Abstract
Cryptococcus neoformans is an important fungal pathogen, responsible for over 140,000 deaths per year worldwide. Like other yeasts, C. neoformans relies on ergosterol as its major membrane sterol and carefully regulates its synthesis and distribution. Ergosterol is also targeted by two of the three compound classes currently used to treat cryptococcal infection. We recently reported the discovery and characterization in C. neoformans of a single retrograde ergosterol transporter of the LAM family, Ysp2. Here we review these findings and discuss directions for future research, including the connections between processes that are perturbed by the absence of Ysp2 (which also abrogates cryptococcal virulence) and possible roles for Ysp2 and other, as yet unknown, lipid transport proteins in this organism.
Cryptococcus neoformans is an opportunistic fungal pathogen that causes over 140,000 deaths worldwide each year, including 19% of all AIDS-related deaths. Cryptococcal meningitis has a mortality rate of 10–75% and is the most common type of meningitis in adults living with HIV in sub-Saharan Africa (Denning, 2024). Effective treatment of this infection is hindered by factors that include limited therapeutic options, adverse effects, and high costs. Given these epidemiological trends and treatment limitations, it is crucial to understand cryptococcal basic biology and pathogenesis in order to develop new therapeutic approaches.
Ergosterol is the most abundant sterol in fungal membranes. This non-polar lipid is critical for defining membrane fluidity and permeability, regulating protein sorting, and modulating the activity of membrane-associated enzymes (Rodrigues, 2018). Ergosterol is also a key therapeutic target for C. neoformans infections. Current treatment for cryptococcosis is limited to three drugs: amphotericin B (AmB), fluconazole (FLC), and flucytosine. AmB and FLC target ergosterol and its biosynthetic pathway, respectively (Iyer et al., 2021).
Ergosterol biosynthesis is a complex and highly regulated process that occurs mainly in the endoplasmic reticulum (ER). Although the biochemical reactions of ergosterol synthesis have been well-defined, less is known about the mechanisms of sterol distribution. After synthesis in the ER, most sterols are transported to other organelles, primarily the plasma membrane (PM), which contains up to 90% of cellular sterols (Sokolov et al., 2019). This movement is mainly mediated by sterol-specific lipid transport proteins, which are independent of the secretory pathway (Baumann et al., 2005). Most of our information about these proteins in yeast comes from Saccharomyces cerevisiae. This model yeast expresses two families of them: oxysterol-binding proteins (OSH) and lipid transfer proteins anchored at membrane contact sites (LAM). The seven cytosolic OSH proteins localize at membrane contact sites and move sterols to and from the ER in exchange for other lipids (Sokolov et al., 2019). The recently identified LAM proteins are anchored by transmembrane domains and possess characteristic StART (Steroidogenic Acute Regulatory Transfer)-like domains which modulate sterol binding, as well as GRAM domains, members of the pleckstrin-homology (PH) domain superfamily that are involved in sensing sterols (Gatta et al., 2015; Luo et al., 2019; Naito et al., 2019; Ercan et al., 2021). In S. cerevisiae, the LAM family is represented by six proteins, Lam1–Lam6, which occur at various membrane contact sites (Gatta et al., 2015).
We recently identified a C. neoformans retrograde sterol transporter, Ysp2, which is a LAM homolog. This protein has a similar domain organization to S. cerevisiae LAM proteins, with a single transmembrane domain, a StART-like domain, and a PH domain of the GRAM family. We showed that under growth conditions that mimic the host environment, the absence of Ysp2 leads to abnormal accumulation of ergosterol at the PM, invagination of the PM, and malformation of the cell wall, all of which may be functionally rescued by inhibiting ergosterol synthesis with the antifungal drug FLC (Choy et al., 2023). We further observed altered architecture of the cryptococcal polysaccharide capsule, mislocalization of a PM-resident protein and, interestingly, a slight increase in total sterol synthesis. Importantly, as a result of perturbed ergosterol distribution and its consequences, cells lacking Ysp2 cannot survive in environments relevant to infection (such as host phagocytes (Gaylord et al., 2020)) and are dramatically attenuated in virulence in an animal model.
The drastic consequences of disrupting retrograde sterol transport demonstrate the fine-tuning of ergosterol content and distribution that is needed for fungal cell survival, especially in the demanding context of an infected host. They also lead to several interesting new questions. For example, our experiments indicate a novel connection between distribution of ergosterol and elaboration of the capsule, which is a key virulence factor (Agustinho et al., 2018). The link between lipid distribution and underlying mechanisms of capsule synthesis and regulation could be a fruitful area of study. Our analysis of lipid species and gene expression also supports crosstalk between ergosterol distribution and synthesis, and suggests that the level of intracellular, rather than total, ergosterol regulates sterol production via the Sre1 signaling pathway (Choy et al., 2023). This is another topic that merits further investigation.
We have examined the role of Ysp2 exclusively in C. neoformans. Notably, while this organism encodes only a single LAM homolog, S. cerevisiae has six such proteins and Aspergillus fumigatus has two (Wong, 2017). It may be that cryptococcal Ysp2 has multiple functions (Figure 1) or that other sterol transport mechanisms remain to be identified in C. neoformans; these possibilities are not mutually exclusive. The first hypothesis is supported by several lines of evidence. First, we observed cryptococcal Ysp2 not only at the cell periphery, likely at PM–ER contact sites, but also intracellularly, presumably at ER contact sites with other organelles (Figure 2) (Choy et al., 2023). PM–ER contact site localization is consistent with previous work on Ysp/Lam proteins in S. cerevisiae (Gatta et al., 2015), while localization to ER-Golgi contact sites has been reported for human GRAM domain-containing proteins (Naito et al., 2023). Second, we noted changes in mitochondrial morphology upon the loss of Ysp2, suggesting it has a role in maintaining mitochondrial membranes (Choy et al., 2023). Third, a study by Wong showed that expression of the StART-like domain from cryptococcal Ysp2 rescues the AmB sensitivity of a S. cerevisiae strain lacking Lam1, Lam2, and Lam3 (Wong, 2017).

Known (solid arrow) and putative (dashed arrow) transport processes mediated by cryptococcal Ysp2.
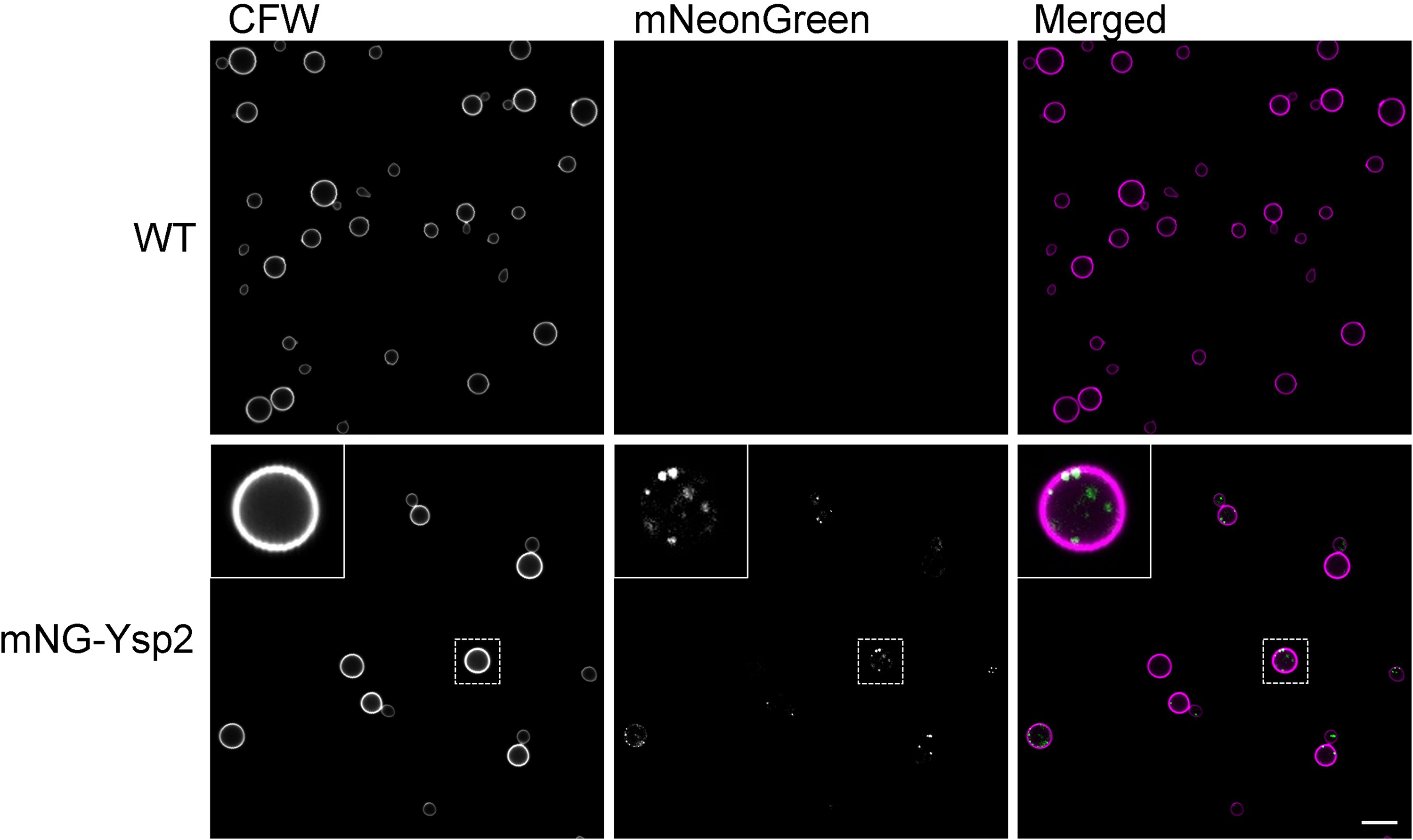

Localization of cryptococcal Ysp2. Confocal microscopy images of unmodified wild-type cells (top) or the same cells expressing mNeonGreen-tagged Ysp2 (mNG-Ysp2, bottom), stained with the cell wall-binding dye Calcofluor White (CFW) to demarcate the cell periphery. Inset, an example cell (boxed) enlarged three-fold. Bar, 10 µm. Figure modified from Choy et al. (2023).
It is also possible that other sterol transport proteins or mechanisms exist in C. neoformans (Figure 3). LAM proteins are characterized by one or more StART-like domains and fall within the StARkin superfamily. Another predicted cryptococcal protein (encoded by CNAG_02308) does not have transmembrane or membrane-targeting PH (pleckstrin homology) domains but does contain a DUF3074 domain, which is a lipid-transfer motif similar to StARkin domains (Wong, 2017). This protein did not rescue the AmB sensitivities of a S. cerevisiae strain lacking Lam1, Lam2, and Lam3, although it conferred “marginal” growth benefit on wild-type and single deletion strains (Wong, 2017). A second protein, encoded by CNAG_05588, contains a StART domain but no PH domain; its function as a potential sterol transporter has not been investigated. Future studies in this area could include in vitro experiments to determine the binding specificity of these proteins and more detailed functional studies to assess the cellular consequences when these proteins are absent. More broadly, large-scale screens may help discover novel sterol transport proteins that cannot be identified through sequence or structural homology.
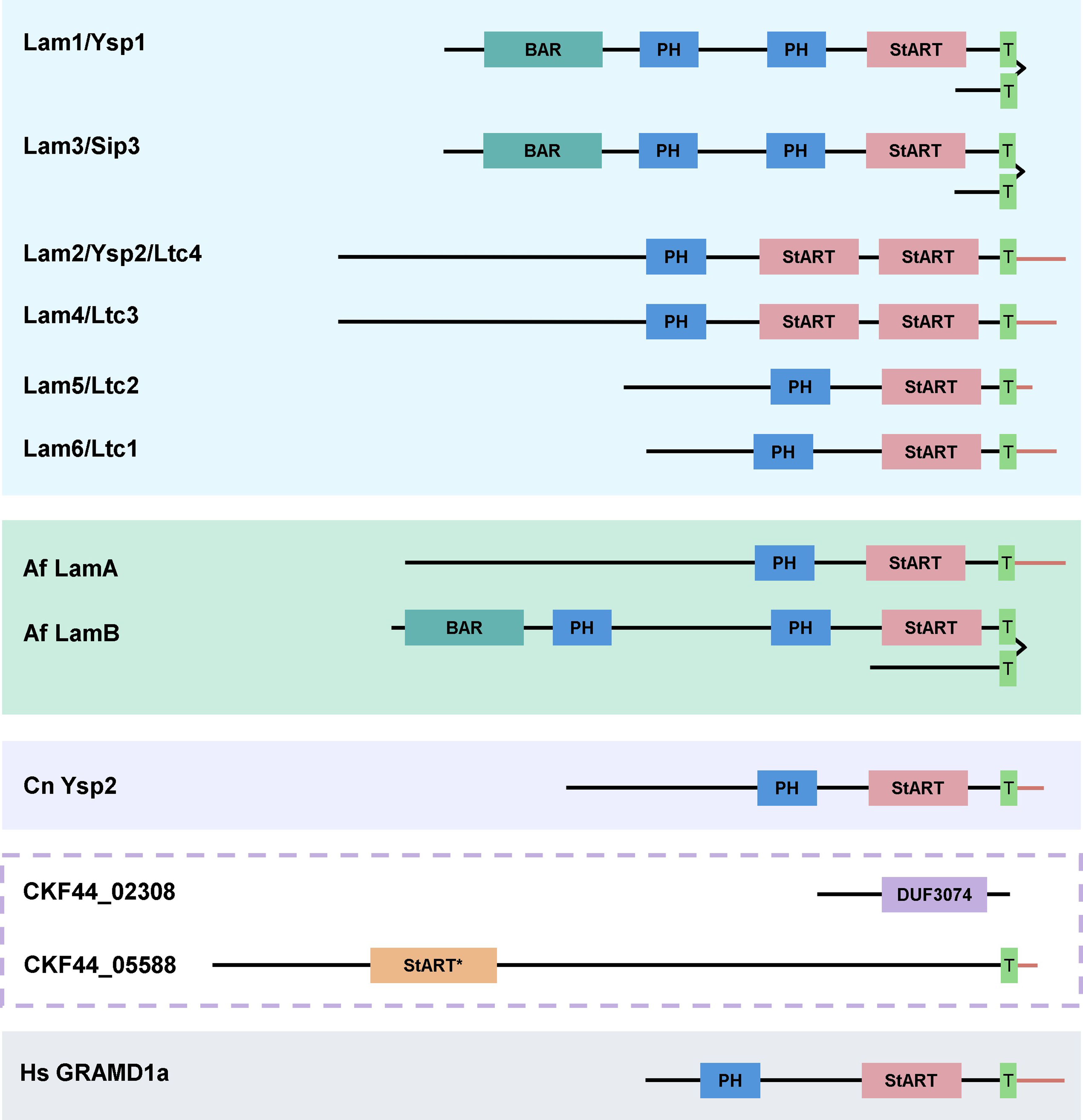
Schematic alignment of cryptococcal Ysp2 with other fungal homologs and a representative mammalian cholesterol transfer protein. Large colored rectangles demarcate LAM family proteins in S. cerevisiae (blue), A. fumigatus (green), and C. neoformans (purple); gray rectangle demarcates the Homo sapiens cholesterol transporter. A broken purple line surrounds two potential C. neoformans lipid transfer proteins discussed in the text. Domain abbreviations are BAR = Bin/amphiphysin/RVS; PH = pleckstrin-homology; StART = Steroidogenic Acute Regulatory Transfer-like; StART* = Steroidogenic Acute Regulatory Transfer, and T = transmembrane. Figure modified from Choy et al. (2023).
While this comment is limited to fungal ergosterol transport, a wealth of knowledge is also available from the study of mammalian cholesterol transporters. Lipid transfer proteins from the two kingdoms share conserved domains and domain organization (Figure 3), and further comparisons between them may yield important insights into the mechanisms of ergosterol sensing and transport in C. neoformans (Gatta et al., 2015; Luo et al., 2019; Naito et al., 2019).
We demonstrated the impact of a single LAM family sterol transporter on C. neoformans cell morphology and ultimately virulence. Our results highlight how ergosterol distribution is deeply intertwined with other factors critical to cryptococcal success, including the polysaccharide capsule and cell wall. Because of ergosterol's importance in fundamental biology and as a therapeutic target, it is imperative to define this pathogen's full range of tools for regulating sterol homeostasis, from known lipid transfer proteins to novel mechanisms.
Footnotes
Acknowledgments
We thank Liza Loza and Dong-gyu Kim for helpful discussions and comments on the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Division of Intramural Research, National Institute of General Medical Sciences (grant numbers R01 AI135012, R21 AI136688, R21 AI140979, R21 AI178330, T32 GM007067).
