Abstract
Methodological developments and software implementations are progressing at an increasingly fast pace. The introduction and widespread acceptance of preprint archived reports and open-source software have made state-of-the-art statistical methods readily accessible to researchers. At the same time, researchers are increasingly concerned that their results should be reproducible (i.e., the same analysis should yield the same numeric results at a later time), which is a basic requirement for assessing the results’ replicability (i.e., whether results at a later time support the same conclusions). Although this age of fast-paced methodology greatly facilitates reproducibility and replicability, it also undermines them in ways not often realized by researchers. This article draws researchers’ attention to these threats and proposes guidelines to help minimize their impact. Reproducibility may be influenced by software development and change over time, a problem that is greatly compounded by the rising dependency between software packages. Replicability is affected by rapidly changing standards, researcher degrees of freedom, and possible bugs or errors in code, whether introduced by software developers or empirical researchers implementing an analysis. This article concludes with a list of recommendations to improve the reproducibility and replicability of results.
Novel statistical methods are being developed, implemented, and applied at an increasingly fast pace. Two recent trends may in part underlie this increase: (a) the use of preprint archives, which make methodological research directly and openly available before work is peer reviewed, and (b) the availability of relatively easy-to-use and powerful open-source programming languages, such as R and Python, which allow researchers with limited programming experience to use new statistical methods. At the same time, there are growing concerns regarding the trustworthiness of the conclusions drawn from psychological studies (Open Science Collaboration, 2015), that is, whether the same conclusions would be drawn if the data were reanalyzed or similar data were collected and analyzed again at a later time. Although the age of fast-paced methodology greatly facilitates the ability to reexamine results over time (e.g., materials are now often shared on the Open Science Framework), it also undermines this ability in ways that are not often realized by researchers. In this article, I attempt to make researchers aware of these threats and offer guidelines to minimize them.
First, I outline threats the fast-paced methodological world poses for the reproducibility and the replicability of results and suggest ways in which researchers can limit those threats. I use the term reproducibility to refer to whether the original numeric results can be exactly reproduced given the same data and analysis code or process used by the original author. I use the term replicability to refer to whether the original conclusions can again be drawn when the data are reanalyzed or new data are analyzed. 1 Reproducibility, therefore, is a basic requirement for replicability; if results are not reproducible, we cannot expect them to be replicable. In this article, I focus primarily on analyses performed using the statistical programming language R—because of its popularity among psychological researchers and because it is one of the main platforms that researchers use to develop new methods—but the arguments also extend to other software frameworks (e.g., Python). Although in principle reproducibility can be achieved, some threats to replicability will be almost impossible to overcome. I conclude this article with recommendations for ways in which methodologists and software developers can aid researchers in achieving reproducibility and replicability.
Disclosures
The code used in all analyses in this article is stored at the Open Science Framework (https://osf.io/u8sdh/). The files there include supplementary materials explaining how this code was adapted to ensure long-term reproducibility; these materials are also available at http://journals.sagepub.com/doi/suppl/10.1177/2515245919847421.
Reproducibility
Reproducibility of results refers to the ability to obtain the exact same numeric results at a later time, using the same data. Reproducibility of results is crucial for two reasons: (a) Reproducible results indicate that the reported numeric results are accurate and (b) allow researchers to use the same analysis procedure in future research, either to build on the analysis methods or to attempt to replicate the conclusions drawn in the study. Note that errors or suboptimal choices in an original analysis do not make that research nonreproducible as long as those errors or suboptimal choices themselves are reproducible. In this section, I outline several threats to reproducibility even when the original analysis code and data are shared, and I conclude with a list of recommendations for improving reproducibility.
Threats to reproducibility
Beta versions
Software requires a lengthy period of development before it may be used. Before software is deemed stable, it is typically referred to as being in beta version, a status that is commonly communicated by starting the version number with 0. For example, the dplyr R package (Wickham, Francois, Henry, & Müller, 2017) is, at the time of this writing, in Version 0.7.8. From left to right, the numbers in a version number usually identify the version, major update, and minor update. Although not all developers use the same numbering system, it is common that the further down a number is in the version number, the less important it is. A version number starting with 0 often indicates that the developer does not yet deem the software stable enough to call it a “released” version; input, output, default arguments, and so forth are still open to change. Consequently, future versions may not reproduce the same exact analysis that the current version does. Open-source software developers often keep their software in beta version for a long time. This point is illustrated in Figure 1, which shows that beta versions constitute a substantial proportion of the rapidly increasing number of contributed R software packages. 2 Naturally, researchers should be extra careful when using packages that are in beta version (or packages that rely on beta-version packages).
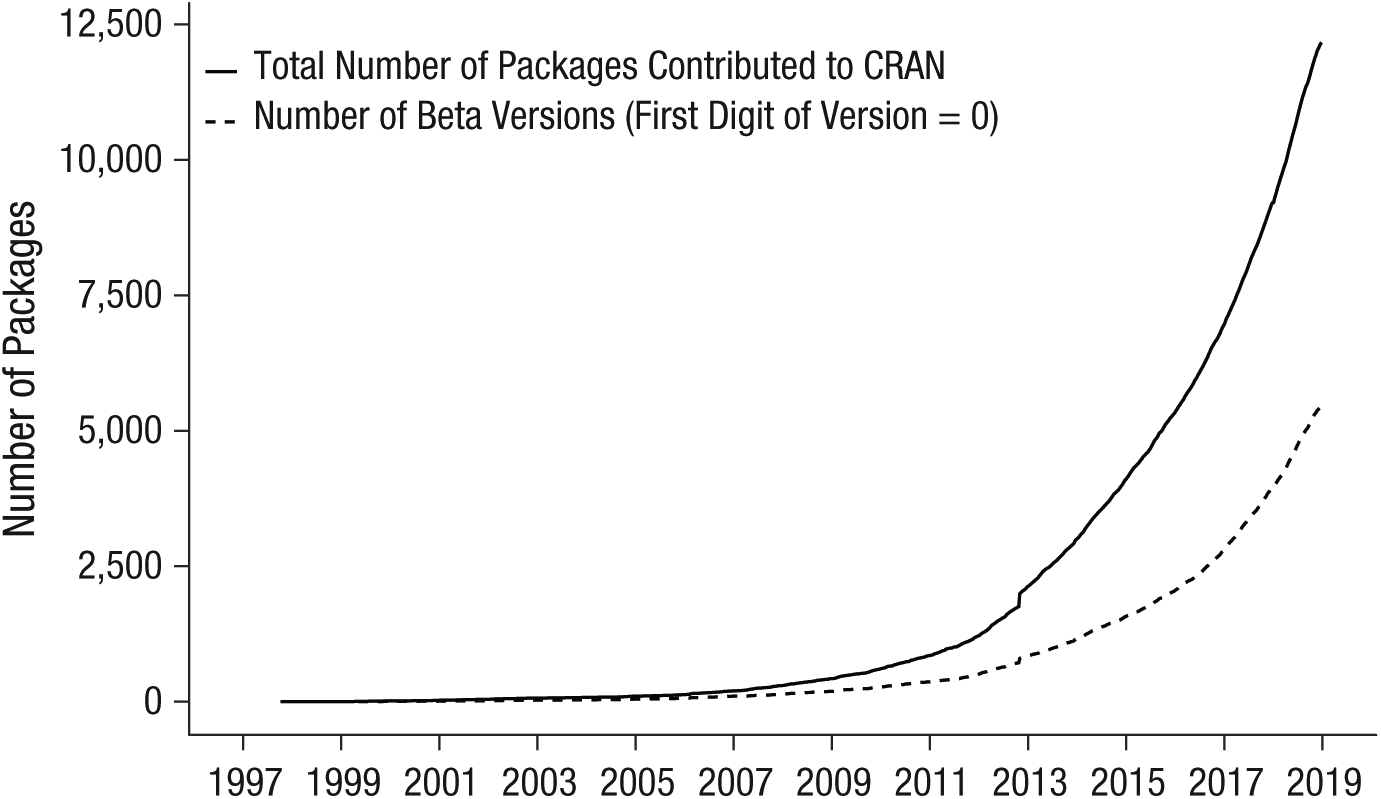
Number of R packages contributed to the Comprehensive R Archive Network (CRAN) as a function of year, up to 2019.
Updates and dependencies
Naturally, a fast-paced methodological world implies that even stable methodologies may change. New research and developments may change the best choices for default arguments or improve estimation methods. In addition, even stable versions of software releases may have bugs that are not yet resolved or newly introduced (see the discussion of bugs in the section on replicability). Results may no longer be reproducible in newer versions of the software originally used. These potential problems are greatly magnified by software packages’ large and intricate dependency trees. For example, Figure 2 shows the dependency tree of the bootnet package (Epskamp, Borsboom, & Fried, 2018), which relies on 140 different packages directly or indirectly. Changes or bugs in any of these other packages may lead to new bugs or different results in the bootnet package. Because of their large dependency trees, R software packages are notorious for their poor backward compatibility. Even if a research report states which version of a software package was used, this does not guarantee reproducibility because the dependencies of the package may have changed.
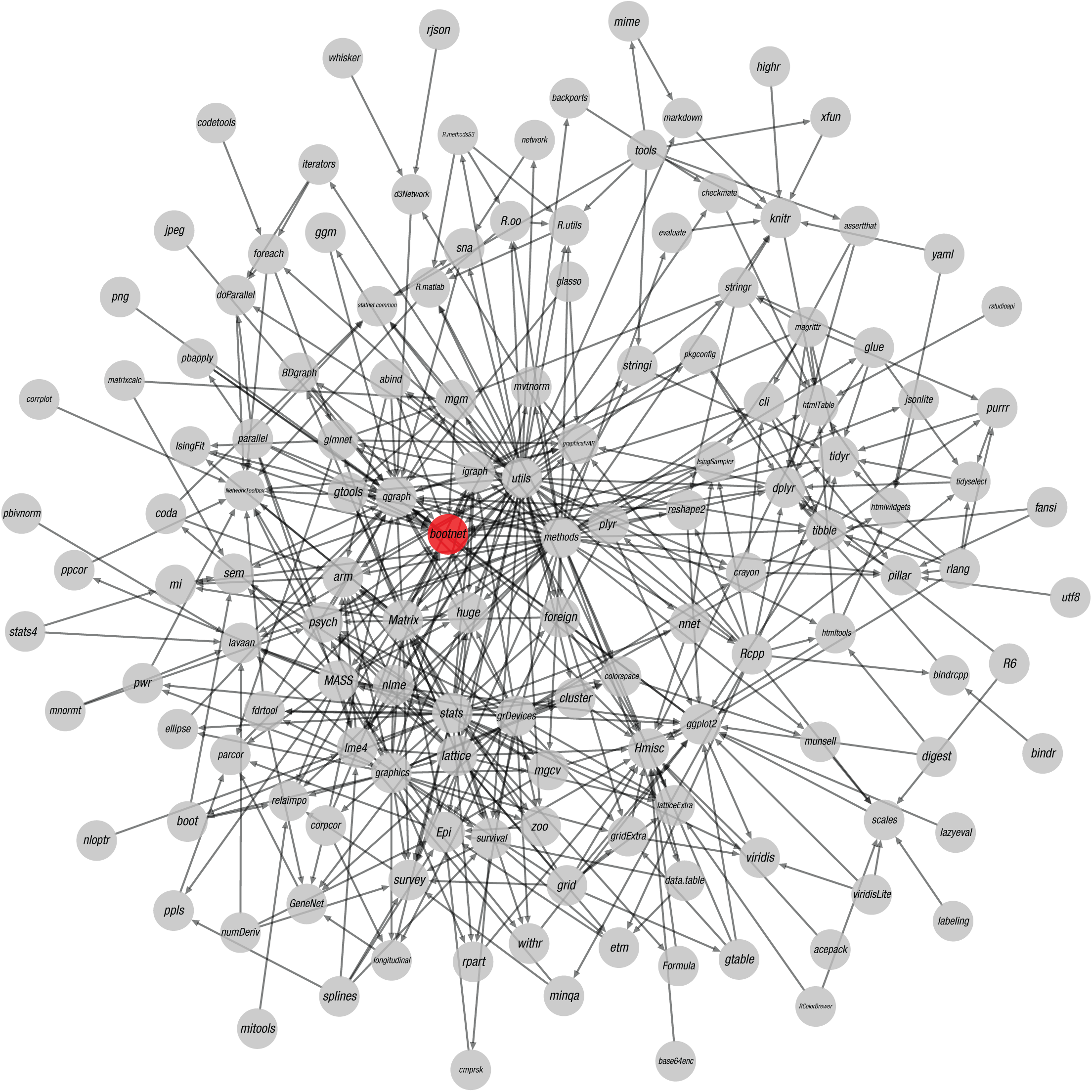

The dependency network of packages required by the bootnet package for R. This visualization was created using the qgraph package (Version 1.6; Epskamp, Cramer, Waldorp, Schmittmann, & Borsboom, 2012).
Recommendations to improve reproducibility
Here, I list several guidelines that can help empirical researchers achieve long-term reproducibility of results. Following these guidelines may also allow future researchers to replicate results using new data. Some of these guidelines are well known and have been tested and discussed in prior work (e.g., Piccolo & Frampton, 2016), but I have included them in order to provide a complete overview.
Use crystallized software if possible
When empirical researchers apply new software routines that implement novel statistical methods, they should consider that the likelihood of changes in a software package’s inputs and outputs increases the less crystallized the software is, and such changes will affect the reproducibility of results (and may also affect replicability, as I discuss later). To help researchers judge the level of crystallization of software packages, I present a checklist in Table 1. Many software packages will not meet these criteria; therefore, it is not feasible to require that all the criteria be met.
Checklist for Determining if Software Is Crystallized and Not Likely to Change
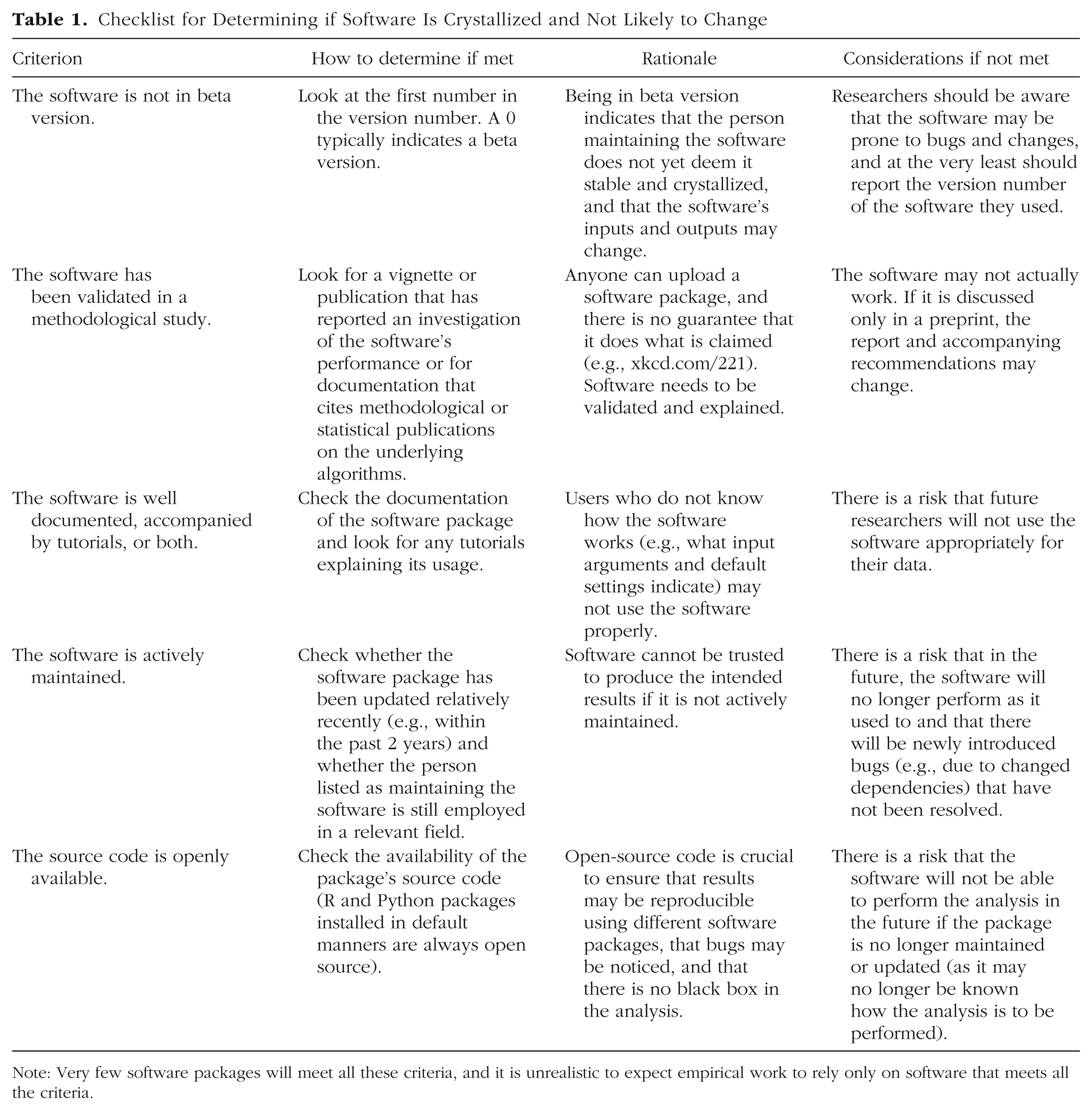
Note: Very few software packages will meet all these criteria, and it is unrealistic to expect empirical work to rely only on software that meets all the criteria.
Provide analysis code and data together with the publication
In order for other researchers to reproduce analyses, it is vital that analysis code be made available together with the publication. Preferably the code would be accompanied by the data or a summary of the statistics needed from the data in order to run the main analyses. It is not sufficient to merely report the analysis used because different software packages may make different choices in the default steps of the analysis (e.g., an analysis of variance will lead to slightly different results with default SPSS options than with default R options). Even if an analysis no longer is reproducible, because of changing software code, investigating the steps taken by the researchers to arrive at their results still provides insight. If a software program with a graphical user interface, such as SPSS or JASP (Wagenmakers, Love, et al., 2018; Wagenmakers, Marsman, et al., 2018), is used to perform the analyses, it is best to supply the analysis syntax (if available) or a saved file that contains the analyses. Alternatively, a researcher may opt to simply provide screenshots of the selected menu items in addition to the version number of the software package. It is advisable to store code in an online repository, such as the website of the journal in which the research is reported (i.e., supplementary materials), the Open Science Framework (osf.io), GitHub (github.com), or some other repository. These repositories are likely to be available for a longer period, compared with a personal website.
Allow researchers to reproduce the state of software packages used
Supplying code so that analyses can be reproduced is a first step to ensure reproducibility, but it is not a guarantee (by itself) because of changing software packages and dependencies. At the very least, researchers should supply the version number of the software they used; in many cases, if the software has been updated, that older version can still be downloaded. For R code, supplying the output from the sessionInfo() command in supplementary materials will provide a good overview of the R version, machine, and package versions that were used.
3
Subsequently, the install_version function from the devtools package (Wickham, Hester, & Chang, 2019) can install the outdated software package. However, dependencies may not revert to their previous versions, so the outdated version may not work properly. The checkpoint package (Microsoft Corp., 2018) can install R packages as if they were installed on a particular date, by relying on daily snapshots of the Comprehensive R Archive Network (CRAN). For example, the following code forces packages to be installed as though it were February 4, 2019: library("checkpoint") checkpoint("2019-02-04")
This code will check the current working directory and all subdirectories for packages used and restore them to the versions they were on the checkpoint date (in this case, February 4, 2019). This command applies only to the current R session; restarting R will bring back the previously installed versions. Researchers using R can also apply the packrat package (Ushey, McPherson, Cheng, Atkins, & Allaire, 2018), which is integrated in the RStudio platform and can store the source code of all package versions used at the time of analysis, thereby allowing users to recompile these packages when they open a project that uses packrat. (RStudio, 2013–2014, has provided a tutorial for packrat at https://rstudio.github.io/packrat/.)
The solutions I have mentioned are specific to R and work from within R. The R architecture is not changed and does not revert back to a previous iteration, which may be problematic if an outdated R package is not supported on the current R version. In addition, these methods will not allow users to reproduce the state of external programs used in an analysis. A more general solution is to perform the entire analysis on an emulated computer, termed a virtual machine (Piccolo & Frampton, 2016), and to share the image of that virtual machine such that other researchers can boot up an exact copy of the computer used at the time of analysis. For example, the VirtualBox program (Oracle, 2019) can be used to run a virtual computer with a Linux distribution (e.g., Ubuntu), in which users can install all software needed to perform their analyses. Such a virtual machine, however, can be complicated to run and is large in size. A simplified solution, which has grown popular in computer sciences, is to use a more lightweight container, such as Docker (Boettiger, 2014; Merkel, 2014) 4 or Singularity (Sochat, Prybol, & Kurtzer, 2017), both of which can function as a lightweight virtual machine that creates a snapshot of software used at the time of analysis.
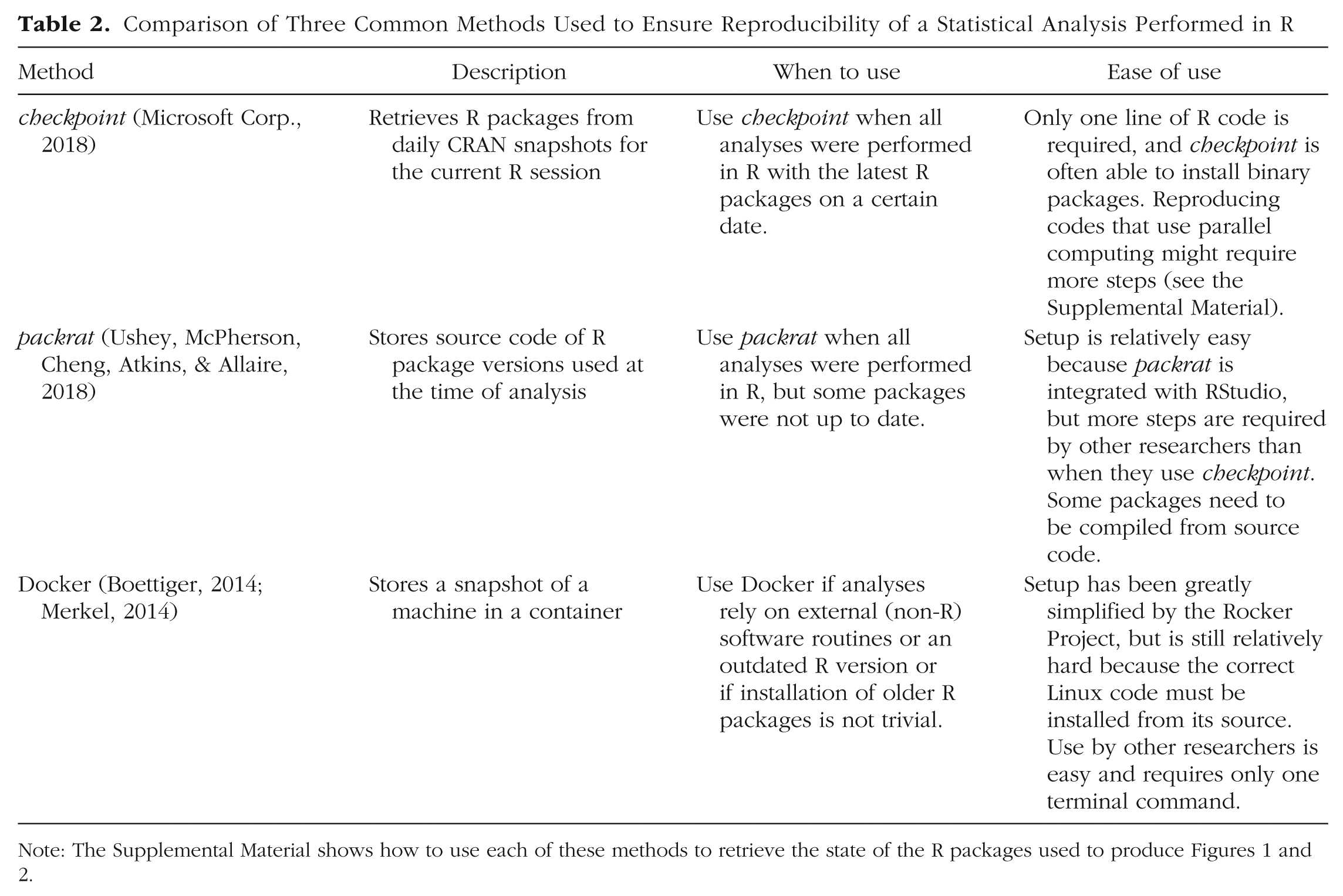
The R code to reproduce Figures 1 and 2, using the checkpoint package, the packrat package, and Docker, is available on the Open Science Framework (https://osf.io/u8sdh/). The Supplemental Material describes how these packages can be loaded and how the Docker container was constructed. 5 Table 2 provides further information on these three solutions, including their ease of use. Note that these solutions should be seen only as ways to reproduce old results and are not recommended for new research. Likewise, an outdated R package should be installed only to reproduce results, and the outdated package should be updated again to the newest version after the analysis has been completed. (When using checkpoint, make sure to restart R before performing new analyses.) Although, for example, Docker can simulate a 10-year-old machine, performing a new analysis on that machine entails ignoring 10 years of methodological progress. During those years, software may have changed for good reasons, for example, to solve bugs in the code, to implement the latest state-of-the-art methods, or simply to adapt to newly accepted standards. If a study is reproducible only under a certain snapshot of software, it is very likely not replicable, which brings me to the next topic.
Comparison of Three Common Methods Used to Ensure Reproducibility of a Statistical Analysis Performed in R

Replicability
Although reproducibility of numeric results may be ensured even in light of rapidly changing software packages, the replicability of results relying on novel methodologies may be harder (if not impossible) to ensure. In this section, I discuss several threats to the replicability of results and propose some recommendations to limit the impact of these threats. As noted earlier, I use the term replicability to refer to whether the original conclusions can be supported at a later time, such as by analyzing the original data using updated techniques. If analysis of the original data according to current standards does not support the prior conclusions, it is unlikely that new data will support those conclusions.
Threats to replicability
Implementation may not be straightforward
The most important methodological advances are found in publications in methodological and statistical journals, in which the methodology is substantiated through mathematical work and simulation studies. Not all such publications come with openly available software code that implements the analyses. Presenting a novel statistical method solely through mathematical representations essentially leads to a reproducible methodology, but it is often not straightforward or easy to duplicate these analyses. It is also common for details to be missing in the manuscript, and this results in ambiguity with respect to implementation. When a method is not implemented exactly in the intended way, there is no guarantee that the method will perform adequately. Furthermore, a subsequent correct implementation of the method may lead to different results that may no longer support the conclusions based on the original analysis.
Bugs and unexpected results
Methodologists typically lack formal training in programming and engineering software. As a result, the software supplied alongside methodological studies may contain bugs or lead to unexpected results when a user inserts unexpected input. These problems are further compounded when only one individual researcher actively developed or maintains the software. In addition, it is common for peer-reviewed research to remain in the publication process for a long time after the original submission (e.g., 1 year), but it is not uncommon for researchers to change institutes or work priorities. Methodological articles may therefore be published even after the main authors of the corresponding software packages have changed positions. Furthermore, even when the main authors have not changed positions, they may have limited time available for work outside their active research and teaching duties. As a result, the package may not be actively maintained. When packages are not actively maintained, new state-of-the-art methods or better default choices may not be implemented in them, and newly introduced bugs may not be resolved. This problem can be greatly amplified when several software packages rely on one another, as I discussed earlier.
Mistakes in the analysis
Not only may software contain bugs, but also the analysis used by empirical researchers may be erroneous, which may lead to results that can be replicated only when the same mistakes are made. For example, Nuijten, Hartgerink, van Assen, Epskamp, and Wicherts (2016) found that a substantial proportion of reported p values might be inaccurate. If novel methodology is implemented in software at all, it might be implemented only in a not-so-user-friendly software package. Furthermore, this novel methodology might not be clearly described in software documentation, tutorials, or textbooks. Applied researchers implementing such novel methods might therefore make mistakes, and analyses conducted after resolving these mistakes might not support the original conclusions.
Researcher degrees of freedom
When performing a relatively novel analysis, empirical researchers must make several choices regarding both how to perform the analysis and how to interpret the results. However, the particular estimation technique that works best in a particular setting or how to interpret results may not yet be established. Such researcher degrees of freedom (Simmons, Nelson, & Simonsohn, 2011) may be common when methodology is so new that detailed aspects of it have not yet been thoroughly investigated. Researcher degrees of freedom may affect replicability in two ways: (a) Certain results may be replicable only under the specific choices made by the original researcher and not be replicable when different (but not necessarily worse) choices are made, and (b) subsequent research may show that the original methodological choices were suboptimal, and results may not be replicated when future state-of-the-art methods are used. A possible remedy for the first problem is to perform a multiverse analysis (Steegen, Tuerlinckx, Gelman, & Vanpaemel, 2016). However, such an analysis cannot overcome the second problem: The results may still not be replicated when future state-of-the-art methods are used.
Limited validation studies
Researchers typically validate novel methods using both mathematical proofs and simulation studies, relying on certain assumptions or validating performance only in certain settings (e.g., “asymptotically, the estimation will converge to the true model, given that the assumptions are met”). For example, a methodology might be validated only under the assumption of multivariate normality, a limited number of variables, no missing data, data missing only at random, or a certain structure of the true model. It is unreasonable to expect researchers to immediately investigate all possible violations of assumptions when introducing novel methods or to investigate all possible forms of data on which the methods can be applied. Violations of assumptions, however, are likely common in empirical data sets, and data sets can differ vastly from one another in many ways. Therefore, even if a method is validated, it may not perform as expected on a particular data set, which introduces the same threat to replicability I discussed earlier: Analyses using future state-of-the-art methods may not replicate the original conclusions.
Preprint reports
It is now common for research articles to be shared online via preprint archives—even before these articles are peer reviewed. Although this process greatly facilitates the use of novel methods (e.g., a tutorial explaining pros and cons of a methodology can be shared and freely accessed through a preprint archive long before it is published), there is a drawback. Anyone can upload a manuscript to a preprint archive, and the people maintaining the archive typically do not review a manuscript beyond checking for basic readability (e.g., is an abstract available, and does the file compile?). Thus, though manuscripts on preprint archives may appear visually similar to published articles, they may not have been read or checked by external experts. It is not expected that the main messages of an empirical manuscript will change much when it is peer reviewed or further adapted for publication. Ideally, authors carefully consider hypotheses and analysis plans before gathering data, and it may be infeasible to collect a completely new data set after peer review. Methodological work, however, may change vastly during the review process. Reviewers may suggest rerunning analyses, adding simulation conditions, or changing descriptions of how results should be interpreted. As a result, the eventual publication or an update to a preprint may contain strikingly different recommendations regarding when and how to use the method. Indeed, this is the fundamental purpose of the peer-review process. Empirical work based on an early version of a preprint may contain outdated analyses, or the conclusions may no longer be relevant. Published peer-reviewed manuscripts come with the assurance that external experts have judged them and that they are no longer expected to change (with the exception of necessary errata).
Recommendations to improve replicability
Some of these threats to replicability are not easy to overcome. Empirical researchers wishing to apply a novel method cannot be expected to check software code for bugs, reduce researcher degrees of freedom, or inherently know that future methodological work will lead to different recommendations. Instead, these threats should be seen as risks that a researcher accepts when applying novel methods, and the researcher should weigh the strength of his or her conclusions appropriately. However, there are certain steps a researcher can take to limit the risk of mistakes and to facilitate replication of the original analyses.
Use crystallized software and methods
The less crystallized a methodology, the more likely it is that recommended best practices for it and recommended interpretation of its results will change. There is no easy way to define when a methodology is crystallized (it is likely that no methodology will ever be regarded as fully crystallized), but good indicators are multiple published methodological articles on the topic from different research groups and several stable software implementations that lead to similar results. Using crystallized software (see Table 1) will also reduce the risk of results being based on suboptimal default options or bugs in the software.
Document analysis code
In addition to providing code, researchers should aim to make the code understandable, such that other researchers can adapt the code to their purposes. This will make it easier for other researchers to check if the original analyses were performed correctly and to correctly implement the analyses themselves in order to see if the findings are replicable. The minimal form of documentation is to augment code liberally with comments (in R, everything that follows the symbol # is a comment). Another option is to make use of literate programming (Knuth, 1984; Piccolo & Frampton, 2016), in which the researcher writes a text document with code chunks that either automatically perform the analysis when the document is compiled or can be compiled to visually show pieces of the code that the user can freely run or adapt. Prime examples of software for literate programming are Sweave (Leisch, 2002) and knitr (Xie, 2016) for R and Jupyter notebooks (Kluyver et al., 2016) for Python.
Check for mistakes in code
As I discussed earlier, a potential source of limited replicability is that errors may be made in the analysis. Therefore, I recommend that researchers double-check their code for potential errors. For example, it is advisable to check if the analysis code runs after the workspace is cleared and if it runs on a different computer. Some tools may aid researchers in this endeavor. For example, the R package statcheck (Nuijten et al., 2016) can be used to check if reported p values are accurate. Automated tools will go only so far in checking your analyses for possible errors, however, so it is always advisable to have colleagues or other researchers check the analyses. For example, Wicherts (2011) discussed a “co-pilot” model in which colleagues routinely check each other’s work to prevent “embarrassing errors” (para. 11). As noted earlier, supplying analysis code (and preferably data as well) will also greatly help reviewers and independent researchers double-check the analyses.
Recommendations for methodologists and software developers
In the previous sections, I have described threats to reproducibility and replicability of results and included guidelines for empirical researchers to help them minimize the impact of these threats. Methodologists and software developers can also aim to minimize these threats. Here, I present a short list of ways in which they may contribute to improved reproducibility and replicability of results.
Provide code together with methodological publications
As I discussed earlier, reimplementation of a method may not be trivial and may result in mistakes. Therefore, it is crucial to supply code alongside methodological publications. This not only allows researchers to implement the reported methods as intended but also allows researchers to replicate the methodological studies (e.g., with simulations studies) themselves. Methodologists should aim to make at least basic code or software routines, regardless of what programming language is used, available to other researchers. The more flexible the code, the better. For example, values specific to the input, such as sample size or number of variables, should not be hard-coded in the analysis but stated in the beginning of the code (and easily changeable).
Provide documentation to the code
In addition to making code available, methodologists should clarify how their code ought to be used (e.g., what do the objects indicate? how should a function be used?). When code is presented as a script, comments should be used to indicate the purpose of each line and where a user is expected to change input. When code is presented in the form of a function or program, methodologists should provide some information on what each argument or option indicates and which default values were used. An ideal means of presenting code written in R is to contribute an R package to CRAN, as the automated checking procedure required for CRAN packages ensures that the developer has passed numerous checks regarding basic documentation.
Use a clear coding style
Open-source software code allows other researchers to learn from the code, spot the locations of bugs, and suggest new developments and changes. These benefits, however, are possible only if the code is understandable. To this end, methodologists should aim to program clearly, for example, by roughly following a style guide (e.g., Google’s R Style Guide, n.d., available at https://google.github.io/styleguide/Rguide.xml). There is no definitive and clear optimal choice of coding style, and style guides sometimes recommend impractical routines; however, it helps when coders follow a clear and consistent way of indenting code and writing object names. Other ways in which the readability of code can be improved are by using informative object names and inserting comments that explain the purpose of the code.
Use a version control system
A version control system helps keep track of changes in code, allowing researchers to backtrack and see what the software did at any point in its development and why an updated version may lead to different results. GitHub provides an excellent version control system (using Git) that shows code in an easy-to-read online environment. Another benefit of using a website such as GitHub is that it provides a centralized platform on which people can ask questions about the software or propose solutions (via pull requests). This not only makes the job of the methodologist easier (someone else might solve a problem for him or her) but also makes it easier for users to see if someone else is experiencing the same problem they are.
Offer backward compatibility
In principle, it should be possible to reproduce earlier analyses using newer versions of the software routines without relying on tools such as checkpoint or Docker. To this end, I recommend that researchers offer backward compatibility as much as possible. For example, when replacing a function with a differently named function, it may be useful to retain the original function as well (and to add a warning stating that the function is deprecated). When a coder changes a function, it may be useful to include a mimic option, which can mimic the exact behavior of an earlier version of the software.
Conclusion
In this article, I have highlighted hazards of applying state-of-the-art statistical methods, especially when they have been discussed only in preprint manuscripts and implemented only in beta versions of software packages: One runs the risk of not being able to reproduce the results (i.e., to obtain the same numeric results at a later time) or to replicate the results (i.e., to draw the same conclusions at a later time) because the use and interpretation of the method and software inputs and outputs may change substantially. I have proposed several guidelines that can help empirical researchers and methodologists minimize these threats to reproducibility and replicability.
The goal of the current article is not to propose solutions to all the problems raised. Indeed, some of the problems I have mentioned are inherently unsolvable. For example, given rapidly changing methodology and software implementations, coupled with large and complicated dependency trees, there is a possibility that a code will no longer yield the same exact results (or run at all) after some years have passed. A possible solution is to use one of several methods that allow a user to take (or use) snapshots of the software at the time the original analysis was run. This method should help users reproduce numbers exactly, but it is highly questionable whether the same conclusion will be drawn under such circumstances because the underlying software packages may have changed for good reasons (e.g., to solve bugs). A second problem that may be insurmountable is that it may not be possible to use fully crystallized software and methods. Very few current software packages will pass all the criteria listed in Table 1. However, statistical methods must be applied at some point, and applications of methods further drive methodological research. In fact, many statistical methods and software routines are developed in response to applied-research questions, and the development of new methods may in turn inspire new research questions. Therefore, it is neither reasonable nor desirable to require researchers to use only crystallized software; however, I do advise that researchers familiarize themselves with the threats of applying novel methods so that they may interpret their results with due care.
Supplemental Material
EpskampOpenPracticesDisclosure – Supplemental material for Reproducibility and Replicability in a Fast-Paced Methodological World
Supplemental material, EpskampOpenPracticesDisclosure for Reproducibility and Replicability in a Fast-Paced Methodological World by Sacha Epskamp in Advances in Methods and Practices in Psychological Science
Supplemental Material
EpskampSupplementalMaterial – Supplemental material for Reproducibility and Replicability in a Fast-Paced Methodological World
Supplemental material, EpskampSupplementalMaterial for Reproducibility and Replicability in a Fast-Paced Methodological World by Sacha Epskamp in Advances in Methods and Practices in Psychological Science
Footnotes
Acknowledgements
I would like to thank Adela-Maria Isvoranu, Katharina Jorgensen, and Ivan Flis for comments, feedback, and discussion on earlier versions of this manuscript.
Action Editor
Frederick L. Oswald served as action editor for this article.
Author Contributions
S. Epskamp is the sole author of this article and is responsible for its content.
Declaration of Conflicting Interests
The author(s) declared that there were no conflicts of interest with respect to the authorship or the publication of this article.
Open Practices
Preregistration: not applicable
All data and materials have been made publicly available via the Open Science Framework and can be accessed at https://osf.io/u8sdh/. The complete Open Practices Disclosure for this article can be found at http://journals.sagepub.com/doi/suppl/10.1177/2515245919847421. This article has received badges for Open Data and Open Materials. More information about the Open Practices badges can be found at ![]() .
.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
