active enhancer regulatory region is enabled to interact with the promoter region of its target gene by formation of a chromosome loop. This can initiate messenger RNA (mRNA) synthesis by RNA polymerase II (RNAP II) bound to the promoter at the transcription start site of the gene. The loop is stabilized by one architectural protein anchored to the enhancer and one anchored to the promoter and these proteins are joined together to form a dimer (red zigzags). Specific regulatory transcription factors bind to DNA sequence motifs on the enhancer. General transcription factors bind to the promoter. When a transcription factor is activated by a signal (here indicated as phosphorylation, shown by a small red star on a transcription factor on the enhancer) the enhancer is activated and can now activate its target promoter. The active enhancer is transcribed on each strand of DNA in opposite directions by bound RNAP IIs. Mediator (a complex consisting of about 26 proteins in an interacting structure) communicates regulatory signals from the enhancer DNA-bound transcription factors to the promoter.
Transcription factor proteins
26
are associated with enhancer DNA sequences.
27
Such transcription factors are essential to activate RNA polymerases that are bound to promoters of the target genes. In the case of at least 6 neuronal immediate early genes (IEGs), the enhancer appears to become persistently attached to the promoter by an architectural protein dimer, CTCF,
28
which also inhibits transcription. As described in Sections titled Topoisomerase II beta DNA double-strand breaks can initiate transcription and Repair of the IEG signal-reponsive double-strand breaks below, upon neuronal activation, a topoisomerase II beta protein bound to the DNA adjacent to CTCF creates a temporary DNA double-strand break that liberates the promoter from CTCF inhibition. Then the neuronal IEG genes are expressed. One IEG,
Sequence of events from initial activation to neuronal reprograming
Signals from the outside environment can cause immediate phosphorylation (activation) of a protein in the cytoplasm of a neuron. The RAF-MEK-ERK kinase cascade is a chain of kinases (proteins that phosphorylate other proteins) in the neuronal cytoplasm that communicates a signal from the surface of a neuron to its nucleus. Upon receiving a cell surface signal, RAF kinase phosphorylates and activates MEK. MEK phosphorylates the protein designated “extracellular signal-regulated kinase (ERK)” Increased phosphorylated ERK (pERK) is found in the nucleus within 2 minutes following both brief and sustained neuronal activity. 29 Once in the nucleus, pERK rapidly phosphorylates the transcription factor protein ELK-1. 29
As indicated by Li et al, 30 ELK-1 primarily controls immediate-early genes (IEGs), all of which have low levels of basal transcription but can reach high levels after stimulation. ELK-1 constitutively binds to immediate-early response gene enhancers without requiring activation. When pERK phosphorylates 2 serines in ELK-1 (Ser383 and Ser389), this allows ELK-1 to activate transcription of the enhancer DNA sequence to which it is bound.
Within 7.5 minutes after neural activation, enhancers that target promoter regions of IEGs begin transcribing their enhancer RNAs (eRNAs) (Figure 3) (see Section below titled Activated enhancers produce eRNAs). The enhancers with attached eRNAs interact with their targeted IEGs and promote their transcription (see Section titled Activated enhancers produce eRNAs). For 6 of the IEGs whose functions are important in learning and memory, the enhancer regions of the genome that target those IEGs usually have associations with the promoters of their target genes that were established prior to neuronal activation (pre-formed associations). These associations appear to be maintained by loops anchored by the dimer-forming protein CTCF (see Section titled Enhancers and chromatin loops). The enhancer regions have regulatory DNA sequences to which transcription factors bind (see Section Transcription factors (TFs) and the mediator complex). These transcription factors are essential for activating the RNA polymerase II (RNAP II) bound to the promoters of the IEG genes. However, CTCF proteins not only form and anchor loops but are able to insulate the transcription factors on the enhancers from activating the RNAP II on the IEG promoters. At this point, about 15 minutes after neural activation, topoisomerase II beta (TOP2B), which is bound to the promoters adjacent to the CTCF proteins, makes temporary (up to 2 hour) DNA double-strand breaks in the promoter regions of 6 memory-related IEGs (see Section Topoisomerase II beta DNA double-strand breaks can initiate transcription). The double-strand break physically frees the promoters of these IEGs to move and to circumvent the insulation function of CTCF. The IEG promoters, with their bound RNAP II, can now closely associate with the transcription factors on their enhancers, resulting, at 15 minutes after neural activation, in initiation of RNA transcription of those IEG genes.
For proper formation of memory, the IEG proteins must not be over-produced. Adjacent to the TOP2B protein on the promoter of the IEG genes, there are also proteins of the DNA repair pathway called non-homologous end joining (NHEJ) (see Section 6.7 Repair of the IEG signal-responsive double-strand breaks). Within 2 hours of formation of TOP2B-induced breaks in IEG promoters, NHEJ seals the breaks and IEG transcriptions return to their low basal levels.
The next step in memory formation is initiation of demethylation of cytosines at many genes, carried out by TET1 enzymes (see Section Alteration or removal of the methyl group on cytosine). TET1 can carry out 3 steps on cytosine methylated at its 5 position if the cytosine is followed by a guanine (a 5-mCpG site). TET1 can form 3 sequential intermediate products. The latter 2 products can be removed by base excision repair enzymes to produce un-modified cytosine.
17
The first product of TET1 action on a 5-mCpG site is 5-hydroxymethylcytosine (Figure 1) at that site (5-hmCpG). 5-hmCpG is a stable molecular configuration and may be the main product of TET1 in neurons. The presence of 5-hmCpG allows the genes with initiated demethylation to have long-term upregulated expression. TET enzymes only bind specifically to 5-methylcytosine if they are recruited. When the IEG
At the same time, in addition to conversion of 5-mCpG to 5-hmCpG at many genes, new methylations at CpG sites occur at many other genes.
Contextual fear conditioning, a memorable event, occurs when a rat or mouse receives an electrical shock of their foot when they have been recently placed in a novel location. The rat or mouse then shows fear in that novel location for a long time afterward. Within 1 hour of contextual fear conditioning and lasting to 24 hours after the event, about 500 genes with newly formed 5-hmCpG and about 1500 genes with newly methylated 5-mCpG sites are found in the hippocampus of such rats or mice (see Section DNA methylation alterations upon the creation of long-term memories). Overall, more than 9% of the genes in the genomes of rat hippocampus neurons are differentially methylated after contextual fear conditioning. These methylations and initiated demethylations in the hippocampus are the basis of the short term memory of the event. However, over a period of 4 weeks, the methylation and initiated demethylation patterns are transferred, by a process not yet understood, to the anterior cingulate cortex (Figure 2 bottom) where long term memories are stored.
There is also another mechanism for bringing TET1 enzymes to genes to allow demethylation initiation of those genes in memory formation. Reactive oxygen species (ROS) also have a role in forming long-term memories (see Section Another pathway: the role of reactive oxygen species in demethylation and memory).
Activated enhancers produce eRNAs
As reviewed by Hou and Kraus 31 enhancers are DNA sequences that act as nucleation sites for the binding of sequence-specific transcription factors. Enhancers associate with target genes that they regulate through enhancer–promoter contacts facilitated by chromatin looping (Figure 3). Most enhancers are bound by multiple RNA polymerase IIs (RNAP IIs) and the enhancer DNA sequence is actively transcribed on each strand, producing eRNAs, long non-coding RNA molecules complementary to their enhancer DNA sequences. The roles of eRNAs are somewhat unclear. Hou and Kraus 31 suggest that the 3 primary roles of eRNAs are: (i) promoting enhancer–promoter looping; (ii) promoting the recruitment of transcription factors and co-regulators, and regulating their activities; and (iii) facilitating RNAP II pause–release at promoters to promote transcription elongation of target genes. Arnold et al 32 suggested that eRNAs contribute to (i) enhancer function by interacting with nuclear proteins to promote enhancer-promoter looping; (ii) chromatin modification; and (iii) regulation of transcriptional machinery. It is possible that eRNAs assume configurations that allow them to act catalytically as ribozymes, to carry out such functions.
In neurons from primary neuron cultures, Carullo et al
33
found 28 492 putative enhancers that were transcribing their DNA (generating eRNAs). These eRNAs were often transcribed from both strands of the enhancer DNA, acting in opposite directions (presumably with 2 different RNA polymerases). Carullo et al
33
used these cultured neurons to examine the timing of enhancer eRNAs compared to mRNAs of the target genes of the enhancers. The cultured neurons were depolarized (activated) and RNA was isolated from those neurons at 0, 3.75, 5, 7.5, 15, 30, and 60 minutes after treatment. Under these conditions 2 of the 5 enhancers of the IEG
Enhancers and chromatin loops
Enhancers are regulatory DNA sequences in the genome that cooperate with promoter DNA sequences, usually on the same chromosome, to control target gene transcription. For example, RNAP II is bound to the promoter region of the
Enhancer regions include regulatory DNA sequences to which transcription factors and mediator complex proteins bind (Figure 3), and several RNAP IIs and other proteins also bind. When an enhancer is activated, its bound RNAP IIs usually transcribe enhancer-expressed non-coding RNAs (eRNAs) (Figure 3). The 2 complementary strands of the enhancer DNA genome region are each ordinarily transcribed by RNAP IIs
33
as indicated in Figure 3. The transcription factors, other proteins and eRNAs at the enhancer all have roles in facilitating the action of RNAP II on the target gene promoter in transcribing the target gene.
35
The IEG
Chromosome loops facilitate the interactions of enhancers with promoters. Chromosome loops are formed by one end of one member of a dimer-forming architectural protein being bound to a DNA sequence at its binding motif in an enhancer, and then the other end of this protein adhering to the other member of the dimer-forming protein that is bound to its binding motif in a target gene promoter (Figure 3). Most chromosome loops are formed by pairs of Yin Yang 1 (YY1) proteins or CCCTC-binding factor (CTCF) proteins. 37 These proteins are highly conserved in evolution and have multiple zinc finger DNA binding domains.28,37
There are more than 24 000 constitutive loops that associate enhancers with promoters in mouse embryonic brain cortex stem cells and a similar number in the human cortex. However, more than 10% of the loops surrounding IEGs, secondary response genes, and synaptic genes are induced de novo during cortical neuron activation. IEGs
Transcription factors (TFs) and the mediator complex
Transcription of genes is carried out by RNA polymerase II (RNAP II). However, transcription factor (TF) proteins must associate with RNAP II at a DNA sequence upstream of the gene, the promoter, in order for RNAP II to begin transcription. 39
TFs are defined as proteins capable of binding DNA in a sequence-specific manner and which also have an effect on regulating transcription. 26 There are about 777 TFs. 40
TFs are usually associated with a mediator complex that forms the bridge between gene-specific TFs and the RNAP II machinery. 41 The mediator complex is comprised of about 26 proteins in mammals. 42 The mediator complex is found at enhancers rather than at core promoters, and different TFs interact with distinct mediator proteins, leading to gene-specific physiological effects. 41 Different TFs are activated in response to different signaling pathways. Different TFs activate transcription, in part, by interacting with different mediator proteins. Thus specific proteins of the mediator complex are able to convert the biological input (eg, neuron activation) to specific transcriptional outcomes. 42
Topoisomerase II beta DNA double-strand breaks can initiate transcription
Madabhushi et al
43
found that some memory-associated IEG genes, including
Activation of cortex neurons (Figure 2 top) (though the cortex neurons studied by Madabhushi et al were in culture) caused the TOP2B associated with some memory-related IEG promoters to create a temporary (up to 2 hours), double-strand break in the promoter DNAs. This break allowed the promoters of these IEGs to physically move to circumvent the insulation function of CTCF. The promoters physically moved to closely associate with their enhancers, resulting in initiation of RNA transcription of those IEG genes during the 2 hour temporary breaks. 43
The double-strand breaks that occur after activation of cortex neurons in culture are found at only 21 locations on the genome.
43
These include breaks at 6 IEG promoters of the memory-related genes
After Madabhushi et al 43 showed that activation of neurons for 20 minutes in tissue culture caused DNA double-strand breaks, Madabhushi et al 43 examined the brains of mice that had new associative memories. These memories were induced when the mice were subjected to contextual fear conditioning. New memories form temporarily in the hippocampus 46 (Figures 2 and 4). Within 15 minutes of contextual fear conditioning, the mouse hippocampal neurons were found to have DNA double-strand breaks. This conditioning caused hundreds of gene-associated DSBs in the medial prefrontal cortex (mPFC) and hippocampus (Figure 2) that are important for learning and memory. 47 Similarly, Suberbielle et al 48 allowed mice to explore a novel environment, developing new memories. They found that DNA double-strand breaks occurred in such mice in their dentate gyrus (Figure 4). The dentate gyrase is involved in spatial learning and memory.

Drawing by Camillo Golgi of a hippocampus and dentate gyrus. Public domain.
As described above, Madabhushi et al
43
found that the formation of persistent double-strand breaks at IEGs facilitated mRNA transcription of these IEGs. The transcription of these genes begins within about 15 minutes of receiving the neuronal signal because RNAP II is already present but paused at the transcription start site. There appears to be maximum expression at about 50 minutes and reduced expression by 70 minutes.
49
The transcript levels of
Repair of the IEG signal-responsive double-strand breaks
Most of the neuronal signal-responsive, persistent promoter double-strand breaks are repaired by non-homologous end joining (NHEJ).43,50 Ju et al 50 carried out an extensive study of initiation of mRNA transcription after TOB2B-induced double-strand breaks. In particular, they determined the DNA repair enzymes bound along with TOP2B at the genomic location that could respond to stimulation with the rapid occurrence of a double-strand break. Using estrogen stimulated cells from the MCF-7 breast cancer cell line, they found that TOP2B, PARP-1, DNA-PKcs, and KU86 and KU70 are closely associated with an estrogen-responsive promoter. These proteins are all present simultaneously on a single promoter nucleosome (there are about 147 nucleotides in the DNA sequence wrapped around a single nucleosome). Thus, DNA-PKcs, and KU80 and KU70, which carry out the first steps of NHEJ DNA repair, appear to be in close association with TOP2B at the promoter of signal-responsive genes (Figure 5).

Paused RNA polymerase and limited, short-term TOP2B-induced DNA double-strand break.
As indicated by Szlachta et al, 51 in order to undergo NHEJ or microhomology mediated end joining (MMEJ) DNA repair, the covalently bound TOP2B that initiated the double-strand break may be first degraded by a nuclear proteasome (some proteasomes are located in the nucleus. 52 ) and then the remaining tyrosine-linked end can be freed by tyrosyl-DNA phosphodiesterase 2 (TDP2). The conversion of the covalently bound TOP2B DNA end to a protein-free end creates the persistent break that stimulates repair through the NHEJ or MMEJ DNA repair pathways. The NHEJ process is initiated by the binding of the KU70/80 heterodimer to both ends of the broken DNA molecule, and the KU70/80 heterodimer is attached to the kinase DNA-PKcs (the catalytic subunit of DNA-PK). The ends of the TOP2B-induced DSB are already suitable for ligation. Then the DNA LIGASE IV/XRCC4 complex can catalyze ligation of the DNA ends. 53
If the NHEJ DNA repair of the double-strand breaks is blocked in neurons, expression of the IEGs do not return to basal levels. 43 As shown by Colón-Cesario et al, 54 blocking NHEJ at the time of contextual fear conditioning prevents long-term memory formation. Blocking NHEJ at 2 hours after contextual fear conditioning, however, allows long-term memory of the event to occur. Thus, NHEJ to reconstitute intact DNA is needed in the first 2 hours of an event for formation of long-term memory of the event. Presumably, NHEJ is needed so that the IEGs are controlled and not excessively expressed. Inhibition of NHEJ in the hippocampus (Figure 2) at the time of the conditioning event reduced long-term memory while inhibition of NHEJ in the cortex (Figure 2) at the time of conditioning did allow long-term memory to form. 54 Kim and Jung 46 reviewed the evidence that the hippocampus region of the brain is where contextual fear memories are first stored, though this storage is transient and does not remain in the hippocampus. They point out that, in rats, contextual fear conditioning is abolished when the hippocampus is subjected to hippocampectomy just 1 day after conditioning. However, the rats retain a considerable amount of contextual fear when a long delay of 28 days is imposed between the time of conditioning and the time of hippocampectomy. Using localized lidocaine injections to impede brain functions, Frankland et al 55 showed that much of the long term storage of contextual fear conditioning memory takes place in the anterior cingulate cortex (Figure 2).
Alteration or removal of the methyl group on cytosine
At least one memory associated IEG encoded protein, EGR1, that was expressed in response to neuronal activation, next acts, in conjunction with a TET enzyme, to alter expression of target genes involved in memory formation.
Methylations of cytosines in promoter regions of genes inhibit expression of those genes, but those methylations can be oxidatively altered or removed by 10-11 translocation methylcytosine dioxygenases (TET enzymes). The alteration or removal can allow gene expression to occur. The 3 related
TET enzymes are dioxygenases in the large family of alpha-ketoglutarate (α-KG)-dependent hydroxylases. The activity of a TET enzyme is illustrated in Figure 6.
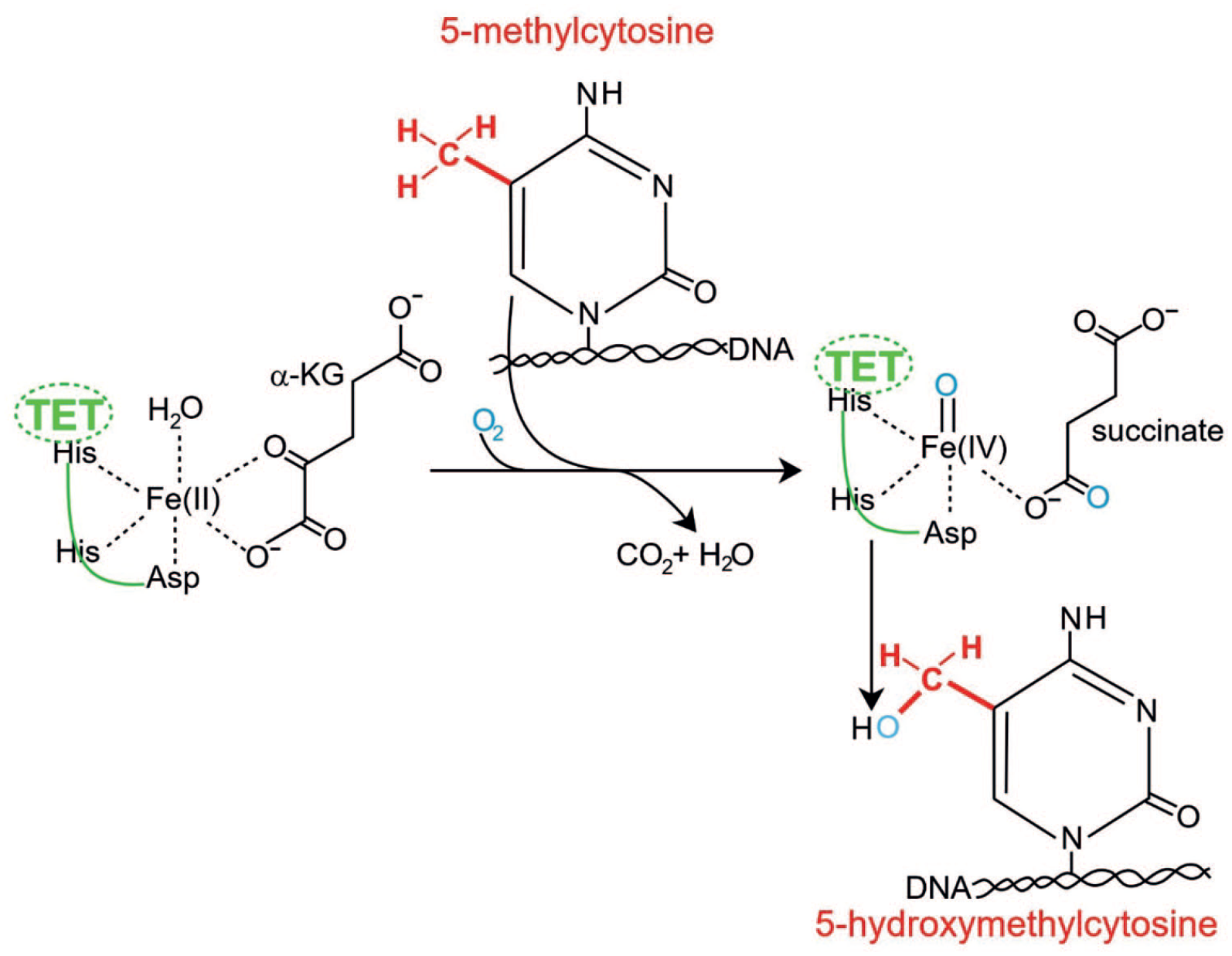
Conversion of 5-methylcytosine (5-mC) to 5-hydroxymethylcytosine (5-hmC) by a TET enzyme.
A TET enzyme has a typical hydroxylase active pocket (composed of histadine-histadine-aspartic acid) that catalyzes, in the presence of a ferrous ion, an oxidation reaction by incorporating a single oxygen atom from molecular oxygen (O2) into its substrate, 5-methylcytosine in DNA (5-mC). This reaction produces the product 5-hydroxymethylcytosine (5-hmC) in DNA. 57 This conversion is coupled with the oxidation of the co-substrate α-KG to succinate and carbon dioxide. The TET enzymes can also oxidize 5-hmC to further products which can then be converted to unmodified cytosine by DNA repair enzymes. However, the 5-hmC initial product appears to be fairly stable and may be the main product of TET enzyme activity in neurons.
Recruitment of TET1 to methylated cytosines
The TET enzymes only bind specifically to 5-methylcytosine if they are recruited. Without recruitment or targeting, TET1 predominantly binds genome-wide to promoters with high levels of CpGs and CpG islands by its CXXC domain that can recognize un-methylated CpG islands. 58 TET2 does not have an affinity for 5-methylcytosine in DNA. 59 The CXXC domain of the full-length TET3, the predominant form expressed in neurons, binds most strongly to CpGs where the C was converted to 5-carboxycytosine (5caC). However, it also binds to un-methylated CpGs. 60
The TET1 enzyme occurs in 2 isoforms, a full length version (TET1FL) and a shorter truncated form TET1s. 60 The short isoform TET1s is expressed in brain neurons while TET1FL is expressed in brain glial cells. 61
Sun et al
62
showed that the early growth response 1 protein EGR1 and TET1s form a complex mediated by the C-terminal regions of both proteins, independently of association with DNA.
EGR1 recruits TET1s to genomic regions flanking EGR1 binding sites. Sun et al 62 showed that TET1s binds to 1450 sites in the wild type mouse brain genome, but if EGR1 is absent then TET1s only binds to 855 sites in the genome. Thus when EGR1 proteins are expressed, they appear to bring TET1s to 595 sites in the genome. TET1s can then initiate demethylation of those 595 sites and likely activate the expression of those downstream genes in the mouse frontal cortex. Presumably some other regulatory protein(s) bring(s) TET1s to other sites in the brain. Some specific memory-related promoter sites are known to be binding sites for EGR1. If those promoter sites are methylated, EGR1 alone is not able to increase expression of the associated genes. However, if TET1s is also present then EGR1 is able to bring TET1s to the sites and this results in initiation of demethylation at those sites and expression of the associated genes. 62
DNA methyltransferases (DNMTs) methylate targeted genes
In addition to initiating demethylation at hundreds of sites when a memory is formed, an even higher number of methylations at specific sites are needed in formation of a memory (see Section DNA methylation alterations upon the creation of long-term memories).
The 3 mammalian DNA methyltransferases (DNMT1, DNMT3A, and DNMT3B) catalyze the addition of methyl groups to cytosines in DNA. Of these, DNMT1 is considered to act primarily as a “maintenance” methyltransferase by copying the CpG methylation patterns from parental DNA strands to daughter strands during DNA replication. In contrast, DNMT3A and DNMT3B are de novo methyltransferases that can produce novel methyl marks independently of DNA replication and are therefore the enzymes principally responsible for altering DNA methylation patterns in response to external events.
As pointed out by Bayraktar and Kreutz,
63
there are 2 splice isoforms for DNMT3A: DNMT3A1 and DNMT3A2. DNMT3A2 lacks the N-terminal 219 amino acids of DNMT3A1 and is transcribed from an alternative promoter.
DNMTs bind to DNA and methylate cytosines at particular locations in the genome. Where the DNA methyltransferases DNMT3A1 and DNMT3A2 bind appears to be determined by histone post translational modifications.65-67
The basic unit of chromatin, the nucleosome, is composed of 147 DNA base pairs wrapped around a histone octamer consisting of 2 copies of each of the following proteins: H2A, H2B, H3, and H4. The histone proteins have core regions and also have tail extensions, with the tails constituting up to 30% of the proteins (Figure 7). 68

A nucleosome showing 4 pairs of histones (H2A, H2B, H3, H4), each pair with the same color. The aminoterminal (N-terminal) tails of one of each pair of histones is shown, labeled with the positions of the lysine amino acids (“K” is the single letter code for lysine) that are subject to acetylation or methylation. The number at each lysine indicates its position counting from the amino end of the protein chain. Acetylations (Ac) and methylations (Me) are shown in some positions susceptible to these epigenetic alterations. The DNA, wound around the histone core, is indicated by the dark line.
Each histone protein can undergo post translational modifications in which molecules, such as an acetyl group or up to 3 methyl groups, are covalently added to (or removed from) lysine residues of their amino terminal (N-terminal) tail. If a single methyl group is added to a lysine located as the fourth amino acid from the N-terminal tail end of histone 3, this is designated H3K4Me1. The single letter K designates lysine. These modifications not only alter the structure of the nucleosome but also change the interaction of DNA with the associated histones. The modified tails also interact with DNMTs at enhancers and promoters to increase or reduce DNA methylation at the sites of particular histone post translational modifications.
Active enhancer DNA is commonly associated with the histone modifications of H3K4Me1 or H3K4Me3 as well as H3K27Ac.31,33 The promoter DNA of actively expressing genes is associated with the histone modifications H3K4Me2 or H3K4Me3 as well as H3K9Ac. 69
DNA methylation alterations upon the creation of long-term memories
An intense learning situation can be applied to rats in which a rat is placed in a new location for a short time and then an electric shock is applied to the rat’s foot. The rat forms a bad association with this new location and freezes in its motions if placed in this new location again, even without the foot shock. This is referred to as contextual fear conditioning. 46 This can result in a life-long fearful memory after a single training event. 46 While the long term memory of this event appears to be first stored in the hippocampus, this storage is transient and does not remain in the hippocampus. 46 Much of the long term storage of contextual fear conditioning memory appears to take place in the anterior cingulate cortex (Figure 2). 55 Contextual fear conditioning applied to a rat induces more than 5000 differentially methylated regions (of 500 nucleotides each) in the rat hippocampal neuronal genome as measured both 1 and 24 hours after the conditioning. 4 This causes about 500 genes to be up-regulated (possibly due to hypomethylation of CpG sites) and about 1000 genes to be down-regulated (observed to be correlated with newly formed 5-methylcytosine at CpG sites in promoter regions). Overall, 9.17% of the genes in the genomes of rat hippocampus neurons are differentially methylated after the conditioning event. The pattern of induced and repressed genes within neurons appears to provide a molecular basis for forming this first transient memory of this training event in the hippocampus of the rat brain. 4 When similar contextual fear conditioning is applied to a mouse, 1 hour after contextual fear conditioning there were 675 demethylated genes and 613 hypermethylated genes in the hippocampus region of the mouse brain. 3 These changes were transient in the hippocampal neurons, and almost none were present in the hippocampus after 4 weeks. However, in mice subjected to contextual fear conditioning, after 4 weeks there were more than 1000 differentially methylated genes and more than 1000 differentially expressed genes in the anterior cingulate cortex, 3 where long-term memories are stored in the mouse brain. 55
Another Pathway: The Role of Reactive Oxygen Species in Demethylation and Memory
In addition to the roles of IEGs in learning and memory, a large body of research (reviewed by Massaad and Klann and Beckhauser et al70,71) shows that another pathway involving reactive oxygen species (ROS) is also required for normal learning and memory functions. ROS have a role in demethylation of cytosine.
Oxidation of guanine in DNA by reactive oxygen species preferentially occurs at CpG sites in which the cytosine is methylated, to form 5mC-p-8-OHdG (Figure 8).

Initiation of DNA demethylation at a CpG site. Image of a CpG site where cytosine is methylated to form 5methylcytosine (5mC) and the guanine is oxidized to form 8hydroxy-2′-deoxyguanosine (8-0HdG). The base excision repair enzyme OGGI targets and binds to 8-0HdG without immediate excision. OGGI recruits TETI and TETI oxidizes the 5mC adjacent to the 8-0HdG. This initiates demethylation of the 5mC.
This preference is due to a lowered ionization potential of guanine bases adjacent to 5-methylcytosine. 72 Steady-state levels of endogenous oxidative alterations of guanosine represent the balance between formation of 8-OHdG and its repair. 8-OHdGs are among the most frequent DNA damages present in the steady-state, with about 2400 8-OHdG damaged nucleotides in the average mammalian cell. 73 The steady state 8-OHdG level in the brain is similar to that in other tissues. 74
The enzyme primarily responsible for the excision of 8-OHdG by the process of base excision repair is 8-oxoguanine glycosylase (OGG1). However, OGG1, which targets and associates with 8-OHdG, has also been shown to have a role in mouse adaptive behavior. This implies a physiologically relevant role for 8-OHdG combined with OGG1 in cognition in the adult brain. 75 For example, OGG1+/− mice, with about half the protein level of OGG1, exhibit poorer learning performance in the Barnes maze compared to wild-type animals. 76 OGG1 protein finds 8-OHdG by sliding along the linear DNA at about 1000 base pairs of DNA in 0.1 seconds. 77 OGG1 proteins bind to oxidatively damaged DNA with a half maximum time of about 6 seconds. 77 When OGG1 finds 8-OHdG it changes conformation and complexes with 8-OHdG in its binding pocket. 78 OGG1 does not immediately act to remove the 8-OHdG. Half maximum removal of 8-OHdG takes about 30 minutes in HeLa cells in vitro, 79 or about 11 minutes in the liver cells of irradiated mice. 80
When OGG1 is present at a 5-methylcytosine-8-OHdG site, it recruits TET1 to the 8-OHdG lesion (Figure 8). 81 This allows TET1 to demethylate an adjacent methylated cytosine. As an example, when human mammary epithelial cells were treated with H2O2, 8-OHdG increased in DNA by 3.5-fold and this caused about 80% demethylation of the 5-methylcytosines in the genome. 81
It appears that in addition to the rapid expression of IEGs when a memorable event occurs (Section Changes in gene expression are central to memory), some oxidative alterations to neurons occur as well. Upon oxidative alteration of neuronal DNA, OGG1 appears to have a role in selectively activating oxidatively altered neuronal genes through attracting TET1 to demethylate the 5mCs in their promoters.
Different Neurons for Different Memories
Sun et al
82
examined neurons in the dentate gyrus (located close to one end of the hippocampus) (Figure 4) of mice that had undergone contextual fear conditioning. About 1%-2% of the neurons in the dentate gyrus were up-regulated for expression of the IEGs
Transfer of mRNA From Nucleus to Dendritic Spine
After a memorable event occurs many mRNAs are up-regulated or down-regulated in the nuclei of neurons within the hippocampus, anterior cingulate cortex and lateral amygdala. As reviewed by Anderson and Kedersha, 84 nuclear mRNA transcripts acquire a protein coat composed of cap- and RNA-binding proteins that allow nuclear export to the cytoplasm. Neurons have mechanisms that enable transcription events initiated in the nucleus to rapidly trigger protein synthesis at distant synaptic surfaces. This involves packaging mRNA into neuronal granules for transport to sites of translation. Neuronal granules harbor translationally silenced mRNAs that are transported to dendritic synapses at dendritic spines (Figure 9), where they are released and translated in response to specific exogenous stimuli.
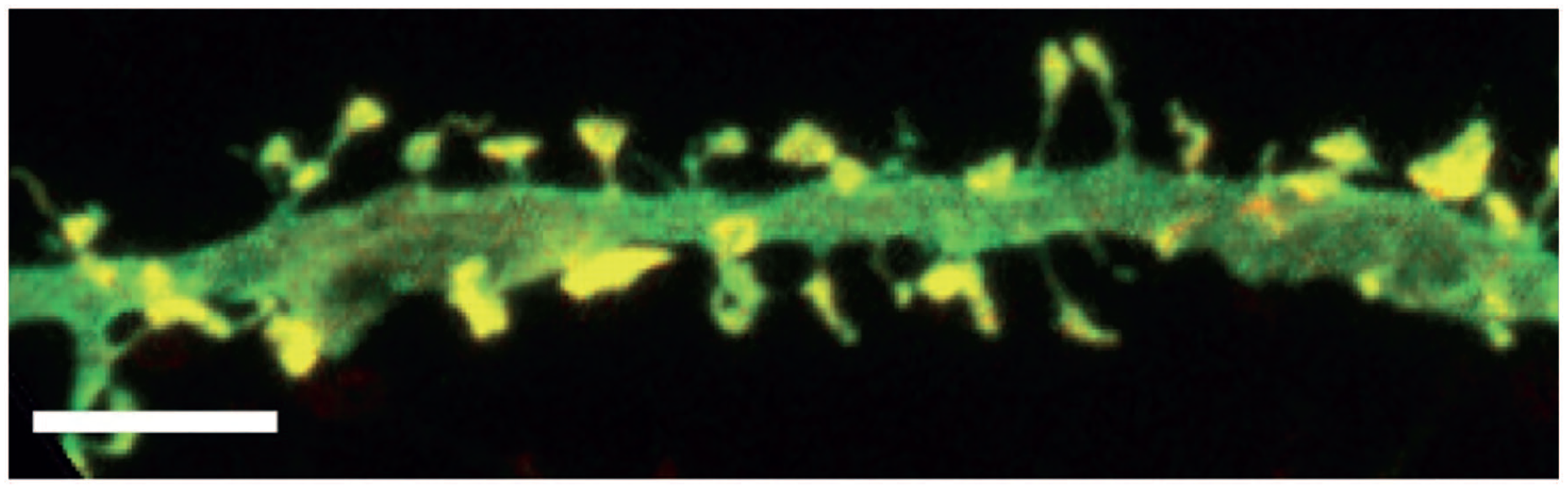
Dendritic spines (yellow) along a cultured hippocampal neuron. The bar is 5 µm.
These granules contain mRNA, small and large ribosomal subunits, translation initiation factors and RNA-binding proteins that regulate mRNA function. Despite the presence of intact ribosomes, mRNAs associated with neuronal granules are not translated until they arrive at their synapses, where their translation may require an activating stimulus.
The neuronal granules travel down the dendrite at about 5 µ/s and are directed to their synapses by the 3′ untranslated regions of the mRNA in the granules. 85 At the synapse (a dendritic spine on one neuron contacting a dendritic spine on a second neuron), the mRNA can be released and translated in response to specific exogenous stimuli. 85
About 2500 mRNAs are localized to the dendrites and axons of hippocampal pyramidal neurons and more than 450 transcripts are enriched in excitatory presynaptic nerve terminals (dendritic spines). 86
When a particular messenger RNA was evaluated by Yiu et al
87
in the lateral amygdala of mice undergoing fear training, the messenger RNA (mRNA) of the immediate early gene a
As described by Doyle and Kiebler, 88 “in the mature brain mRNA localization into dendrites of fully polarized neurons serves a distinct function. The presence of a specific set of transcripts and the entire translational machinery at dendritic spines suggests that local translation could be regulated in an activity-dependent manner.”
There can be over 10 000 spines on a single neuron, each representing a point of synaptic contact at which the cell receives input from another neuron. 89 Spines on one neuron interact with spines in other neurons forming multiple synaptic junctions. These synaptic junctions are the basis of synaptic plasticity which is the ability of synapses to strengthen or weaken over time, in response to increases or decreases in their activity. Since memories are postulated to be represented by vastly interconnected neural circuits in the brain, synaptic plasticity is considered to be one of the important neurochemical foundations of learning and memory. The review of Campbell and Wood 90 indicates that the field of neuroepigenetics has just begun to investigate how epigenetic control of synapse-related genes is regulated during memory processes.
Conclusions
An abbreviated version of the information detailed in this review, from initiating event to long-term memory, is illustrated in Figure 10.

Elements of a pathway from memorable event to long-term memory in the cortex.
When a hippocampus neuron is activated by a memorable event, the TOP2B dimer located at the promoter of the immediate early gene
Also, in activated hippocampus neurons, the immediate early gene
The altered methylation pattern in the hippocampus neuron generates a corresponding methylation pattern in the anterior cingulate cortex that changes the transcription patterns of a set of neurons there, forming a long-term memory. Other immediate early genes, not shown in Figure 10, also undergo up-regulated expression, due to various mechanisms, after a memorable event. One such mechanism could involve OGG1 acting at the site of a 5-methylcytosine-8-OHdG in one or more of their promoters (see Section Another pathway: the role of reactive oxygen species in demethylation and memory). These other immediate early genes control the expression of further later expressing genes. Overall, memorable events alter the methylation patterns in neurons, creating long-term memories in the anterior cingulate cortex of the brain.
Footnotes
Author Contributions
Carol Bernstein provided the research and writing of this review.
Declaration of Conflicting Interests:
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author received no financial support for the research, authorship, and/or publication of this article.
