Abstract
In Nigeria, Annona muricata L. has been used to treat a variety of ailments. The mechanism of the antimalarial activity of ethanolic leaf extract of Annona muricata (EEAML) was investigated using both an in vivo and an in silico approach. The experimental mice were divided into five groups: A-F. The mice in groups B-F were inoculated with Plasmodium berghei NK-65 and treated accordingly. Groups A and B are the negative and positive controls (infected and untreated), respectively. Group C received 10 mg/kg chloroquine (standard drug), whereas groups D-F received 100, 200, and 300 mg/kg body weight of the extract orally respectively. The mice were euthanized eight days after infection, and their liver and blood were collected and used in biochemical tests. Molecular docking was performed using the extract's HPLC compounds and Plasmodium falciparum proteins. In the suppressive, prophylactic, and curative tests, there was a significant decrease (p < 0.05) in parasitemia levels in groups treated with the extract compared to the positive control and standard drug. When compared to the positive control, there was a significant (p < 0.05) reduction in liver MDA, total cholesterol, and total triglyceride levels. The binding energies of luteolin and apigenin-pfprotein complexes were significantly (p < 0.05) higher compared to their respective references. The anti-plasmodial activity of the extract may result from its hypolipidemic effect, which deprives the parasite of essential lipid molecules needed for parasite growth, as well as from the inhibitory effects of apigenin and luteolin on specific proteins required for the Plasmodium metabolic pathway.
Introduction
Malaria is endemic in Africa and is responsible for more death than Coronavirus disease 2019 (COVID-19), especially among pregnant women and children that are less than five of age. Children who have experienced a severe malarial episode may have permanent cognitive and neurological damage, which might leave them with disability.1–4 Plasmodium parasite is the causative agent of malarial infection prevalent disease in the tropics and subtropics. 5 Plasmodium falciparum is the most deadly of the four kinds of Plasmodium parasites that infect people (vivax, ovale, malariae, and falciparum).6,7 Therefore, treating the infection entails reducing the parasite load in the infected humans’ or animals’ bloodstream or tissue. Although medication resistance is a serious obstacle in the fight against the risks associated with this infectious disease. Outside of Africa, reports of resistance to common antimalarial medications like chloroquine, mefloquine, quinine, and artemisinins have been made. However, due to high levels of immunity among the African population and the complexity of African infections, which limit the formation of relatively unsuitable resistance strains, resistance selection may not be as severe in Africa as it is in other areas of the world.8,9 However, one the mechanisms of drug action in malaria treatment include the inhibition of some vital metabolic enzymes in Plasmodium that cause parasite death; these enzymes are antimalarial drug target enzymes.10,11 The major obstacle to the control of malaria in West Africa, however, is the emergence of Plasmodium falciparum resistance to aminoquinolines, which are important allies in artemisinin-based combination therapies (ACTs); the antifolate sulfadoxine-pyrimethamine (SP), which is still a standard option for malarial infection chemoprevention.9,12,13 Further, polymorphisms in the P. falciparum chloroquine resistance transporter (PfCRT) and P. falciparum multidrug resistance protein 1 (PfMDR1) proteins 11 may contribute to P. falciparum resistance to this antimalarial medications. 14 Similar to this, P. falciparum SP resistance is brought on by mutation in the target enzymes dihydrofolate reductase (PfDHFR) and dihydropteroate synthase (PfDHPS).15,16 Since this protozoon experiences constant genomic mutation, a novel drug that targets a different drug target site may be the most effective antimalarial. Therefore, more research is required to find novel antimalarial treatments made from plants like Annona muricata L because the majority of antimalarial medications are sourced from plants. However, various parts of this plant, including the leaves, bark, fruit, and seed, have traditionally been used as a decoction or infusion to treat a variety of diseases.17,18 In the countries where it grows, Annona muricata leaves have been utilized for a variety of purposes. Skin infections have been treated using water from cooked Annona muricata leaves,19,20 and the leaf's decoction has been used as an analgesic.21,22 Additionally, the leaf has been explored for the treatment of cold, flu, asthma-related discomforts,23,24 malarial, and hypertension in tropical countries.18,25–27 In Nigeria, the plant's fruits are consumed as food, and its decoctions are used to treat or manage prostate enlargement and early-stage breast cancer. Similarly to this, locals in some parts of the nation have used plant leaves to treat malaria by consuming boiled leaf water. The fruit may be used as food and juice to treat diarrhea, heart disease, liver disease,18,28 and intestinal parasites in South America. Previous research supported the ethnobotanical use of Annona muricata, leaf for the treatment against a chloroquine-sensitive strain of Plasmodium parasite.29–34 There is a paucity of literature on the antimalarial mechanism and potential target enzyme in the Plasmodium parasite for the antimalarial bioactive constituent of Annona muricata leaf extract. As a result, the current study was designed to assess the antimalarial mode of action of Annona muricata leaf extract in vivo and in silico.
Material and Method
Plant Material and Authentication
The leaves of Annona muricata L. (Figure 1) were collected from the Adeyemi compound at Adewole Estate, Ilorin Kwara State, Nigeria, and were authenticated at the Herbarium of the Department of Plant Biology in a University and was assigned a voucher number; UIH 001/1106.

Photograph of Annona muricata plant.
Procurement of P. berghei NK-65 Parasite
The Institute for Advanced Medical Research and Training (IAMRAT), College of Medicine, in Ibadan, Nigeria provided the chloroquine-sensitive strain of Plasmodium berghei NK-65.
Experimental Animals
Adult Swiss albino mice (Mus musculus) numbering ninety (90) and with an average weight of 18 ± 2 g were procured from the Institute for Advanced Medical Research and Training (IAMRAT) College of Medicine in UTH Nigeria. The mice were allowed free access to mouse pellets (Chikun Feed, Ilorin Nigeria) and tap water ad libitum. The mice were kept in aluminum cages under the standard laboratory conditions (12-h light/dark cycle, 25 ± 2 °C, humidity: 40-45%) for the two weeks of acclimatization and the period of the experiment. University Ethical Review Committee with the Protocol gave the Handling and Use of Animal Approval with Identification Code UERC/LSC/123 and UERC Approval Number: UERC/ASN/2017/902.
Preparation of Ethanolic Extract of Annona muricata Leaf
The method described by Malairajan, Gopalakrishnan, Narasimhan, Veni and Kavimani 35 was used for the ethanolic extract of Annona muricata L. leaves. The leaves were washed in clean water and air-dried to avoid the loss of volatile ingredients in the leaves. The dry leaves were pulverized into powder using a miller (Jenway, Germany). The ethanolic extract of the plant leaf was prepared by maceration of about 100 g of the dry powder of plant leaves in 1000 ml of absolute ethanol at room temperature for 24 h. The extract was then filtered first using a muslin cloth and then with filter paper (125 mm Whatman No. 1). The extract was concentrated at 40 degrees centigrade with a rotary evaporator (Jenway Germany). The dry weight of the extract was produced through drying at 60 degrees with a water bath (Modulyo Freeze Dryer, Edward, England). The extract was kept in a refrigerator in an airtight container until when needed for the experiment.
Qualitative Analysis of Secondary Metabolites
One gram of powdered Annona muricata L. leaves was extracted with 100 ml of distilled water (1% w/v) and was used for the secondary metabolites screening. A small portion of the extract was subjected to standard tests as described for phenolics 36 ; flavonoids and tannins 37 ; alkaloids, terpenoids and cardiac glycosides 38 and saponins. 39
High Performance Liquid Chromatography (HPLC-UV) Analysis
The chromatographic system (N 2000, Korea) used for the HPLC analysis of the ethanolic extract of Annona muricata leaf includes an Autosampler (YL 9150) with a 100 l fixed loop and a UV-Visible detector (YL9120). The separation was carried out at room temperature on an SGE Protocol PC18GP120 (250 mm, 4.6 mm, 5 m) column. The separations were carried out in isocratic mode with an elution flow rate of 1 ml/min using methanol to water as the mobile phase (70:30 v/v). A UV detector operating at 254 nm was used to detect the samples after they had been operated on for 15 min. All chromatographic data were recorded and processed using the autochrome −2000 software. As reference materials for the analysis, chlorogenic acid, benzoic acid, gallic acid, luteolin, and apigenin were utilized.
Infection of Experimental Animals with Parasites
Blood from infected mice (donor mice) with parasitemia levels of 20–30% was collected in heparinized tubes via cardiac puncture. The blood was then diluted with phosphate-buffered saline (PBS) to the parasitemia level of each donor and the RBC count of normal mice, resulting in a 1 ml blood sample containing 5 × 107 parasites. Each of the mice was inoculated through intraperitoneal (IP) injection of 1 × 107 of parasitized red blood cells (PRBCs). 40
Microscopic Test and Determination of Percentage Parasitemia
The mouse tail was venesection and a drop of blood was collected on a glass slide (76 × 26 mm thickness). The blood was evenly spread on the slide to make a thin blood film. The thin smear was allowed to air dry for ten minutes. The smear was fixed for 5 s by dipping the slide in 90% ethanol. After that, the film was stained with 10% Giemsa stain for 30 min, then washed with distilled water, drained, and allowed to dry before being examined under a microscope.
41
The Leishman positive bodies, which represent parasitized red blood cells, were used to calculate parasitemia:
Test for Suppressive Activity (Peter's 4-day Test)
The suppressive antimalarial activity of P. berghei infected albino mice was evaluated using the method described by Knight and Peters. 42 On the first day, P. berghei was inoculated into the mice (D0). The infected mice were treated for four days (D0–D3). A thin blood film was produced from each experimental animal's tail blood on the fifth day (D4), fixed in methanol, and stained with Giemsa to reveal parasitized erythrocytes. Parasitaemia was determined using light microscope.
Test for Prophylactic Activity
The prophylactic antimalarial activity was evaluated by the method described by Peters’. 43 The mice were given the antimalarial agent for three days in a row (D0 - D2). The mice were inoculated intraperitoneally with P. berghei PRBCs on the fourth day (D3), and the level of parasitaemia was measured 72 h later using a blood smear (In falciparum (P.berghi) infection, only one-third of parasitized red cells circulate freely within 48-h of the asexual cycle. Therefore the remaining parasites are sequestered in the venules and capillaries. 44 Therefore at this point the peripheral blood parasite count is a variable underestimate of the total parasite burden45,46 hence, at 72 h, the parasite count would represent an accurate evaluation of the parasitemia).
Test for Curative Activity (Rane's Test)
According to Ryley and Peters’ method, 47 the curative test was conducted. P. berghei PRBCs were injected intraperitoneally into mice (D0) and 72 h later, antimalarial treatment began. For five days, the treatment was given once a day (D3). Thin smears were used to monitor parasitemia levels.
Protein Preparation
Protein preparation was undertaken to harvest and prepare the proteins into dockable format before the commencement of the molecular docking studies. In the absence of deposited crystallographic structures for most of the Plasmodium berghei, the proteins crystal structures of Plasmodium falciparum proteins were adopted: phoosphoethanomaline methyltransferase (PDB ID: 3UJB), Spermidine synthase (PDB ID: 4CWA), plasmepsin II (PDB ID: 4YA8), falciparin 2 (PDB ID: 6JW9), Lactate dehydrogenase (PDB ID: 1U4O) and phosphocholine cytidylyltransferase (PDB ID: 4ZCR) were retrieved from the protein databank (http://www.rcsb.org). The removal of water molecules and initially bound ligands was followed by the addition of the hydrogen atom required in the protein structure using the Autodock v4.2 program, Scripps Research Institute. The proteins were saved in the Protein Data Bank, Partial Charge, and Atom Type (PDBQT) format and later used for molecular docking.
Preparation of Ligand
Ligand preparation was undertaken to harvest and prepare the ligands into dockable format before the commencement of the molecular docking studies. The identified phytochemicals (chlorogenic acid, benzoic acid, gallic acid, luteolin, and apigenine) were obtained in structure data file (SDF) format from http://www.pubchem.ncbi.nlm.nih.gov, PubChem database, and converted to PDB format using Discovery Studio Visualizer, BIOVIA, 2016. The original ligands that were bound to the selected target Pfproteins were removed from the protein, which was then used as the reference compound. Polar Gasteiger-type hydrogen charges were introduced, and the nonpolar hydrogens were merged with the carbons to determine internal degrees of freedom and torsions. Using Autodock tools, the prepared ligand was also saved in the usual format (PDBQT) as the protein.
Active Site Targeted Molecular Docking
A targeted molecular docking experiment was conducted in silico, to simulate the docking pattern of the ligands to the selected protein targets. The identified phytochemicals and reference compounds were docked into the actives site region of the respective Pfproteins. Vina 48 incorporated into PyRx 0.8 was used for docking the ligands to the protein targets, and determination of binding energies. The individual Pfproteins (in PDB format) were imported into PyRx 0.8 and converted to PDBQT, while the ligands were imported through Openbabel. 49 Openbabel was used for the minimization using Universal Force Field (UFF) as the energy minimization parameter and conjugate gradient descent as the optimization algorithm. The grid boxes parameter (Table 1) displayed the amino acids and interacting residues in the active site region of the protein where interactions occurred to form the protein ligand complex. Individual binding energies of the test compounds for the six Pfproteins were measured and graded using affinity scores. Molecular interactions between lead phytochemicals and Pfproteins were investigated using BIOVIA's Discovery Studio Visualizer.
Binding Site Coordinates of Plasmodium Falciparum Proteins.
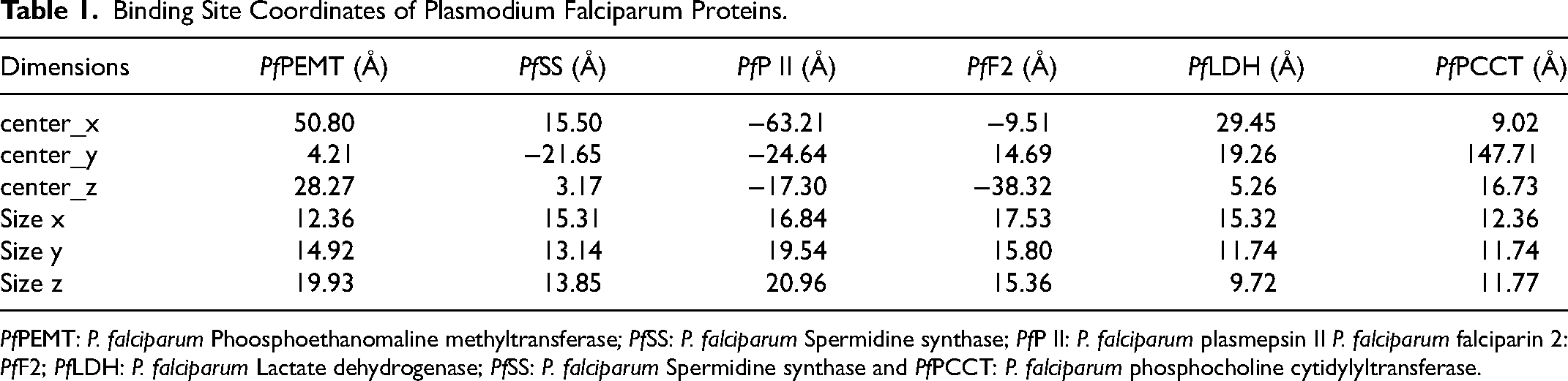
PfPEMT: P. falciparum Phoosphoethanomaline methyltransferase; PfSS: P. falciparum Spermidine synthase; PfP II: P. falciparum plasmepsin II P. falciparum falciparin 2: PfF2; PfLDH: P. falciparum Lactate dehydrogenase; PfSS: P. falciparum Spermidine synthase and PfPCCT: P. falciparum phosphocholine cytidylyltransferase.
Molecular Dynamics Simulation
Molecular dynamics was undertaken to study the stability and flexibility of the docked phytochemicals in the active sites of the selected proteins. The stability of the bound structures was compared with the unbound proteins. Plasmodium falciparum proteins such as phoosphoethanomaline methyltransferase (3UJB), Spermidine synthase (4cwa), plasmepsin II (4YA8) and falciparin 2 (6JW9) were selected for molecular dynamics (MD) simulation analysis. The unbound PfProteins and the complexes with the lead phytochemicals (LP): apigenine and luteolin were submitted for MD simulation employing GROMACS 2019.2 and GROMOS96 43a1 forcefield.50–52 The topology files for apigenine and luteolin were obtained using the PRODRG webserver (http://davapc1.bioch.dundee.ac.uk/cgi-bin/prodrg). 53 With periodic boundary conditions and a physiological concentration of 0.154 M set by neutralized NaCl ions inside the cubic box of the transferable intermolecular potential, the PfProteins and LP - PfProteins complex systems were solvated. This was done using a four point (TIP4P) water model. Using a constant ensemble of atoms, volume, and temperature, the system was condensed to 1000 steps at 0.3 nanosecond using the steepest descent algorithm (NVT). Equilibration occurred in an ensemble with a constant number of atoms, pressure, and temperature at 0.3 nanosecond as well (NPT). Using velocity rescale and the Parrinello-Rahmanbarostat, the temperature and pressure were kept constant at 310 K and 1 atm, respectively. A leap-frog integrator with a time step of 2 femtosecond was used. For each system, 100 ns of simulation were performed, and a snapshot was saved every 0.1 nano, for a total of 1000 frames from each system 54 to calculate the thermodynamic parameters (RMSD, RMSF, SASA, RoG, and number of H-bond).
Biochemical Assays
A digital UV/VIS spectrometer (Jenway, England) was used to investigate the biochemical parameters in the liver homogenate. The body weight of mice is the weight of the mice in g. 55 Gornall, Bardawill and David 56 described the method that was used to estimate total protein concentration. The methods described by Wright, Leathwood and Plummer 57 were used to estimate the activities of ACP and ALP. Malondialdehyde, a byproduct of lipid peroxidation, was estimated using measurements of thiobarbituric acid reactive substance (TBARS) as described by Satoh. 58 β-Hydroxyl-β-methylglutaryl coenzyme A (HMG-CoA) reductase was also estimated according to Rao and Ramakrishnan. 59 Lipids in the freshly harvested liver were extracted following the method described by Folch, Lees and Sloane Stanley. 60 The extracted mixture was further washed with 0.05 M KCl solution. Cholesterol and triglycerides concentrations were estimated in aliquots of chloroform-methanol extract using kit. Gidez, Miller, Burstein, Slagle and Eder 61 method was adopted in the extraction of HDL. Beckman Coulter automated software was used to estimate hematological parameters (Beckman Coulter Inc. Brea CA, USA).
Animal Grouping and Treatment
The same experimental design was applied to curative, prophylactic, and suppressive test. A total of thirty (30) mice were used for each test. The mice were randomly grouped into six of 5 mice each.
Group A: Negative control: administered 0.2 ml of 2% DMSO Group B: Positive control: Infected and untreated Group C: Infected mice treated with 10 mg/kg body weight of chloroquine Group D: Infected mice treated with 100 mg/kg body weight of ethanolic extract of Annona muricata leaf Group E: Infected mice treated with 200 mg/kg body weight of ethanolic extract of Annona muricata leaf Group F: Infected mice treated with 300 mg/kg body weight of ethanolic extract of Annona muricata leaf.
Mice infected with P. berghi NK-65 had their hearts pierced to remove their blood. The blood was diluted using physiological saline. A 0.2 ml intraperitoneal dose of the diluent was administered to each animal. A blood smear was created using the blood from the mice's cut tail tip. To determine infection, the blood smear was fixed and stained with Leishman stain. The parasitemia level was investigated under a microscope. As soon as it was determined that the infected mice had the parasite, treatment with a conventional drug (standard) and dosages of the extract were begun. Checks were made on the parasitemia levels at 4, 6, and 8 days after infection.
Histopathological Examination
The formalin preserved liver tissue was embedded by a standard laboratory protocol. The tissue was sliced into 4 mm size and then fixed on the slides. After the slide was de-paraffinized with p-xylene and rehydrated in graded ethanol (100, 80, 70% and 50%), it was rinsed with water. Furthermore, the slide will be stained with hematoxylin for 5 min and rinsed with water, and subsequently counter stained in eosin, that was mounted in DPX, cover-slipped and viewed with Leica slide scanner (SCN 4000, Leica Biosystems, Wetzlar, Germany),
Data Analysis
Data were analyzed statistically and the results were expressed as mean ± SEM (standard error of mean). With one-way analysis of variance (ANOVA), and Duncan Multiple Range Test (DMRT) post hoc comparison was used to evaluate of significant level at p < 0.05. Statistical evaluation of the data was executed with SPSS version 21.0 (IBM Corporation, NY USA) and Graph pad Prism (version 9, Graph Pad Software Inc., La Jolla, CA).
Results
Anthocyanin, saponins, terpenoids, triterpenes, flavonoids, and tannins were all present in the ethanolic extract of Annona muricata leaf, with flavonoids having the highest concentration of these phytochemicals (Table 2). Similarly, the flavonoids gallic acid, luteolin, apegenine, benzoic acid, and chlorogenic acid were all identified, with benzoic acid being the most prevalent (Table 3). The HPLC chromatogram showed peaks for the identified compounds of flavonoids (Figure 2).

HPLC chromatogram of flavonoids component of Annona muricata leaf extract.
Secondary Metabolites Profile of Ethanolic Extract of Annona Muricata Leaf.

Data are expressed as means ± SEM of triplicate determinations.
HPLC Flavonoids Compounds of Ethanolic Extract of ANNONA MURICATA LEAF.

There was a significant increase (p < 0.05) in the mice's weight on the eighth day in the groups treated with standard drugs and the 100 and 300 mg/kg BW of extract compared to the mice infected and untreated in a suppressive test (Figure 3A). Similarly, the weight increased significantly on the eighth-day post infection at 300 mg/kg body BW compared to the infected and untreated group in a prophylactic test (Figure 3B). In the curative test, there was a significant increase (p < 0.05) in the mice weight at 300 mg/kg BW and standard drug compared to the infected and untreated group (Figure 3C). For the curative, prophylactic, and suppressive tests, there was a significant decrease (p < 0.05) in parasitemia levels at the fourth, sixth, and eighth post-infection in mice infected with Plasmodium berghei NK-65 and treated with ethanolic extract of Annona muricata leaf and chloroquine, respectively, compared to mice infected with Plasmodium berghei NK-65 and untreated (Figure 4A,B & C). Similarly, In the curative, prophylactic, and suppressive tests, there was a significant increase (p < 0.05) in the percent parasite growth inhibition at the fourth, sixth, and eighth post-infection in mice infected with Plasmodium berghei NK-65 and treated with ethanolic extract of Annona muricata leaf and chloroquine, respectively, compared to mice infected with Plasmodium berghei NK-65 and untreated (Figure 5A, B, &C). The packed cell volume (PCV), hemoglobin (HGB), and red blood cell (RBC) levels in mice infected and treated with EEAML and chloroquine respectively were significantly higher (p < 0.05) than in infected and untreated mice (Table 4). Similarly, PCV, HGB, and RBC levels in infected mice treated with EEAML increased significantly (p < 0.05) when compared to the control (Table 4).

Body weight of mice infected with Plasmodium berghei NK-65 and treated with ethanolic extract of Annona muricata leaf following suppressive (A), prophylactic (B) and curative (C) test. Values are means of five (5) replicates, BW; body weight, standard drug (10 mg/kg body weight of Chloroquine). Values with different superscript are significantly different at p < 0.05.

Parasitaemia levels of suppressive (C) prophylactic (B) and curative (A) test at fourth, sixth, and eighth post infection with Plasmodium berghei NK-65 in treated with ethanolic extract of Annona muricata leaf. Values are means of four (5) replicates. BW; body weight, CQ; Chloroquine, EEAML: ethanolic extract of Annona muricata leaf.
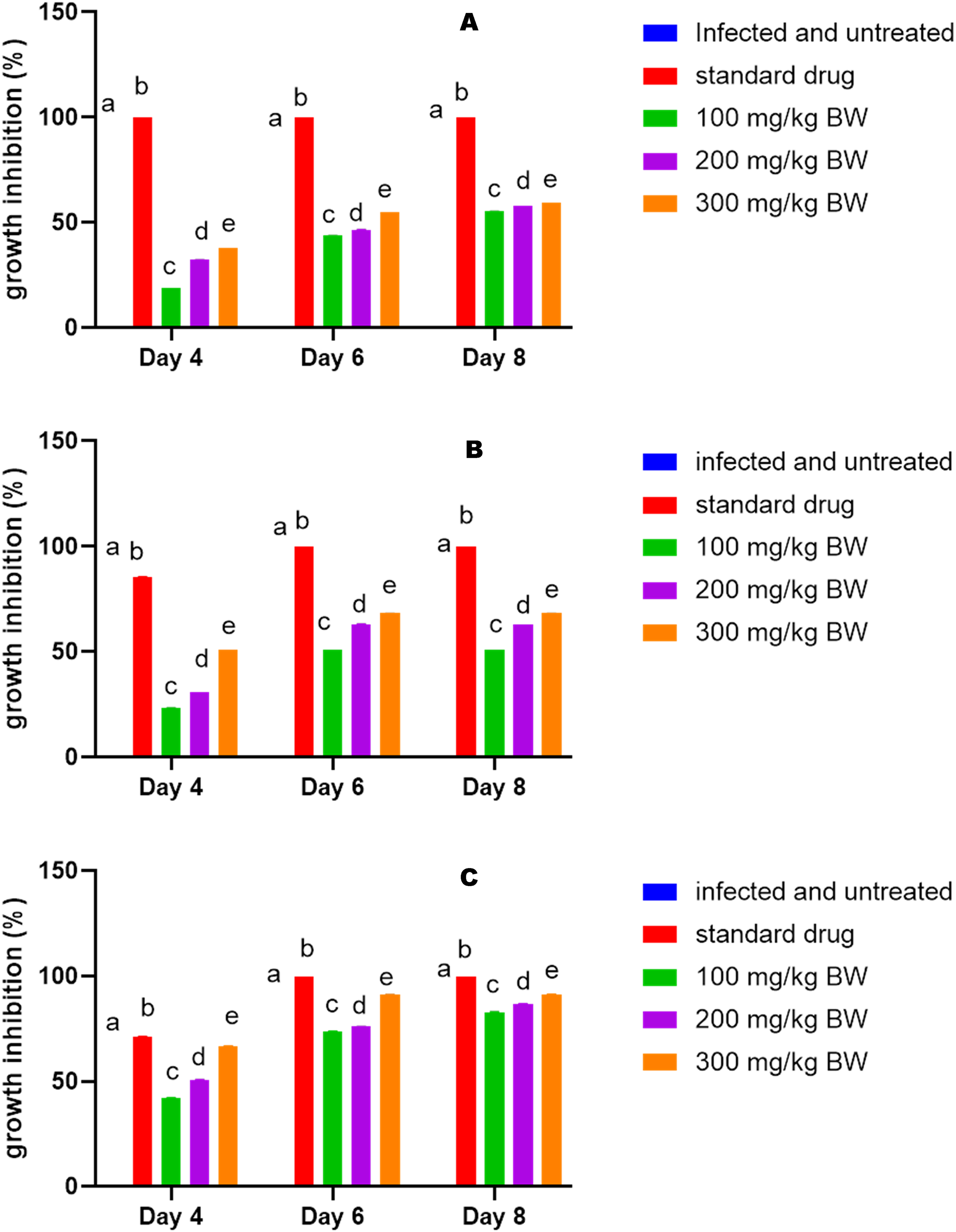
Parasitemia (A) and parasite growth inhibition (B) at fourth, sixth, and eighth day post infection with Plasmodium berghei NK-65 in mice treated with ethanolic extract of Annona muricata leaf (suppressive test, prophylactic test and curative test,). Values are means of four (5) replicates. BW; body weight, CQ; Chloroquine,. EEAML: ethanolic extract of Annona muricata leaf.
Haematological Indices of Plasmodium Berghei NK-65- Infected (Eighth Day Post Infection) Mice Treated with Ethanolic Extract of Annona Muricata Leaf (Curative Test).
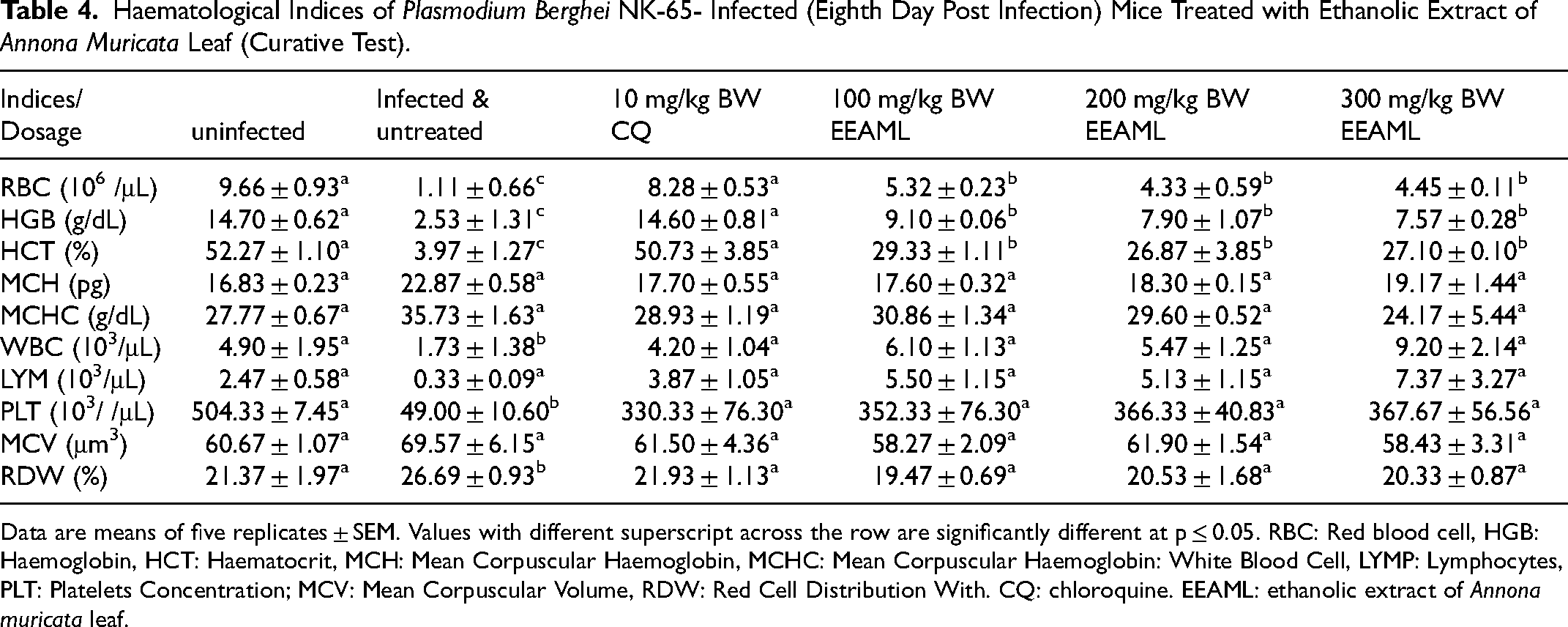
Data are means of five replicates ± SEM. Values with different superscript across the row are significantly different at p ≤ 0.05. RBC: Red blood cell, HGB: Haemoglobin, HCT: Haematocrit, MCH: Mean Corpuscular Haemoglobin, MCHC: Mean Corpuscular Haemoglobin: White Blood Cell, LYMP: Lymphocytes, PLT: Platelets Concentration; MCV: Mean Corpuscular Volume, RDW: Red Cell Distribution With. CQ: chloroquine. EEAML: ethanolic extract of Annona muricata leaf.
Relative to the reference compounds used in this study, two phytocompounds (apigenin and luteolin) exhibited the least minimal binding energies to the six Pfproteins (Table 5). The analysis of the docking scores shows that except for the reference compounds to plasmepsin II (binding energy of −8.8 kcal/mol), apigenin and luteolin demonstrated higher binding affinities to the five other Pfproteins (Table 5). The amino acid interactions of the Pfproteins with apigenin and luteolin are represented in Table 3. The interaction of residues of the proteins with respective ligand groups of the lead phytochemicals (LPs) was mainly through Hydrogen bonds (bonds below 3.40 Å were selected), hydrophobic interactions, and a few other bonds (Figures 6–11 and Table 6). The results of the interactive analysis show that the LPs (apigenin and luteolin) were orientated in the active site of Plasmodium falciparium falcipain 2 (PfF 2) in a similar binding pose as the reference inhibitor (E64301), though they interacted with a different amino acid of the active site. The 4-hydroxyphenyl moiety of apigenin formed a hydrogen bond with Ile85 and 2 hydrophobic interactions: Pi-sigma bonds with Leu84 and Pi-alkyl bonds with Ala175 of PfF 2. The chromen-4-one ring made a Pi-sulphur and amide-Pi stacking with Cys42 and Asn173 of PfF 2 respectively (Figure 6a). In the case of luteolin the 5,7-hydroxy groups that are attached to the chromen-4-one ring formed 2 hydrogen bonds with Asn81 and His174 of PfF 2, while the first ring of the chromen-4-one and 3, 4-dihydroxy phenyl ring made Pi-alkyl and Pi-sigma contacts with Cys42 and Leu84 of PfF 2 respectively (Figure 6b). Apigenin and luteolin were docked into the catalytic site of PfP II in a region accommodated by the reference hydroxyethyl amine-based inhibitor (49W400). Apigenin made 2 hydrogen bonds to Thr114 and Ser118 of PfP II through the hydroxyl group of the 4-hydroxyphenyl moiety. The 4-hydroxyphenyl ring formed Pi-sulphur and Pi-pi T-shaped contact with Met15 and Phe120. The chromen-4-one ring interacted through Pi-anion and Pi-alkyl with Asp214 and Val78 of PfP II respectively (Figure 7a). The 3-hydroxyl group of the 3,4-dihydroxy phenyl ring of luteolin made the only hydrogen bond with Asp34 and Pi-Pi T-Shaped and Pi-Alkyl contact with Tyr77 and Ala78 (Figure 7b). The 5-hydroxyl group of the chromen-4-one ring of apigenin formed a hydrogen bond with Thr97 and a Pi-sigma and Pi-alkyl contact with Ile31, while the 4-hydroxyphenyl ring made 3 Pi-alkyl contacts with Ala23, Pro250, and Pro246 of PfLDH (Figure 8a). The 5-hydroxyl and the carbonyl group of the chromen-4-one ring of luteolin formed 2 hydrogen bonds to Ser245 while the 7-hydroxyl group of the same ring formed a hydrogen bond with Met30. The chromen-4-one ring made Pi-alkyl contact with Ile31 and Pro250 of PfLDH (Figure 8b). The 5-hydroxyl group of chromen-4-one ring and the hydroxyl group of the 4-hydroxyphenyl ring made 2 hydrogen bonds with Ser239 and Tyr19 of PfPEMET. The chromen-4-one ring formed Pi-cation bonds in contact with Arg179 and Lys247. In the case of luteolin the all hydroxyl group on the 3,4-dihydroxy phenyl and chromen-4-one formed hydrogen bonds with the active residues of PfPEMET (Figure 9a), while the chromen-4-one ring made several Pi-cation with Lys247 and Arg179 of PfPEMET (Figure 9b). The 5-hydroxyl and the carbonyl group of the chromen-4-one ring of apigenin made hydrogen bonds with Val625 and His709 of PfPCCT respectively. The second chromen-4-one ring of apigenin formed Pi-cation and Pi-anion to His633 and Asp710 of PfPCCT respectively (Figure 10a). In the case of luteolin, the carbonyl and 5-hydroxyl group of the chromen-4-one moiety formed 2 hydrogen bonds with Thr756, while the hydroxyl group of 4-hydroxyl group of the 3,4-dihydroxy phenyl ring formed a hydrogen bond with Asp710. 2 Pi-cation interactions were made between the chromen-4-one ring of luteolin and Arg755 and His630 of PfPCCT (Figure 10b). Apigenin and luteolin were accommodated in the binding cavity of the 1H-Benzimidazole-2-pentanamine reference inhibitor of PfSS. The 4-hydroxyl of the 4-hydroxyphenyl ring, 7-hydroxyl, and the carbonyl group of the chromen-4-one ring form hydrogen bonds with Asp199, Glu147, and Ala204 respectively, the rings also formed several Pi-alkyl contacts with Val91, Pro203, Val152 and Leu88 of PfSS (Figure 11a).

Amino acid interactions of reference compound and lead phytochemicals from the docking studies in the active site of Plasmodium falciparum falciparin 2. Stick representations of the ligands are shown by colours (A) Green: reference compound (B) Red: Apigenine (C) Gold: Luteolin while, (D) carton representation of ligands in binding pocket of the protein.

Amino acid interactions of reference compound and lead phytochemicals from the docking studies in the active site of Plasmodium falciparum plasmepsin II. Stick representations of the ligands are shown by colours (A) Green: reference compound (B) Red: Apigenine (C) Gold: Luteolin while, (D) carton representation of ligands in binding pocket of the protein.

Amino acid interactions of reference compound and lead phytochemicals from the docking studies in the active site of Plasmodium falciparum lactate dehydrogenase. Stick representations of the ligands are shown by colours (A) Green: reference compound (B) Red: Apigenine (C) Gold: Luteolin while, (D) carton representation of ligands in binding pocket of the protein.

Amino acid interactions of reference compound and lead phytochemicals from the docking studies in the active site of Plasmodium falciparum phoosphoethanomaline methyltransferase. Stick representations of the ligands are shown by colours (A) Green: reference compound (B) Red: Apigenine (C) Gold: Luteolin while, (D) carton representation of ligands in binding pocket of the protein.
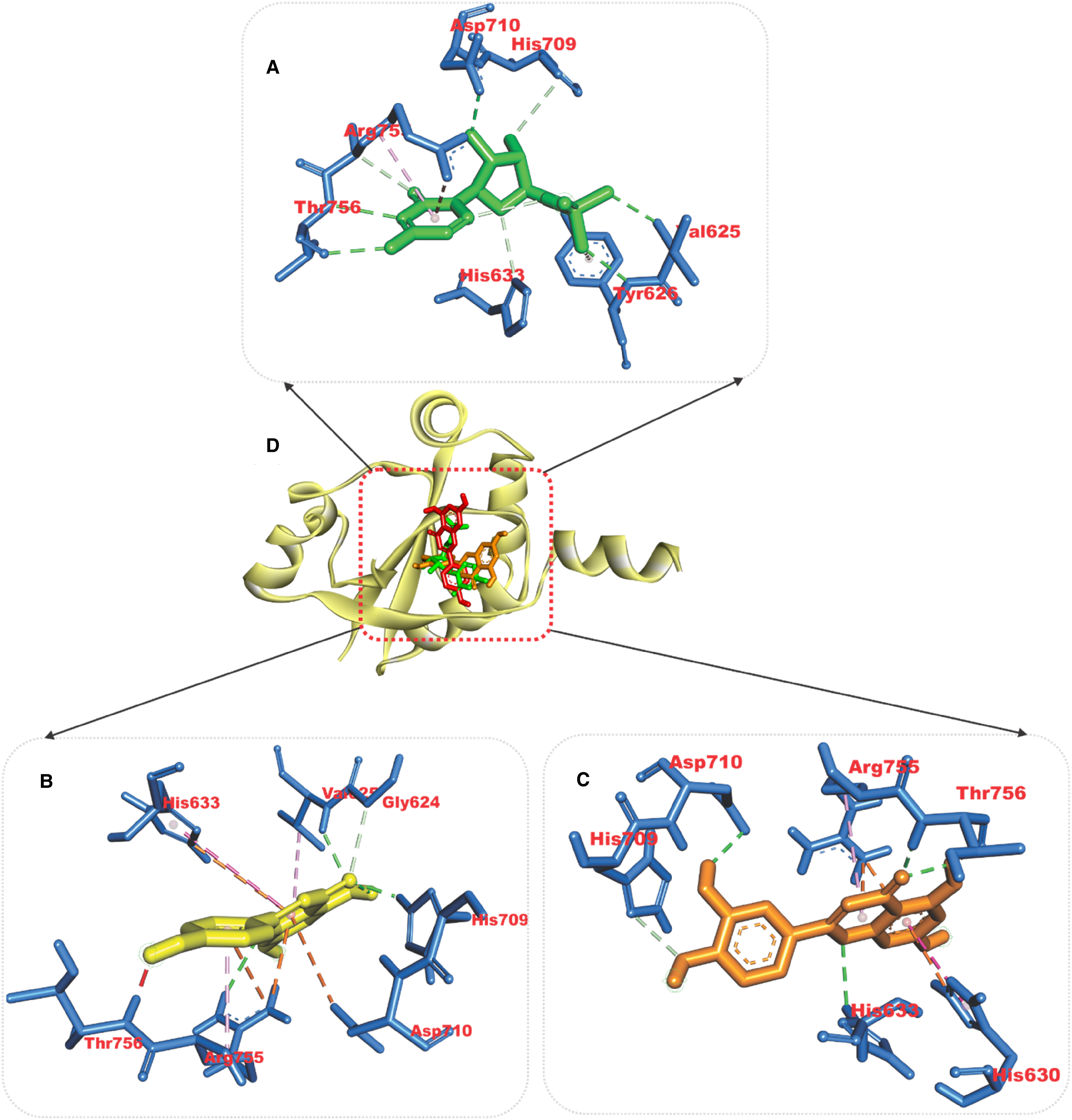
Amino acid interactions of reference compound and lead phytochemicals from the docking studies in the active site of Plasmodium falciparum phosphocholine cytidylyltransferase. Stick representations of the ligands are shown by colours (A) Green: reference compound (B) Yellow: Apigenine (C) Gold: Luteolin while, (D) carton representation of ligands in binding pocket of the protein.
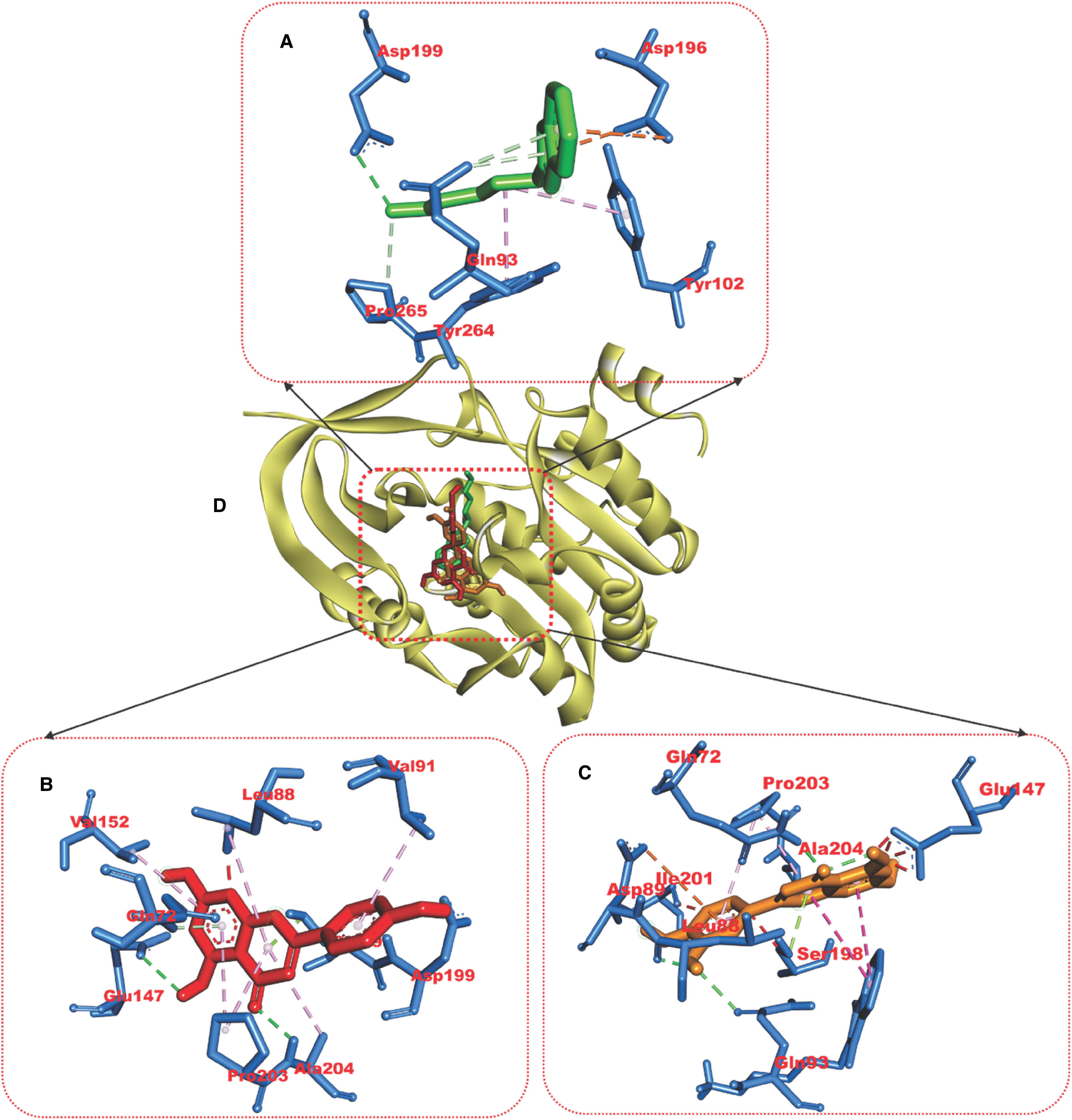
Amino acid interactions of reference compound and lead phytochemicals from the docking studies in the active site of Plasmodium falciparum Spermidine synthase. Stick representations of the ligands are shown by colours (A) Green: reference compound (B) Red: Apigenine (C) Gold: Luteolin while, (D) carton representation of ligands in binding pocket of the protein.
Binding Affinities of Identified Phytochemicals and Reference Compounds to the Active Site Residues of Plasmodium Falciparum Proteins.
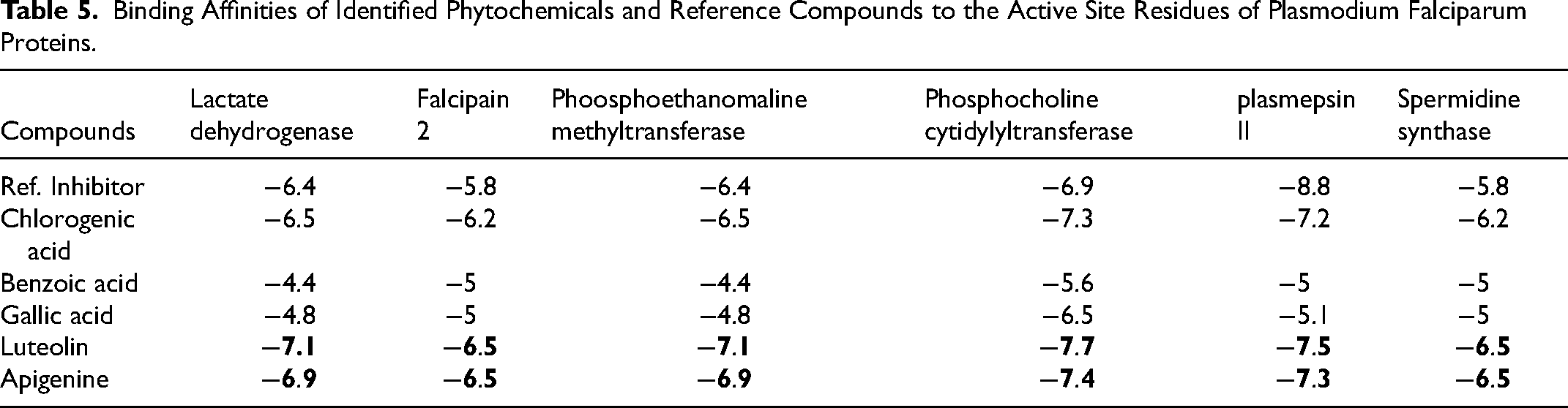

Interactions of Top Docked Phytochemicals with Active Site Residues of Plasmodium Falciparum Proteins.

The structural integrity of the unbound PfProteins was compared with the complexes of LP in a full atomistic MD simulation. From the MD simulation trajectories obtain for each system, the following thermodynamic parameters (RMSD, RMSF, SASA, RoG, and number of H-bonds) were computed. The parameters of the complexes were plotted as a function of the time frame in nanoseconds (ns) and compared with the unbound PfProtein as the reference structure. The average values for the five calculated parameters are presented in table 7. The stability and structural integrity of the protein structures were measured using The RMSD plots, by observing the rate of the deviation of the spikes from the reference structure. The four Pfproteins were equilibrated around 12, 20, 7, and 11 ns for the phosphoethanolamine methyltransferase, spermidine synthase, plasmepsin II and falcipain 2 systems respectively. The RMSD plots (Figure 12) and computed average RMSD values (Table 7) for the four Pfprotein systems show that the unbound protein expressed higher fluctuation than the LPs-Pfprotein complexes, indicating a more stable structure upon the binding of the LPs. The level of compactness of Pfproteins upon binding of the LPs was evaluated from the RoG plots and mean values. A stably folded protein structure exhibits a steady RoG plot. Figure 13 shows the RoG plots of the four Pfproteins systems. The plots for the Pfproteins systems show a steady progression with minimal fluctuations that is less than 3 Å. The mean RoG values computed for the Pfproteins systems show that the binding of the LPS to the active site of the proteins did not strain the structural conformation of the Pfproteins. The lower average RoG values for the LPs Pfproteins complexed systems suggest a more compacted confirmation upon the binding of the LPs. The SASA plots and calculated mean values for the systems are a complimentary analysis of the RoG that assesses the accessibility of solvent by the surface of the Pfproteins. Both RoG and SASA plots indicate the level of the structural unfolding of proteins regarding their original structure. A similar pattern of progression was observed for both the unbound and the complexed systems with minimal fluctuation. A slight reduction in the mean SASA values of the LPs-Pfproteins complexed as compared to the unbound systems was observed, this further shows that the binding of the LPs did not distort the structural conformation of the proteins (Figure 14). The RMSF plots show the flexibility of the individual amino acid residues of the Pfprotein. Because of terminal motions, higher fluctuations were observed at the N and C terminal ends of the proteins (Figure 15). A. Based on the computed average RMSF values, the LPs-Pfprotein complexed systems had lower mean RMSF values except for all of the Pfprotein. This also suggests that the binding of LPs resulted in more compacted structures. In the case of phosphoethanolamine methyltransferase systems, the spike in the fluctuation around amino acid residues 93 and 217 was minimized by the binding of the LP as observed in Figure 5a; while the binding of apigenin increased the fluctuation of amino acid residue around 64 and 241. Figure 16 depicts the average number of hydrogen bonds for the unbound and complexed systems of the four Pfproteins. The phosphoethanolamine methyltransferase complexes systems had a slight increase in the average number of hydrogen bonds when compared to the unbound protein (Table 7).
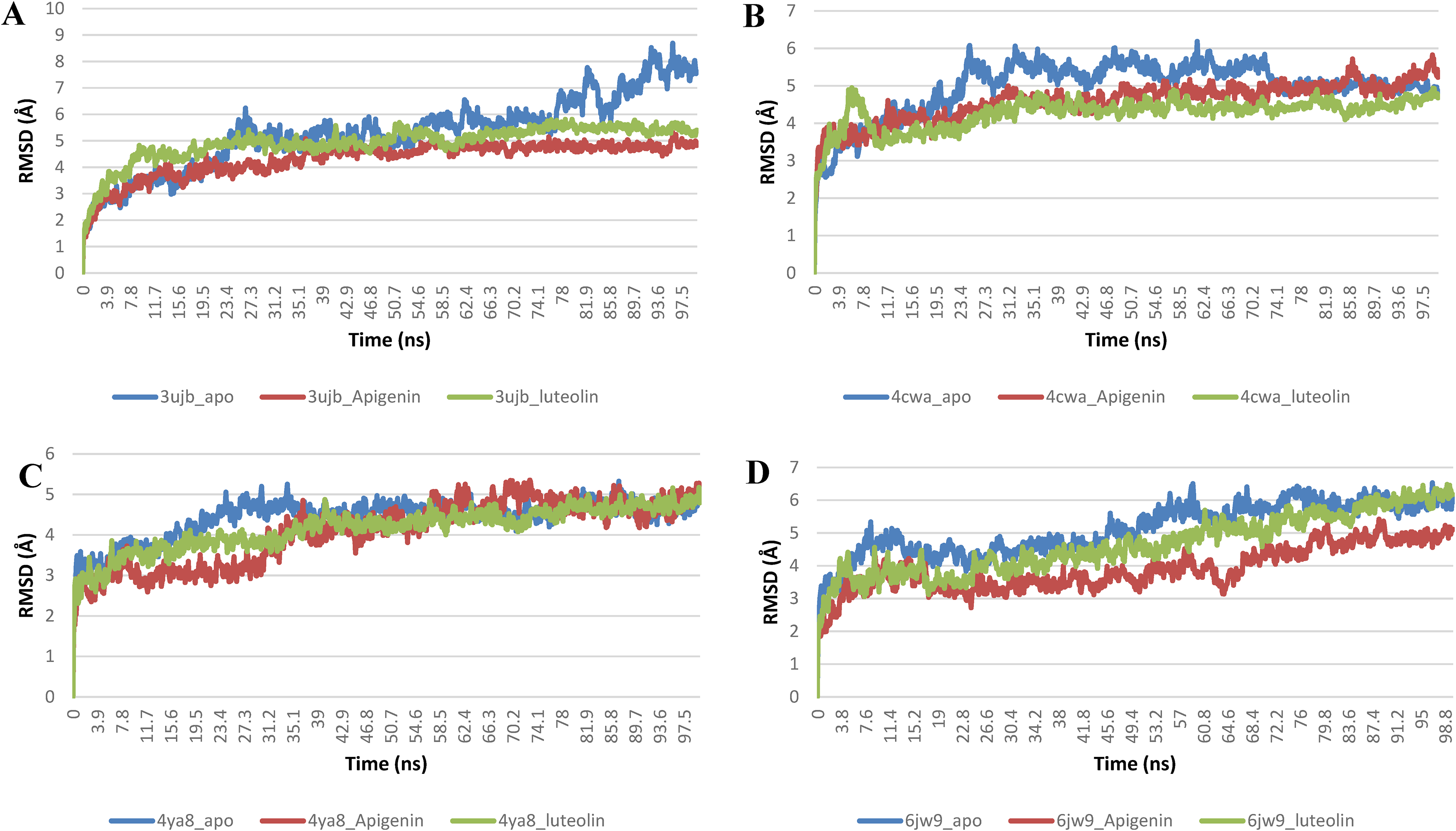
The Backbone-Root Mean Square Deviation (RMSD) plots of molecular dynamics (MD) simulation of (A) phoosphoethanomaline methyltransferase (3UJB) (B) Spermidine synthase (4cwa) (C) plasmepsin II (4YA8) (D) falciparin 2 (6JW9).

The Radius of gyration (RoG) plots of molecular dynamics (MD) simulation of (A) phoosphoethanomaline methyltransferase (3UJB) (B) Spermidine synthase (4cwa) (C) plasmepsin II (4YA8) (D) falciparin 2 (6JW9).

The Surface Accessible Surface Area (SASA) plots of molecular dynamics (MD) simulation of (A) phoosphoethanomaline methyltransferase (3UJB) (B) Spermidine synthase (4cwa) (C) plasmepsin II (4YA8) (D) falciparin 2 (6JW9).

Per residue Root Mean Square Fluctuations (RMSF) plots of molecular dynamics (MD) simulation of (A) phoosphoethanomaline methyltransferase (3UJB) (B) Spermidine synthase (4cwa) (C) plasmepsin II (4YA8) (D) falciparin 2 (6JW9).

The changes in the number of H-bonds during the MDS trajectory of (A) phoosphoethanomaline methyltransferase (3UJB) (B) Spermidine synthase (4cwa) (C) plasmepsin II (4YA8) (D) falciparin 2 (6JW9).
The Mean Values for the Computed Thermodynamic Parameters from the MD Simulation Trajectories of Plasmodium Falciparum Proteins.

The total cholesterol concentration in the liver of Plasmodium berghei NK-65-infected mice treated with ethanolic extract of Annona muricata leaf decreased significantly (p < 0.05) when compared to the untreated infected mice (Figure 17A). Furthermore, there was no significant (p > 0.05) reduction in cholesterol concentration in the liver of uninfected mice or those treated with the standard drug (chloroquine) compared to the EEAML group. There was a significant increase (p < 0.05) in liver total triglyceride concentration in Plasmodium berghei NK-65- infected mice compared to the uninfected mice. There was a significant (p < 0.05) increase in the liver total triglyceride concentration in the infected mice treated with 100 mg/kg body weight and significantly decreased in the groups treated with 200 and 300 mg/kg body weight compared to Plasmodium berghei NK-65- infected mice and untreated (Figure 16B). There was a significant increase (p < 0.05) in triglycerides levels in the group treated with the standard drug compared to the uninfected mice (Figure 17B). The plasma HDL-c of the infected untreated mice was significantly decreased (p < 0.05) compared to the uninfected mice (negative control). The EEAML when administered to Plasmodium berghei NK-65-infected mice significantly increased (p < 0.05) the plasma HDL-cholesterol concentration compared to the untreated infected mice. There was a significant increase (p < 0.05) in the plasma HDL-c in the infected mice treated with the standard drug compared to the infected and untreated (Figure 17C). There was a significant increase (p < 0.05) in HDL-triglyceride concentration in Plasmodium berghei NK-65- infected mice and untreated compared to the negative control (uninfected). There was a decrease (p < 0.05) in plasma triglycerides in the group treated with 100 mg/kg body weight of the extract but was increased significantly (p < 0.05) in the groups treated with 200 and 300 mg/kg body weight of extract compared to P. berghei NK-65- infected and untreated mice (Figure 17D). The P. berghei NK-65- infected mice treated with standard drug recorded a significantly increased (p < 0.05) in triglycerides compared to uninfected mice (Figure 17D). There was a significant decrease (p < 0.05) in liver protein in Plasmodium berghei NK-65- infected mice and untreated compared to the uninfected mice. There was a significant increase (p < 0.05) in the liver HMG-CoA/mevalonate ratio in the mice infected groups compared to the mice in the control. However, the mice treated with 200 mg/kg body weight of EEAML showed a significant decrease (p < 0.05) in the HMG-CoA/mevalonate ratio compared to the untreated Plasmodium berghei NK-65- infected mice (Figure 18A). Plasmodium berghei NK-65-infected mice treated with EEAML showed a significant increase (p < 0.05) in liver protein in a dose-dependent manner compared with untreated Plasmodium berghei NK-65- infected mice. The liver total protein concentration in mice treated with EEAML also showed a significant increase (p < 0.05) compared to untreated control and the group treated with chloroquine (Figure 18B).

Liver total cholesterol (A), triglyceride (B), Plasma HDL-cholesterol (C) and Plasma HDL-triglyceride (D) concentration of Plasmodium berghei NK −65- infected mice (8-day post infection) treated with ethanolic extract of Annona muricata leaf (EEAML). CQ: chloroquine

Liver β-Hydroxy β-methylglutaryl-CoA (HMG-CoA)-mevalonate (A) and total protein concentration (B) of Plasmodium berghei NK-65- infected mice (8-day post infection) treated with ethanolic extract of Annona muricata leaf. CQ: chloroquine. Means ± SEM of five (5) replicates. Bars with different superscripts are significantly different (p < 0.05).
There was a significant (p < 0.05) increase in acid phosphatase activity in Plasmodium berghei NK 65- infected mice (8-day post-infection) untreated compared to the mice in the control group. Similarly, the activity of the enzyme also increased (p < 0.05) significantly in the infected mice treated with doses of EEAML and chloroquine compared to mice in the control (Figure 19A). Alkaline phosphatase activity in Plasmodium berghei NK- 65- infected mice (8-day post-infection) treated with EEAML at 200 and 300 mg/kg BW was significantly increased (p < 0.05) compared with ALP in uninfected mice and untreated Plasmodium berghei NK-65- infected mice (Figure 19B). ALP activity in infected mice treated with chloroquine also increased significantly compared to infected untreated mice. The liver MDA concentration of Plasmodium berghei NK-65-infected mice (8-day post-infection) untreated showed a significant (p < 0.05) increase compared to the control group. Meanwhile, the groups treated with doses of EEAML and the standard drug showed a significant decrease (p < 0.05) in MDA concentration compared to Plasmodium berghei NK-65- infected mice (8-day post-infection) untreated group (Figure 20). Neutrophil infiltration and liver cell degeneration were visible in the infected mice's histology; (Figure 21 II). However, those who received the extract in addition to the conventional medication displayed normal hepatic cell architecture (Figure 21 I, III-VI)

Liver acid phosphatase (A) and alkaline phosphatase (B) activity of Plasmodium berghei NK-65- infected mice (8-day post infection) treated with ethanolic extract of Annona muricata leaf. CQ: chloroquine. Means ± SEM of five (5) replicates. Bars with different superscripts are significantly different (p < 0.05).

Liver MDA concentration of Plasmodium berghei NK-65- infected mice (8-day post infection) and treated with ethanolic extract of Annona muricata leaf. CQ: chloroquine. Means ± SEM of five (5) replicates. Bars with different superscripts are significantly different (p < 0.05).

Photomicrograph of liver of Plasmodium berghei NK-65- infected mice (eighth day post infection) (H&E X 40): Group I normal liver architecture, Group II, liver cell degeneration, infiltration with neutrophils, Group III, normal hepatic cell, Group IV, normal hepatic cells Group V, hepatic cell with no infiltration, Group IV, mild infiltration of the liver cell, Group V, normal hepatic cell, Group VI, normal cell architecture.
Discussion
Flavonoids, triterpenes, terpenoids, anthocyanin, saponins, and tannins present in the extract are attributed to the biological activities exhibited by medicinal plants.62,63 Medicinal plants have been used for the treatment of ailments and the management of disease conditions long before the onset of modern medicine and the use of the synthetic drug. Due to bioactive ingredients present in herbal plants; they have been beneficial in the treatment of malarial infections. Malaria infection caused by Plasmodium species leads to the destruction of human red blood cells and associated pathogenic conditions. These bioactive components of medicinal plants inhibit the proliferation of the parasites through various mechanisms. The antimalarial effect of saponin is linked to a reduction in blood cholesterol, which plasmodium scavenges for its proliferation. This finding supports a prior study that found saponin to have cholesterol-lowering properties. 64 Saponins isolated in some plants have the potential to cause hemolysis which will result in parasite death. 65 Similarly, alkaloid compounds from several plants exhibit antiplasmodial activity.66–69 It has also been documented that terpenes or terpenoids have bioactive ingredients that are used as an active antiprotozoal and antimalarial agents in many pharmacological studies. 70 Artemisinin, the antimalarial bioactive ingredient in the traditional Chinese antimalarial qinghaosu is a terpenoid. Antimalarial activity against strains of the Plasmodium parasite has been demonstrated in previous studies using plant extract.71,72 Derivatives of 9,10-anthraquinone include many important drugs such as antimalarials like rufigallol. 73 The antimalarial activity observed by the crude extract in this study could be attributed to the individual activity of identified phytochemicals or to their synergistic influence on each other. Similarly, these compounds may have a different mode of action or drug site of action that is responsible for the observed antimalarial activity. The weight decrease in infected mice observed in this study at the commencement of treatment may be attributed to the inhibitory effect of the disease on the system metabolism or the suppressive effect of the infection on the appetite control center in the brain. 74 The increased percentage of the average parasitemia in P. berghei inoculated mice was attributed to the rapid proliferation of the Plasmodium parasite, which also correlated with the high degree of infection in the mice; this eventually resulted in the death of the mice. 75 Fidock, Rosenthal, Croft, Brun and Nwaka 76 and Ene, Atawodi, Ameh, Kwanshie and Agomo 77 also observed a high percentage of parasitemia in mice infected with P. berghei after 5 days and the death of mice after 7 days of inoculation. In this investigation, the extract showed a steady decline in parasitemia from day 4 to 8 day-post infection, demonstrating its efficacy for both therapeutic and preventive antimalarial activity. Similarly, previous experiments reported that the antimalarial activity of plant extract was due to the presence of certain secondary metabolites. Since EEAML is rich in flavonoids, terpenes, and terpenoids, it could be deduced that the antimalarial activity of this medicinal plant was due to the antioxidant activity of these secondary metabolites or the synergistic effect.78–80 Furthermore, the antioxidant capabilities of the extract may be beneficial in combating the generated oxidative stress caused by Plasmodium parasite activity and infection. This may be one of their modes of action in malaria therapy. The erythrocytic stage of malaria infection is responsible for hematological changes that have been reported to be the most common malaria complications and are responsible for the pathogenesis of the infection. Red blood cells, White blood cells, and thrombocytes are the major cells that are changed by the infection.81–84 Patients diagnosed with malaria tended to have substantially lower platelets, WBCs, lymphocytes, eosinophils, RBCs, and Hb levels while monocytes and neutrophils were significantly higher relative to patients infected with non-malaria.82,83,85–87 The most common complication during malaria infection is thrombocytopenia.86,88 The low RBC levels observed in the untreated animals in this study could be due to Plasmodium destruction of red blood cells; RBC being the Plasmodium parasite's primary target. Furthermore, several other investigations have linked low RBC to situations such as rapid clearance of both parasitized and non-parasitized RBC, 89 bone marrow dysfunction89,90 and the level of parasitemia. 91 Finally, low RBC volume will result in low pack cell volume (HCT). However, the extract's enhanced RBC volume in this study could be related to its anti-plasmodial activity, which resulted in a decrease in parasitemia level. Similarly, malarial infection resulted in a large decrease in WBC volume, which was attributable to WBC deployment to fight the parasite. This also explains the decrease in lymphocyte volume caused by malarial infection. The extract's antimalarial action is also linked to the restoration of WBC volume in treated rats. Further, this study also showed that Plasmodium infection resulted in high levels of MCV, MCH, MCHC, and RDW. This may be an adaptive mechanism due to assault by terpenoids on the pathogenicity of the infection. 82 The decrease in the level is associated with malarial infection hence the finding in this study corroborated what was reported. However, these biomarkers were reversed by the treatment with the extract and standard drug. The same effect reported on hematological parameters of mice infected with Plasmodium parasite was also observed with the n-hexane fraction of the extract. Malaria parasite infection is associated with certain biochemical changes such as lipid profile changes. Some researchers reported that the magnitude of the change correlates to the severity of the infection.92,93 Therefore the transient change in the lipid profile which is also dependent on the parasitaemic stage has been suggested to be a potential adjuvant diagnostic parameter for malarial.94–96
Furthermore, docking experiment in this study was motivated by the previous study which reported the anti-parasitic activity of flavonoids.97,98 The antimalarial drug action/mechanism can be elucidated through prediction with the binding energies of molecular interactions between plasmodium drug target proteins and chemical ligands of plant extract. Further information from the in silico study will further consolidate and provide a molecular explanation of in vivo experimental findings. 99 In this present study, the interaction between six P. falciparum drug targets proteins and the HPLC ligand of ethanolic leaf extract of Annona muricata were explored through molecular docking and molecular dynamics simulation. The aspartate proteases (Plasmepsins I-V) and the cysteine proteases (falciparum 1, 2, and 3) are potential targets for antimalarial. 100 Plasmodium species utilize these proteases to degrade hemoglobin for its amino acids. Peptides that are already cleaved by plasmepsins are further broken down into smaller fragments and amino acids by falcipain-2, a cysteine protease, of the parasite. 101 Hence compounds that inhibit these proteases ultimately will interfere with the parasite's ability to degrade hemoglobin, which will consequently limit the availability of amino acids necessary for parasite proliferation which will then result in the death of the parasite. 102 Apigenin and luteolin when docked in the protease active site of plasmepsin II interacted with the catalytic Asp34 and Asp214 side chains just like the hydroxyethyl amine-based reference inhibitor 103 and this may suggest that the antimalarial activity of the extract could be to obstruct the parasite potential to degrade hemoglobin in RBC. Similarly, Apigenin and luteolin high binding affinity with P. falciparum falcipains than the native ligand is also a prediction that the exhibited antimalarial by the extract could be through the obstruction of parasite hemoglobin degradation which will limit the availability of amino acids needed for parasite growth.100,104 It is worthy of note that, the LPs interacted with several Cys residues of the active site of the falcipain-2 active site through alkyl hydrophobic interaction. Nitrites are known inhibitors of cysteine proteases. They form a reversible thioimidate intermediate from the nucleophilic attack of the catalytic cysteine residue105,106 and have been reported to display excellent inhibitory activity against falcipain-2 and cultured P. falciparum. 107 Thus, the interaction with cysteine residues of falcipain-2 could be one of the mechanisms by which the LPs exhibited their antiplasmodial activity. Because of the vital role in energy production in P. falciparum lactate dehydrogenase (PfLDH) in the parasite, it has been considered a potential drug target enzyme for some antimalarial drugs of the future. 108 Chloroquine interacts with PfLDH in NADH pocket by binding at a similar position as the adenyl ring of the coenzyme where it blocks glycolysis by competitively inhibiting this important enzyme of this pathway. 108 Apigenin and luteolin like the reference drug interacted with lactate protein in the hydrophobic pockets created largely by Pro246, Tyr247, and Pro250. A similar feature of competitive inhibition with NADH binding was reported for gossypol and 8-deoxy hemigossylic acid derivatives.109,110 Due to the absence of functional cytochrome necessary for oxidative phosphorylation in Plasmodium species, they depend mainly on glycolysis for energy supply. 111 Furthermore, NAD+ required for glycolysis in the plasmodium parasite comes from lactate dehydrogenase because the erythrocyte which houses the erythrocytic developmental cycle of the parasite lacks mitochondria. 112 Therefore, inhibition of PfLDH by the LPs may affect the production of ATP and consequently result in energy deprivation and death of the organism. 110 A multifunctional enzyme, P. falciparum phosphoethanolamine methyltransferase (PfPEMT) which catalyzes the methylation of phosphoethanolamine (pEA) to phosphocholine which is needed for membrane biogenesis is vital for the parasite growth, survival of the parasite and can be a potential an antiparasitic drug target enzyme. 113 The LPs like the reference inhibitor interacted with Tyr-19 and His-132 of the “catalytic” latch that locks ligands in the active site and orders the site for catalytic reaction behind phosphobase methylation. 113 In addition, each of the tyrosine residues in the site (Tyr-27, Tyr-160, Tyr-175, and Tyr-181) has been demonstrated to be important for the activity of PfPEMT. 113 Phosphocholine cytidylyltransferase PfPCCT, the rate-limiting enzyme in the Kennedy pathway for the biosynthesis of phosphatidylcholine, a vital and abundant phospholipid for membrane formation in plasmodium parasites, is an important drug target enzyme for antimalarial drugs. 114 The LPs docked into the substrate-binding site and interacted with the D623, Y626, Q636, K663, W692, H709, Y714, I740, Y741, and R755 active site residues responsible for ligand accommodation. This inhibits the enzyme's catalytic mechanism by preventing further interactions with the nucleotide and choline moieties of the enzyme-substrate. 115 The aminopropyl moiety of decarboxylated S-adenosylmethionine is transferred to putrescine by the transferase PfSS, resulting in the formation of spermidine and 5-methylthioadenosine, an important polyamine biosynthetic intermediate. 116 Interactions with LPs at the active site of the enzyme, as shown by the docking result into the enzyme's ligand-binding domain, could inhibit the enzyme and kill the parasite. The plots of the various thermodynamic parameters (RMSD, RMSF, RoG, SAS, and H-bonds number) calculated from the MDs trajectory files of the apo and complexed Pfproteins show that the complexes were stable throughout the simulation time. 117 LPs-Pfproteins complex when compared to the apo Pfproteins exhibited low average values for every parameter. When bound with apigenin and luteolin, the integrity of the enzyme structure was preserved but formed a more compact structure.118–120 Therefore further experimental validation must be performed with these complex systems.
The therapeutic effect of antimalarial agents, which includes total parasite removal or reduction of parasite load in the host, will also result in the reversal of the biochemical changes caused by infection to that of a healthy state. The extract's induction of xenobiotic-metabolizing enzyme production may be responsible for the increased liver tissue total protein found in this study. Furthermore, the HMG-CoA / mevalonate ratio assesses HMG-CoA reductase activity, which is the rate-limiting step in the cholesterol biosynthetic pathway. The inhibition of HMG-CoA reductase activity observed in this study could be attributed to feedback inhibition caused by an increased level of cholesterol in the host hepatocyte. Also, the increase in hepatic cholesterol caused by infection, as demonstrated in this study, could be due to an increase in the expression of host genes involved in metabolite transport and anabolism.119,121 The parasite lacks the enzymatic pathways to synthesize the cholesterol. 122 Hence, the decrease in cholesterol in this present study may indicate a reduction in parasitaemia due to the growth inhibitory effect of the extract on parasite proliferation; hence a decrease in hepatic cholesterol could indicate a decrease in parasite levels. Similarly, the decrease in hepatic triglycerides could lead to decrease in the parasite levels in the liver. This is because of the depravation of vital molecule for the parasite to divide because the parasites’ ability to synthesize fatty acids is limited since they must scavenge for them from their hosts. The parasite replication in host cells necessitates a high demand for lipids, which the parasite requires for the development of organelle membranes and the accumulation of lipids in lipid bodies. 123 In addition to support the findings in this study, hypertriglyceridemia has been described in several diseases with haemophagocytosis. 124 At the blood-stage development of the infections disease, the Plasmodium relies mostly on the host HDL as the source of cholesterol. The decrease in HDL-cholesterol observed in this study aligns with some other studies which showed that a major lipid source for Plasmodium is HDL which transfers their lipid cargo to the erythrocyte membrane. 125 The same study also demonstrated that the serum replacement by different fractions of lipoprotein showed that only the HDL fraction is capable of ensuring total schizogony as well as erythrocyte reinvasion. 126 The increase in liver ACP activity by malarial infection reported in this study corroborated the previous findings which reported an increase in serum ACP. 127 Increased production of ROS associated with the malarial infection will lead to an imbalance in antioxidants with resultant peroxide lysis and consequently trigger the release of a certain enzyme such as ACP. 128 The infection did not increase the ACP activity in this present activity however the increase in the activity reported in the group treated with the extract could be due to assault on the liver cell by the extract. Most infection diseases cause significant inflammation and oxidative stress.129,130 As a result, the increase in MDA in the liver caused by infection could be due to increased ROS production by the innate immune cell and parasite metabolism activity in the host. Innate immune cells detect pathogens and launch an inflammatory response to combat them. The pathogen that the cells phagocytose is eliminated by a rapid increase in ROS production in their phagosome, a process known as an oxidative or respiratory burst. 131 The liver cells are affected by the plasmodium parasite. Previous work has shown that the infection of Plasmodium parasite cause increase in the kupffer cell with marked congestion. 132 The normal cell free of infiltration implied that the extract was able to clear the liver of the parasite by the extract.
Conclusion
The plant leaves extract demonstrated antimalarial activity in mice infected with Plasmodium berghei NK −65. The bioactive elements present in the plant that have antiplasmodial and hypolipemic actions were the mediators of this effect. The in-silico analysis also showed that the flavonoid components apigenin and luteolin reduced the activity of some Plasmodium proteins necessary for the parasite's adaptability and survival in the host.
Footnotes
Acknowledgements
Not Applicable.
Authors’ Contributions
C.O.N and E.A.B conceive the study. C.O.N carried out the literature search. C.O.N and G.A. G performed the statistical and docking analysis. C.O.N and G.A. G reviewed the final draft.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Trial Registration
This is not applicable since it is purely an animal study and not clinical trial.
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Informed Consent
This is not applicable because it is not clinical study.

