Abstract
Phosphatidylinositol 4-monophosphate (PtdIns(4)P) is one of the key membrane components which mark the membrane contact sites. In the mammalian Golgi complex, PtdIns(4)P is produced at various subregions via specific mechanisms for each site. Particularly, PtdIns(4)P pools generated at the distal Golgi regions are pivotal for the determination of membrane contacts between the endoplasmic reticulum (ER) and Golgi, at which inter-organelle lipid transport takes place. In this short review, we will focus on C10orf76 (or ARMH3), which we propose to rename as DGARM after a
Introduction
Eukaryotic cells have various membrane-bound compartments called organelles. In addition, each organelle seemingly has various sub-regions (or zones) with different functions (Sasaki and Yoshida, 2019; Nishitoh, 2019). Membrane contact sites are the subcellular regions consisting of two different organelle membranes within ∼30 nm distance and the cytosol fraction between them. Although organelle contacts have been recognized early in 1960s in cell images by classic electron microscopy, their functions have been clarified since 2000s. As a pivotal function of membrane contact sites, inter-organelle transport of lipids by various lipid transfer proteins (LTPs) has been extensively studied in this decade; providing mechanistic insights into the long enigmas of how cell properly accomplishes intracellular trafficking of lipids or of how a specific lipid type is picked up from a specific organelle and then delivered to another specific organelle in a cell (reviewed in Hanada, 2018; Balla et al., 2019; Wong et al., 2019). Phosphatidylinositol 4-monophosphate (PtdIns(4)P) is a well-studied membrane component for its role in specifying various contact sites. We recently found that C10orf76 (alias ARMH3), a PtdIns 4-kinase-binding protein, is recruited to the distal Golgi and plays a pivotal role in the ER-to-distal Golgi trafficking of ceramide for the synthesis of sphingomyelin (Mizuike et al., 2023). Together with the predicted structure of C10orf76 as an armadillo (ARM)-like fold (Figure 1), consisting of α-helices arranged into a characteristic superhelix (Tewari et al., 2010), we proposed to rename it to DGARM after a

DGARM, a predicted armadillo-fold protein. A predicted structure of DGARM by alphafold (Jumper et al., 2021; Varadi et al., 2022; ver. 2022-11-01). ARM-type fold region (InterPro IPR016024) and regions involved in PI4KB-binding (McPhail et al., 2020) are colored green and magenta, respectively. DUF1741; a domain of unknown function, conserved only in primary sequences of DGARM orthologs. DUF1741 is colored cyan, but the colors of ARM-type fold or PI4KB-binding regions are prioritized.
The Architecture of the Golgi complex
The Golgi complex is an organelle with a distinctive structure (Figure 2A and B): multiple cisternae stacking in a polarized manner. Golgi cisternae can be classified as cis, medial, or trans, with the cis side being proximal (or early) in the anterograde membrane traffic starting from the endoplasmic reticulum (ER), and the trans side being distal (or late). Additionally, the ER-Golgi intermediate compartment (ERGIC) and trans-Golgi network (TGN) are also thought to be subregions of the Golgi complex (Pantazopoulou and Glick, 2019). Although the Golgi apparatus in several fungi, including the model yeast Saccharomyces cerevisiae, do not form a stacking structure under conventional growth conditions, the punctate cisternae are classified into cis-, medial-, and trans-Golgi (Suda and Nakano, 2012). The Golgi complex is regarded to be compartmentalized in two ways, separation of cisternae and intra-cisterna segregation (Tie et al., 2018); thereby allowing accurate sequential processing/modification of proteins and lipids. Particular lipids show heterogeneous distribution among the cisternae: for instance, sphingomyelin, cholesterol and PtdIns(4)P levels are higher at the distal Golgi than the proximal (Moreau and Cassagne, 1994; D’Angelo et al., 2008; Del Bel and Brill, 2018). These distribution gradients of lipids are generated by local lipid transport coupled with lipid metabolism.

Distal Golgi–ER contact enables bypass of the proximal Golgi. (A) The intracellular distribution of ceramide-metabolizing enzymes in eukaryotes. For de novo ceramide synthesis machinery, mammalian enzymes are shown as a representative. (B) Golgi–PtdIns(4)P is generated by PI4KB or PI4K2A. While PI4K2A is palmitoylated at the TGN in a cholesterol-dependent manner for membrane association, PI4KB utilizes multiple recruitment systems to localize at various regions within the Golgi complex. PtdIns(4)P-phosphatase Sac1 cycles between the ER and the proximal Golgi, keeping the proximal PtdIns(4)P levels low. ER-resident Sac1 also catalyzes PtdIns(4)P turnover at the ER-TGN contacts. (Lower) Chemical structures of lipids discussed in this review. Only the polar headgroups are shown (enclosed in cyan boxes) for complex sphingolipids. Note that the major ceramide backbone in plants and fungi is phytoceramide, which has a hydroxyl group at the C4 position. SPT, serine-palmitoyltransferase; KDSR, 3-ketodihydrosphingosine reductase; CERS, ceramide synthase; DES, sphingolipid delta(4)-desaturase; IPCS, inositol phosphorylceramide synthase in fungi and plants; GCS, glucosylceramide synthase in eukaryotes; CPES, (insect-specific) CPE synthase; SMS, sphingomyelin synthase in vertebrates.
PtdIns(4)P as a Determinant of the Lipid Transport Zones at the Distal Golgi
In the de novo synthetic pathway of sphingolipids, ceramide is synthesized in the ER and then transported to the Golgi complex for further conversion to obtain complex sphingolipids (Figure 2A). The major phosphosphingolipids, which we focus on in this review, varied in evolution (Figure 2A): sphingomyelin in vertebrates, ceramide phosphoethanolamine (CPE) in invertebrates, and phosphoinositol-containing sphingolipids in fungi and plants, while some invertebrates synthesize sphingomyelin (Panevska et al., 2019). Whereas ceramide trafficking for inositol phosphorylceramide (IPC) synthesis in the budding yeast S. cerevisiae, depends both on vesicular transport and LTPs Nvj2 and Svf1 (Kajiwara et al., 2014; Liu et al., 2017; Limar et al., 2023), the synthesis of sphingomyelin and CPE highly depends on an LTP, the ceramide transport protein CERT. CERT and oxysterol-binding protein (OSBP) are representative LTPs working at the Golgi complex for ER-to-Golgi transport of ceramide and cholesterol, respectively (Hanada et al., 2003; Mesmin et al., 2013). These LTPs share similar domain construction: a pleckstrin homology (PH) domain which binds PtdIns(4)P, a “two phenylalanines in an acidic tract” (FFAT) motif that binds ER-resident protein VAPs, and a lipid transfer domain (i.e., a steroidogenic acute regulatory protein-related lipid transfer [START] domain in CERT and an OSBP-related domain [ORD] in OSBP; Goto et al., 2020). Dual targeting via interactions between PH–PtdIns(4)P and FFAT–VAP enables these LTPs to localize at the ER–Golgi contacts. A previous study suggested that OSBP displays only a weak or indirect interaction with CERT in CHO cells (Perry and Ridgway, 2006), and a recent study showed that TMED2/10 form a supercomplex with OSBP, CERT, VAP-A in a PtdIns(4)P-, and ARF1-dependent manner in HeLa cells (Anwar et al., 2022). Thus, a partial, even if not all, population of Golgi-bound CERT, may act with OSBP at the same contact sites.
Reconstruction of the 3D Golgi structure from electron microscope tomography showed that specific regions of the ER adhere with the trans-cisternae or TGN (Mogelsvang et al., 2004; Venditti et al., 2019), suggesting that the ER–Golgi contacts are mainly formed at the distal Golgi in mammalian cells. Within the Golgi complex, the site for de novo synthesis of sphingomyelin is more distal than the site for the synthesis of glucosylceramide (Halter et al., 2007; Huitema et al., 2004; D’Angelo et al., 2007; Hayashi et al., 2018). The enrichment of PtdIns(4)P at the distal Golgi leads CERT and OSBP to preferentially localize at this region. This enables an efficient influx of ceramide to the site of sphingomyelin synthesis, which should provide a rationale for the intra-Golgi gradient of sphingomyelin. However, it is not clear whether PtdIns(4)P is produced at contact sites or PtdIns(4)P production induces formation of organelle contacts. Meanwhile, both could happen; PtdIns(4)P could be necessary for contacts to form but might also be needed to be produced more to maintain contacts. Note that the ability of OSBP to counter-transport cholesterol and PtdIns(4)P confers PtdIns(4)P to serve as the “fuel” to drive the ER-to-Golgi uphill movement of cholesterol by coupling with the Golgi-to-ER downhill movement of PtdIns(4)P (de Saint-Jean et al., 2011; Mesmin et al., 2013).
PtdIns(4)P synthesis in the Golgi complex principally depends on two PtdIns 4-kinases, PI4KB and PI4K2A (also known as PI4KIIIβ and PI4KIIα, respectively; Figure 2B). PI4K2A is palmitoylated at the TGN in a cholesterol-dependent manner and is regarded to be localized at cholesterol-rich membrane domains in the TGN and endosomes (Balla et al., 2002; Lu et al., 2012). On the other hand, PI4KB is mostly dispersed in the cytosol and only a portion is recruited to the Golgi membrane (Blomen et al., 2015; McPhail et al., 2020). PI4KB adopts several types of Golgi-localization machinery in mammalian cells; that is, via interaction with ACBD3, DGARM (or C10orf76, ARMH3), or GGA2 (Sasaki et al., 2012; Mizuike et al., 2023; Daboussi et al., 2017). This allows PI4KB to localize at various regions within the Golgi complex (Figure 2B): since ACDB3 predominantly localize at the proximal regions (Tie et al., 2018; Mizuike et al., 2023), DGARM at the distal regions (Mizuike et al., 2023), and GGA2 at the TGN (Doray et al., 2002). Although further observation of PI4KB using a super-resolution microscopy is demanded to prove this model. These PI4KB-binding proteins further interact with other proteins and will hence establish characteristic and functional differences in PtdIns(4)P pools. Amongst these, DGARM was shown to interact with GBF1 (Chan et al., 2019), which likely leads to the activation of PI4KB by ARF1, allowing vigorous PtdIns(4)P production at the distal Golgi.
A pool of PtdIns(4)P generated by the DGARM–PI4KB axis was shown to be important for defining the site at which CERT works to provide ceramide for sphingomyelin synthesis (Mizuike et al., 2023), as discussed above. In this study, ACBD3 was also indicated to partially compensate for this function. In addition, DGARM–PI4KB axis-dependent PtdIns(4)P production and the downstream lipid transport were suggested to be critical for the activation of the cGAS–STING pathway, a DNA-triggered innate immunity system (Fang et al., 2023). STING translocates from the ER to the Golgi upon ligand binding and is palmitoylated and phosphorylated for activation (Mukai et al., 2016; Zhang et al., 2019; Ritchie et al., 2022). STING-DGARM interaction enables the local PtdIns(4)P production for recruiting LTPs (mainly CERT and ORPs including OSBP), thereby fine-tuning the environment of distal Golgi membrane into that suitable for STING activation. In this connection, ACBD3 was reported to be involved in the upstream step of STING activation: the ER-to-cis-Golgi transport of STING (Motani et al., 2022). ACBD3 concentrates ligand-bound STING in the face of ERGIC or cis-Golgi cisternae for efficient export from the ER, although PI4KB may be dispensable in this process. Still, it is of interest that PI4KB-binding proteins work sequentially in the same pathway at different regions.
Phylogenetic Relevance of PtdIns(4)P Production Machinery at the Distal Golgi
PtdIns(4)P is an essential lipid for eukaryotic life (Dickson and Hille, 2019; Posor et al., 2022), and PI4KB orthologs are widely conserved throughout eukaryotes (Figure 3A). Concurrently, ARF1 and GBF1, involved in PI4KB activation, are also widely conserved. However, genes involved in recruiting PI4KB to the Golgi complex are not so widely conserved (Figure 3A). ACBD3 is conserved only in Holozoa while DGARM is more widely conserved across the eukaryotes including fungi, plants and protists (Figure 3A and B), suggesting that the DGARM–PI4KB axis occurred earlier than the ACBD3–PI4KB axis in the evolution of life (Chan et al., 2019).
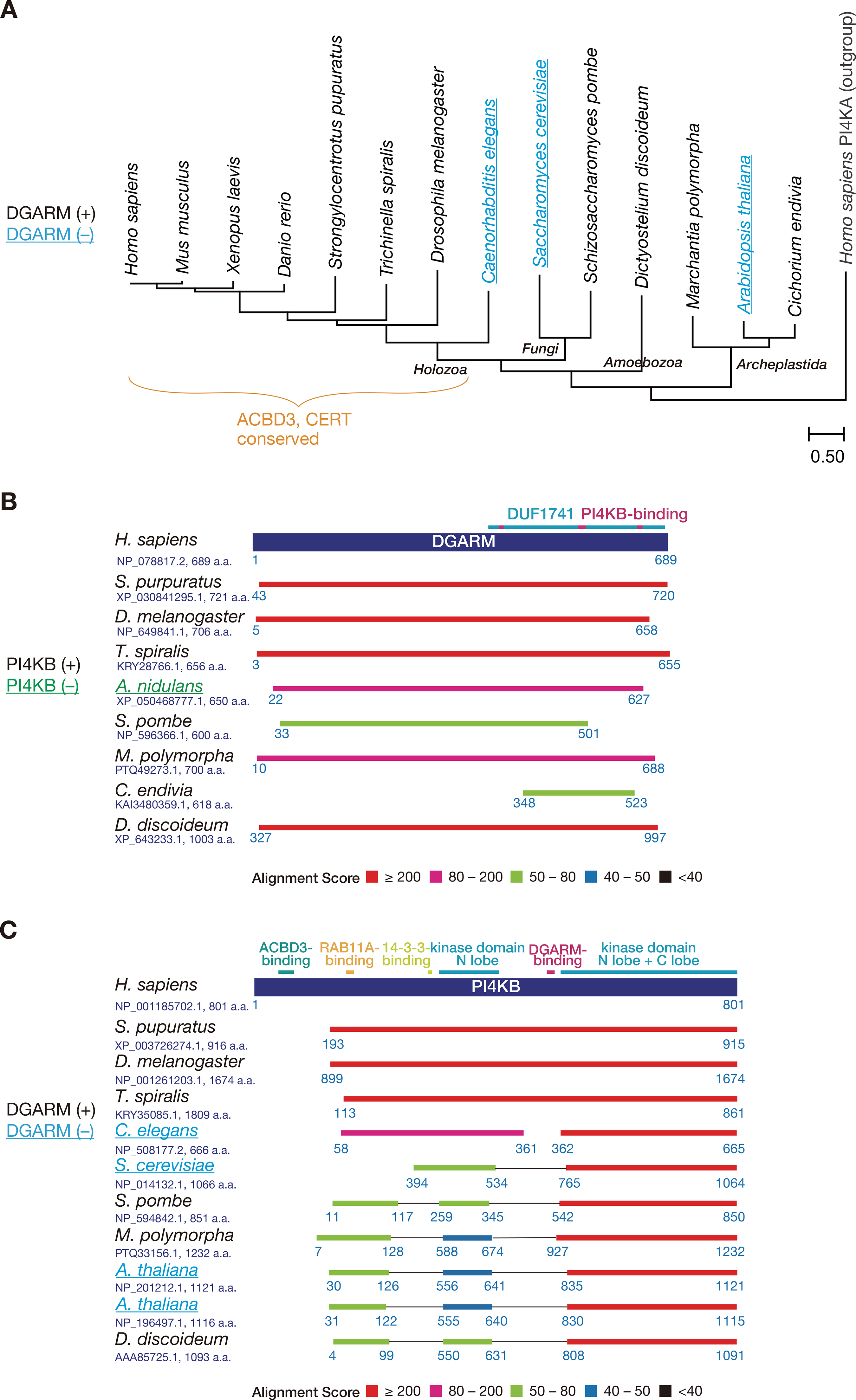
Evolutionary conservation of the DGARM–PI4KB axis. (A) The molecular phylogenetic tree of PI4KB is shown. Primary sequences of PI4KB orthologs (and human PI4KA as an outgroup) registered in UniProt were aligned by MUSCLE program and subjected to generate the phylogenetic tree by maximum likelihood method using MEGA software (ver. 11). Scale bar of the branch length represents the number of changes per sites. The analysis of DGARM phylogeny was extended from Chan et al. (2019), while PSI-BLAST was used instead. It should also be noted that, while ortholog search suggested that some plant species do conserve DGARM ortholog, most of the plant species have lost it. (B and C) Distributions of the BLAST hit sequences on human DGARM (B) and PI4KB (C) in non-vertebrates. The accession number and length (a.a.) of protein sequences are shown below the species name. The numbering represents the amino acid number. Domains of the query proteins are shown above as colored lines. Note that A. thaliana genome encodes two copies of PI4KB homolog. For ortholog search, PSI-BLAST was executed against the non-redundant protein sequences database for selected organisms with the profile-inclusion threshold of e-value set to 0.005. Species that lack DGARM or PI4KB are underlined and colored in cyan or green, respectively.
Nonetheless, DGARM was lost in several species including major model organisms Caenorhabditis elegans, S. cerevisiae, and Arabidopsis thaliana. FASTA search (https://www.genome.jp/tools/fasta/) for DGARM (seq. ID hsa:79591) suggested that DGARM is highly conserved in Holozoa, however, six of seven nematode species (registered in KEGG GENES) have lost DGARM. Notably, the PI4KB homolog in Trichinella spiralis (DGARM-positive) possesses the N-lobe kinase linker, which was suggested to contain the DGARM-binding region (McPhail et al., 2020), while that in C. elegans (DGARM-lost) does not (Figure 3C). For a detailed review of the DGARM-binding region of PI4KB, see (McPhail and Burke, 2023). It is conceivable that organisms which had lost DGARM have developed other (unique) systems for recruiting PI4KB orthologs to the distal or late Golgi regions. For instance, S. cerevisiae utilizes Frq1–Arf1 for recruiting Pik1, the PI4KB homolog in fungi, to the late Golgi (Strahl et al., 2005; Highland and Fromme, 2021). On the other hand, the Frq1 ortholog in another model yeast Schizosaccharomyces pombe, which has DGARM, is dispensable for Golgi localization of Pik1 (Willet et al., 2023). However, whether DGARM interacts with Pik1 in S. pombe (and other fungal, plant, and ameba species) needs to be analyzed, as the DGARM-binding region is not conserved in Pik1 of these species. It should also be noted that some of the filamentous fungi (e.g., Aspergillus nidulans) are unlikely to have a Pik1 ortholog, whereas they have a DGARM. Other PtdIns 4-kinase(s) could be responsible for PtdIns(4)P production at the late Golgi although it remains unclear whether DGARM orthologs contribute to the generation of Golgi PtdIns(4)P pools in these species.
One of the pivotal roles of PtdIns(4)P in the distal Golgi membrane is to determine the contact sites with the ER, as mentioned above. The ER-distal-Golgi contacts provide the cell with a new pathway for the direct delivery of lipids from the ER to the distal Golgi regions, bypassing the proximal regions. The pathway that explicitly demands to bypass the proximal Golgi is ceramide delivery for de novo synthesis of phosphosphingolipids in vertebrates and insects (Figure 2A). Sphingomyelin synthase and insect-specific CPE synthase localize at the distal Golgi (Halter et al., 2007; Vacaru et al., 2013), whereas glucosylceramide synthesis, which competes for the precursor ceramide, occurs predominantly at the proximal Golgi (Allan and Obradors, 1999; Hayashi et al., 2018; Ishibashi et al., 2018). Therefore, vertebrates and insects might require direct ceramide supply from the ER for robust phosphosphingolipid synthesis (Hanada et al., 2003; Rao et al., 2007). Notably, the evolutionary emergence of CERT is nearly coincident with that of SM and its close relatives including CPE (Hanada, 2014; Sato et al., 2019).
Perspectives
Local PtdIns(4)P production at the distal Golgi was suggested to be essential for growth and development since homozygous knockout of DGARM caused embryonic lethality in mice (Perez-Garcia et al., 2018). However, there remain unsolved questions about the regulatory mechanism of DGARM. (a) What factor(s) does DGARM recognize to focally localize at sites appropriate for lipid transport in the distal Golgi? (b) DGARM was suggested to cycle between the cytosol and Golgi membrane (Blomen et al., 2015; Chan et al., 2019), but what switches the localization of DGARM? Future studies on these issues would help us comprehend the details of the regulation system of the ER–distal Golgi contacts, which are presumed to be highly dynamic and short-lived (David et al., 2021; Subra et al., 2023). Evolutional analysis for the function of DGARM orthologs, especially in fungi and plants that do not possess CERT, would also broaden our understanding of the significance of constructing the transport pathway which bypasses the proximal Golgi.
A number of studies utilizing genome-wide knockout screens for various cellular events, including pathogen infection, have identified DGARM as a candidate (Blomen et al., 2015; Jeng et al., 2019; Yang et al., 2023, Cigler et al., 2023; Sasaki et al., 2023). Studies on the functional roles of DGARM in these cellular events would also attract interest in the pathologic and therapeutic aspects.
Footnotes
Abbreviations
Acknowledgement
We would like to deeply thank the collaborators for their invaluable contribution on the study of DGARM and CERT.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Japan Agency for Medical Research and Development, Japan Society for the Promotion of Science (grant number JP22fk0108561, JP21H02630).
