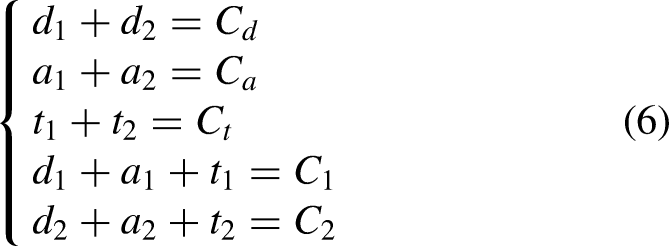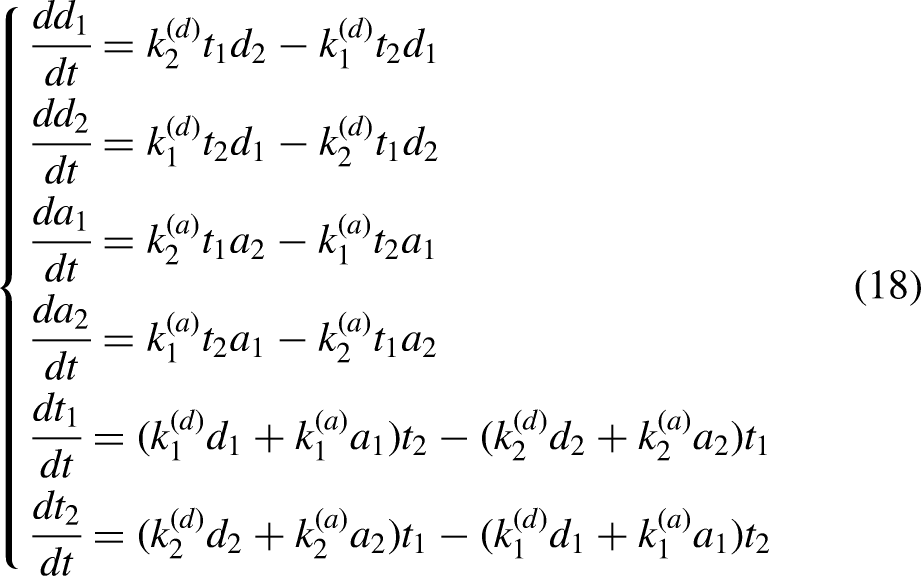Abstract
Lipid transfer proteins (LTPs) transfer lipids between different organelles, and thus play key roles in lipid homeostasis and organelle dynamics. The lipid transfer often occurs at the membrane contact sites (MCS) where two membranes are held within 10–30 nm. While most LTPs act as a shuttle to transfer lipids, recent experiments reveal a new category of eukaryotic LTPs that may serve as a bridge to transport lipids in bulk at MCSs. However, the molecular mechanisms underlying lipid transfer and MCS formation are not well understood. Here, we first review two recent studies of extended synaptotagmin (E-Syt)-mediated membrane binding and lipid transfer using novel approaches. Then we describe mathematical models to quantify the kinetics of lipid transfer by shuttle LTPs based on a lipid exchange mechanism. We find that simple lipid mixing among membranes of similar composition and/or lipid partitioning among membranes of distinct composition can explain lipid transfer against a concentration gradient widely observed for LTPs. We predict that selective transport of lipids, but not membrane proteins, by bridge LTPs leads to osmotic membrane tension by analogy to the osmotic pressure across a semipermeable membrane. A gradient of such tension and the conventional membrane tension may drive bulk lipid flow through bridge LTPs at a speed consistent with the fast membrane expansion observed in vivo. Finally, we discuss the implications of membrane tension and lipid transfer in organelle biogenesis. Overall, the quantitative models may help clarify the mechanisms of LTP-mediated MCS formation and lipid transfer.
Keywords
Introduction
Lipid transfer proteins (LTPs) specifically transfer lipids between cellular membranes more rapidly than vesicular transport. Consequently, LTPs play key roles in lipid homeostasis, organelle biogenesis, metabolite or ion exchange, and signaling between organelles (Wong et al., 2017; Wong et al., 2019; Leonzino et al., 2021; Reinisch and Prinz, 2021). For example, numerous LTPs mediate cholesterol transfer in humans. Dysfunction of LTPs has been associated with various diseases or disorders, including atherosclerosis caused by cholesterol deposition in blood vessels, obesity, and neurodegeneration (Hamilton and Deckelbaum, 2007; Potter et al., 2009; Sandhu et al., 2018; Ugur et al., 2020).
So far, hundreds of LTPs have been discovered, and more are being identified (Wong et al., 2019; Hanna et al., 2021). Most LTPs contain hydrophobic lipid-binding cavities that specifically bind one or more species of lipids. These LTPs extract lipids from one membrane, hold them in the cavities, diffuse across the cytosol, and release the lipids to another membrane, thus acting as a shuttle to transfer lipids. A representative shuttle LTP is the oxysterol-binding homology protein 4 (Osh4p) in yeast that transfers ergosterol from the endoplasmic reticulum (ER) to the trans-Golgi and phosphatidylinositol-4-phosphate (PI4P) backward in a coupled manner (de Saint-Jean et al., 2011). Structural, biophysical, and biochemical studies reveal that Osh4p binds ergosterol and PI4P in overlapping cavities and exchanges the two lipids upon binding to membranes (Delfosse et al., 2020). Intriguingly, Osh4p can transfer ergosterol against its gradient at the expense of a downgradient transfer of PI4P (de Saint-Jean et al., 2011; Moser von Filseck et al., 2015b). This counter-exchange activity has been observed both in the cell and in the reconstituted systems. Importantly, the activity is also shared by other Osh proteins and oxysterol-binding protein-related proteins (ORPs) in higher eukaryotes. For example, Osh6, ORP5, and ORP8 counter-transfer phosphatidylserine (PS) and PI4P at the ER-plasma membrane (PM) MCSs (Chung et al., 2015; Ghai et al., 2017; Sohn et al., 2018; Lipp et al., 2019). Yet, the molecular mechanism of LTP-mediated counter-transfer is not well understood.
Recently, a new category of eukaryotic LTPs has been discovered that potentially act as bridges to move lipids from ER to other organelles for their biogenesis (Kumar et al., 2018; Osawa et al., 2019; Valverde et al., 2019; Li et al., 2020; Melia et al., 2020; Hanna et al., 2021; Leonzino et al., 2021; Reinisch and Prinz, 2021; Wang et al., 2021a). These LTPs include ATG2, VPS13, and Ship164, all of which comprise a characteristic chorein motif at their N-termini. Structural studies revealed that these LTPs are club-like and contain a continuous groove 10–20 nm long capped with the chorein domain at one end (Valverde et al., 2019; Li et al., 2020; Hanna et al., 2021). The groove is lined with hydrophobic residues and can bind more than ten lipids in nearly nonspecific manner. Substituting the hydrophobic residues in the middle of the VPS13 bridge with hydrophilic residues impair yeast sporulation (Li et al., 2020). In addition, these LTPs contain domains that interact with other proteins residing in the two membranes to help engage the two ends of bridges to membranes (Leonzino et al., 2021). All these observations imply that ATG2 and VPS13 may act as a bridge to transport lipids in bulk between membranes. However, a direct test of bulk lipid flow remains missing. A key open question is what drives such directional lipid flow (Reinisch and Prinz, 2021). It has been hypothesized that certain ATPases may pump lipids into the bridges in a manner similar to many ATP-dependent lipid translocases, such as the lipopolysaccharide-transport protein complex in bacteria (Li et al., 2019; Owens et al., 2019). While this hypothesis is appealing, any candidate for such ATPases remains elusive. In addition, these ATPases only slowly translocase lipids with a maximum rate of ∼5 lipids per second observed in vitro (Mi et al., 2017; Li et al., 2019; Shukla and Baumgart, 2021), which is orders of magnitude smaller than the speed expected for the fast membrane expansion observed in vivo (∼5,000 lipids per second, see the forthcoming section). Therefore, other potential driving forces need to be explored.
Most LTPs transfer lipids at the MCSs (Saheki and De Camilli, 2017a; Wong et al., 2019; Prinz et al., 2020; Reinisch and Prinz, 2021). Many LTPs contain membrane-binding domains, such as C2, PH, and PX domains, that help tether two membranes to form MCSs (Lemmon, 2008). These domains bind membranes by recognizing specific lipids, Ca2+, or other ligands. Consequently, MCSs are highly dynamic and triggered by many signaling clues in the cell, particularly PI(4,5)P2 (Prinz et al., 2020). Eukaryotic extended-synaptotagmins (E-Syts), with three isoforms E-Syt1, E-Syt2, and E-Syt3, are representative LTPs and membrane tethers (Min et al., 2007; Chang et al., 2013; Giordano et al., 2013; Saheki and De Camilli, 2017b). They contain an N-terminal hydrophobic hairpin anchored to the ER membrane and a synaptotagmin-like mitochondrial lipid-binding protein (SMP) domain as the lipid transfer module (Kopec et al., 2010; Saheki et al., 2016; Yu et al., 2016) (Figure 1A). Two SMP domains form a stable dimer that mediates homo- or hetero-dimerization between E-Syts. The crystal structure of an E-Syt2 fragment reveals two lipids bound by each SMP domain (Schauder et al., 2014). A defining feature of E-Syts is an array of C2 domains at the C-termini, five in E-Syt1 and three in E-Syt2 and E-Syt3, connected by disordered polypeptides (Min et al., 2007; Tunyasuvunakool et al., 2021). Cellular imaging shows that, at a low cytosolic Ca2+ level, E-Syt2 and E-Syt3 are constitutively localized at the ER-PM MCSs, whereas E-Syt1 appears diffusive in the entire ER membrane (Giordano et al., 2013). An elevation of the cytosolic Ca2+ level triggers the recruitment of E-Syt1 to the MCSs, a decrease in membrane separation (Fernandez-Busnadiego et al., 2015), and accompanying lipid transfer mediated by all E-Syts.

Schematics of E-Syt structures and experimental setup to study C2-membrane binding. A: Domain organization of E-Syts and their roles in forming membrane contact sites (MCSs) and transferring lipids. The synaptotagmin-like mitochondrial-lipid-binding (SMP) domain mediates dimerization of E-Syts and lipid transfer. The numbers near the disordered polypeptide linkers indicate their length in amino acids. B: Experimental setup to pull the E-Syt1 C2 array (E-Syt1 C2ABCDE) using optical tweezers. The array is attached to the lipid bilayer supported on a silica bead at its N-terminus and conjugated at its C-terminus to a 2,260 bp DNA handle that is immobilized on a polystyrene bead at another end. The extension of the protein-DNA tether is measured to probe dynamic C2-membrane binding and unbinding in the presence of controlled force. Both diagrams are not drawn to scale. For comparison, E-Syt1 C2E is located ∼76 nm away from the ER membrane as estimated by the contour length of the protein. Adapted from (Ge et al., 2022).
In the following, we first review recent progress in understanding the membrane binding of E-Syts, its role in membrane contact formation, and the mechanism of lipid transfer by the SMP domain. Then, we propose quantitative models of lipid transfer by shuttle LTPs based on the lipid exchange mechanism and test the model with experimental data. Finally, we discuss the roles of canonical and osmotic membrane tension in bulk lipid transfer mediated by bridge LTPs.
Single-Molecule Measurements of E-Syt-Membrane Binding and Modeling of MCSs
While the membrane-binding properties of individual C2 domains have been extensively studied (Corbalan-Garcia and Gomez-Fernandez, 2014), it was challenging to characterize the membrane binding of E-Syts containing an array of protein domains that may bind membranes and each other (Bian et al., 2018). Furthermore, the membrane binding in trans generates a pulling force to the C2 domain, which in turn modulates its membrane binding (Ma et al., 2017). To characterize the force-dependent interactions, we applied optical tweezers to dissect the membrane binding of E-Syt1 and E-Syt2 as a function of Ca2+ and lipid composition (Ge et al., 2022). A fragment of the C2 containing region of E-Syt1, e.g., E-Syt1 C2ABCDE, is tethered to a lipid bilayer coated on a silica bead at its amino terminus and to a polystyrene bead at its carboxy terminus via a 2,260 bp DNA handle (Figure 1B). The two micron-sized beads are optically trapped with high-resolution dual-trap optical tweezers (Zhang et al., 2013) and pulled to apply force to the C2 repeat. The extension of the protein-DNA tether is detected to report dynamic binding and unbinding of the C2 domains to the membrane or other C2 domains (Figure 2A). The E-Syt1 protein undergoes fast and three-state transitions at a constant mean force in the range of 3.6–4.6 pN (Figure 2B), indicative of sequential membrane binding and unbinding of the C2E and C2CD domains (Figure 2A). The membrane binding of the C2AB domain is weak and only observed in the presence of the higher 20 mol% DOPS. The C2CD domain binds to the membrane in a Ca2+-dependent manner, while the C2E domain shows moderate membrane affinity in a PI(4,5)P2-dependent but Ca2+-independent manner. Pulling a single monomer within the cytoplasmic domain of the E-Syt1 dimer reveals similar membrane binding kinetics, suggesting minimum interactions between the SMP dimer and the membrane. The single-molecule assay did not reveal any significant interactions between different E-Syt1 domains, indicating that these interactions are weak. Similarly, the membrane binding kinetics of E-Syt2 is measured.
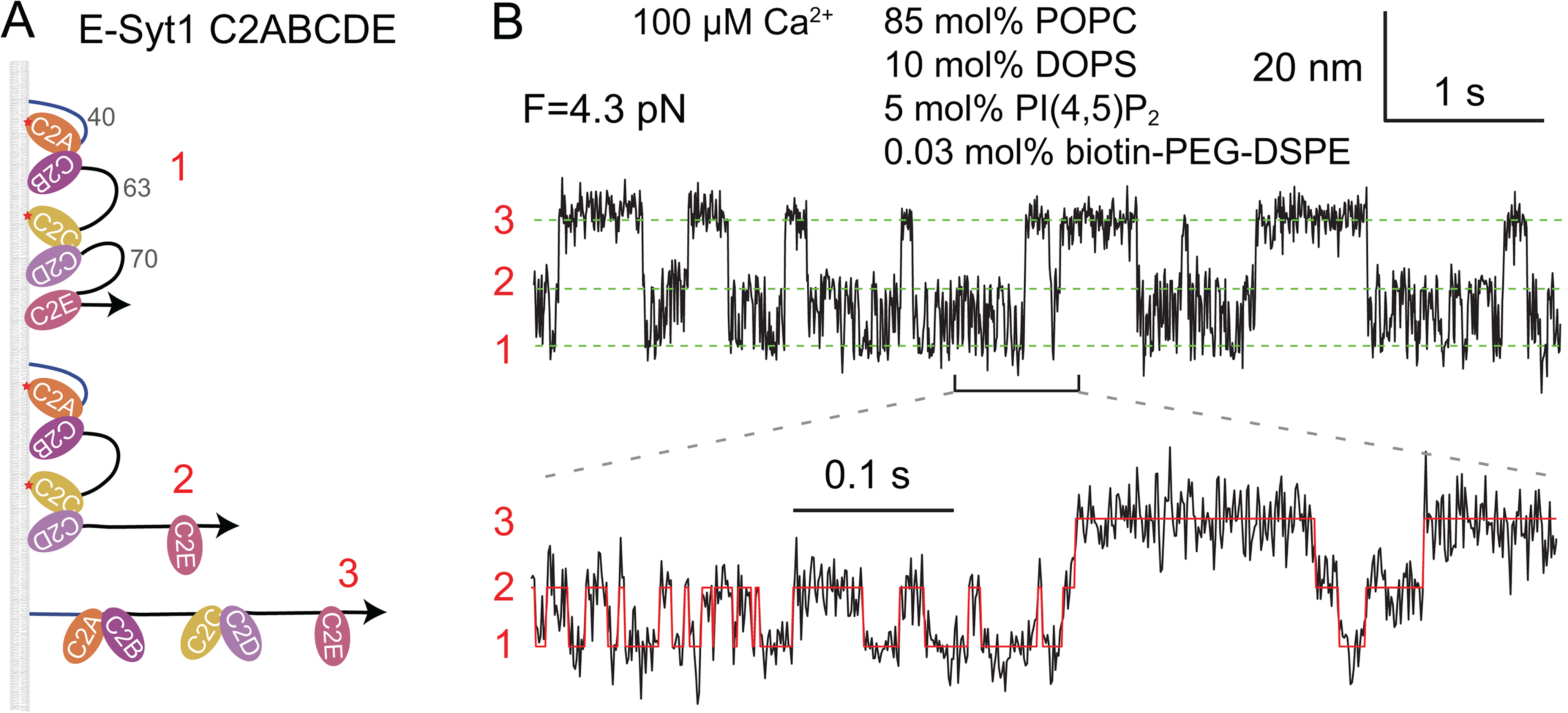
C2 domains sequentially bind to and unbind from the membrane detected by optical tweezers. A: Three C2 binding states with distinct extensions. The corresponding state numbers are shown in red. B: Time-dependent extension measured at the indicated constant mean force showing transitions of the C2 array among the different binding states. The green dashed lines mark the average positions of different states. The red curve indicates the idealized state transitions derived from hidden Markov modeling (Zhang et al., 2016). Adapted from (Ge et al., 2022).
We develop an analytical theory to predict the trans-membrane binding of E-Syts required for MCS formation based on the cis-membrane binding parameters measured by us (Ge et al., 2022). The membrane binding of different C2 modules depends on the lengths of the flexible polypeptide linkers between these C2 modules. We use the Gaussian chain model to describe the conformation of each linker and extracted the membrane-binding energy and rate corresponding to each isolated C2 module. This leads to binding energy of 10.4 (±0.9) kBT for C2CD, 6 (±1) kBT for C2E in E-Syt1, 7.1 (±0.4) for C2AB, and 13.0 (±0.7) for C2C in E-Syt2 under the experimental conditions indicated in Figure 2B. Next, we calculate the trans-membrane binding probability of each C2 module as a function of the distance between the ER- and PM-like membranes. The trans-membrane binding is a tug of war between the affinity of each C2 module for the PM-like membrane and the opposing entropic force that stretches the linker between the C2 module and the ER anchor (Figure 1A). Here, the force and energy needed to stretch the linker are determined using a worm-like chain model for the linker (Marko and Siggia, 1995; Rebane et al., 2016). The repulsive interactions between the ER and PM membranes provide another opposing force for the trans-membrane binding. With an empirical potential for the membrane interactions (Zorman et al., 2014), the total energy of the system can be calculated to predict the equilibrium membrane distance and the probability of MCS formation.
Several salient features of the E-Syt-mediated MCSs are predicted (Ge et al., 2022). In the absence of Ca2+, E-Syt1 has less energy in the unbound state than in any C2-bound state, indicating that E-Syt1 alone cannot mediate an MCS. In contrast, E-Syt2 has minimal energy in a C2C-bound state at an equilibrium distance of ∼19 nm due to the high affinity of C2C for the PM-like membrane. The free energy of this state is −2.2 kBT relative to the unbound state, consistent with a stable E-Syt2-mediated MCS. In the presence of Ca2+, E-Syt1 starts to tether the PM membrane at ∼16 nm due to binding of both C2CD and C2E, while E-Syt2 shows a metastable state at ∼11 nm due to binding of both C2AB and C2C. All these predictions, including the Ca2+-dependent equilibrium membrane separation, are consistent with the cellular imaging results obtained by electron and fluorescence microscopy (Chang et al., 2013; Giordano et al., 2013; Fernandez-Busnadiego et al., 2015; Idevall-Hagren et al., 2015).
The SMP Domain Acts as a Shuttle to Transfer Lipids
It is puzzling how the SMP domain transfers lipids between membranes. While many shuttle LTPs contain relatively small hydrophobic cavities that accommodate one lipid at a time (Delfosse et al., 2020), the SMP dimer in E-Syts forms a hydrophobic groove along its 9 nm length that can hold four phospholipids (Schauder et al., 2014). These observations suggest that the SMP dimer might serve as a bridge for lipid transfer. SMP belongs to the tubular lipid binding protein (TULIP) family of LTPs with conserved structural features, including long hydrophobic grooves (Kopec et al., 2010; Reinisch and De Camilli, 2016). One of the well-studied TULIP LTPs is cholesterol ester transfer protein (CETP) that transfers cholesterol esters and triglycerides between high-density lipoproteins (HDL) and low-density lipoproteins (LDL) in the blood (Qiu et al., 2007; Liu et al., 2012). Compelling evidence suggests that CETP binds both HDL and LDL and acts as a bridge to transfer cholesterol ester or triglycerides between them (Zhang et al., 2012; Lei et al., 2016). The structural similarity between the SMP dimer and CETP also implies a bridge mechanism for lipid transfer by SMP. However, such an inference about SMP is not supported by experimental observations (Reinisch and De Camilli, 2016). The measured ER-PM distance at the E-Syt-mediated MCSs is ∼20 nm in a rest state with a low cytosolic Ca2+ concentration and ∼15 nm in an activated state with an elevated Ca2+ concentration (Fernandez-Busnadiego et al., 2015); both are significantly greater than the 9 nm length of the SMP dimer. In contrast, a recent study suggests that tricalbins, the E-Syt homologs in yeast, mediates MCSs with an ER-PM distance of 7–8 nm that are essential for lipid transfer (Collado et al., 2019), making it possible for the SMP dimers to form bridges at the MCSs. In summary, more evidence is required to resolve the lipid transfer mechanism by E-Syts.
It was technically challenging to distinguish whether an LTP transfers lipids as a shuttle or a bridge, due to the difficulty in controlling the membrane separation with nanometer resolution in vitro. Recently, Bian et al. combined the DNA-origami technique and the FRET-based lipid transfer assay to investigate the mechanism of lipid transfer by the SMP dimer (Bian et al., 2019) (Figure 3). They created rigid DNA rings as a scaffold to form two types of liposomes ∼28 nm in diameter. The isolated SMP dimer was attached to one type of scaffolded liposomes via a disordered polypeptide linker with a specific length. These liposomes also contained NBD- and rhodamine-labeled lipids. When mixed, the two types of liposomes were joined by a rigid DNA pillar and held at a specific membrane distance. The potential lipid transfer mediated by the SMP dimer was detected via an increase in NBD fluorescence.

SMP dimer can act as a shuttle to transfer lipids between membranes, with their separation controlled by DNA origami. A: The SMP dimer attached to one liposomal membrane can transfer lipids to another liposomal membrane 26 nm apart that cannot be bridged by the SMP dimer (∼9 nm in length). B: The short linker attaching the SMP dimer to the membrane prevents the SMP dimer from reaching another distant membrane for lipid transfer.
Using this new assay, Bian et al. demonstrated that the SMP domain of E-Syt1 transfers lipids between the two membranes even when they are separated by 26 nm (Figure 3A), significantly greater than the length of the dimeric SMP domain. Shortening the contour length of the linker from ∼30 nm to ∼8 nm abolishes lipid transfer, while lengthening the linker to ∼85 nm rescues the lipid transfer activity, even when the two membranes are further separated to ∼67 nm (Figure 3B). All these observations are consistent with a shuttle mechanism for the SMP dimer and inconsistent with the bridge mechanism under these experimental conditions. However, it remains to test whether the SMP domain or other LTPs containing SMP domains (Reinisch and De Camilli, 2016) can also act as a bridge to transfer lipids at a short membrane distance close to the length of the SMP dimer, given its structural resemblance to other proposed bridge LTPs.
Lipid Exchange Model for Shuttle LTPs
Suppose an LTP transfers lipid species 1 and 2, referred to as lipid 1 and lipid 2 for simplicity, between two membranes. Based on the lipid exchange mechanism of the Osh/ORP-family LTPs (de Saint-Jean et al., 2011; Mesmin et al., 2013; Chung et al., 2015; Moser von Filseck et al., 2015a; Moser von Filseck et al., 2015b; Ghai et al., 2017; Dong et al., 2019; Wang et al., 2019; Kawasaki et al., 2022) (Figure 4A), we proposed a reaction scheme to quantify lipid transfer (Figure 4B). To simplify our model, we assume that the LTP is always bound by either of the lipids. The assumption appears to be justified for most LTPs, as they show high affinities for lipids with estimated dissociation constants less than 0.1 µM (Petrescu et al., 2001; Schauder et al., 2014; Yu et al., 2015; Sandhu et al., 2018), compared to much higher lipid concentrations in typical lipid transfer assays (> 6 µM) (Moser von Filseck et al., 2015b; Saheki et al., 2016; Yu et al., 2016; Horenkamp et al., 2018) or at MCSs (up to 0.1 M) (Ma et al., 2015; Ge et al., 2022). In addition, we assume that the LTP can freely diffuse in the solution without being tethered to any membrane. When a lipid 2-bound LTP binds a membrane, the LTP exchanges for lipid 1 with a bimolecular rate constant

Lipid exchange scheme for lipid transfer by shuttle LTPs. A: Osh4p transfers ergosterol from the ER to the trans-Golgi upgradient and PI4P downgradient in a reverse direction via a lipid exchange mechanism. B: Proposed scheme of lipid exchange between a shuttle LTP and a membrane characterized by two bimolecular reaction constants. The lipids 1 and 2 exchange between the LTP and the membrane. C: LTP-mediated lipid transfer between two membranes via two lipid exchange reactions. The concentrations of lipids 1 and 2 in the membranes and the LTP are indicated.
Direct lipid exchange between LTPs and membranes is often measured with fluorescence approaches (de Saint-Jean et al., 2011; Moser von Filseck et al., 2015b). In these experiments, LTPs are loaded with lipid 1 and tested its exchange with an excessive of lipid 2 in a membrane by measuring the change in the fluorescence of the LTPs bound by different lipids. As a result, the effects of lipid composition and LTP mutation on the exchange rate are widely investigated. Suppose at time t, the membrane has molar concentrations of
In the presence of another membrane, the LTP mediates lipid exchange with both membranes (Figure 4C). For the convenience of our forthcoming description, one membrane is called a lipid donor membrane and the other acceptor membrane. The two exchange rate constants may depend on membranes, as the LTP and lipids may differentially interact with proteins or non-exchangeable lipids in different membranes. Consequently, the two sets of rate constants associated with the donor and acceptor membranes may be similar or different. In the following, we will consider the two cases separately.
Our model of lipid exchange differs from the previous model developed by Drin and coworkers (Moser von Filseck et al., 2015b; Ikhlef et al., 2021), in which several intermediate states of Osh4p-mediated lipid transfer are included, for example, the states in which Osh4p is bound to the membrane but not loaded with any lipid. Consequently, the model contains six coupled reactions with more than 10 rate constants, which make it difficult to derive an analytical solution for the lipid transfer rate and fit the model to experimental data. In contrast, our model only contains one reaction with two rate constants for the two membranes with similar lipid and protein composition or two reactions with four rate constants for the membranes with distinct composition. Consequently, our model yields analytical solutions and convenient fitting to experimental data, as shown in the forthcoming sections. It must be noted that, to our knowledge, the binding affinity of Osh4p to any lipid remains to be measured. Thus, the assumption adopted in our model that all LTPs are bound by lipids awaits further experimental tests.
Lipid Exchange Between Membranes with Similar Lipid and Protein Composition
Suppose at time t, a donor membrane has molar concentrations of

Simulations of Osh4p-mediated lipid transfer between membranes of similar or distinct lipid and protein compositions. A,C: Initial and equilibrium concentrations of lipids accessible to Osh4p (black numbers in the unit of µM) and molar percentages (mol%) within the transferable lipids (red numbers) in donor and acceptor membranes with similar (A) or distinct (C) lipid and protein compositions. The simulations were performed under similar conditions as in the representative lipid transfer assay with 0.2 µM Osh4p (Moser von Filseck et al., 2015b). The lipid transfer kinetics and equilibrium lipid compositions depend on the rate constants of the lipid exchange at the two membranes (in a unit of 105 M−1s−1). A gradient of the molar percentage of the exchangeable lipid drives lipid mixing, as highlighted by the red box. B: Calculated fractions of DHE in the donor and acceptor membranes and of DHE-bound Osh4p with equal (solid lines) or unequal (dashed lines) exchange rate constants (see A and C). The approximate analytic solutions (dotted lines) calculated based on Eqs. (12) and (22) overlap the corresponding numerical solutions. The equilibrium lipid concentrations and molar percentages within the exchangeable lipids are also shown in A and C.
To better understand the kinetics of lipid transfer, we derived an approximate analytic solution to the system of equations in Eqs. (10). We noticed that the fraction of DHE-bound Osh4p (Osh4p-DHE) reaches a constant within ∼0.2 s, much earlier than the moment when the fractions of DHE and PI4P significantly change (Figure 5B, solid blue curve). Indeed, the two lipids equilibrate among Osh4p and both membranes with a time constant
The analytical solution leads to interesting observations or predictions. First, despite the steady-state approximation, the analytical solutions are indistinguishable from the numerical solutions with overlapping time courses (Figure 5B), suggesting a high accuracy of the analytical solution. Second, the lipid transfer follows single-exponential kinetics characterized by a single rate constant k as shown in Eq. (13). This rate constant is proportional to the LTP concentration
The analytical formula provides new insights into the mechanism of the transfer of lipids against their gradients. The solutions in Eqs. (12) give the identical fractions of the two lipids in both the donor membrane (
Lipid Exchange Between Membranes of Distinct Composition
The LTP-mediated lipid exchange depends on the curvature and composition of membranes (Moser von Filseck et al., 2015b; Ikhlef et al., 2021). For example, Osh4p-PI4P extracts DHE more rapidly in DOPC liposomes than in POPC liposomes. In addition, small liposomes promote DHE extraction. To simulate lipid transfer between the membranes of distinct composition and morphology, we use two sets of rate constants:
We again derived an approximate analytic solution to Eqs. (18) or (19), i.e.,
In conclusion, we developed a quantitative model for LTP-mediated lipid transfer. We found two nonexclusive mechanisms by which LTPs transfer lipids against a lipid gradient – selective lipid mixing between membranes of similar composition and lipid partition between membranes of distinct composition with different energy. Both mechanisms not only are consistent with experimental results (Moser von Filseck et al., 2015b) but also lead to several interesting predictions that may be tested in future experiments to better understand the lipid transfer process.
Lipid Exchange Model fit to the Experimental Data
To test whether the lipid exchange scheme shown in Figure 4B is consistent with the experimental results, we analyzed the time courses of Osh4p-medicated DHE and PI4P exchange measured by Drin and coworkers (Moser von Filseck et al., 2015b). As shown in Figure 5A, the lipid transfer experiments began with 10 µM DHE in the donor liposomes and 0–5 µM transferable PI4P but no DHE in the outer leaflets of acceptor liposomes with the addition of 0.2 µM Osh4p. Because DOPC constitutes the majority of the total 200 µM lipids in both types of liposomes, the donor and acceptor membranes are approximately equal, and the lipid exchange reaction is characterized by two rate constants to be determined. We first calculated the time- and PI4P-dependent DHE amount transferred by Osh4p and fit the time courses with both single- and double-exponential functions (Figure 6A). We found that the transfer kinetics is not single-exponential (Figure 6A, green dashed line), but a double-exponential function fits each time course well. The double-exponential fitting revealed a fast phase with a rate (

Comparisons of the measured and predicted DHE transfer amount and rates. A: Time-dependent DHE amount transferred by Osh4p (symbols) in the presence of different concentrations of PI4P in the acceptor membrane. The time courses are well fit by a double-exponential function
The lipid exchange scheme is consistent with the measured lipid transfer kinetics. Considering the transferable lipids in the outer leaflets of both donor and acceptor membranes, we have the total lipid concentration
Energetics and Kinetics of Lipid Exchange by Shuttle LTPs
The desorption free energy needed to transfer a POPC lipid from membranes to the solution is 68 kcal/mol (27 kBT), with an even higher energy barrier of 96 kcal/mol (38 kBT) (Mclean and Phillips, 1984; Grafmuller et al., 2013). To efficiently solubilize the lipids required for lipid transfer, LTPs need to tightly bind lipids with unbinding energy commensurate with the desorption energy. Thus, although LTPs free of lipids can be purified in vitro, most LTPs are probably always bound by lipids in the cell or at MCSs with abundant lipids. Indeed, lipids are generally found in the purified LTPs and are widely seen in their high-resolution structures (Liu et al., 2012; Schauder et al., 2014; Kumar et al., 2018; Valverde et al., 2019). Consequently, these LTPs can only exchange their bound lipids with the lipids in the membrane. Since no net lipid transfer occurs if an LTP only recognize a single lipid species and exchanges another lipid in the same species, these LTPs are expected to bind two or more species of lipids. These lipids bind to partly overlapping cavities in the LTP to accommodate both common and distinct structural features of these lipids (de Saint-Jean et al., 2011; Lipp et al., 2020). A lipid may first latch onto the distinct binding region and then displace another lipid in the overlapping binding cavity to facilitate lipid exchange. This lipid binding mode allows different lipids to exchange like toehold-mediated DNA strand displacement, which overcomes the high energy barrier for the exchange processes (Zhang and Winfree, 2009). Because the exchange process is reversible, the difference in the binding energy of both lipids for the LTP is expected to be similar to the difference in their desorption energy. Otherwise, a lipid with an excessive binding affinity for the LTP would be trapped in the binding cavity and block the exchange reaction. Consistent with this view, the LTP-lipid binding affinity appears to be tuned to a similar level for efficient exchange of the cognate lipids (Ikhlef et al., 2021). Therefore, based on the energetics and kinetics of lipid extraction and solubilization, we hypothesize that lipid exchange may be a common mechanism for shuttle LTPs that transfer lipids with long acyl chains. Consequently, our model for LTP-mediated lipid transfer may be broadly applicable to LTPs. However, some LTPs appear to bind only one species of lipids and might transfer lipids via a lipid-unbound LTP state without lipid exchange (Wong et al., 2017). To equilibrate this state with the lipid-bound state, these LTPs may undergo large conformational changes to shield their hydrophobic lipid binding cavities to reduce the energy of the lipid-unbound state. More experiments are needed to test these hypotheses.
Driving Forces for Directional and Bulk Lipid Transfer Through Bridge LTPs
Experiments so far have revealed a slow lipid transfer via shuttle LTPs (<1 s−1) (Moser von Filseck et al., 2015b; Wong et al., 2019). It is generally believed that bridge LTPs transport lipids much faster than shuttle LTPs to allow rapid membrane expansion observed in vivo (Kumar et al., 2018; Melia et al., 2020; Leonzino et al., 2021). Evidence suggests that ATG2 mediates lipid transfer from the ER to the autophagosome, a double-membrane organelle essential for macroautophagy (Maeda et al., 2019; Osawa et al., 2019; Valverde et al., 2019). The lipid transfer allows autophagosomes to grow from small seed vesicles into large organelles a few hundred nanometers to one micron in diameter (Sawa-Makarska et al., 2020; Ghanbarpour et al., 2021). The growth takes ∼10 min (Axe et al., 2008; Xie et al., 2008; Schutter et al., 2020), requiring roughly three million lipids for a typical 400 nm autophagosome and a total lipid transfer rate of ∼5,000 lipids per second (Melia et al., 2020). Although many LTPs could cooperate to meet the speed requirement, the limited size of the MCSs might not accommodate so many LTPs, for example, at the ER-autophagosome contact site, which necessitates a fast bridge LTP like ATG2.
Surprisingly, all bridge LTPs are found to transfer a few lipids per minute per LTP in vitro (Kumar et al., 2018; Maeda et al., 2019; Osawa et al., 2019; Valverde et al., 2019; Li et al., 2020; Hanna et al., 2021). The discrepancy may be caused by several limitations of current assays to detect lipid transfer by bridge LTPs, which lead to underestimations of their lipid transfer rates. First, experiments so far utilize the same FRET- and liposome-based assays to measure lipid transfer by both bridge and shuttle LTPs. These assays cannot distinguish lipid mixing and bulk lipid flow. Thus, it is possible that the lipid transfer rates measured for bridge LTPs may be mainly caused by lipid mixing rather than bulk lipid flow. Second, the bulk lipid flow requires the LTP to tether and bridge two membranes. However, the membrane bridging under in vitro experimental conditions may be unstable and transient, especially in the absence of other factors that facilitate the membrane tethering in the cell. Therefore, the measured lipid transfer rate is probably scaled by the limited duty cycle of the LTPs. Third, the actual number of bridge LTPs involved in membrane tethering is unknown. Consequently, the measured lipid transfer rate of a bridge LTP may no longer be proportional to the concentration of the LTP as the shuttle LTP, depending upon how the LTPs bridge the two membranes (Von Bulow and Hummer, 2020). Finally, and most importantly, current assays lack essential driving forces for bulk lipid flow present in vivo, as discussed below. New experimental and modeling methods are required to quantitatively understand lipid flow through bridge LTPs.
In the absence of any lipid synthesis, the bulk lipid flow through bridge LTPs causes an increase in the area of the acceptor membrane and an accompanying decrease in the area of the donor membrane. As a result, the bulk lipid flow is energetically coupled to the difference in membrane tension of the two membranes. By analogy with the roles of pressure in the water flow through a pipe and the transmembrane potential in the current through an ion channel, we propose that the membrane tension difference or gradient is a general driving force for the bulk lipid flow through a bridge LTP (Figure 7A). Membrane tension is defined as positive when the tension stretches the membrane and negative when the tension compresses the membrane. Thus, lipids flow from a membrane with lower tension to a membrane with higher tension. The lipid flow rate (
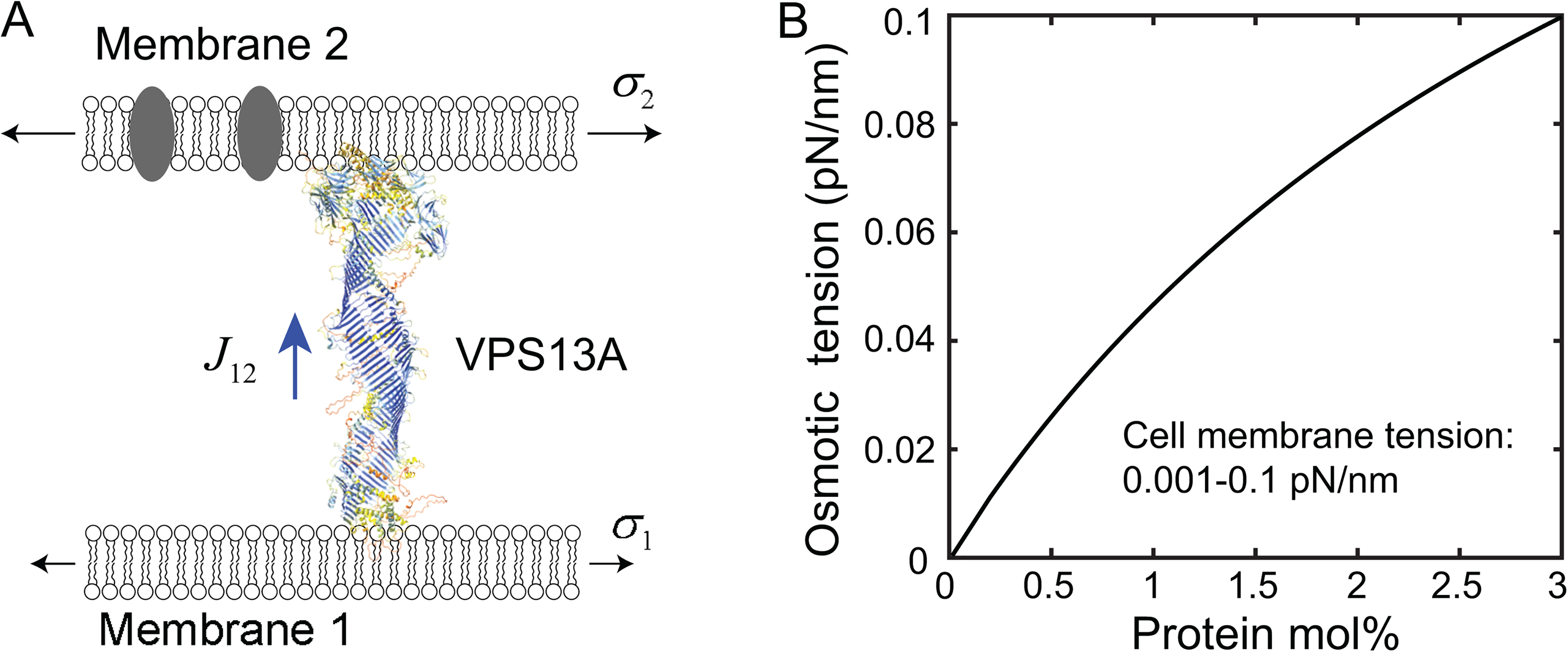
Potential driving forces for bulk lipid flow through bridge LTPs. A: Canonical membrane tension and the hypothetical osmotic membrane tension may drive lipid flow through bridge LTPs. The structure of human VPS13A, one of four isoforms of VPS13 proteins, shown here is derived from AlphaFold (Tunyasuvunakool et al., 2021). B: The predicted osmotic membrane tension as a function of the concentration of membrane proteins. The concentration of the membrane protein
The lipid flow through bridge LTPs may also be driven by different lipid and protein compositions of membranes. Integral and peripheral membrane proteins are unlikely to pass through the narrow bridges (Li et al., 2020; Tunyasuvunakool et al., 2021). Consequently, an LTP bridge may act like a semipermeable membrane to generate directional lipid flow in analogue to osmosis (Figure 7A). Therefore, we term this predicted new phenomenon mediated by bridge LTPs as membrane osmosis. In this case, the large molar amount of transferable lipids serves as a solvent, while the small molar amount of mobile membrane proteins and non-transferable lipids acts as a solute that cannot pass through the bridges. In parallel, we define the membrane osmotic tension (
We estimate the lipid flow rate in the presence of a membrane tension gradient. A lipid in the LTP bridge experiences a pulling force f towards the acceptor membrane with higher membrane tension. This force can be estimated from the energy gain
Implications of the Driving Forces in Membrane Biogenesis
Bridge LTPs are widely involved in biogenesis of organelles (Kumar et al., 2018; Valverde et al., 2019; Melia et al., 2020; Leonzino et al., 2021). Here we focus on the membrane expansion during autophagosome biogenesis. The dramatic membrane expansion occurs in multiple stages and is mediated by dozens of conserved proteins (Hurley and Schulman, 2014; Gomez-Sanchez et al., 2021) (Figure 8A). Recent experiments suggest that the de novo autophagosome biogenesis starts from formation of the MCS between a small, ATG9-containing seed vesicle and the ER membrane with the help of ATG2 and ATG9 (Yamamoto et al., 2012; Guardia et al., 2020; Sawa-Makarska et al., 2020; Ghanbarpour et al., 2021). ATG9 directly binds to ATG2 and acts as a lipid scramblase to equilibrate lipids between the two leaflets of membranes at the MCS during the flow of lipids only in the outer leaflets mediated by ATG2 and other bridge LTPs (Guardia et al., 2020; Maeda et al., 2020; Matoba et al., 2020; Ghanbarpour et al., 2021). The directional lipid flow initially enlarges the seed vesicle, then likely induces its transition into a cup-shaped phagophore, a morphologically distinct precursor of the autophagosome. Further lipid transfer expands the phagophore to engulf the cytoplasmic materials to degraded, which then matures into the double-membrane autophagosome by sealing its highly curved rim. The size of phagophores is highly dynamic and adapts to the size of its cargo (Xie et al., 2008; Melia et al., 2020). Thus, the molecular mechanism of autophagosome biogenesis is key to understanding autophagy.
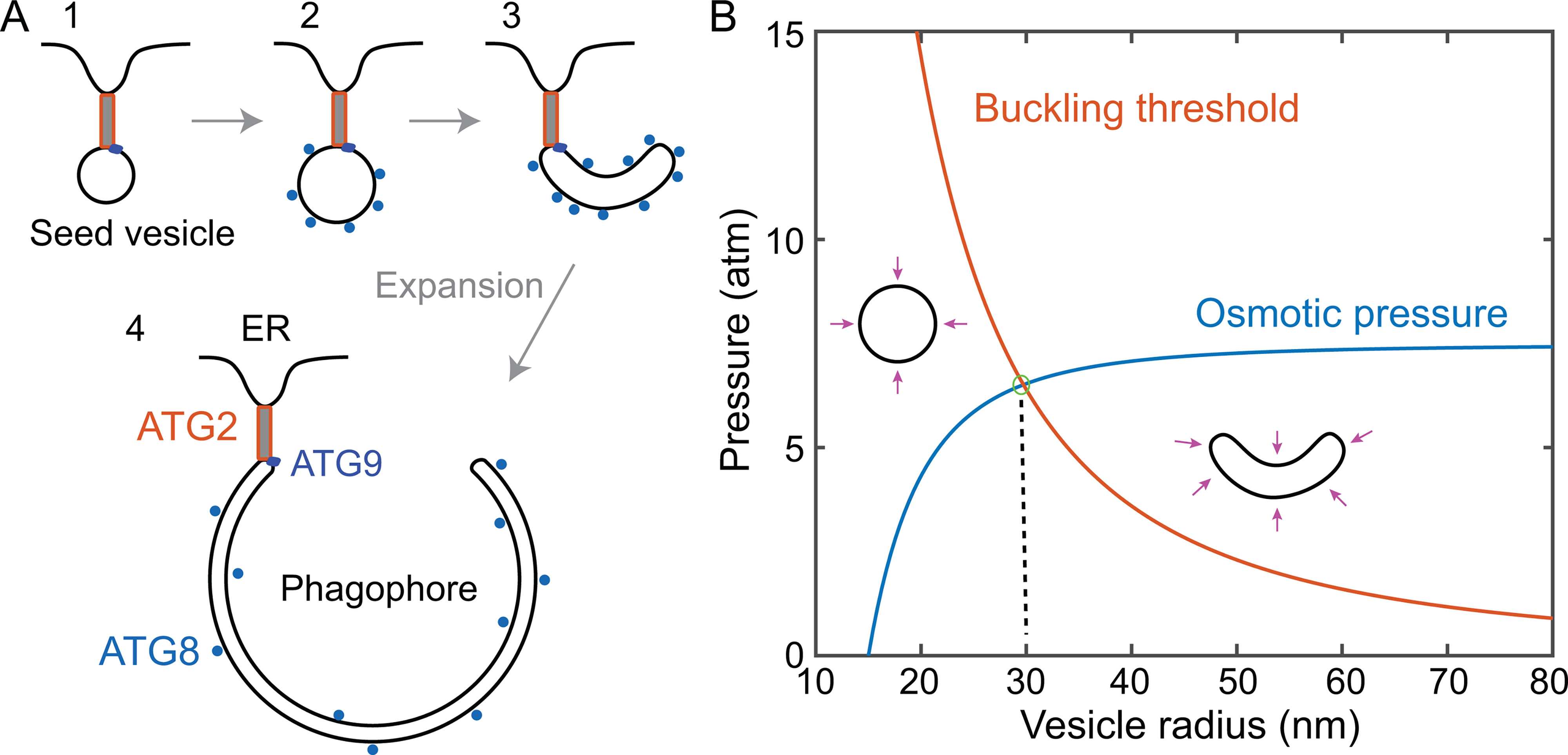
Lipid transfer, membrane conformational transitions, and osmotic pressure. A: Membrane precursors involved in autophagosome biogenesis, including the Atg9-containing vesicle tethered to the ER membrane by ATG2, ATG9, and other proteins (state 1), the growing vesicle containing lipidated ATG8 (state 2), the cup-shaped phagophore (state 3) with its convex side attached to cytoplasmic materials to be degraded (not shown), and the expanded phagophore (state 4) engulfing cytoplasmic materials. Sealing of the top rim of the expanded phagophore via membrane fission leads to a mature phagophore. B: Comparison of the vesicle buckling pressure and osmotic pressure as a function of the vesicle radius. A growing spheric vesicle buckles into a cup-shaped vesicle as the compression osmotic pressure reaches the buckling pressure at 6.5 atm and a vesicle radius of ∼30 nm (marked by a green circle). The pressure unit is atmospheric pressure (atm). For the calculations, the osmolarity of cytosol is chosen as 300 mOsm. The osmolarity inside the seed vesicle 15 nm in radius is the same but decreases as its radius increases.
Various factors in the cell may generate the canonical and osmotic membrane tension to regulate autophagosome biogenesis. Cytoskeletons and their associated motors play wide roles in biogenesis and remodeling of organelles, for example, in endocytosis, and help regulate membrane tension of the PM (Shi et al., 2018). Likewise, cytoskeletons are essential for nearly every stage of autophagosome biogenesis (Kruppa et al., 2016; Kast and Dominguez, 2017). Agents that block actin or microtubule polymerization inhibit phagophore growth. Although the underlying mechanisms are generally unclear, it is possible that cytoskeletons and their associated molecular motors attach to and pull on growing phagophores, which induce tension in phagophore membranes that draw lipids from the ER for phagophore membrane expansion via ATG2.
The ATG2-mediated lipid flow may also be driven by the osmotic membrane tension induced by many peripheral proteins essential for autophagosome growth, especially ATG8 (Figure 8A) (Melia et al., 2020; Sawa-Makarska et al., 2020). ATG8 is covalently and reversibly conjugated to phosphatidylethanolamine (PE) in phagophore membranes during autophagosome biogenesis (Xie et al., 2008; Abdollahzadeh et al., 2017). The lipidated ATG8 is the most abundant protein on phagophore membranes and long serves as a defining marker of autophagosomes (Geng et al., 2008). Importantly, the amount of lipidated ATG8 correlates with the size of autophagosome for an unknow mechanism (Xie et al., 2008). We hypothesize that the concentration of lipidated ATG8 contributes mostly to the osmotic membrane tension of phagophores, which in turn regulates lipid flow to phagophore by ATG2, in addition to many other functions of ATG8 lipidation in autophagy (Shpilka et al., 2011). In yeast, each autophagosome has an average radius of ∼150 nm and contains 272 lipidated ATG8 proteins under growth conditions (Xie et al., 2008). It appears that these ATG8 proteins accumulate in the early phase of phagophore growth. Assuming the same number of lipidated ATG8 on a spheric vesicle with a radius of 30 nm before or 150 nm after membrane expansion, we estimated ATG8-generated osmotic membrane tension at 0.1 pN/nm or 0.002 pN/nm, respectively. Therefore, the smaller vesicle probably has much higher osmotic membrane tension than the ER, driving fast lipid flow from the ER to the vesicle. In contrast, the larger mature autophagosome bears osmotic membrane tension similar to or even lower than that of the ER, which prevents further growth of the autophagosome. Thus, the dilution of the lipidated ATG8 accompanying autophagosome growth may provide a simple mechanism to regulate the autophagosome size via the osmotic membrane tension.
Directional lipid flow is intimately coupled to membrane configurations. The lipid flow may help generate the characteristic cup-shape phagophore essential for engulfing the cytoplasmic materials to be degraded. This coupling between lipid transfer and membrane configurations is recently proposed by Ghanbarpour et al. based on conservation of the vesicle volume (Ghanbarpour et al., 2021), as is demonstrated in previous studies on membrane mechanics and morphology (Seifert, 1997; Flatt and Bruce, 2009; Bahrami et al., 2017). Here we clarify this important concept based on vesicle buckling induced by the osmotic pressure. Except for ATG9, the phagophore membrane is essentially devoid of any integral membrane proteins (Fengsrud et al., 2000), including ion channels and transporters that may regulate the osmotic pressure of phagophores. The cytosol of a cell typically has an osmolarity of ∼300 mOsm, which could generate a maximum compression pressure of 7.5 atmospheric pressure (atm) on a membrane bound organelle corresponding to zero osmolarity in its interior. If the mechanical strength of the organelle could not withstand the pressure, the organelle would flatten, eventually turn into a cup-shape with smaller volume to minimize both the osmotic pressure and membrane bending energy (Seifert, 1997).
We predict a transition in membrane configuration from a sphere to a cup-shape during phagophore expansion. To this end, we calculated the osmotic compression pressure of a spherical vesicle and its buckling pressure threshold as a function of the vesicle radius (Figure 8B). Any thin spherical shell buckles at a maximum compression pressure
In conclusion, the gradients of canonical and osmotic membrane tension may drive the directional flow of bulk lipids via bridge LTPs required for organelle biogenesis. The lipid flow is intimately coupled to membrane configuration and osmotic pressure and is facilitated by various proteins that remodel membranes, regulate the volumes or osmotic pressure, and equilibrate the lipids between the two leaflets of bilayers (Owens et al., 2019; Matoba et al., 2020; Ghanbarpour et al., 2021).
Conclusions and Perspectives
Despite extensive studies, there have been no tractable quantitative models for lipid transfer mediated by LTPs. We have developed an analytic theory for the lipid transfer process mediated by shuttle LTPs based on a simplified lipid exchange scheme. The theory is consistent with some experimental results and accounts for the widely observed transfer of lipids against their gradients via two nonexclusive mechanisms: selective lipid mixing between membranes of similar composition and partitioning between membranes of different composition. The theory also leads to many interesting predictions awaiting further experimental tests. First, we found that the initial lipid transfer rate often used to characterize an LTP does not represent an intrinsic property of the LTP and may not be compared among different LTPs tested under different experimental conditions. Instead, the lipid exchange rate constants better characterize the intrinsic properties of an LTP. Second, the lipid transfer rate is proportional to the LTP concentration. Finally, the lipid transfer rate depends on the relative concentrations of the two exchangeable lipids (
We proposed two potential driving forces for the bulk flow of lipids through bridge LTPs. Despite compelling evidence for VPS13 and ATG2 as bridge LTPs, direct support for the bridge model has been lacking. Improved assays will be required to monitor bulk lipid flow and the accompanying membrane area changes. These assays may need controlled canonical or osmotic membrane tension, membrane tethering, and osmotic pressure. In addition, new assays may be needed to better distinguish between shuttle and bridge mechanisms of an LTP. Some LTPs might transfer lipid as both shuttles and bridges, at least in vitro, depending upon experimental conditions. An N-terminal domain of ATG2 transfers lipids and can substitute the wild-type ATG2 in autophagy in the cell (Valverde et al., 2019). An ATG2 mutant that fails to tether membranes still transfers lipids, albeit with one-third of the activity of the wild-type (Maeda et al., 2019). These observations seem to corroborate a shuttle mechanism of lipid transfer by at least the mutant ATG2. Finally, the quantitative models for lipid transfer and membrane contact formation should be combined to simulate lipid transfer at the MCSs.
In conclusion, continuing discoveries of new LTPs and their biological functions call for quantitative models for lipid transfer and membrane contact formation regarding their kinetics and thermodynamics. These models will also improve our understanding of the thermodynamics of membranes and potential new mechanisms that govern organelle biogenesis and stability.
Footnotes
Acknowledgments
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Institute of General Medical Sciences (grant number R35GM131714).