Abstract
Background:
Increased levels of the potent vasodilator calcitonin gene-related peptide (CGRP) have been found in ipsilateral jugular vein blood during the active phase of cluster headache (CH) and this is hypothesized to cause distinctive vasodilation. The receptor activity-modifying protein 1 (RAMP1) is part of the CGRP receptor complex responsible for ligand binding and specificity and therefore constitutes a promising candidate gene for CH. The aim of this study was to investigate the possible genetic association of RAMP1 with CH in Sweden, with focus on two RAMP1 single nucleotide polymorphisms, rs3754701 and rs7590387, and quantify RAMP1 mRNA expression levels in biological tissue from CH patients and controls.
Methods:
rs3754701 and rs7590387 were genotyped by quantitative polymerase chain reaction (qPCR) in 542 CH patients and 585 control subjects. RAMP1 mRNA expression was determined by reverse transcription qPCR in tissue from 12 CH patients and 12 controls.
Results:
We identified a significant difference between the CH patient and control groups for rs3754701 (p = 0.0088). In addition, RAMP1 mRNA expression was enhanced in primary fibroblasts from CH patients compared to controls (p = 0.0073).
Conclusion:
The association between rs3754701 and CH and the enhanced RAMP1 mRNA expression in CH patients support the hypothesis that CGRP and its receptor component RAMP1 are involved in CH pathophysiology.
Keywords
Introduction
Cluster headache (CH) is characterized by bouts or clusters of recurring excruciating, unilateral pain attacks lasting up to 3 h accompanied by ipsilateral autonomic symptoms, including nasal congestion, rhinorrhoea, lacrimation, ptosis, and/or a feeling of restlessness. The more common episodic subtype of CH is marked by active bouts persisting from 7 days to 1 year and interrupted by pain-free remission periods of 3 months or longer according to the newly revised International Classification of Headache Disorders (ICHD-III) compared to more than 1 month in the previous beta version. 1,2 The chronic subtype of CH occurs in 10–15% of the patients according to studies based on previous diagnostic classifications with a duration of bouts for more than a year and remissions lasting 1 month or less. ICHD-III defines chronic CH as recurring attacks without remission periods or with remissions lasting less than 3 months for at least 1 year. 1,2 Some patients previously classified as episodic may with ICHD-III have to be reclassified as chronic. The disease is about three to four times more common in men than in women. 3,4 The average lifetime prevalence is 0.1–0.2%. 5,6 A heritable component, defined as having at least one first- or second-degree relative diagnosed with CH, is present in 7–20% of the cases and points toward a genetic contribution. 7,8
During CH attacks, the trigeminal-autonomic reflex is activated provoking vasodilation of cranial arteries by the release of vasodilatory molecules, including calcitonin gene-related peptide (CGRP). 9,10 Injection of CGRP has been previously shown to induce a delayed, migraine-like headache in 57–75% of migraineurs with and without aura. Because CGRP did not have this effect on healthy controls, it was suggested that migraineurs have increased sensitivity to CGRP. 11,12 Interestingly, a recent study shows that CGRP infusion in CH patients triggers attacks when in active period, but not in remission. 13 Further evidence for CGRP having a central role in headache has been demonstrated by data from studies showing that successful treatment of migraine headache pain with the serotonin 5-HT1B/1D receptor agonist sumatriptan, as well as other “triptan” drugs, resulted in the normalization of CGRP levels. 14
CGRP signal transduction is mediated by a functional CGRP receptor consisting of the calcitonin receptor-like receptor, the CGRP receptor component protein, and the receptor activity-modifying protein 1 (RAMP1). 15 RAMP1 is a single-transmembrane protein pivotal for receptor function, as it provides specificity for ligand binding. 16 Transgenic mice in which RAMP1 is overexpressed in the nervous system have been shown to display light-aversive behavior and enhanced central CGRP-induced mechanical allodynia. 17,18 In addition, there are reports indicating that RAMP1 gene expression might be affected by age in rodents. 19,20 CGRP and its receptor components play a major role in a wide range of diseases, including vasodilatory, inflammatory, and neurological conditions. 9,21 –23 More recently, they have taken center stage as therapeutic targets for primary headaches. 24 Multiple CGRP and CGRP receptor antibodies, as well as antagonists, are on their way constituting a new frontier of CH and migraine medication. 25 –27 On May 17, 2018, the U.S. Food and Drug Administration approved the first CGRP receptor antibody, Aimovig® (erenumab-aooe), for the preventive treatment of migraine in adults. 28 Hence, RAMP1 is a promising target for genetic characterization in relation to CH.
The single nucleotide polymorphism (SNP) rs3754701 is located in the 5′-end of RAMP1 on chromosome 2. Previous studies have reported a weak association between rs3754701 and migraine in an Australian male migraine population. 29 Rs3754701 has also been shown to be associated with cerebral infarction and essential hypertension through haplotype analysis in Japan. 30,31 Similarly, the SNP rs7590387 located 1.4 kb downstream of the RAMP1 gene was found to be associated with cerebral infarction and transformation of migraine into medical overuse headache in an Italian study. 30,32 In addition, rs3754701 has been reported to be linked to a change in RAMP1 expression in whole blood on the GTEx portal website (www.gtexportal.org). However, the expression quantitative trait locus (eQTL) association of this variant did not withstand the multiple hypothesis correction in an increased whole blood sample set in the GTEx v7 release. There are no reports for significant eQTLs for rs7590387 in different tissues. Our objectives were to investigate if any of these two RAMP1 SNPs are linked to CH in Sweden and if general RAMP1 gene expression differs between CH patients and controls using our Swedish biobank, and further to investigate if potential disease-associated RAMP1 SNPs lead to changes in RAMP1 gene expression.
Material and methods
Case-control material
The case-control material consisted of 542 CH patients and 585 controls living in the central part of Sweden (see Table 1 for details). Whole blood samples and cutaneous biopsies were obtained at the Karolinska University Hospital between 2014 and 2017 after approval of the Regional Ethical Review Board in Stockholm, Sweden, and informed consent from all participating individuals according to the Declaration of Helsinki. CH patients are part of a Swedish CH biobank that has been previously described and characterized. 4 CH patients were diagnosed on the basis of the ICHD third edition (ICHD-III beta) criteria. 2 The CH diagnosis was validated by the authors (CS, AS, and EW) from case records and a diagnostic questionnaire. 4 Five hundred seventy-three control individuals were anonymous healthy blood donors recruited from the Stockholm area. Together with 12 additional neurologically healthy control individuals from the Swedish CH biobank, they represent a general healthy adult Swedish population (https://geblod.nu/english-engelska/).
Demographic and clinical characterization of CH patients and controls.

CH: cluster headache; SD: standard deviation; NA: not applicable; /n: number of individuals for whom information was available.
a At least one first-, second-, or third-degree relative with CH diagnosis.
Genotyping
DNA was extracted from whole blood using standard protocols. Genotyping of CH patients and controls was performed by quantitative real-time polymerase chain reaction (qPCR) with an ABI 7500 fast real-time PCR instrument (Applied Biosystems, Foster City, California, USA) programmed according to the manufacturer’s instructions with increased cycle number. TaqMan® Master Mix and predesigned TaqMan® SNP genotyping assays were used as reagents (C_27496443_10 for genotyping rs3754701 and C__26481962_10 for genotyping rs7590387; Applied Biosystems, Carlsbad, California, USA). Call rate for both assays was >99%.
Cell culture and gene expression
Cutaneous biopsies were obtained at the Karolinska University Hospital from 12 CH patients and 12 control individuals from our cohort, see Table 1 for demographic information. Primary fibroblast cultures were established according to the following protocol: the biopsy was cut in five to six pieces, placed under a sterile glass coverslip in a culture dish and grown in DMEM GlutaMAX™ (supplemented with 25 mM
RNA was extracted from frozen cells using the QIAGEN RNeasy Mini Kit (QIAGEN, Hilden, Germany). cDNA was prepared with the QuantiTect Reverse Transcription Kit (QIAGEN). Primers were designed with the web-based software Primer3. 34 Specificity was tested with NCBI’s alignment tool BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi) and folding energy was assessed with the web-based software Mfold v.2.3. 35 Gene expression studies were performed with PowerSYBR® Green PCR Master Mix (Thermo Fisher Scientific, Waltham, Massachusetts, USA). Primers were manufactured for the target gene RAMP1 (F: 5′ GTAGACATGGAGGCCGTCG 3′, R: 5′ ATGCACTGCCAGGAAGAACC 3′), the fibroblast housekeeping gene pyruvate dehydrogenase beta (PDHB; F: 5′ GGTTTCCCATTCAAGACCTG 3′, R: 5′ TGGTTTCCATGTCCATTGGT 3′; all from Thermo Fisher Scientific). A pool of all samples was used as a reference. The ABI 7500 fast real-time PCR instrument was programmed for quantitation by relative standard curve according to the manufacturer’s recommendations with increased cycle number.
Statistical analysis
The software Online Encyclopedia for Genetic Epidemiology studies was used to ensure agreement of the observed genotype frequencies with the Hardy–Weinberg equilibrium. 36 Power analysis was performed with the sample size software PS v3.1.2. 37 With our sample size and an estimated minor allele frequency of 0.35 (rs3754701) and 0.50 (rs7590387; https://www.ncbi.nlm.nih.gov/snp), we have 80% power to detect true odds ratios (ORs) for CH below 0.693 or above 1.414 and below 0.715 or above 1.4, respectively. Genotyping was analyzed with the 7500 Software v2.0.6 (Applied Biosystems). Genetic associations were evaluated by logistic regression with sex as a covariate, using the PLINK whole-genome association toolset v1.0.7. 38 Bonferroni correction for multiple testing (two tests) was applied.
Gene expression analysis was performed with qBase v1.3.5 39 and statistical analysis with GraphPad Prism v5.04 (GraphPad Software Inc., La Jolla, California, USA) and GraphPad QuickCalcs online tools. Gene expression was normalized to a validated housekeeping gene (PDHB) and to a reference sample consisting of equal amounts of cDNA from all 24 individuals. 40 Relative gene expression between two groups was compared using the Mann–Whitney U-test, and the Kruskal–Wallis test was used for three groups. Linear regression was used to evaluate a potential correlation between expression levels and age (see Online Supplemental Data). One control sample was identified as an outlier and removed from all gene expression analyses. We used two separate statistical methods to discern outliers, Grubbs test (critical Z value: 2.41) and a boxplot analysis. The tolerated interval was set as follows: upper limit = (75th percentile) + (1.5*(75th percentile − 25th percentile)), lower limit = (25th percentile) − (outlier coefficient*(75th percentile − 25th percentile)). All statistical tests were performed with two-tailed p-values, significance level was set to 0.05.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article, its supplementary materials, or on request from the corresponding author (ACB).
Results
At the time when blood samples from 542 individuals had been collected, a total of 855 CH patients had been contacted to participate in the study. Thirteen patients were deceased before this follow-up and excluded, 74 did not wish to participate, and 226 had not replied yet at the specific time point.
Genotypic and allelic analysis
The two RAMP1 SNPs rs3754701 (A > T) and rs7590387 (C > G) were screened for association with CH in 542 CH patients and 585 controls (Table 2). The minor allele of rs3754701 was observed more frequently in patients than in controls. Genetic analysis using logistic regression confirmed a significant association between this SNP and an increased risk for CH (OR = 1.28, p c = 0.0088). We used sex as a cofactor in the analysis in order to avoid any bias introduced by the higher proportion of men in the CH group, 68.5% as compared to 57.1% in the control group. When performing a subgroup analysis in order to explore these data further, we found that the effect of the association was stronger in episodic patients than in the entire CH population (Table 3). The disease associated allele, T, was slightly more common in episodic CH patients than in the whole patient-group, while chronic CH patients had allele frequencies highly similar to control individuals. Statistical analysis controlling for gender confirmed the association between the minor T allele and episodic CH (p = 0.0021). When comparing allele distribution in episodic patients with chronic patients, we could not identify a difference (OR = 1.34, 95% confidence interval: 0.89–2.04, p = 0.16), probably due to the small number of individuals in the chronic subgroup. Four CH patients were removed from the subgroup analysis due to missing data on subtype. No significant association between the RAMP1 variant rs7590387 and CH was identified in Swedish CH patients (Table 2).
Results from the analysis of two RAMP1 SNPs in CH patients and controls.a

SNP: single nucleotide polymorphism; CH: cluster headache; OR: odds ratio; CI: confidence interval; p c value: p-value after correction for multiple testing.
a Data was analyzed with logistic regression with gender as a covariate.
b Statistically significant (α < 0.01).
Analysis of rs3754701 in chronic and episodic patients compared to controls.a

ECH: episodic cluster headache; CCH: chronic cluster headache; OR: odds ratio; CI: confidence interval.
a Data was analyzed with logistic regression with gender as a covariate.
b Statistically significant (α < 0.01).
Gene expression
RAMP1 mRNA expression was evaluated in primary fibroblast cultures from 12 CH patients and 12 healthy controls. Gene expression was normalized to a housekeeping gene (PDBH) and to a reference sample and these relative quantities were further log2 transformed since the sample was not normally distributed. PDHB expression levels did not differ between CH cases and controls and showed a normal distribution (data not shown). We found that RAMP1 mRNA expression was enhanced in CH patients as compared to healthy controls (Figure 1(a), p = 0.0012). CH subgroup analysis showed that there is a tendency for chronic patients to have a higher RAMP1 expression (Figure 1(b)), although there is no significant difference (p = 0.86) between episodic (n = 9) and chronic patients (n = 3).
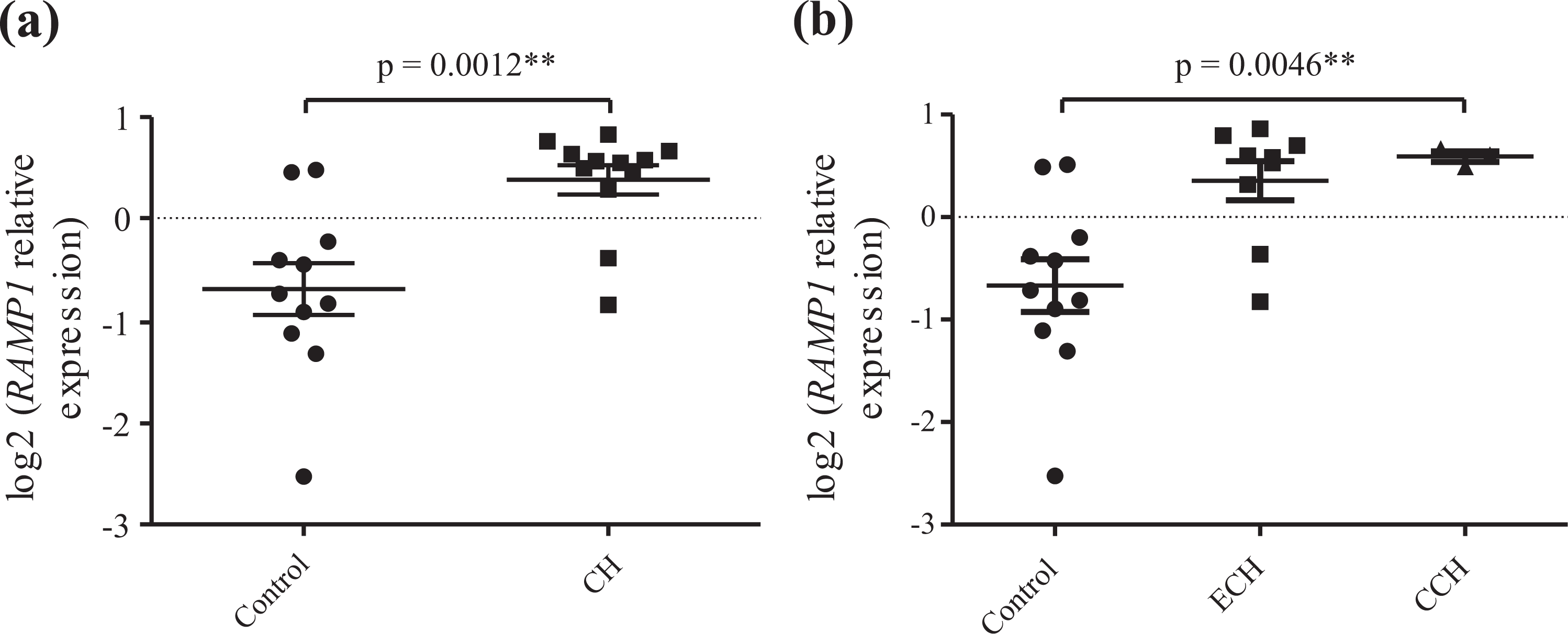
Relative quantification of RAMP1 expression in fibroblast cell cultures from cluster headache (CH) patients (n = 12) and healthy controls (n = 11) normalized to the endogenous control PDHB grouped by (a) disease status and (b) to subtype of CH; episodic CH (ECH) and chronic CH (CCH). PDHB: pyruvate dehydrogenase beta.
To further assess a potential effect of the two RAMP1 variants, rs3754701 and rs7590387, RAMP1 mRNA expression was analyzed with respect to genotype of each SNP, independently of diagnosis (Figure 2(a) and (b)). We were surprised to discover no significant difference in the relative mRNA levels in fibroblasts from individuals with the three possible genotypes (AA/AT/TT) at rs3754701 (Figure 2(a)). Analysis of rs7590387 revealed a weak trend for association between this SNP and RAMP1 mRNA levels (Figure 2(b)). Group comparison between individuals with the three possible genotypes (CC/CG/GG) at rs7590387 and relative mRNA levels showed that individuals with the reference genotype CC had slightly higher RAMP1 levels than the other groups (p = 0.069). Detailed comparison between the three genotypes further showed that RAMP1 mRNA was significantly lower in individuals with the CG genotype (p = 0.043) and GG genotype (p = 0.05) when compared to individuals with CC at rs7590387, mRNA levels between CG and GG carriers were comparable (p = 0.89). Because of these results, we ran one additional analysis, grouping CG and GG carriers together. This confirmed that RAMP1 gene expression was significantly lower in individuals with one or more G alleles as compared to CC carriers (p = 0.023) (Figure 2(c)). There is no difference in age between patients and controls using an unpaired t-test (p = 0.331). Furthermore, our data showed that RAMP1 expression is stable with increasing age in patients and controls and, therefore, we choose not to consider age in our analysis (Online Supplemental Figure 1A to 1C).

Analysis of the effect of genetic variants on RAMP1 expression. Relative quantification of RAMP1 expression in fibroblast cell cultures from CH patients (n = 12) and healthy controls (n = 11) analyzed according to rs3754701 and rs7590387 genotype. (a) rs3754701 genotypes AA (n = 7), AT (n = 12), and TT (n = 4) and (b) rs7590387 genotypes CC (n = 6), CG (n = 8), and GG (n = 9), and (c) RAMP1 mRNA analyzed by presence or not of an rs7590387 G allele (CC vs. CG + GG). CH: cluster headache.
To validate the relation between gene expression and genotype in another tissue, we analyzed RAMP1 mRNA levels in lymphocytes immortalized by Epstein–Barr virus (EBV) transfection (see Online Supplemental Data). Cells were obtained from individuals with Parkinson’s disease (PD), a neurological disorder with different pathology from CH, and healthy controls. 41 Similar to the fibroblast analysis, RAMP1 expression was normalized to a reference gene TBP (TATA-box binding protein) and to a reference sample and then log2 transformed. The RAMP1 gene expression in cells from PD patients corresponded to that observed in control individuals and was used as controls to increase power for the EBV transfected lymphocytes. One sample was identified as an outlier and removed, and a t-test was used to verify that expression did not differ between patients and controls (p = 0.84; Online Supplemental Figure 2A). RAMP1 mRNA expression was then analyzed with respect to the genotypes for rs3754701 and rs7590387 (Online Supplemental Figure 2B and 2C), which showed no significant difference between the different groups (p rs3754701 = 0.74, p rs7590387 = 0.86). These results indicate that RAMP1 expression does not vary with genotype for either SNP in lymphocytes.
Discussion
In our Swedish CH case-control material, the RAMP1 rs3754701 T allele was more common in CH patients compared to controls. The test statistics were significant and indicative of a small effect attributed to the risk-associated allele: the OR from the allele analysis was 1.28 with a confidence interval well-separated from zero. Small ORs are expected in association studies of common SNPs, which are present in both patients and healthy individuals. This association is in the order of magnitude of our power analysis. We performed a stratified analysis to test whether the association was influenced by CH subtype. Interestingly, the stratification revealed an association of rs3754701 with episodic, but not with chronic CH. The clinical features of chronic CH are well characterized and the phenotype is more common in women and in patients with later onset. In a recent report, chronic CH patients were discovered to have lower plasma levels of CGRP and pituitary adenylate cyclase-activating polypeptide-38 than episodic patients, but very little is known on how the pathophysiology differs between these two conditions. 42 Our finding of a genetic RAMP1 marker enriched in episodic patients suggests that there might be a genetic basis for susceptibility of these phenotypes. It is also a possibility that different pathological events are responsible for a clinically similar phenotype. These findings are particularly interesting considering the fact that monoclonal anti-CGRP antibodies have been proven ineffective in patients with chronic CH.
One potential limitation of our study is the control cohort consisting of anonymous blood donors with no information about age, and so on. There is a faint possibility that CH occurs in the control population, but despite these concerns, we consider this an excellent control group representing a healthy Swedish population between the ages of 18 years and 60 years (https://geblod.nu/english-engelska/). The proportion of individuals which had not agreed to participate at the time of the study (26.4%) is lower compared to previous CH studies and therefore not regarded as a limitation of this study. 43
The hypothesis that CGRP and its receptor component RAMP1 are involved in CH pathophysiology is additionally supported by our finding that RAMP1 mRNA expression was enhanced in CH patient fibroblasts compared to controls. Chronic and episodic patients have very similar mRNA levels, indicating RAMP1 is dysregulated in both subgroups. There are three published studies on broad screenings for genes differentially transcribed between individuals with CH and controls. 44 –46 Neither of these three studies report differences in RAMP1 expression in their samples. Important to note is that all three studies have low power to detect gene expression differences of the magnitude we reported after correction for multiple testing due to low sample size in relation to high number of tests (Costa et al., n = 8 (CH) and n = 10 (controls) vs. 36,079 transcripts; Sjöstrand et al. n = 3 (CH) and n = 3 (controls) vs. 54,000 gene transcripts; and Eising et al. n = 19 (episodic CH) and n = 20 (chronic CH) patients and n = 20 (controls) vs. 13,416 genes).
Moreover, we could confirm the GTEx portal data suggesting that rs3754701 and rs7590387 do not affect RAMP1 expression in whole blood using EBV-transfected lymphocytes. Since we did not observe any effect of rs3754701 on RAMP1 gene expression in neither fibroblasts nor blood, we hypothesize that rs3754701 could be in linkage disequilibrium with the real disease-causing variant or that this genetic variant affects RAMP1 in some other way than mRNA transcription. Genotypic and allelic frequencies of rs7590387 did not differ significantly between CH cases and controls in our cohort. However, we observed a weak association between the CC genotype and higher gene activity in fibroblasts compared to the CG and GG genotypes, indicating the rs7590387 is an eQTL in certain tissues.
RAMP1 is the part of the CGRP receptor complex responsible for specific binding of CGRP, the most potent vasodilator in the brain. 47,48 Numerous studies have shown a clear link between abnormal CGRP levels and active primary headache bouts, including CH. 9,42,49 –52 Colocalization of the vasodilator with all three CGRP receptor components is prominent in many parts of the trigeminovascular system, which has been shown to be activated during CH attacks. 9,53 Due to its localization in perivascular nerve terminals innervating the vascular smooth muscle, CGRP is further thought to be responsible for the painful dilation of cranial arteries during headache attacks. 54 This effect can potentially be attributed to a malfunctioning receptor altering CGRP binding and activity, making RAMP1 a highly promising drug candidate.
In summary, we have found the T allele of a SNP (rs3754701) located upstream of the RAMP1 gene to be a susceptibility factor for CH in our cohort. However, we call for genetic studies in independent cohorts to further confirm our findings. In addition, increased RAMP1 mRNA expression was identified in fibroblasts from CH patients compared to controls. A higher receptor activity could potentially either be an effect of increased CGRP levels and/or lead to altered release of CGRP. This is in line with observations of other types of receptors, for example, dopamine (DA) D2 receptor (D2 R) overexpression in the striatum has an impact on DA levels and rates of DA turnover. 55 Our findings suggest the involvement of the RAMP1 gene in the pathophysiology of CH and thus, further studies are required.
Supplemental material
Supplementary_material_RAMP1_clean_version_(1) - Involvement of CGRP receptor RAMP1 in cluster headache: A Swedish case-control study
Supplementary_material_RAMP1_clean_version_(1) for Involvement of CGRP receptor RAMP1 in cluster headache: A Swedish case-control study by Julia M Michalska, Caroline Ran, Carmen Fourier, Anna Steinberg, Christina Sjöstrand, Elisabet Waldenlind and Andrea Carmine Belin in Cephalalgia Reports
Footnotes
Acknowledgement
We thank Katrin Wellfelt and Margareta Widing for excellent technical assistance, as well as Ann-Christin Karlsson for help with patient recruitment.
Author contribution
JMM and CR contributed equally.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Swedish Brain Foundation, the Swedish Research Council, Karolinska Institute Funds, and the Swedish Headache Society.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
