Abstract
The endoplasmic reticulum (ER) and lipid droplets (LDs) are essential organelles involved in lipid synthesis, storage, and transport. Physical membrane contacts between the ER and LDs facilitate lipid and protein exchange and thus play a critical role in regulating cellular lipid homeostasis. Recent research has revealed that ER-LD membrane contact sites are targeted by pathogens seeking to exploit host lipid metabolic processes. Both viruses and bacteria manipulate ER-LD membrane contact sites to enhance their replication and survival within the host. This review discusses the research advancements elucidating the mechanisms by which pathogens manipulate the ER-LD contacts through protein molecular mimicry and host cell protein manipulation, thereby hijacking host lipid metabolic processes to facilitate pathogenesis. Understanding the crosstalk between ER and LDs during infection provides deeper insight into host lipid regulation and uncovers potential therapeutic targets for treating infectious diseases.
Keywords
Lipid Droplet Function
Lipid droplets (LDs) are single membrane organelles that serve as a reservoir of neutral lipids, such as triacylglycerol (TAG) and sterol esters (SE). LDs control lipid flow between organelles to fulfill cellular energy requirements and supply fatty acid precursors for synthesis of membrane and signaling lipids (Olzmann and Carvalho, 2019). As a result, LDs play key roles in regulating intracellular signaling, the innate immune response, and metabolic processes such as autophagy. While found ubiquitously in most eukaryotic cells, LD number and size vary widely among cell types and metabolic state. For example, excess cellular carbon sources such as fats or glucose promote neutral lipid production, leading to increased LD number and size. In contrast, fewer and smaller LDs occur when carbon sources are limited due to increased LD lipolysis and mobilization of stored fatty acids.
While LDs were once considered inert storage organelles, recent research has demonstrated that LDs actively interact with various cellular organelles, including mitochondria, endosomes, lysosomes, peroxisomes, plasma membrane, nuclear envelope, and the endoplasmic reticulum (ER) (Gao and Goodman, 2015; Olzmann and Carvalho, 2019; Herker et al., 2021). These interactions occur at specialized membrane contact sites (MCS), where the membranes are in close proximity (less than 30–40 nm apart) but do not fuse (Scorrano et al., 2019). Lipid transfer proteins (LTPs) play a key role at MCS by facilitating the movement of lipids between organelles. LTPs contain a hydrophobic cavity that shields lipids during transport, thereby improving the efficiency and specificity of lipid transfer between organelles (Wong et al., 2019). Thus, LD contact sites are crucial for maintaining cellular lipid homeostasis, energy metabolism, and membrane synthesis. Among these various LD interactions, the crosstalk between LDs and the ER is particularly significant. As the primary site for lipid synthesis and the origin of LD biogenesis, the close physical and functional association between the ER and LDs ensures efficient management of lipid stores and regulation of lipid metabolism (Xu et al., 2012; Kassan et al., 2013).
Lipid Droplet Biogenesis
LD biogenesis is initiated by the synthesis of neutral lipids, primarily TAG and SE, in specialized ER subdomains enriched with proteins critical for LD formation (Joshi et al., 2018; Salo et al., 2019) (Figure 1). This includes Acyl-CoA Synthetase Long-chain family member 3 (ACSL3), which synthesizes acyl-CoA for transfer to excess fatty acids by the diacylglycerol acyltransferase enzymes DGAT1 and DGAT2 (Kassan et al., 2013; Farese and Walther, 2023). TAG can also be generated in a DGAT-independent pathway by the acyltransferase DGAT1/2-Independent Enzyme Synthesizing Storage Lipids (DIESL, also known as TMEM68) (McLelland et al., 2023). DIESL is a multi-pass transmembrane protein located in the ER which utilizes phospholipids as acyl donors for TAG synthesis. Cholesterol esters are synthesized by Acyl-CoA Cholesterol O-Acyltransferases 1 and 2 (ACAT1 and ACAT2) (Anderson et al., 1998; Cases et al., 1998). TAG and SE are synthesized at specialized ER subdomains where the increasing neutral lipid concentration triggers LD biogenesis. At low concentrations, neutral lipids freely diffuse within the ER phospholipid bilayer (Thiam and Ikonen, 2021). However, once the neutral lipid concentration exceeds a certain threshold, the neutral lipids spontaneously phase separate to form a lipid lens which subsequently buds into the cytosol as an LD (Thiam and Ikonen, 2021). While lipid phase separation can occur spontaneously, the ER integral membrane protein Seipin and its interacting partners can also regulate these events to promote LD biogenesis (Sui et al., 2018; Walther et al., 2023). Seipin oligomerizes into a ring-like structure at ER subdomains containing nascent LDs and facilitates neutral lipid packaging for LD growth (Wang et al., 2016; Sui et al., 2018; Yan et al., 2018; Cao et al., 2019; Zoni et al., 2021). Seipin is regulated by Lipid Droplet Assembly Factor 1 (LDAF1), where LDAF1 promotes Seipin binding to TAG and induces membrane bending, a crucial step in formation and efficiency of LD biogenesis (Chung et al., 2019).

Lipid droplet biogenesis.
While LDs are still in contact with the ER, protein and lipids are exchanged between the ER and LDs to promote LD growth. Acetyltransferases, such as GPAT4, DGAT2, and AGPAT3, are recruited to the LD surface through ER-LD membrane bridges, where they facilitate localized neutral lipid synthesis (Wilfling et al., 2013). Interestingly, LDs can still form and grow even without LD-localized lipid synthesis, as enzymes such as DGAT1 can effectively target neutral lipids to LDs through ER-LD membrane bridges (Nguyen et al., 2017).
ER-LD Contacts Regulate Lipid Droplet Metabolism
ER-LD interactions exhibit two primary morphological features: (i) ER-LD membrane bridges that are a continuum of the LD monolayer with the ER bilayer and formed during LD biogenesis, and (ii) ER-LD membrane contact sites between released LDs and the ER, where the membranes are connected by tether proteins and enriched with functional and regulatory proteins (Hugenroth and Bohnert, 2020) (Figure 2). Tether proteins can extend from one organelle membrane to the other or consist of two interacting proteins present on adjacent organelle membranes. For example, ER-integral membrane proteins Vesicle-Associated Membrane Protein-Associated Proteins (VAPs) interact with FFAT motif (two phenylalanines in an acidic tract)-containing proteins, such as ORP2 and VPS13A/C, on the opposing organelle membrane through the VAP Major Sperm Protein (MSP) domain. Functional proteins such as LTPs, ion channels, and metabolite transporters also function at the membrane contact sites. Regulator proteins can modulate the contact function by changing the redox or phosphorylation state of membrane contact site proteins. These different categories of proteins are not mutually exclusive; for example, ORP5 (discussed below) acts as both tether and LTP (Scorrano et al., 2019).

Protein participants in ER-LD and MAM-LD membrane contact sites.
Multiple tethers or tethering protein complexes have been identified at LD-ER contact sites. The Sorting ER protein nexin 14 (Snx14) enriches at ER microdomains containing the fatty acyl-CoA ligase ACSL3 (Figure 2A). At these sites, Snx14 promotes LD maturation by stabilizing ER-LD membrane bridges by anchoring to the ER membrane through transmembrane helices while also binding to LDs via an C-terminal amphipathic helix. Importantly, Snx14 promotes LD maturation independent of Seipin (Datta et al., 2019). The Rab GTPase Rab18 establishes ER-LD membrane contact sites through the NAG-RINT1-ZW10 (NRZ) tethering complex and its associated SNARE proteins (Syntaxin18, Use1, BNIP1) (Xu et al., 2018). Rab18-NRZ/SNARE-mediated ER-LD contacts facilitate transfer of ER-synthesized lipids to LDs and promote LD growth (Ozeki et al., 2005; Xu et al., 2018). Another ER-resident protein, Double FYVE-Containing Protein 1 (DFCP1) also interacts with Rab18-ZW10 proteins on LDs and mediates ER-LD contacts. The Rab18-DFCP1-ZW10 complex modulates the formation and stability of ER-LD membrane contact sites to promote LD biogenesis (Li et al., 2019). Motile Sperm Domain-containing Protein 2 (MOSPD2) is a member of the VAP family of proteins involved in ER-LD membrane contact sites. Unlike other VAPs, MOSPD2 has an additional cytoplasmic domain CRAL-TRIO (Cellular Retinaldehyde-Binding Protein (CRALBP) and Triple Functional Domain Protein (TRIO)) at its amino-terminus. MOSPD2 mediates ER-LD contacts through the CRAL-TRIO domain (Zouiouich et al., 2022). While the lack of MOSPD2 shows smaller LDs, whether MOSPD2 has the lipid transport ability at ER-LD interface is not known (Zouiouich et al., 2022).
The LTPs of the oxysterol-binding protein-related (ORP) family - including ORP2, ORP5, and ORP8 - play critical roles at LD-ER MCS (Figure 2B). These three ORPs have a lipid-binding domain known as the OSBP-related domain (ORD). ORP2 is primarily cytosolic, while ORP5 and ORP8 are anchored to the ER by a transmembrane domain (Laitinen et al., 2002; Chung et al., 2015). ORP2 is a cholesterol transporter involved in movement of cholesterol from the plasma membrane to the ER and LDs (Jansen et al., 2011). In addition, LD-localized ORP2 can facilitate ER-LD membrane contact sites, where the ORP2 FFAT motif binds VAP proteins in the ER (Olkkonen et al., 2019). LD-associated ORP2 regulates SE and TAG synthesis, and promotes LD lipolysis and turnover by interacting with COPI Coat Complex Subunit Beta 1 (COPB1) to recruit the Adipose Tissue Triglyceride Lipase (ATGL) to LDs (Wang et al., 2020). It is not clear whether ORP2 interacts with any specific lipids or influences transfer of lipids at ER-LD membrane contact sites. However, sterol binding by ORP2 abrogates ORP2 localization to LDs and increases its association with the plasma membrane (Hynynen et al., 2009).
ORP5 and ORP8 have 80% sequence similarity and were first established as proteins at PM-ER and mitochondria-ER contact sites (Galmes et al., 2016; Ghai et al., 2017). However, ORP5 has a clearer role in ER-LD contacts. Upon oleate treatment, ORP5 enriches at ER-LD contacts and encircles LDs (Du et al., 2020; Guyard et al., 2022) (Figure 2C). The ORP5 ORD interacts with LDs and transfers phosphatidylinositol-4-phosphate (PI4P) and phosphatidylserine (PS) between the ER and LD by a counter-transport mechanism (Du et al., 2020). On the other hand, ORP8 only localizes to a subset of ER-LD contacts, where the Mitochondria-Associated ER Membrane (MAM) associates with LDs (MAM-LD contacts). ORP8 localization to MAM-LD contacts depends on the presence of ORP5 (Galmes et al., 2016; Guyard et al., 2022) (Figure 2C). While the function of ORP8 at MAM-LD contact sites is not clear, knockdown experiments suggest that ORP5 and ORP8 each promote LD biosynthesis and enrich Seipin at MAM-LD contacts (Du et al., 2020; Guyard et al., 2022).
Vacuolar Protein Sorting-associated 13 (VPS13) protein family members VPS13A and VPS13C are involved in mediating ER-LD membrane contact sites in addition to ER-mitochondria and ER-endosome contact sites (Kumar et al., 2018). VPS13A/C localizes to LDs through their amphipathic helix and interact with ER resident VAPs using FFAT motifs to establish ER-LD membrane contact sites (Kumar et al., 2018) (Figure 2B). The N-terminal region of the VPS13 family contains a well-conserved repeating β-groove (RBG) motif, which is a hydrophobic channel that binds and transfers glycerolipids between the membranes (Kumar et al., 2018).
In summary, ER-LD membrane contact sites are maintained by a diverse set of proteins which regulate cellular LD metabolism. Further, membrane contact site protein localization and function is influenced by the nature and metabolic state of the cells. Therefore, manipulating one or multiple ER-LD membrane contact site proteins can have a significant effect on LD metabolism, including disrupting lipid trafficking to or from LDs, altering LD turnover, and LD composition.
Pathogens Hijack Host ER-LD Membrane Contact Sites
LDs harbor innate immune complexes and contribute to antimicrobial defense. Acting as platforms for pro-inflammatory cytokine and type-I interferon production, LDs help establish an antiviral state and support immune cells, such as macrophages and lymphocytes, in pathogen recognition and response (Coulombe et al., 2014; Bosch et al., 2020; Castoldi et al., 2020; Jarc and Petan, 2020; Monson et al., 2021). In macrophages, LDs enhance phagocytic capacity by regulating inflammatory gene expression and enabling the release of bactericidal histones (Anand et al., 2012). In adaptive immunity, LDs influence T-cell activation, proliferation, and cytokine release, though these roles remain less thoroughly investigated (Berod et al., 2014; Howie et al., 2017; Schmidt et al., 2021). Additionally, LDs affect macrophage polarization and modulate pro- and anti-inflammatory states through fatty acid metabolism and mTOR signaling pathways (Wu et al., 2019). LDs also serve as reservoirs for lipophilic antibiotics, thereby enhancing drug efficacy within macrophages (Sandoz et al., 2014; Greenwood et al., 2019).
Despite LDs strong contribution to antimicrobial defense, intracellular pathogens have developed mechanisms to exploit host LDs to meet their lipid requirements and evade host immune responses. For viral pathogens, LDs can serve as the site of replication or assembly as well as a lipid source for the viral replication cycle. Bacterial pathogens, both those that inhabit the cytoplasm as well as membrane-bound vacuoles, target LDs to influence the host immune response and direct cellular lipid flow. To date, there is little known about mechanisms behind pathogen-driven manipulation of LDs, despite clear evidence that pathogens alter LD metabolism. The role of LD metabolism during viral and bacterial infection has been discussed in detail in recent review articles (Herrera-Moro Huitron et al., 2023; Husler et al., 2023; Tan et al., 2024). Hence, we limit our focus to two pathogens, SARS-CoV-2 and Coxiella burnetii, which specifically manipulate ER-LD membrane contact sites as a mechanism to alter host LD homeostasis.
SARS-CoV-2
SARS-CoV-2 induces LD formation in infected monocytes and upregulates key genes involved in lipid metabolism, including CD36, PPAR-γ, Sterol regulatory element-binding protein-1 (SREBP-1), and DGAT1 (Dias et al., 2020). The SARS-CoV-2 accessory protein ORF3a is both necessary and sufficient to induce LD accumulation by inhibiting autophagic flux (Wang et al., 2023). To fulfill viral lipid requirements, SARS-CoV-2 relies on de novo fatty acid and TAG synthesis as well as LD lipolysis (Farley et al., 2022). The SARS-CoV-2 nucleocapsid (N) protein upregulates DGAT1/2 expression, leading to LD biogenesis. Additionally, N protein interacts with ADRP on the LD surface and facilitates viral replication (Yuan et al., 2021; Ricciardi et al., 2022). Collectively, these findings suggest that LDs play a pivotal role in SARS-CoV-2 replication. Recent studies have shed light on the molecular mechanisms by which SARS-CoV-2 modulates host LDs by targeting ER-LD membrane contact sites. These mechanisms will be explored in detail in the following sections.
SARS-CoV-2 Utilizes ER-LD Membrane Contact Sites for Development of the Viral Replication Organelle
SARS-CoV-2 replicates within specialized double membrane vesicles (DMVs) inside host cells in a process which requires host lipids (Ricciardi et al., 2022). The viral nonstructural proteins NSP3 and NSP4 are responsible for creating DMVs, while NSP6 plays a crucial role in linking DMVs to LDs using ER membrane connectors (Figure 3A). NSP6 homodimerization and the amphipathic helix generates zippered regions of ER membranes known as ER membrane connectors (Ricciardi et al., 2022). ER membrane connectors lack luminal space, which promotes lipid exchanges while restricting ER luminal proteins from entering DMVs. NSP6 at ER membrane connectors also interacts with the LD-tethering complex DFCP1-Rab18 to establish DMV-LD contacts. The DMV-LD contacts, through ER membrane connectors, establish a supply of LD-derived lipids which are required for viral replication. Proper formation of NSP6-mediated ER membrane connectors and functional LD contacts are necessary for the virus to replicate effectively within the DMVs (Ricciardi et al., 2022).
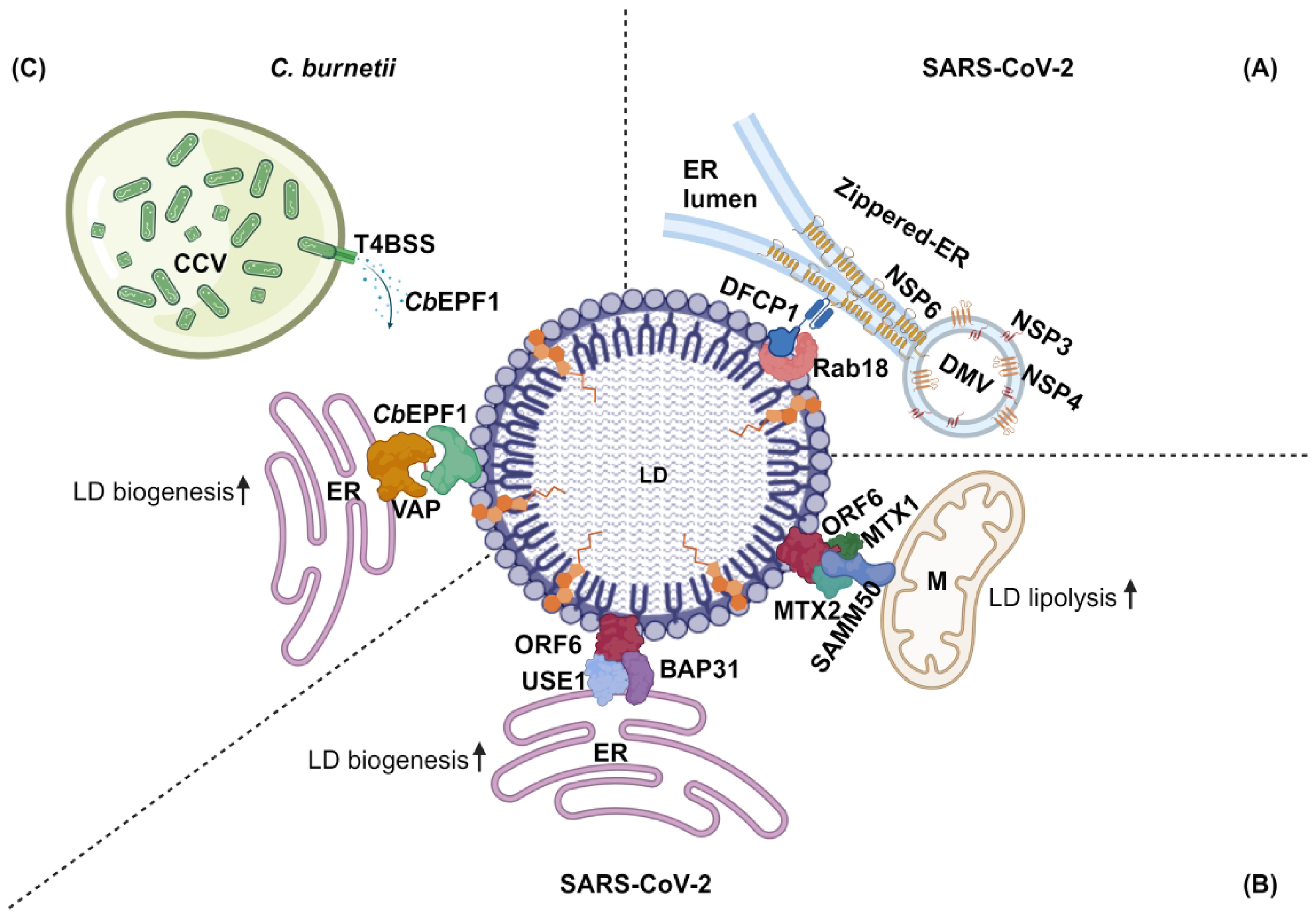
Pathogen proteins induce ER-LD and mitochondria-LD membrane contact sites to manipulate host LD metabolism.
SARS-CoV-2 ORF6 Mediates ER-LD and LD-Mitochondria Membrane Contact Sites
Besides exploiting the host cell ER-LD membrane contact site proteins DFCP1-Rab18, SARS-CoV-2 deploys its own protein, ORF6, to mediate ER-LD and LD-mitochondria contact sites (Yue et al., 2023) (Figure 3B). The SARS-CoV-2 genome encodes 29 proteins, where ORF6 and ORF9c increase cellular TAGs, ceramide, and polyunsaturated fatty acids (Farley et al., 2022). However, only ORF6 localizes to LDs (Yue et al., 2023). ORF6 homologs in other coronaviruses, including SARS-CoV, bat SARS-CoV, and bat coronavirus, also localize to LDs, suggesting a conserved function for ORF6 among SARS coronaviruses (Yue et al., 2023).
ORF6-mediated ER-LD contacts promote LD biogenesis in SARS-CoV-2 infected cells, where ORF6 localizes to LDs through two amphipathic helix (AH) domains (Yue et al., 2023). Once on the LD surface, ORF6 interacts with the host cell ER proteins B-cell receptor-associated protein 31 (BAP31) and Unconventional SNARE in the ER 1 (USE1) to form ER-LD contacts (Yue et al., 2023). B-cell receptor-associated protein 31 (BAP31) is an ER transmembrane protein that forms ER-mitochondria contacts through interactions with Tom40, a protein that serves as the core component of the translocase of the outer mitochondrial membrane (TOM complex). BAP31 regulates both lipolysis and LD biogenesis in adipose tissue (Wei et al., 2023), while USE1 forms LD-ER contacts as part of an ER SNARE complex that interacts with Rab18 on LDs (Xu et al., 2018). Though ORF6 mediated ER-LD contacts require both BAP31 and USE1, LD number and viral titer only decrease after double knockdown of BAP31 or USE1, suggesting SARS-CoV-2 may utilize redundant mechanisms for ER-LD contacts (Yue et al., 2023).
On the LD surface, ORF6 also interacts with and enhances ATGL binding to its coactivator CGI58. ORF6 stabilization of ATGL-CGI58 interaction promotes LD lipolysis (Yue et al., 2023). In addition, ORF6 interacts with mitochondrial outer membrane SAM complex to generate LD-mitochondrial contacts (Yue et al., 2023). ORF6 mediated LD-mitochondrial contacts promote fatty acid transfer from LDs to mitochondria for β-oxidation and ATP generation required for SARS-CoV-2 replication. By mediating membrane contact sites between multiple organelles, ORF6 is crucial for both LD formation and lipolysis (Yue et al., 2023). SARS-CoV-2 ORF6 is an example of how viral proteins evolve to manipulate more than one inter-organelle contact for their proliferation.
Coxiella burnetii
Coxiella burnetii, which replicates in a specialized vacuole known as the Coxiella Containing Vacuole (CCV), has been observed in foamy macrophages of Q fever endocarditis patients (Brouqui et al., 1994; Madariaga et al., 2004). In vitro studies further confirmed that C. burnetii induces LD formation in macrophages through the action of effector proteins secreted by the C. burnetii Type IVB Secretion System (T4BSS) (Mulye et al., 2018). LDs have also been detected within the CCV during infection in human alveolar macrophage cells (Graham et al., 2013). Interestingly, while blocking de novo lipid synthesis involved in LD biogenesis significantly increases C. burnetii growth, inhibiting LD catabolism decreases C. burnetii growth (Mulye et al., 2018). This suggests that fatty acids and/or cholesterol derived from LD breakdown could be required for C. burnetii growth. While C. burnetii manipulates host LDs, the underlying mechanism(s) of how C. burnetii manipulates host LDs, as well as the role of LD-derived fatty acids or sterols, is not known.
C. burnetii Effector Protein CbEPF1 Induces ER-LD Membrane Contact Sites
To understand how C. burnetii manipulates host LDs, a recent study identified a novel T4BSS effector protein, CbEPF1, that influences host LD metabolism (Angara et al., 2024). CbEPF1 localizes to LD biogenesis sites on the ER as well as the LD surface. CbEPF1 enrichment at LD biogenesis sites suggests possible interactions between CbEPF1 and host proteins which regulate LD biogenesis. In support of this, CbEPF1 expression increases the number of LDs in host cells. However, how CbEPF1 is recruited to LD biogenesis sites and whether CbEPF1 interacts with any of the host proteins involved in LD biogenesis needs further investigation. CbEPF1 contains two signals involved in LD localization: a hydrophobic domain in the middle of the protein and an amphipathic helix at the C-terminus. While CbEPF1 lacks sequence homology with other proteins, it shares functional resemblance to eukaryotic proteins that relocate from host ER to the LD surface. This includes GPAT4, DGAT2, DFCP1, LDAF1, and Rab18, which are involved in LD biogenesis and translocate from the ER to LDs (McFie et al., 2011; Wilfling et al., 2013; Xu et al., 2018; Chung et al., 2019; Li et al., 2019). While sequence analysis does not identify any conserved enzymatic or regulatory domains in CbEPF1, further structural and biochemical investigations may reveal how CbEPF1 influences host LD biogenesis.
On the LD surface, CbEPF1 interacts with ER-associated VAPs (VAPA, VAPB, and MOSPD2) and mediates ER-LD membrane contact sites (Angara et al., 2024) (Figure 3C). CbEPF1 contains two FFAT motifs, and at least one functional FFAT motif is required and sufficient for VAP interaction. Interestingly, the two CbEPF1 FFAT motifs show nonredundant binding with VAPs. While both FFAT motifs can bind VAPA and VAPB, only the second FFAT motif binds MOSPD2. The presence of two FFAT motifs indicates the importance of interactions between CbEPF1 and VAP proteins, while the preferential interaction of the two FFAT motifs with different VAPs suggest complex events behind evolutionary selection. Further, MOSPD2 is the only VAP known to mediate ER-LD membrane contact sites and influence LD metabolism (Zouiouich et al., 2022). Interesting future directions include investigating CbEPF1 structural features and whether CbEPF1 interaction with VAPA/B and MOSPD2 differentially influences LD metabolism. The presence of two FFAT motifs in a bacterial protein was previously observed in the Chlamydia trachomatis protein IncV, which is secreted by the bacteria and inserted into the Chlamydial inclusion (Murray et al., 2017). The IncV FFAT motifs interact with VAPA/VAPB in the ER using its FFAT motifs and establishes membrane contact sites between the Chlamydial inclusion and ER. Interestingly, one of the IncV FFAT motifs is regulated by phosphorylation (Ende et al., 2022). However, whether either of the two FFAT motifs interacts with MOSPD2 is unknown. Considering the nonredundant nature of FFAT motifs in CbEPF1, it is possible that the IncV FFAT motifs show preferential interaction with VAP proteins.
Oleate treatment of HeLa cells expressing CbEPF1 leads to larger LDs compared to the control cells in a process that requires at least one functional FFAT motif. This suggests that CbEPF1 generates larger LDs through ER-LD contacts, perhaps through lipid transfer activity. It is also possible that CbEPF1 recruits LTPs to the site of ER-LD contacts. A CbEPF1-FFAT double mutant that lacks both functional FFAT motifs not only loses ER-LD contacts but leads to significantly smaller and clustered LDs. While the mechanism behind CbEPF1-induced LD clustering in unknown, it could be due to recruitment of host proteins Rab40c, DFCP1, AUP1, and CG9186 to the LDs or depletion of LD associated proteins like Atg2A, Atg2B, and Seipin or through changes in LD surface lipids such as phosphatidic acid, phosphotidylcholine and squalene (Szymanski et al., 2007; Fei et al., 2011; Krahmer et al., 2011; Ta et al., 2012; Velikkakath et al., 2012; Lohmann et al., 2013; Tan et al., 2013; Thiel et al., 2013; Li et al., 2019; Salo et al., 2019). Therefore, the CbEPF1-FFAT double mutant localized LDs that lack ER-LD membrane contact sites may show changes in LD proteome and/or lipidome that influence LD clustering.
CbEPF1-induced LD accumulation could be a strategy to reduce toxic cholesterol in the CCV membrane. While the CCV is rich in sterols, excess CCV cholesterol causes CCV hyperacidification and bacterial degradation (Mulye et al., 2017). Consequently, C. burnetii has evolved multiple molecular mechanisms to regulate CCV cholesterol (Justis et al., 2017; Clemente et al., 2022; Schuler et al., 2023). One notable mechanism involves recruitment of the host sterol transporter ORP1L to the CCV membrane (Justis et al., 2017). ORP1L on the CCV membrane interacts with VAPs in the ER to mediate ER-CCV membrane contact sites and facilitate cholesterol efflux from the CCV to the ER (Justis et al., 2017; Schuler et al., 2023). In this context, CbEPF1 mediated ER-LD contacts and larger LDs supports an active mechanism by C. burnetii to redirect cholesterol away from the CCV to the ER, where it is then stored in LDs. Future studies using C. burnetii CbEPF1 mutants will help elucidate the significance of cholesterol and LDs during Coxiella pathogenesis.
Conclusion and Future Directions
While many pathogens induce LD accumulation within host cells and utilize them for energy or structural needs, whether these pathogens encode proteins to directly manipulate lipid transfer to or from LDs is largely unknown. Studies discussed in this review reveal that intracellular pathogens encode proteins capable of mimicking host tether proteins to induce ER-LD membrane contact sites, thereby influencing lipid trafficking and LD metabolism. However, it remains unclear whether the pathogen proteins function as LTPs. Given the largely unexplored secretome of obligate intracellular bacteria or proteome of viruses, these discoveries open new avenues for research into pathogen-induced inter-organelle contact sites. It is likely that additional effector proteins mediate membrane contact sites at other organelles beyond the ER-LD and LD-mitochondrial interface, further influencing host lipid metabolism to benefit pathogen survival and replication. Future studies aimed at understanding how disruptions in inter-organelle communication contribute to pathogenesis could offer new therapeutic opportunities for targeting these infections.
Footnotes
Acknowledgements
We thank members of the Gilk Lab for feedback and helpful discussions.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by NIH grants AI173990 and AI139176 (S.D.G).
