Abstract
Lipid droplets (LDs) are the primary lipid-storage compartments of eukaryotes and feature a unique architecture distinct from membrane-bound organelles. Observed for over a century but initially ignored, recent cell biological, structural, and modeling studies provide new insights into how LDs are made, how they form connections with other organelles, and the roles they play in human physiology and disease. In this brief review, we highlight some key insights that helped define the LD and its roles in the cell. We also summarize new studies of molecular tethers that facilitate the LD interorganelle crosstalk and emerging technologies that provide key insights into the functions and organization of LDs.
Lipid droplets (LDs) are ubiquitous organelles and the primary lipid-storage compartments of eukaryotes. Unlike membrane-bound organelles that are encapsulated by a lipid bilayer, LDs feature a unique architecture defined by a lipid monolayer that segregates their lipid stores away from the cytoplasm (Walther et al., 2017; Henne et al., 2018; Olzmann and Carvalho, 2019). LD biogenesis is also remarkable in that nascent LDs emerge from the surface of the endoplasmic reticulum (ER), which donates the cytoplasmic leaflet of its own membrane for the creation of the LD phospholipid surface. Although classically defined as lipid-storage compartments, recent studies have revealed diverse and unexpected roles for LDs in signaling, cellular homeostasis, and development (Welte and Gould, 2017; Olzmann and Carvalho, 2019). LDs are also now respected as key mediators of adaptive responses to cellular and organismal stress, providing both a reservoir for excess lipids that may otherwise contribute to lipotoxicity and sources of energy-rich fatty acids (FAs) that may be harvested by oxidative organelles during times of energetic crisis.
LDs were originally observed by light microscopy in the late 19th century, and later by Keith Porter and others in the 1940s and 1950s using electron microscopy, although these sightings were initially ignored compared with other organelles (Altmann, 1890; Porter et al., 1945; Porter and Palade, 1957). Early biochemical work pioneered the identification of key enzymes that are necessary for creating the neutral lipids including triglyceride (TG) that are stored within LDs. However, the major site of TG synthesis was found to be the ER, and it gradually became clear that these lipids could subsequently be shunted into cytoplasmic oil droplet inclusions (Stein and Stein, 1968; Lutas and Zucker-Franklin, 1977).
While classic microscopy and enzymology continue to play key roles in the study of droplets, recent genetic, cell biological, and microscopy developments have provided new molecular insights into the mechanisms that govern LD biogenesis, maturation, and homeostasis. These studies include large-scale genetic screens, atomic-resolution structures, and cutting-edge microscopy approaches that characterize the molecular machinery regulating LD production, and how LDs interact with other organelles.
The purpose of this review is to briefly highlight just a few of these recent discoveries and how they have added to the molecular understanding of LD biogenesis and function. Due to space constraints not all findings can be covered, and the author apologizes in advance for omitted studies. For more comprehensive reviews on LD biogenesis, function, and roles in organismal physiology, the author directs the reader to other reviews (Welte and Gould, 2017; Henne et al., 2018; Olzmann and Carvalho, 2019).
Discovery and Characterization of LD-Associated Proteins
Following their initial observation by light and electron microscopy, LDs were relatively understudied compared with other organelles like the lysosome that became associated with digestive enzymatic activity. LDs were known to store the neutral lipid TG, whose enzymatic synthesis was characterized in the 1950s and 1960s, but whether other proteins coordinated LD biogenesis were unclear (Kennedy, 1957). The discovery and cloning in the early 1990s of two perilipin proteins, which copurify with LDs and coat the LD exterior, was a key finding that initiated a still-ongoing era of characterizing LD-associated proteins (Greenberg et al., 1991, 1993). In the early 2000s, the budding yeast Saccharomyces cerevisiae became an enabling model system for the identification of LD regulating proteins. The fact that LDs can be easily labeled with vital dyes such as BODIPY and Nile Red, combined with the commercial availability of the yeast knockout collection, allowed large-scale genetic screening for genes whose loss affected the morphology or the ability to produce LDs. Two such genome-wide screens were conducted by the Goodman and Yang labs and identified many genes whose loss affected LD morphology and abundance (Szymanski et al., 2007; Fei et al., 2008). Of particular note, both studies identified the yeast protein seipin/Sei1p/Fld1p, the homolog of human protein BSCL2, named for the lipodistrophy disease Berardinelli–Seip Congenital Lipodistrophy Type 2 that was initially characterized in the 1950s (Berardinelli, 1954; Seip, 1959; Magre et al., 2001; Figure 1). This protein is now extensively studied, and most researchers have settled on its name simply as seipin.

A Timeline of Significant Discoveries in the Study of LDs. Relevant events in the discovery and characterization of LDs are denoted on this timeline. Key references: 1890s: first observations of LDs (Altmann, 1890); 1954/1959: Berardinelli and Seip first describe BSCL2 disease (Berardinelli, 1954; Seip, 1959); 1991: perilipins isolated from LDs, perilipins 1 and 3 are cloned (Greenberg et al., 1991, 1993); yeast seipin characterized (Szymanski et al., 2007; Fei et al., 2008); structures of human and Drosophila seipin oligmers (Sui et al., 2018; Yan et al., 2018). LD = lipid droplet; ER = endoplasmic reticulum.
Although seipin-deficient yeast could still generate LDs, LD morphology was altered and LDs appeared as either small grape-like clusters or a single super-sized LD. Seipin was also identified as an ER-resident protein, further connecting LD homeostasis to the mother organelle the ER where most TG is synthesized. Subsequent studies in mammalian cells confirmed that mutations or genetic ablation of human seipin indeed altered LD morphology and proper LD biogenesis, although the synthesis of neutral lipids was itself not altered (Salo et al., 2016; Wang et al., 2016). Using live-cell imaging and fluorescence microcopy, seipin is now recognized as an ER-resident protein that directly interacts with nascent LDs and acts as a tether to promote LD maturation from the ER, although the molecular details of seipin’s role in LD biogenesis are still being resolved (Fei et al., 2008; Grippa et al., 2015; Han et al., 2015; Salo et al., 2016; Wang et al., 2016; Salo et al., 2019). A key insight from several of these studies was that seipin enriches at junctions between the ER surface and the maturing LD, indicating that ER-LD junctions serve as critical sites for proper LD growth (Grippa et al., 2015; Salo et al., 2016). This observation also indicated that LDs maintain contact with the ER for extended stages of their maturation and in yeast appear to maintain contact with the ER throughout their existence (Jacquier et al., 2011).
LD Tethers and Connections to Other Organelles
ER-LD Connections
The observation that seipin localized to ER-LD junctions reinforced the long-held belief that protein tethers could mediate direct contact between LDs and other organelles (Grippa et al., 2015; Han et al., 2015; Salo et al., 2016). Indeed, classic transmission electron micrographs from Porter and Palade indicated that LDs made intimate contact with the ER as well as with mitochondria, but the machinery that mediated these contacts was unclear (Palade, 1952). The past decade has seen an explosion of LD interorganelle contact site research, with specific tethers being identified that connect LDs to almost every other cellular compartment. In addition to seipin, other tethering proteins and complexes including the Rab18-NRZ-SNARE complex, DFCP1, Snx14, and Promethin can localize to ER-LD contacts and influence LD maturation (Figure 2; Datta et al., 2019; Li et al., 2019; Xu et al., 2018). Furthermore, the enzymes DGAT2 and FA transport protein 1 (FATP1), both of which contribute to TG synthesis, have been proposed to form heterodimeric complexes at ER-LD junctions to couple ER-LD tethering with local TG generation (Xu et al., 2012). Why cells encode so many ER-LD tethers is unclear, but studies indicate functional specificity for each tether. Like seipin, Snx14 is also an ER-anchored transmembrane protein that can interact with LDs in trans, but Snx14 appears to be recruited to LDs after their formation, suggesting it may function in LD growth (Datta et al., 2019). Snx14 knockout cells display perturbed LD morphology following addition of the FA oleate, and multitimepoint imaging reveals that Snx14 is recruited to LDs following oleate addition, consistent with Snx14 functioning in FA-stimulated LD expansion (Datta et al., 2019). Promethin is also an ER-associated protein that is recruited to seipin-positive LDs, suggesting it may regulate seipin activity at ER-LD contacts (Castro et al., 2019).
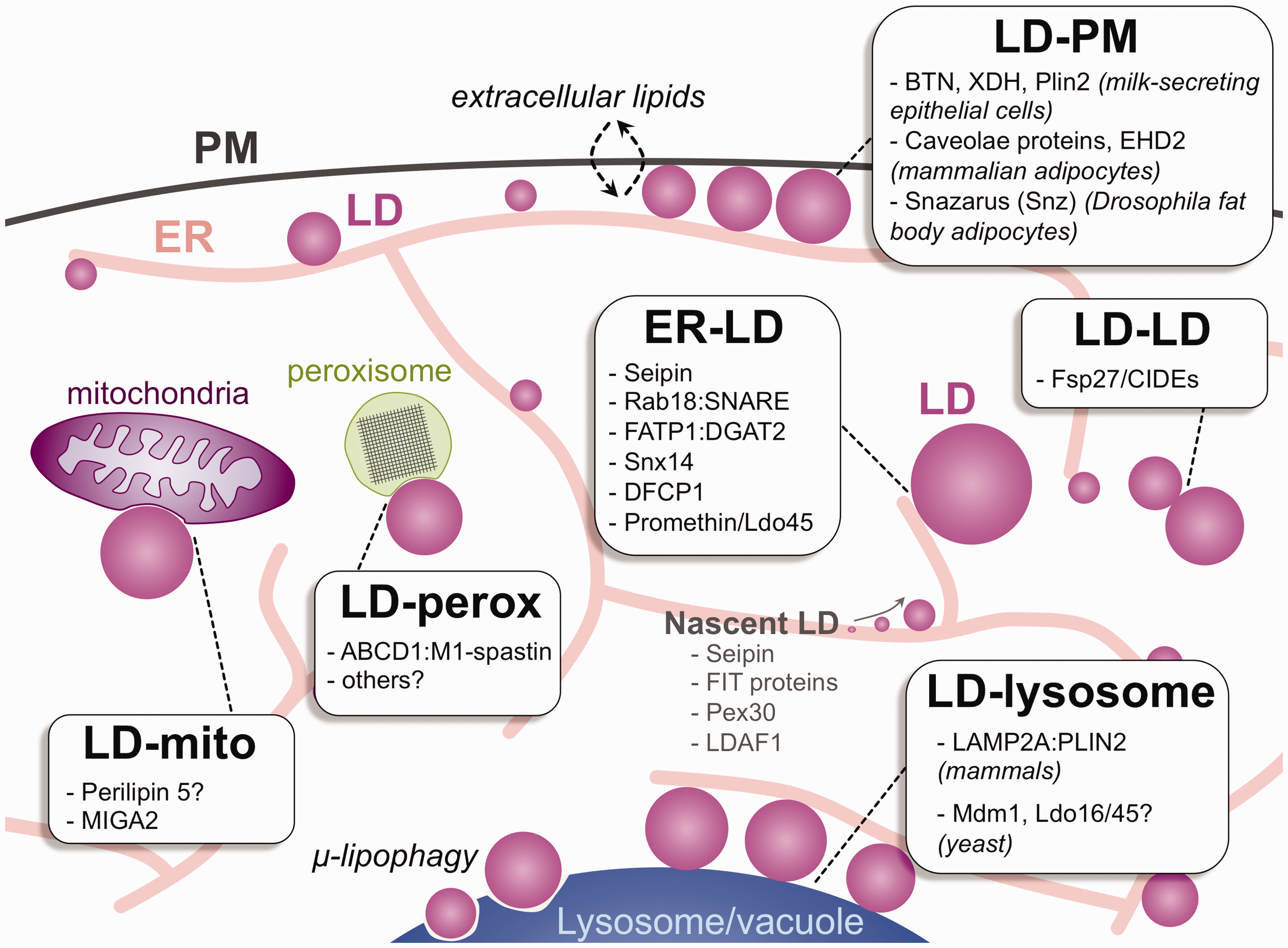
Interorganelle Contacts Involving LDs. Diagram of eukaryotic cell displaying numerous interorganelle contacts involving LDs. LDs are produced from the ER and can make contact with the mitochondria (mito), peroxisomes (perox), PM, other LDs, and lysosomes (the budding yeast vacuole). Proteins identified as being localized to each of these interorganelle junctions are denoted. Proteins that enrich at nascent LD lenses are also denoted. LD = lipid droplet; ER = endoplasmic reticulum; PM = plasma membrane; BTN = PM transmembrane protein butyrophilin; XDH = cytoplasmic protein xanthine dehydrogenase/oxidoreductase; Plin2 = perilipin 2.
LD Connections With Themselves and Oxidative Organelles
Similar to their connections to the ER, a few tethers connecting LDs to oxidative organelles have been identified. Perilipin 5 has been proposed to act as an LD-mitochondrial tether, but can also indirectly influence FA oxidation and mitochondrial biogenesis, so its role as a direct tether remains debated (Wang and Sztalryd, 2011). Very recent work with white adipocytes indicates that MIGA2, a mitochondrial surface protein, can directly interact with the LD surface via a LD-targeting helix. In line with this, MIGA2 overexpression in adipocytes enhances mitochondria-LD contacts (Freyre et al., 2019).
FAs can also be oxidized by peroxisomes, although the molecular tethers this facilitate LD-peroxisome contacts have been unclear. Recent work proposes that the peroxisomal FA transporter ABCD1 can bind to M1 spastin, an AAA ATPase associated with spastic paraplegia that binds to the LD monolayer surface, thus establishing a heterodimer that stabilizes LD-peroxisome contacts (Chang et al., 2019). As ABCD1 is a peroxisome FA importer, this implies that LD interorganelle tethers may facilitate the lipid exchange between LDs and other cellular compartments. In yeast, LDs and peroxisomes have been observed to form extensive interorganelle contacts and potentially even coalesce or fuse together, although the yeast proteins that facilitate this remain unclear (Binns et al., 2006). LDs can also sometimes fuse with other LDs, and the Fsp27/CIDE protein family has been proposed to act as LD-LD tethers that regulate this homotypic fusion and lipid exchange between LDs (Gong et al., 2011; Jambunathan et al., 2011). Identifying additional tethers that connect LDs to each other and to oxidative organelles remains a major question in the field.
LD-Lysosome Contacts
LDs can also be delivered into lysosomes to be digested in order to harvest FAs for oxidative metabolism or membrane biosynthesis. In mammals, LD-lysosome contacts formed between the lysosome protein LAMP2A and the perilipin Plin2 can facilitate the uncoating of LDs for subsequent LD breakdown in chaperone-mediated autophagy (Kaushik and Cuervo, 2015). In budding yeast, LDs are directly delivered into the vacuole (the yeast lysosome) during prolonged nutrient starvation by micro-autophagy on the vacuole surface, a process known as µ-lipophagy (Wang et al., 2014; Seo et al., 2017). This requires the formation of visually striking sterol-containing liquid-ordered domains on the vacuole membrane surface, which serve as platforms for the docking and eventual envelopment of LDs (Toulmay and Prinz, 2013; Seo et al., 2017). Several proteins appear to be involved in the targeting of LDs to the vacuole for µ-lipophagy. The process is dependent on many of the autophagy-related genes machinery (Wang et al., 2014; Seo et al., 2017). The interorganelle tethering protein Mdm1 may also contribute to LD delivery into the vacuole. Mdm1 is an ER-anchored tethering protein that localizes to the nucleus–vacuole junction (NVJ) by binding the phospholipid PtdIns(3)P on the vacuole in trans (Henne et al., 2015). It can bind directly to LDs that accumulate at the NVJ prior to µ-lipophagy (Hariri et al., 2018). Intriguingly, NVJ-associated LDs are also decorated with LD organizing proteins Ldo16 and Ldo45, which promote LD clustering at the NVJ. These proteins also interact with yeast seipin, indicating they play some role in LD biogenesis (Eisenberg-Bord et al., 2017; Teixeira et al., 2017). Remarkably, Mdm1 and Ldo16/45 are all conserved in metazoans. One human Mdm1 homolog is Snx14 that localizes to ER-LD contacts during LD growth (Datta et al., 2019). The human Ldo16/45 homolog is Promethin, which also localizes to ER-LD contacts and interacts with seipin (Castro et al., 2019).
LD-Plasma Membrane Contacts
A more recently appreciated interorganelle contact involving LDs is with the cell surface, and this may influence lipid exchange between LDs and the extracellular space. Milk-producing epithelial cells contain cortical LDs that dock with the plasma membrane (PM) prior to their secretion, and it is thought that these LD-PM contacts are stabilized by a protein complex containing the PM transmembrane protein butyrophilin, the cytoplasmic protein xanthine dehydrogenase/oxidoreductase, and perilipin Plin2 (Figure 2; McManaman et al., 2002; McManaman, 2012). LD-PM contacts have also been observed in adipocytes. Mammalian white adipocytes are coated with surface caveolae that stud the PM and influence receptor organization and lipid absorption (Scherer et al., 1994; Parton, 2018). Caveolae also appear to directly interact with LDs. During elevated lipolysis, small LDs accumulate adjacent to the caveolae of primary human adipocytes, implying that LD-PM contacts somehow influence lipid mobilization (Moren et al., 2019). Indeed, caveolae have long been proposed to serve as metabolic platforms for lipid exchange across the PM and facilitate cholesterol absorption as well as harbor TG stores (reviewed in Ortegren et al., 2007). Consistent with this, FATP1, FA translocase CD36, and FATP4 have all been observed at caveolae, suggesting FA absorption and processing locally occurs at these sites (Vistisen et al., 2004; Ost et al., 2005). As FATP4 is ER localized, this suggests a close functional coupling of the PM and cortical ER in lipid processing and TG storage, although this needs to be more thoroughly investigated (Milger et al., 2006).
Recent work in Drosophila indicates a close functional coupling between the ER, PM, and LDs in adipocyte fat storage. In fruit flies, the primary fat-storing organ is the fat body, a multicellular tissue composed of hundreds of adipocytes each filled with LDs (Arrese and Soulages, 2010). Intriguingly, fat body adipocytes contain LDs that are spatially segregated according to size; there is a clear population of small peripheral LDs (pLDs) adjacent to the cell surface and larger LDs further within the cytoplasm (Ugrankar et al., 2019). The pLDs appear to be a specialized LD subpopulation that makes direct contact with the PM. Remarkably, Drosophila do not encode caveolin genes to generate cavaeolae, but their adipocytes nevertheless form distinct PM invaginations that surround and contact pLDs. Fasting induces the loss of pLDs from larval fat body tissue, whereas the more intracellular LDs are not affected, suggesting the pLD population is preferentially mobilized to supply lipids for extracellular lipid trafficking (Ugrankar et al., 2019). Furthermore, ablation of de novo lipogenesis within the fat body perturbs the larger cytoplasmic LD pool but not pLDs, supporting a model where pLDs are maintained primarily by extracellular lipid trafficking.
How are these surface-associated pLDs maintained? The Drosophila protein Snazarus (Snz), a homolog of yeast ER-vacuole tether Mdm1, has been implicated in tethering pLDs to the cell surface. Originally characterized as a longevity-associated gene (Suh et al., 2008), Snz is an ER integral membrane protein and localizes to ER-PM contacts in Drosophila adipocytes and colocalizes with the ER-PM biomarker MAPPER (Ugrankar et al., 2019). Snz also directly binds to LDs, suggesting it influences pLD homeostasis. Consistent with this, Snz knockout flies are viable but display defects in pLD morphology as well as altered TG storage. Conversely, tissue-specific Snz overexpression in the fat body elevates TG storage, starvation resistance, and extends fly life span. Collectively, this suggests that Snz-mediated crosstalk between the PM, ER, and LDs can alter lipid storage as well as organismal homeostasis and longevity (Ugrankar et al., 2019). Further studies are needed to understand the crosstalk between the PM, ER, and LDs, but this organelle trio appears to be important for coordinating lipid absorption, processing, and storage in adipocytes.
Structural Insights Into Seipin
Recent structural studies of seipin are revealing the molecular details into how it contributes to LD biogenesis at the ER. Two studies have revealed cryo-electron micrograph structures of the oligomeric forms of human and Drosophila seipin (Sui et al., 2018; Yan et al., 2018). Both structures reveal seipin oligomers as rings with a central pore. Intriguingly, the two structures reveal different oligomeric stoichiometries: Human seipin rings are undecamers (11 monomers), whereas Drosophila seipin is a dodecamer. The structures also reveal a hydrophobic helical region in the luminal domain of seipin that targets to LDs when expressed in vitro (Sui et al., 2018). The luminal domain exhibits a protein fold similar to the cholesterol-binding protein Niemann Pick Type 2, although the role of this domain in lipid binding remains unclear. Interestingly, purified seipin was also found to bind to phosphatidic acid (PA) in vitro, suggesting it may play a role in organizing PA in the ER prior to its processing into TG for storage in LDs (Yan et al., 2018). As both structures lack high-resolution structures of the seipin transmembrane region, future structure analysis of this protein will no doubt reveal further molecular details into its function.
Based on this new structural information, an emerging model for seipin function is that seipin monomers or oligomers move across the ER network, surveying for lipids. When TG synthesis pools are encountered, ring-like seipin oligomers assemble and promote the coalescence of TG into a lens that can then matures into a nascent LD (Bohnert, 2018). New work on seipin-associated proteins supports this model. LDAF1 forms a cocomplex with seipin in the ER membrane and stabilizes TG pools and LD budding (Chung et al., 2019). An alternative model is that seipin primarily influences PA conversion into diacylglyceride (DG) at the ER prior to the eventual transfer of TG into LDs, although these models are not mutually exclusive. In addition, seipin likely works collaboratively with ER-anchored fat storage-inducing transmembrane (FIT) proteins to achieve this, and FIT proteins also appear to interact with DG at ER subdomains prior to DG processing into TG (Choudhary et al., 2015). Seipin may also help to maintain contacts between the ER and LD so that TG can efficiently be loaded into LDs during their maturation and growth (Salo et al., 2019).
Microscopy and Modeling Innovations in the Study of LDs
Recent developments in microscopy, biochemical approaches, and molecular modeling have also contributed greatly to our understanding of LD biogenesis and the contacts LDs make with other organelles. The advent of multichannel live-cell fluorescence imaging has allowed multiple organelles to be simultaneously imaged in living cells. This has revealed that LDs are in close proximity to essentially every other organelle in the crowded cytoplasm and can form intimate contacts with these compartments during cell stress and growth (Valm et al., 2017). A current challenge is to apply this methodology to specific cell types such as neurons, which display unique organelle arrangements in line with their unique functions. Raman scattering microscopy allows the identification of specific lipid species within living cells and has recently been adapted to monitor the composition of neutral lipids inside LDs (Fu et al., 2014). Similarly, electron tomography has also revealed new details into the molecular organization within LDs. Striking cryo-electron tomography imaging of LDs in ambiently cultured and metabolically stressed mammalian cells has revealed that cholesterol esters can undergo liquid-crystalline phase separation inside LDs during specific metabolic cues (Mahamid et al., 2019). The physiological reason for this lipid segregation is still unclear but suggests that the LD interior may elicit a previously unappreciated level of subcompartmentalization during nutrient stress.
Emerging biochemical techniques are similarly providing new molecular insights into the protein composition of LDs and their connections with other organelles. Using the proximity-based labeling APEX (engineered ascorbate peroxidase), which facilitates the biotinylation of proteins so they can be subsequently identified by mass spectrometry-based proteomics, elegant studies have revealed the detailed proteonome of the LD surface (Bersuker et al., 2018). This has revealed expected proteins such as the perilipins in addition to key enzymes involved in lipid synthesis and ER homeostasis, suggesting the LD surface is rich in numerous metabolic reactions critical to cell homeostasis. In line with this, transmission electron microscopy-based studies using APEX fusions to ER-LD tethering proteins like Snx14 and seipin reveal that the ER can form extensive contact with LDs, potentially creating a unique microenvironment at the ER-LD interface important for lipid synthesis reactions (Cao et al., 2019; Datta et al., 2019). These studies highlight the intimate relationship the ER has with LDs and how these two organelles can maintain functional contact with one another throughout LD maturation.
In situ molecular modeling is also revealing new mechanisms governing LD biogenesis and maturation. A series of elegant studies from Abdul Thiam’s group has been able to combine molecular modeling with giant unilamallar vesicle-based biochemistry to probe the molecular mechanisms governing the demixing of lipids in bilayers and the emergence of artificial LDs from membranes. An overarching conclusion is that asymmetry in the tension of membrane bilayers can contribute to the initial formation and emergence of LDs (Chorlay and Thiam, 2018). Furthermore, this membrane asymmetry can be modulated by differences in phospholipid abundance or composition between the two leaflets, providing directionality to LD emergence and maturation. This provides a working model for how LDs tend to grow into the cytoplasmic leaflet of the ER as they mature (Chorlay et al., 2019). Collectively, these studies provide a mathematical and experimental framework to reconstitute and dissect LD formation in a membrane bilayer and provide testable models to dissect the relative contributions of membrane composition, tension, and proteins to LD biogenesis.
Conclusions
In closing, over the past few decades, our advancement in the understanding of the LD, from an inert lipid inclusion to a well-regulated organelle critical to cellular homeostasis, has come a long way. We are clearly in a molecular era of LD biology, where key proteins that govern LD assembly, turnover, and interactions with other organelles are continuing to be identified and characterized. New technologies and approaches will continue to reveal unexpected roles for the LD in cellular homeostasis and adaptions to stress. Several major questions remain. What dictates the separation of mature LDs from the ER? How are different LD subpopulations within a cell distinguished from one another and differentially maintained and utilized by the cell? How do LDs contribute to organismal homeostasis and aging? Aberrant accumulations of LDs in neuron-associated astrocytes and other cell types are now well recognized in disease states such as neurodegeneration and Alzheimer’s diseases (Liu et al., 2017); how do LDs relate to these neurological diseases? Do they mostly represent a consequence of pathology or the cell’s attempt to respond to the disease? New imaging approaches, reconstitution assays, and atomic structures of LD-associated proteins will no doubt provide new mechanistic details into how LDs are made and interact with lipids and organelles. The molecular era of the LD will continue to excite and surprise us with new discoveries.
Footnotes
Acknowledgments
The authors want to thank Joel Goodman and members of the Henne and Goodman lab for helpful advice in the preparation of this review.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Welch Foundation (I-1873) and National Institute of General Medical Sciences (GM119768).
