Abstract
Containers have become increasingly popular in computing and software engineering and are gaining traction in scientific research. They allow packaging up all code and dependencies to ensure that analyses run reliably across a range of operating systems and software versions. Despite being a crucial component for reproducible science, containerization has yet to become mainstream in psychology. In this tutorial, we describe the logic behind containers, what they are, and the practical problems they can solve. We walk the reader through the implementation of containerization within a research workflow with examples using Docker and R. Specifically, we describe how to use existing containers, build personalized containers, and share containers alongside publications. We provide a worked example that includes all steps required to set up a container for a research project and can easily be adapted and extended. We conclude with a discussion of the possibilities afforded by the large-scale adoption of containerization, especially in the context of cumulative, open science, toward a more efficient and inclusive research ecosystem.
Together with preregistration, one of the primary solutions to address the replication crisis in psychology has been to encourage open data and materials (Levenstein & Lyle, 2018). Facilitated by the rise of open-source programming languages (R, Python), free repositories (e.g., OSF, GitHub), and incentive journal policies (e.g., open science badges; Kidwell et al., 2016), the field of psychology has made important progress toward reproducible results in recent years. These incentives enable stronger foundations for future research—as of early 2019, about 35% of faculty researchers in psychology embrace open science practices compared with a mere 5% just 5 years earlier (Nosek, 2019). Yet, although necessary, sharing data and materials remains insufficient to fully ensure reproducible results (Epskamp, 2019) because results can differ significantly depending on software versions or operating systems (Glatard et al., 2015; Gronenschild et al., 2012). To alleviate these problems, it has been suggested that software version numbers and details about computing environments (e.g., operating system) should be included in the methods of scientific articles. However, obtaining previous software versions can be cumbersome, especially if the software is proprietary or relies on system dependencies (other software installed on the computer that the current software needs to work) of a specific operating system that might not be at one’s disposal.
More effective solutions have been proposed. For example, the package renv (Ushey, 2021), superseding packrat (Ushey et al., 2018), manages dependencies in R (R Core Team, 2020) by storing the source code of all R packages used in a project. These can then be recompiled for later use, which ensures consistency in R package versions across users. However, this approach does not handle system dependencies or dependencies of other programming languages and software packages. For example, the package rJava (Urbanek, 2020) needs a specific Java version to be installed on the computer; it is therefore possible that rJava works for a given user but not another—even if the project is managed by renv and operating systems are identical.
Ideally, we want to “package up” (isolate) our whole computing environment in a way that anybody on any computer can examine and replicate our work, independently of installed software, drivers, and operating systems (see Fig. 1). A popular way to achieve this goal is with virtual machines (VMs). A VM is a computer program that creates a virtual computer running inside the physical computer with its own operating system, software, and files. Using VMs, all software and their system dependencies (and sometimes scripts and/or data) used for a particular project can be bundled up and then shared with others. Yet VMs can get very large and slow, which can make sharing and using them impractical.
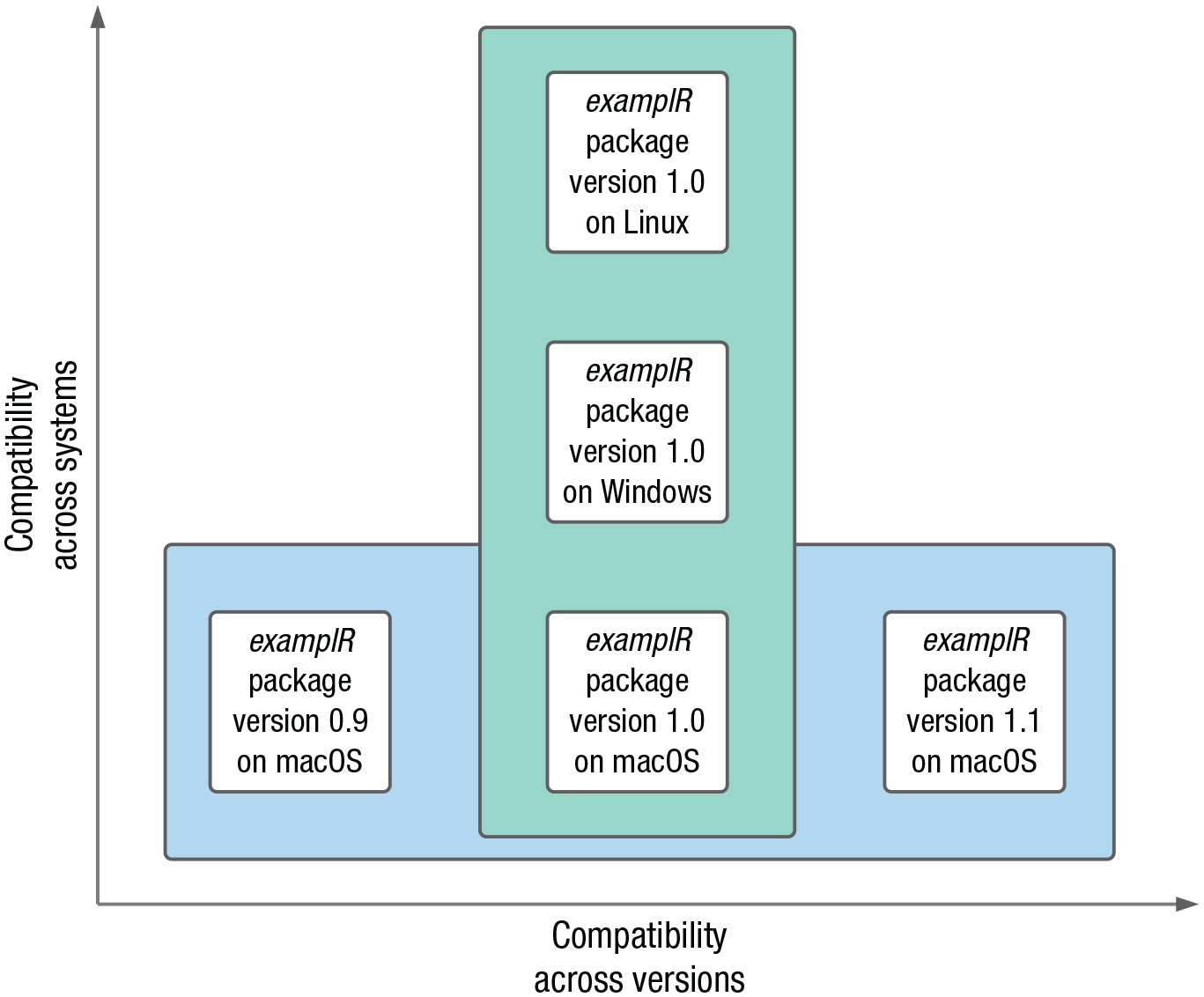
Two dimensions of compatibility enabled by containerization. Compatibility across systems relates to differences in system dependencies within or across operating systems; here, we chose to depict an example of compatibility across operating systems. Compatibility across versions relates to differences between software releases.
An alternative approach that has gained traction recently is containerization. Containers also isolate computing environments but use fewer resources than VMs. Containers are thus a lightweight alternative to VMs that make more efficient use of the underlying computing system and can more easily be shared. Originally mainly used in computer science as a way to develop and test applications in an isolated environment, containers have recently been adopted for scientific computing (Boettiger, 2015). Containerization in this context is a way to not only support reproducibility once a project is completed but also to facilitate working efficiently and collaboratively while the project is ongoing, by ensuring everything related to a research project runs smoothly and identically across all computers used across the life span of a project independently of collaborators’ individual setups.
Despite the advantages of containers and their prevalent use in fields such as computing, software engineering, and, more recently, neuroscience, containerization remains rarely used in psychology, perhaps because of a lack of awareness or because using—and especially building—containers can seem daunting for those of us lacking a computer science background. This tutorial aims to remove this barrier and demystify containerization by providing a step-by-step guide to using, building, and sharing containers within a research workflow. We use the container platform Docker and focus on the R language. After some background information on Docker, we provide an overview of basic Docker commands and of how to run R in Docker containers (Tutorial Part I: Docker Basics and the Rocker Project). In the section Tutorial Part II: Building and Sharing Personalized Docker Containers, we provide a step-by-step worked example of building, using, and sharing a personalized container for a research project. This worked example includes all required steps from start to finish and is designed to be easily adoptable and extendable by readers. We conclude with a brief summary of the tutorial and an overview of more advanced containerization workflows.
Brief Introduction to Docker
Docker (docker.com; Merkel, 2014) is an open-source containerization project based on Linux; that is, Linux is running inside the containers even though we might be on a Windows or Mac computer. Docker consists of three components: the Docker software, Docker objects, and an online Docker registry (see figure in Box 1). The Docker software consists of the Docker client and the Docker daemon. The Docker client is a command-line tool that you use to tell Docker what you want to do. When you type a command, the Docker client talks to the Docker daemon in the background, which then does all the work, such as building, running, and distributing containers. In our case, the Docker client and the Docker daemon are both on our computer, but it is possible to connect a Docker client to a remote Docker daemon.
Box 1.
Glossary of Docker Terminology

The main Docker objects are Docker images and Docker containers. A Docker image is a read-only, unmodifiable blueprint of the desired computing environment. The image contains all instructions to create the computing environment and can be shared if others want to use the same environment. From the image, Docker can create a container—a runnable version (instance) of the image, that is, the actual computing environment that can be used to run applications or conduct analyses. An unlimited number of containers can be created from one image, and, in contrast to images, containers can be modified while running. These changes, however, are not saved back into the image. This desirable property means that each time a container is run from a particular image, it is exactly the same; you cannot “break” the image, no matter what you do in the container. If you want changes you made in the container to be saved, you need to create a new image that incorporates these changes. You can think of a Docker image as a cake recipe and the corresponding Docker container as a finished cake. The recipe contains the instructions to make the cake, it can be used to make as many cakes as you like, and it can be shared to enable others to make the same cake. Everyone using the recipe ends up with the same kind of cake, which can then be modified by adding, for instance, icing or sprinkles. However, your addition of icing and my addition of sprinkles will not change the recipe, and the next time we use the recipe, we get the same reproducible cake as before.
Finally, the third Docker component is an online repository of Docker images called Docker Hub, from which existing images can be downloaded and to which you can upload your own images if you want to share them. We illustrate all these concepts in practice throughout the tutorial, which will further clarify them. Box 1 contains additional details about the Docker architecture and definitions of Docker-related terms.
Even though other containerization platforms exist, such as CodeOcean (described in detail in Clyburne-Sherin et al., 2019) and Singularity (sylabs.io), we chose to use Docker for this tutorial for five main reasons. First, Docker is one of the leading container platforms and has been established as best practice in several research fields (Boettiger, 2015). Second, Docker runs on all major operating systems (Linux, macOS, and Windows). Third, Docker containers are easy to use, very lightweight, and fast. Fourth, Docker images can be stored and shared for free on the central registry Docker Hub. Finally, and thanks to its growing popularity, Docker benefits from a large user community and a rich ecosystem of related tools, such as Rocker (rocker-project.org; Boettiger & Eddelbuettel, 2017), which provides containers with R environments, and Neurodocker (github.com/ReproNim/neurodocker), which facilitates setting up customized containers for neuroimaging projects. Familiarity with Docker is also helpful because major tools in neighboring fields have moved to using the Docker format, such as the standardized functional MRI preprocessing pipeline fmriprep (fmriprep.org; Esteban et al., 2019). In psychology, similar trends are emerging with projects like the Experiment Factory, which allows creating Docker containers to ensure behavioral experiments can run smoothly across platforms (expfactory.github.io; Sochat, 2018).
Disclosures
All materials (Dockerfiles, data, scripts) used in this tutorial are available at osf.io/z85k3; the corresponding Docker images can be found at hub.docker.com/u/kwiebels. A wiki with commonly encountered Docker errors is also available at osf.io/z85k3. We designed this tutorial to be accessible for novices, but we do assume basic knowledge of computers (e.g., knowledge of terms such as path). If you want to learn more about basic command-line usage, see the Software Carpentry lesson on the Unix shell (swcarpentry.github.io/shell-novice). For the interested reader wanting to go beyond the material covered in this tutorial, a comprehensive general Docker guide can be found at docker-curriculum.com.
Tutorial Part I: Docker Basics and the Rocker Project
Preparations
Before starting the tutorial, go to osf.io/z85k3 and download the folder
Installing Docker
Docker is available for a variety of Linux distributions, for Mac, and for Windows. On Linux, you will install Docker directly; if you are on Mac or Windows, Docker is installed through Docker Desktop, an application that includes all features needed to build and share containers. To download and install Docker, follow the detailed instructions listed at docs.docker.com/get-docker in the section for your operating system. If you are using Windows, see Box 2 for Windows-specific considerations.
Box 2.
Windows-Specific Considerations
There are two ways to run Docker on Windows, either using the Hyper-V backend or using the Windows Subsystem for Linux (WSL) 2 backend (available for Windows 10 Version 1903 or higher). WSL is a full Linux kernel built by Microsoft and allows Docker containers to run natively without the need for emulation. Using the WSL 2 backend is recommended because it makes more efficient use of resources and greatly increases speed. If possible on your system, Docker will automatically select this option during the installation process (see screenshot of the Docker settings below, “Use the WSL 2 based engine” tick box). Depending on the computer setup, the WSL 2 feature on Windows might need to be enabled and the Linux kernel update package installed. Docker Desktop will alert you to this if these steps are necessary, in which case you can follow Steps 1 through 5 at docs.microsoft.com/en-us/windows/wsl/install-win10 (for further details about the WSL 2 backend, see docs.docker.com/docker-for-windows/wsl). Older versions of Windows will automatically use the Hyper-V backend (i.e., the box in the screenshot will be unticked), for which no additional steps are required.
We recommend using the Windows PowerShell. If you are using WSL 2 and prefer the WSL 2 bash terminal, you will need to prepend all commands with ‘sudo.’
Paths in Windows are different from the ones on Linux and Mac (which is used in this tutorial), so you will need to adapt the paths slightly. If your path is

Running Docker containers
Once Docker is installed, it is time to launch it! You should see the Docker icon in the taskbar, indicating that Docker is running. Docker is then accessible from the terminal (on Mac, open the terminal by opening the Applications folder, then opening Utilities and double-clicking on Terminal; on Windows, open the PowerShell by clicking Start, typing PowerShell, and then clicking Windows PowerShell). Once a terminal window is open, you can type commands and execute them using the return/enter key.
The general syntax for running a Docker container is:

An IMAGE, from which the container is derived, must be specified; the rest of the terms are optional. OPTIONS can be used to constrain the container’s behavior; we describe a range of them throughout the tutorial. The TAG instructs Docker to use a specific version of an image, and finally, the COMMAND tells Docker which command to execute when the container starts. Most of the time, the Docker image developer specifies this start-up behavior (e.g., a typical command of an R container is simply to start R), but it can be overridden, for example to get the R version instead of starting R (
To test your Docker installation, open a terminal window and run the line below:

If your installation works correctly, you should see the text output shown in Figure 2.

Screenshot of the hello-world container output.
If you see this output, you just ran your first Docker container! The container is called
Once you have downloaded a Docker image, you can run

to get a list of Docker images available on your computer. You will see the

Screenshot of the docker images command output.
In addition to a list of images, you can also get a list of all containers on your computer using

The

Screenshot of the docker ps -a command output.
The container’s status indicates for how long the container has been running or how long ago it was stopped (exited). Some containers stop automatically (as was the case with the hello-world container), but some have to be stopped manually.
By default, after a container is stopped, it is not removed from the computer, and every time you use

Note that you can copy and paste the container ID instead of having to write it out manually (in the terminal on Mac, copying and pasting works as usual; in the Windows PowerShell, highlight what you want to copy and use right click to paste). To remove all containers from your computer at once, the shortcut
Box 3.
Glossary of Common Docker Commands
docker pull <image_name> downloads an image from DockerHub.
docker run <image_name> runs a container.
docker images or docker image ls returns a list of all Docker images stored on the computer.
docker ps or docker container ls returns a list of currently running containers.
docker ps -a returns a list of currently running and stopped containers (i.e., all created containers).
docker rmi <image_ID> removes an image.
docker rmi $(docker images -a -q) is a shortcut to remove all images.
To avoid having to manually remove containers, it is possible to specify at the time a container is created that the container should be removed automatically after it is stopped. This is achieved with the

This time, the container was automatically removed from the computer after exiting. Note that if you use Docker Desktop, you can also start, stop, and remove containers and images in the Docker Desktop interface.
The Rocker project
Thanks to the Rocker project (rocker-project.org), running R inside a Docker container is just as easy. Rocker maintains Docker images with R or RStudio preinstalled, which means that we do not have to create R Docker images from scratch. There are different images suited for different needs (see all images at hub.docker.com/u/rocker). Some images only include R or RStudio, whereas others already have some R packages preinstalled, for example the
Rocker base R containers
To run a container with a base R environment, simply type the line below:


An interactive R session is now running in Docker (see Fig. 5), and you can use R as you usually would. For example, typing

Screenshot of an interactive base R session in Docker running in the terminal.
By default, Docker pulls and runs the latest version of an image from Docker Hub. At the time of writing, the latest Rocker image uses R version 4.0.4 (see Fig. 5). When Docker is used for reproducibility reasons, a versioned—instead of the latest—image should be used to ensure that everyone using the container will get the same R version independently of when the container is created. To use a specific R version, tags can be specified. To use R version 3.6.1, for instance, the previous command needs to be adapted as follows:

Rocker RStudio containers
Rocker also maintains RStudio images, which might provide a more comfortable analysis environment than accessing R through the command line. These containers run an instance of RStudio server, which can be accessed from—and interacted with—a web browser. As with the base R container, you can choose the R version you want to use. Running the following command will start an RStudio container with R version 3.6.1 (note that the command has to be written on one line):

There are three additional flags this time. The
After typing the command, Docker will print the container ID to the terminal. To use the containerized RStudio, open a browser window and enter
Screenshot of the RStudio server login screen.
Upon login, you will be running RStudio in the browser (see Fig. 7). This RStudio instance can be used just like RStudio running locally on your computer. To quit the session, click the red button on the top right and close the browser tab.
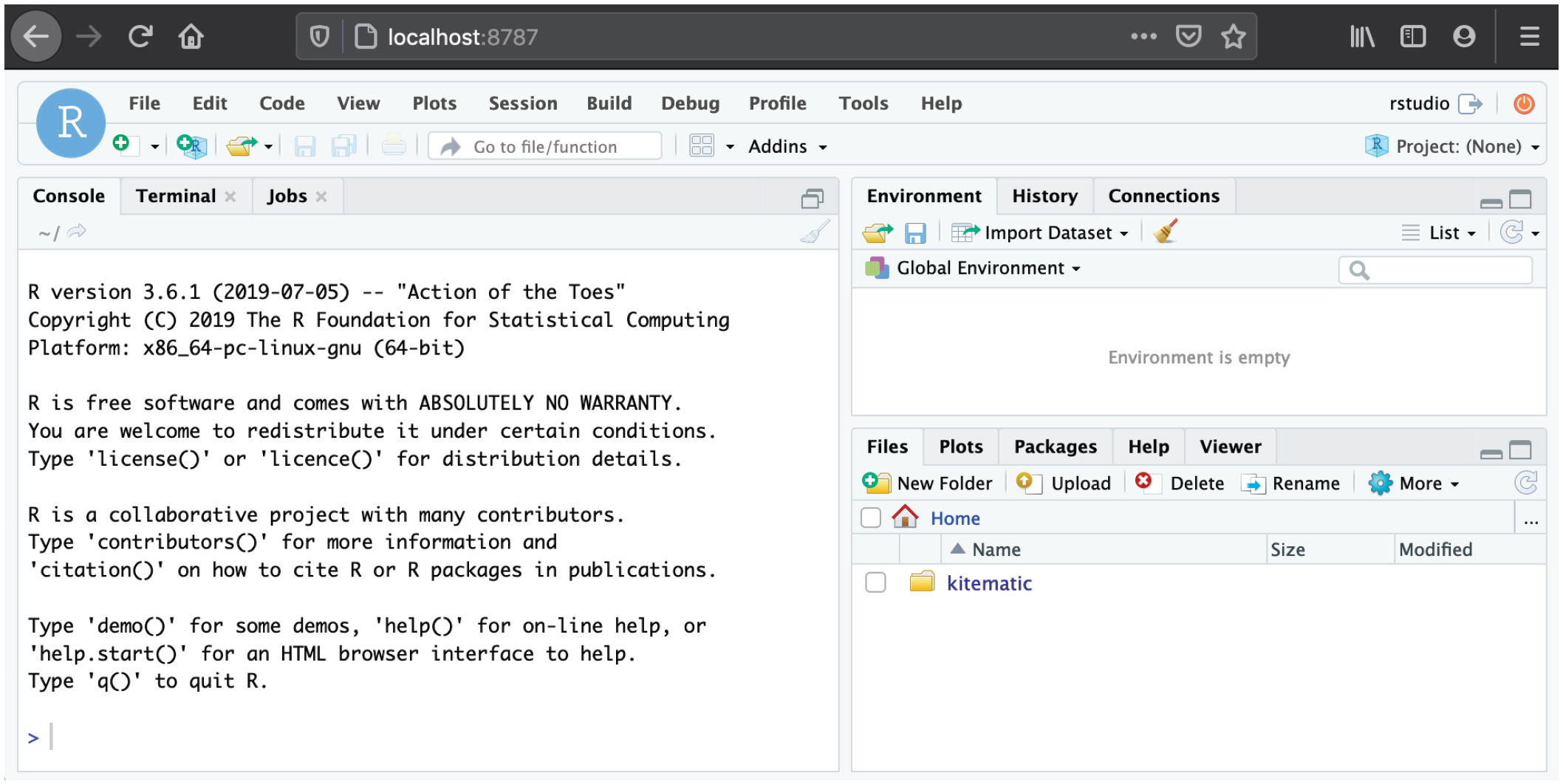
Screenshot of an RStudio session running in Docker.
When you are done, you need to stop the Docker container manually in the terminal because it will still be running in the background (because of the

Once stopped, the container is automatically removed because we used the
Accessing files in Docker containers
By default, Docker does not have access to any files on the computer; however, we might want to load a local data set or write results to a specified folder on the computer. To give access to a local folder, the
The command below will map the folder at

After running the command, open RStudio in the browser, as before. You should now see the content of the folder in the RStudio Files tab (see Fig. 8). Remember to stop the container when you are done using

Screenshot of an RStudio session with access to a local folder.
If your research project does not require packages beyond the ones provided by Rocker, you can use one of the Rocker images without any modifications. If you need additional packages—which will likely be the case—it is possible to install them in RStudio while the container is running. The problem with this approach, however, is that the packages need to be reinstalled every time a new container is created, which also means that they will not be available automatically for other researchers who use your container. To build a personalized computing environment that can be used and shared with collaborators and other researchers, you can build your own Docker image with all required packages. That way, the packages will automatically be available in each container that is created from the image. In the following section, we provide a worked example of how to create and use a personalized container and how to share it. The guide includes all required steps so that it can easily be adopted and extended for your own research.
Tutorial Part II: Building and Sharing Personalized Docker Containers
In this section, we aim to provide a step-by-step guide on how to build, use, and share a personalized container for your research project. Building a personalized container involves several steps but is a straightforward process, especially if only R is required. In the following sections, we demonstrate how to (a) build your own personalized Docker container with RStudio and additional R packages; (b) load a local data set inside your personalized container, create summary statistics and a plot, and write results to a local folder; and (c) make the container available on Docker Hub or OSF (osf.io).
Building a personalized Docker container
Building a personalized container involves two main steps. We first need to write a Dockerfile—a file with a set of instructions in which we specify everything that we want to include in the container. This Dockerfile is then used to build a Docker image, from which we can then run containers.
For this example, we want to build an RStudio container with R version 3.6.1 and use the R packages psych (Revelle, 2011), ggplot2 (Wickham, 2011), and gghalves (Tiedemann, 2020) to generate summary statistics and a plot. As described previously, thanks to the Rocker project, we do not need to build our container from scratch, which would involve installing R on the Linux system inside the container. Instead, we can simply use a suitable Rocker image as a starting point and then add the packages we need. Given that we need ggplot2, which takes quite a long time to install, and remotes (Hester et al., 2019) to install gghalves from GitHub, we will start with the
To start building your container, open the terminal and move into the

You can check that you are in the right location by typing

On Windows, use

The general format of a Dockerfile is

Instructions do not have to be capitalized, but it is convention to do so. A Dockerfile must start with a
For our Docker container, we are going to start with a versioned Rocker tidyverse image. Open the Dockerfile using your favorite text editor, and add the following line:

After this line, we need to specify which additional R packages we want to install. Before adding this information to the Dockerfile, it is usually a good idea—that can save a lot of time—to test installing the packages first. Most packages should install without any issues, but some packages rely on system libraries that have to be installed first and will throw an error at first try. These errors can be challenging to track down if they are encountered only while building the Docker image. To test the installations, we run a container of the tidyverse image and then—instead of opening RStudio—open a terminal inside the running container. This way, we can install the packages from the command line to ensure those commands will work in the Dockerfile.
A running container can be accessed using the
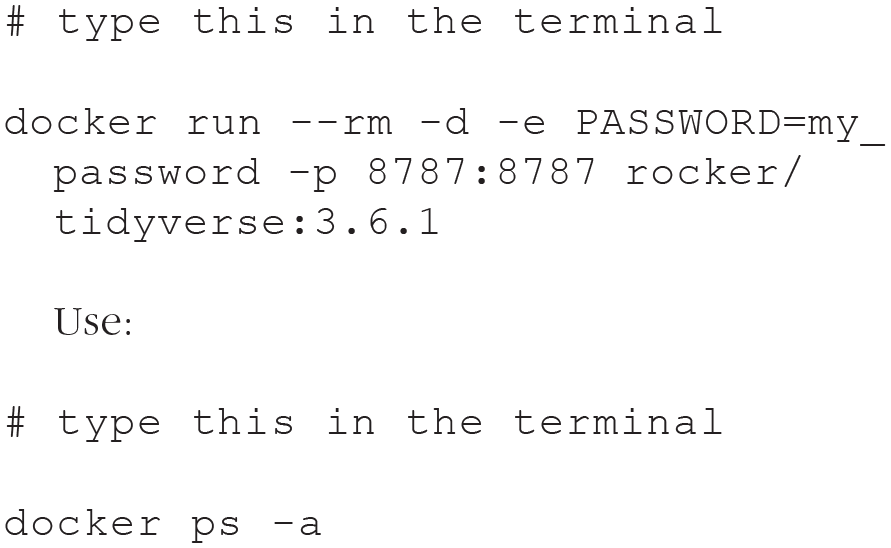
to get the ID of the running container and type:

Bash is the command language used by the terminal in this container. The command will open a terminal inside the container (see Fig. 9). In this case,

Screenshot of a bash prompt inside a running container.
In the terminal inside the container, you can try out installing the packages you need. Running

will use R’s

Once those two packages have installed successfully, you can safely add them to the Dockerfile using RUN instructions:

After saving this Dockerfile, exit the terminal inside the Docker container by typing

Using the container
Having set up the container, we can now use it to load a data set, compute summary statistics, create a plot, and save the output in a folder on the computer. By now, you are familiar with how to run a container; all that is needed is to replace the image name with the name you gave your personalized Docker image. We use the script and the data you downloaded from OSF, so you need to give Docker access to the folder that contains these downloaded files (replace

You are now running your first personalized container! After opening RStudio in the browser, you will see the data file (
When looking at the script, you will see that it loads some packages, computes summary statistics using the psych package, creates a plot using ggplot2 and gghalves,
2
and saves the results in a file. Run the script as you usually would in RStudio, and you will see two new files being created,
In some situations, you—or others—might just want to run the container to reproduce and inspect the results instead of interacting with the data or code inside the container. In this case, a slightly adapted command can be used to start the container, run the script, save the output, and then exit and remove the container:

Sharing the container
When you have created and used a personalized container for a research project, you might want to share it, along with the scripts and data, with your collaborators while the project is ongoing or upon completion of the project to make your analysis reproducible for other researchers. Docker containers can be shared either by uploading the Docker image to Docker Hub so that others can download it or by sharing the Dockerfile on a repository such as OSF so that others can build the corresponding Docker image themselves. We demonstrate both options in the next two sections.
It is good practice to include usage instructions for the container in the Dockerfile and/or in a README file. To do this, open the Dockerfile and add the following information at the bottom of the file:


Sharing the Docker image on Docker Hub
To share the image on Docker Hub, you will need to create a free Docker Hub account. To do this, go to hub.docker.com, click on “Sign up,” and fill out the information. Once your account is created, return to the terminal and log in with your credentials:

For Docker to know to which repository on Docker Hub to upload your image, you need to tag your image with your Docker ID (the username you used when creating the account on Docker Hub) and the image name in the following format:

The image name can remain the same. Given the example above, this would be

You can then upload (push) the container to Docker Hub:

It is important here to use the format

Sharing the Dockerfile on OSF
If you prefer sharing the Dockerfile on OSF, along with any materials or data you want to share, you can achieve this very easily. If you have never used OSF before to share project-related files, follow Soderberg’s (2018) guide. All that is left to do once the OSF repository is set up is to upload the Dockerfile into that repository. Others can then download that Dockerfile and use the
These two approaches can also be combined because they both have advantages. Sharing the image on Docker Hub makes it easier for others to download and use it because they do not need to build the image themselves. Sharing the Dockerfile itself has the advantage that others can inspect the file to see exactly what is included and adapt it for their own purpose if desired. Note that there is a slightly more advanced way to upload Docker images to Docker Hub, which will automatically make the Dockerfile available as well (see Box 5, Automated Builds Using GitHub).
Once you have finished the tutorial, you might want to delete all containers and images we used throughout because they can take up quite a lot of disk space. Use the following two commands to achieve this:

Discussion
In this tutorial, we explained the basics of containerization and provided step-by-step guides for building, using, and sharing Docker containers. The first part of the tutorial introduced basic Docker commands and the Rocker project as a way to run R code in containers. In the second part of the tutorial, we showed how to set up a personalized container for a research project, from writing a Dockerfile to sharing the Docker image.
Containerization is an important step toward making research reproducible by providing a consistent computing environment that can be used by all collaborators over the course of a project and that can be shared along with the publication. Throughout the tutorial, we focused on the R language, but we provide a resource below for setting up a Python environment to illustrate the process for projects that require tools beyond R. We aimed to provide a resource that can be easily adapted and extended to one’s own research projects or workflows. Below, we summarize some Docker uses that are beyond the scope of this tutorial but might be of interest to some readers before concluding the tutorial with some general remarks.
Additional steps
Integrating containerization into the research workflow
Sharing a container after a research project has been finalized ensures that your analyses are reproducible. However, containerization can also greatly benefit your collaborators—and yourself—throughout the development of the project by making sure that code does not break over time and that every collaborator works in exactly the same environment without the need to synchronize all pieces of software manually. This aspect might be especially beneficial for collaborators who are not heavily involved in the data analysis part of the project and need to inspect the results only from time to time or give feedback.
Figure 10 depicts an example workflow with containers being used as the computing environment from the start of a research project. At the beginning of the project (purple box), a shared location (e.g., a network drive) is set up where the data and all analysis scripts will be stored (blue box). A Dockerfile for the anticipated computing environment is also created. Using the Dockerfile, a Docker image is built, which is then shared on Docker Hub. From there, all collaborators download the Docker image and use it as the computing environment for the duration of the project. Given that the Dockerfile might have to be adapted from time to time (e.g., to make additional packages available), the
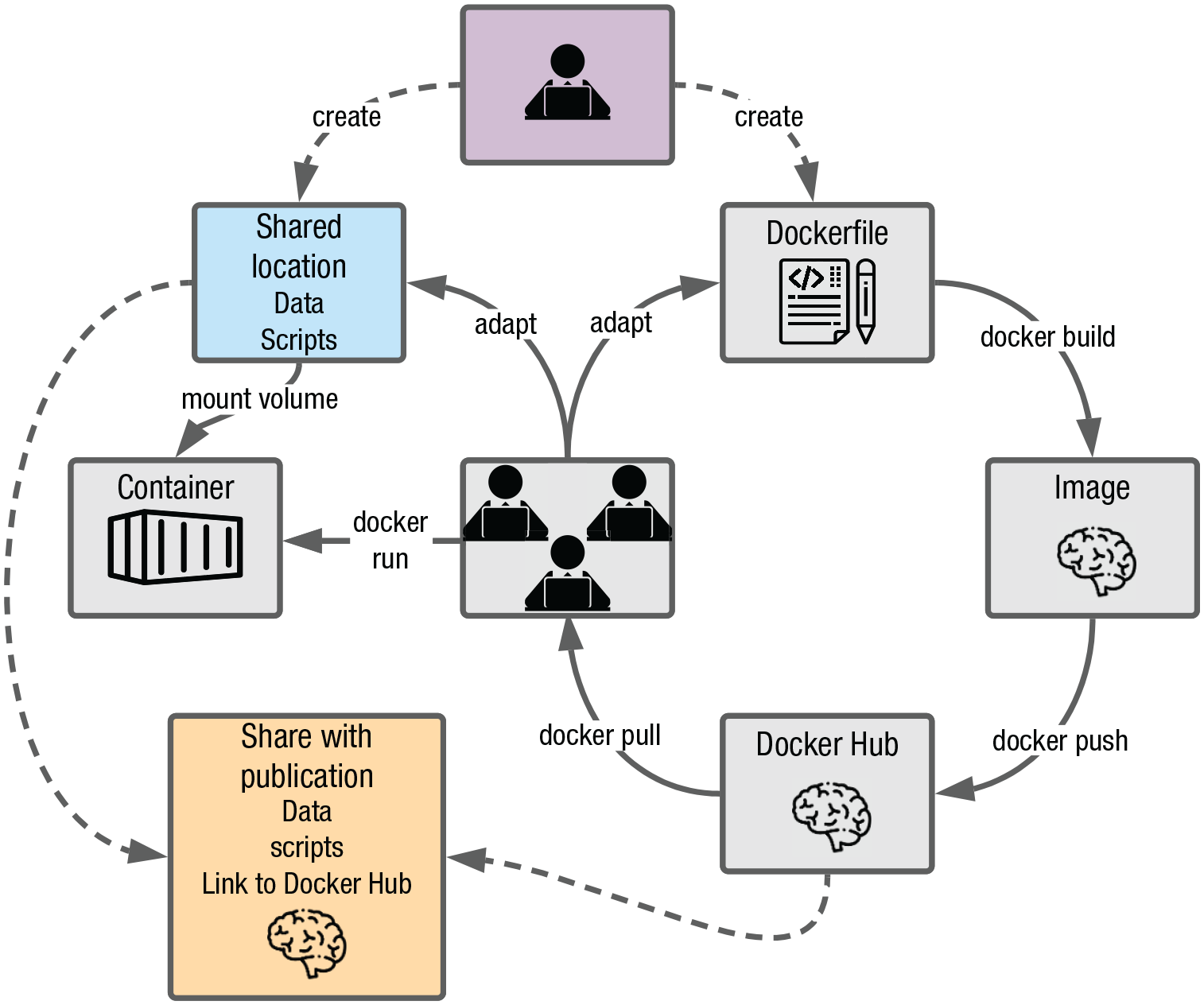
Example research workflow using containerization.
More advanced containers and workflows
The image we have built in this tutorial is relatively simple; it includes only R and a few packages. We chose this example to be accessible and easily extendable to suit your purpose. However, some projects require analysis tools beyond R. For instance, if you have electroencephalography (EEG) data and want to use MNE-Python (Gramfort et al.,2013, 2014) in addition to R for data processing and analysis, an R container needs to be extended with a Python environment and the right Python tools need to be installed. We show an example of a Dockerfile for this advanced scenario in Box 4 and provide a step-by-step guide for building this container at osf.io/z85k3.
Box 4.
Docker Containers Beyond R
Sometimes, languages beyond R are required for data analysis. For this example, we want to build a Docker container with RStudio, Python (Version 3.6 or higher), the mne Python package, and the reticulate (Allaire et al., 2018) and mne (Engemann, 2020) R packages.
An example Dockerfile for this scenario is shown below. See osf.io/z85k3 for a step-by-step guide.
# this is a Dockerfile
# use the Rocker tidyverse image for the R environment
# update Debian package manager
# install Anaconda
wget –quiet https://repo.anaconda.com/archive/Anaconda3-2020.07-Linux-x86_64.sh -O ~/anaconda.sh && \
/bin/bash ~/anaconda.sh -b -p /opt/conda && \
rm ~/anaconda.sh
# set Python path
# configure reticulate to point to the conda Python executable
# install MNE
# install reticulate and MNE-R
Beyond more elaborate containers, there are also a number of additional, more advanced features that can be integrated within the containerization workflow. These are not covered in depth here for simplicity purposes; however, interested readers may refer to Box 5 for examples of more advanced setups and corresponding resources.
Box 5.
Advanced Docker Workflows
There are several more advanced ways Docker can be used to facilitate the reproducibility of the research workflow. Here, we highlight two important ones: automated builds using GitHub and containerization beyond computing environments.
The Docker build process can be automated by storing the Dockerfile in a GitHub repository and by linking this GitHub repository to Docker Hub. The Docker Hub repository can then be configured such that every time the Dockerfile on GitHub is updated, an updated Docker image is automatically built, tested, and pushed to Docker Hub. Automatic builds therefore render the manual build and push steps in Figure 10 redundant, which is especially useful if the Dockerfile is anticipated to change frequently throughout the project. See docs.docker.com/docker-hub/builds for a guide on how to set up automated builds and Vuorre and Curley (2018) for a tutorial on Git and GitHub.
Although the focus of this tutorial is on containerization for the data analysis part of a research project, Docker can facilitate incorporating other parts of the research process into a reproducible workflow, including running experiments and reporting results.
The Experiment Factory (Sochat, 2018) facilitates creating Docker containers for behavioral experiments. Containerizing experiments ensures that they can be run anywhere and on any computer and that they can easily be shared. This is likely especially useful for decentralized research projects, which are run by several labs, to minimize problems with the setup and compatibility issues. See expfactory.github.io for further details.
Reproducible reporting can be achieved with the R package liftr (Xiao, 2019), which uses Docker to containerize and render RMarkdown documents. An RStudio addin is available to facilitate this process. See liftr.me for further details. See also Peikert and Brandmaier (2019) for a suggested comprehensive workflow, including version-controlled data management, dependency management using Makefiles, containerized computing environments using Docker, and dynamic document generation using RMarkdown.
Concluding remarks
Containers are important components to increase reproducibility, but the advantages of containerization have more widespread ramifications: They facilitate collaboration, especially across global research groups, enabling efficient workflows with a common template of the research project. Another exciting prospect brought about by containers is that of truly cumulative science—sharing practices such as open data and materials have helped shape incremental research tremendously, yet cumulative science can still be hindered by compatibility and dependency issues. Containerization is the next step in that process to ensure robustness across users and time and facilitate secondary data analysis (Weston et al., 2019).
Finally, and beyond advancing scientific research, investing time and effort in learning and working with containers may be a wise professional move for researchers, especially at early stages of a career. This idea perhaps seems to run against mainstream thinking about the cost associated with open practices, especially for early career researchers (C. Allen & Mehler, 2019; Nosek et al., 2012; Poldrack, 2019). Yet given job prospects in research and academia, many researchers may likewise question whether in-depth, systematic knowledge about very specific aspects of psychological science remains valuable or, at the very least, transferable. Thorough expertise on the validity of a specific scale, construct, or paradigm may not generalize well to another professional workplace; in contrast, computational or software skills such as fluency in one or more programming languages (e.g., R, Python) or a practical understanding of version control or containerization (e.g., git, Docker) can easily generalize to professional settings outside of academia. In this context, proficiency with containerization, among the broader set of computational tools required of a modern scientist, may prove to be a worthwhile investment for psychological scientists at all career stages.
Footnotes
Acknowledgements
We thank Matti Vuorre, Erin M. Buchanan, and two anonymous reviewers for their constructive and helpful feedback throughout the reviewing process. We also thank Ding-Cheng Peng and Lenore Tahara-Eckl for comments and feedback on an earlier version of this tutorial.
Transparency
Action Editor: Daniel J. Simons
Editor: Daniel J. Simons
Author Contributions
K. Wiebels and D. Moreau developed the idea for the article and wrote the manuscript. Both authors approved the final manuscript for submission.
