Abstract
The psychological and psychiatric communities are generating data on an ever-increasing scale. To ensure that society reaps the greatest utility in research and clinical care from such rich resources, there is significant interest in wide-scale, open data sharing to foster scientific endeavors. However, it is imperative that such open-science initiatives ensure that data-privacy concerns are adequately addressed. In this article, we focus on these issues in clinical research. We review the privacy risks and then discuss how they can be mitigated through appropriate governance mechanisms that are both social (e.g., the application of data-use agreements) and technological (e.g., de-identification of structured data and unstructured narratives). We also discuss the benefits and drawbacks of these mechanisms, particularly as regards data fidelity. Our focus is on de-identification methods that meet regulatory requirements, such as the Privacy Rule of the Health Insurance Portability and Accountability Act of 1996. To illustrate their potential, we show how the principles we discuss have been applied in a large-scale clinical database and distributed research networks. We close this article with a discussion of challenges in supporting data privacy as open-science initiatives grow in their scale and complexity.
Keywords
Data about clinical patients are increasingly important for psychological and psychiatric research. In particular, advances in predictive modeling and machine learning can enable data-driven investigations of a variety of topics, such as the risk of worsening disease, treatment resistance, and suicidal behaviors (Kautzky et al., 2017; Zhao et al., 2017). At the same time, the psychological community is moving toward open science, encouraging researchers to share data and supporting materials from their studies in order to improve reproducibility, transparency, and openness in scientific investigation (Tackett et al., 2017). In many respects, concerns over lack of reproducibility and transparency have catalyzed the push toward open data sharing in psychology and psychiatry. It has also been demonstrated that sharing these data has the potential to facilitate new discoveries and innovations, especially those that require a large data set combining patients’ data from multiple and disparate sources.
Despite these advances, widespread sharing of data from psychological and psychiatric studies is not yet standard practice, and this fact has contributed to changes in editorial and peer-review processes in the academic literature. For instance, some journal editors have threatened to reject any manuscript not accompanied by full study data, no matter how sensitive these data may be or how great the potential is for participants’ privacy to be invaded (Naik, 2017), and initiatives such as the Peer Reviewers’ Openness Initiative (n.d.) encourage authors to share data and to be transparent in situations in which data sharing is not possible. Clinical data in mental health touch on the most intimate aspects of people’s lives and experiences; sensitive details, including information on relationships, past traumas, education, employment, medical history, and more, are all intermingled in clinical narrative. Unlike congestive heart failure, diabetes, or sepsis, mental illness lacks structured and numeric biomarkers. Thus, open data sharing in clinical psychology and psychiatry benefits from the inclusion of unstructured text. The mode in clinical psychology and counseling research remains the often laborious and expensive prospective study, and the output of these investigations may be qualitative or unstructured data, in addition to structured data such as diagnostic coding or scores on quantitative psychological instruments. Qualitative and unstructured data are rich resources for investigation and may beneficially be shared with other investigators, but they are harder than structured data to process in a manner that upholds a patient’s privacy (as we describe later).
Yet as a community, we are moving toward open data not just for transparency, but also to increase the potential sample sizes for studies that have been limited in the past by rare events and low sampling rates. Suicide, for example, is a rare event, and data-driven studies on suicide cannot succeed without increased statistical power enabled by data sharing. The same can be said for research on posttraumatic stress disorders, opioid abuse, treatment resistance in schizophrenia, chronic overlapping pain conditions, and many more topics in psychology. Researchers must be armed with a solid foundation in the risks of data sharing to ensure that they can innovate and collaborate without putting their study participants, their labs, their departments, and themselves at risk.
Failure to attend to privacy and data security can lead to breaches, and simple processes such as equipment upgrades are common sources of risk. For example, in Illinois, a vendor-led upgrade of software at Silver Cross Hospital potentially publicly exposed information pertaining to the mental-health status of approximately 9,000 patients (E. Sweeney, 2017). In another incident, a server breach at the New York State Office of Mental Health affected more than 22,000 individuals in 2016 (Dwyer, 2016). However, unlike the situation in Illinois, in this case, the data were not managed in a readily identifiable form, which provided an added level of privacy protection to affected patients. Although no set of privacy controls is perfectly effective, we outline many risk-mitigation strategies that may help reduce privacy risks in the era of data-sharing initiatives aimed at making data more accessible and widely distributed.
This article is divided into three main sections. First, we describe the privacy risks involved in sharing clinical data. We then consider methods for mitigating those risks. Finally, we discuss practical examples of how privacy risk is managed at Vanderbilt University Medical Center (VUMC), a major academic medical center that has embraced broad-scale and open data sharing. We contend that careful data sharing with perpetual vigilance for privacy risks may improve care and, in some cases, save lives. Attention to risks and potential unintended consequences of breaches can protect collaborations as researchers and clinicians form partnerships across traditionally distinct practice boundaries to create data-sharing environments.
Privacy Risks in Sharing Clinical Psychological and Psychiatric Data
Privacy is a complex and multifaceted concept that often conflates the notions of anonymity, confidentiality, and the right to be left alone (Solove, 2006). In this article, we focus on privacy issues that arise when information about a patient is moved beyond the initial point of collection, that is, in the clinical setting. In essence, this transfer is a disclosure of personal information, and there are questions regarding the risks and potential harms of such disclosures.
First and foremost, there are concerns about identity disclosure. This type of privacy risk corresponds to the notion of anonymity. In effect, an identity disclosure occurs when data that have been stripped of identifiers are re-identified as corresponding to a named individual. This concept is central to many privacy-related statutes and directives in the United States and elsewhere. Such disclosures can happen, for instance, when residual information in a patient’s record are linked to some resource that contains his or her identity (El Emam, Rodgers, & Malin, 2015). These resources may be publicly accessible (e.g., voter registration lists; L. Sweeney, 1997), or they may be available to a select few (e.g., neighbors or family members to whom a patient has disclosed that he or she is part of a research study; El Emam & Arbuckle, 2014). The notion of identity disclosure is often relied upon for representing privacy concerns in the clinical domain. Given the sensitivity of information communicated in this domain, many patients who want their data to be used in research would prefer not to be identified (Hull et al., 2008). This desire for anonymity is notably more cogent in the mental-health setting compared with other clinical settings (e.g., drug-dosing trials) because patients may be concerned that disclosure of their psychiatric diagnoses or treatments could embarrass them or lead to stigmatization or discrimination by family, friends, the broader public, and organizations that might use such information in a manner that infringes upon their welfare (e.g., employers or insurers; Chew-Graham, Rodgers, & Yassin, 2003; Corrigan, 2004; Hayward & Bright, 1997).
Second, there are concerns that go beyond anonymity (Duncan & Lambert, 1989). Of particular note, in addition to establishing the identity of the individual associated with a clinical record, the data recipient may gain sensitive information about that individual. This is called an attribute disclosure. More concretely, imagine that a patient’s potentially identifying features (e.g., the patient’s demographics) are aggregated to look like those of many other people in a study focused on paranoid schizophrenia. If all of the people in the study have the same diagnosis and the recipient of the data can determine that a particular person is in the study, then the recipient will know that this individual has been diagnosed with this disorder. Another type of privacy violation, referred to as a membership disclosure, was brought to popular attention several years ago, when it was shown that the genomic summary statistics of case-control studies could allow someone with only DNA data of a particular individual to detect the inclusion of that individual in a study (Gymrek, McGuire, Golan, Halperin, & Erlich, 2013; Homer et al., 2008). This concern is even more notable because of the growth of consortia studying the genetic bases of psychiatric disorders (Gratten, Wray, Keller, & Visscher, 2017; Psychiatric GWAS Consortium Bipolar Disorder Working Group, 2011); the data sets produced by such investigations are increasingly accessible through resources such as the Database of Genotypes and Phenotypes (dbGaP) at the National Institutes of Health (Mailman et al., 2007).
Although there are many more definitions of privacy, some of which may be relevant in the context of making data more open for psychological and psychiatric research, we want to emphasize that only identity disclosure is formally recognized in federal law. In the United States, sharing of personally identifiable data is limited by both the Common Rule (Federal Policy for the Protection of Human Subjects, 2017), which governs data about human research participants, and the Health Insurance Portability and Accountability Act (HIPAA) Privacy Rule (Standards for Privacy of Individually Identifiable Health Information, 2002), which governs clinical data, including psychiatric records. Yet the Common Rule and the Privacy Rule allow such data to be shared without the consent of the patient when they are de-identified. Similar exemptions can be found in Canada, where de-identified data are outside the scope of Ontario’s Personal Health Information Protection Act (2004), and in Europe, where such information is outside the oversight of the European Union’s Data Protection Directive (European Parliament and Council of the European Union, 1995). Most recently, the European Medicines Agency (2017), which has pushed to make data from clinical trials open to the public, recommended that re-identification risk be assessed and appropriately mitigated. As a result, we focus here on application of de-identification methods to support data privacy in the open-science setting.
Risk-Mitigation Methods
As clinical psychological and psychiatric data move beyond the primary-care environment, privacy risks may be mitigated through appropriate governance mechanisms. It is important to recognize that these mechanisms could be social, technological, or a combination of the two. In this section, we review these mechanisms and summarize their benefits and drawbacks.
Contractual agreements and obligations
A social mechanism for mitigating risk is to require sharers of data to enter into a contract, such as a data-use agreement. Such agreements typically require that data recipients avoid contacting the individuals whose data they receive or, in the event that the data have been de-identified, attempt to re-identify the data. This is a common practice for various data sets in the clinical domain, such as the PsychChip data made available by the Psychiatric Genomics Consortium (see Psychiatric Genomics Consortium, 2016, for their download agreement). Data-use agreements can also specify penalties for users who violate the terms of the agreement. Still, although contracts serve as a deterrent, they do not technically thwart illicit behavior.
The terms of such a contract can be reinforced technologically by requiring data recipients to access the data through computer servers maintained by the organization sharing the data. Unique usernames and passwords can be required for data use, such that all actions are logged for audit at a later point in time if that is deemed necessary.
Data de-identification
Further privacy protection may be accomplished by obscuring certain aspects of an individual’s record. De-identification, as this mechanism is called, needs to comply with privacy laws and regulations. Often, these laws and regulations provide alternative pathways for de-identifying data and specify the conditions that the de-identified data must satisfy given each pathway. For example, the HIPAA Privacy Rule provides two pathways for de-identifying data, as summarized in Figure 1. The first is the safe-harbor model, according to which data are considered de-identified, and thus shareable through open-science initiatives, if 18 types of features are removed. These features include explicit identifiers (e.g., personal names of the individuals and their relatives) and quasi-identifiers (e.g., specific dates of events, full Zip codes if the region formed by combining all ZIP codes with the same three initial digits contains 20,000 people or fewer, and exact ages of people over 89 years old) that, in combination, could lead to the unique re-identification of data. The safe-harbor model was defined after it was shown that hospital discharge records, seemingly devoid of an individual’s identity, could be re-identified by cross-referencing them with publicly available resources, such as voter registration lists, using the combination of date of birth, residential five-digit ZIP code, and gender (L. Sweeney, 2000). In a particularly famous event in the mid-1990s, this approach was applied to re-identify hospital-admission data about the then-governor of Massachusetts (L. Sweeney, 1997).
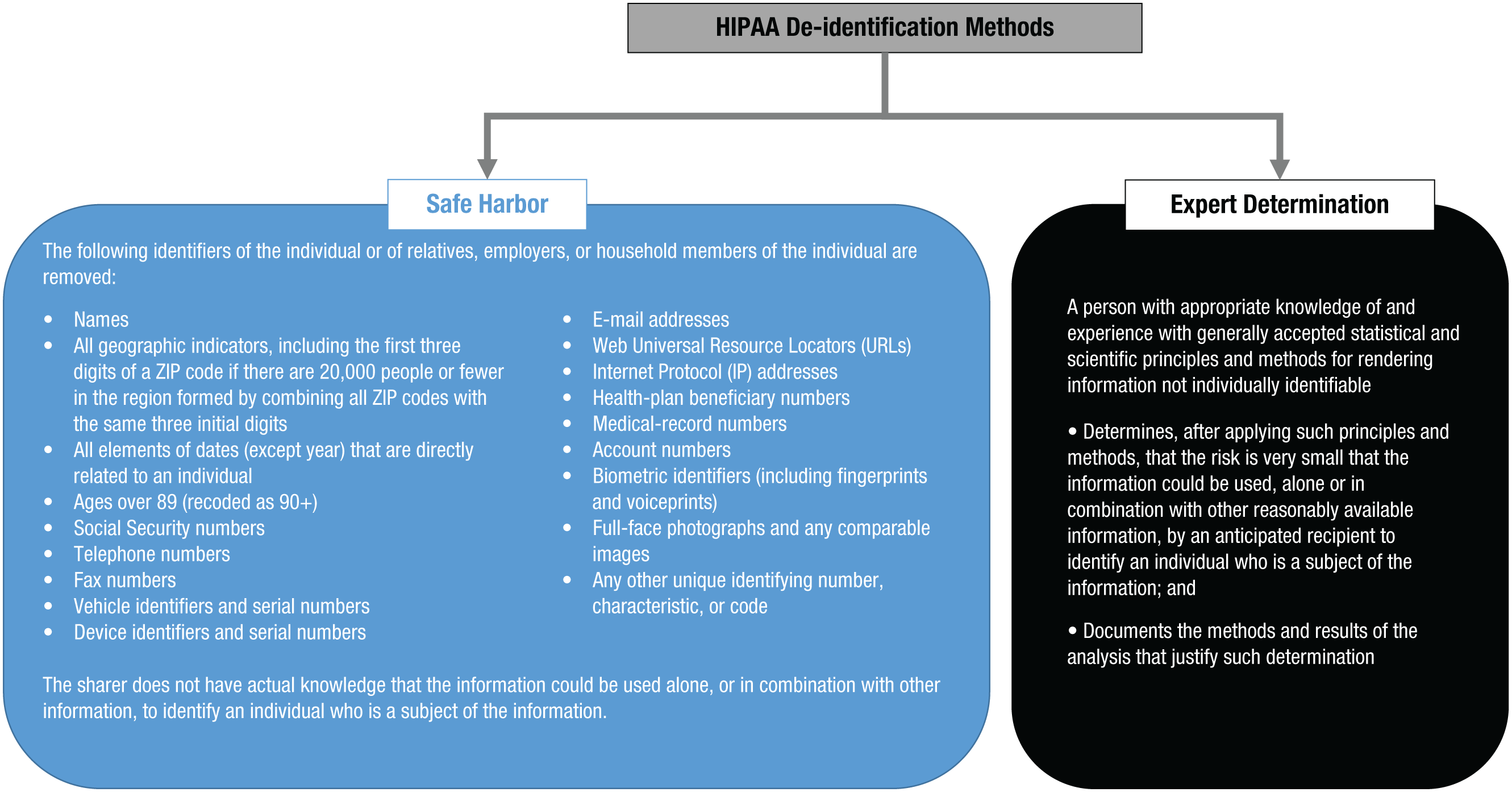
A summary of the two methods for de-identifying data to meet the requirements of the Health Insurance Portability and Accountability Act (HIPAA) Privacy Rule: safe harbor (left) and expert determination (right).
Though popular among institutional review boards, the safe-harbor method can, at times, lead to data sets that are suboptimal for various research projects (Malin, Benitez, & Masys, 2011). For instance, patients in dementia studies are often above the threshold of 90 years old. Thus, attempts to build a model predicting dementia using age as an ordinal value would be hampered by all ages above 89 being forced into a single value. Similarly, there are many research topics, such as seasonal affective disorder, for which timing over the course of the year is critical, but the safe-harbor method prevents meaningful studies on seasonality because no dates would be in the data. In recognition of the limitations of safe harbor, the HIPAA Privacy Rule provides a second de-identification approach called expert determination. In this approach, data can be disclosed in any format provided that an expert has shown that the risk of re-identification of the data is “very small” with respect to an anticipated recipient. However, this approach raises many open questions and challenges, particularly for clinical psychological and psychiatric research. Perhaps most important is the fact that HIPAA regulations provide no guidance on what is “very small” risk. Rather, it is left to the community to determine what is an appropriate level of risk.
De-identification of structured clinical data
Some of the data generated in the clinical psychology and psychiatry setting are highly structured. For example, patients’ demographics and the billing codes for diagnoses and procedures in the electronic medical record are reported according to well-defined taxonomies. An example of structured patient data is shown in Table 1. Because structured data are common in a variety of other domains (e.g., administrative data from the U.S. Census), many mechanisms for de-identifying structured data have been developed (Fung, Wang, Chen, & Yu, 2010; Gkoulalas-Divanis, Loukides, & Sun, 2014). We briefly review several mechanisms that are most relevant to sharing patients’ psychological and psychiatric data and mention how each influences re-identification risk and data utility.
An Example of Structured Clinical Data With Psychiatric Implications
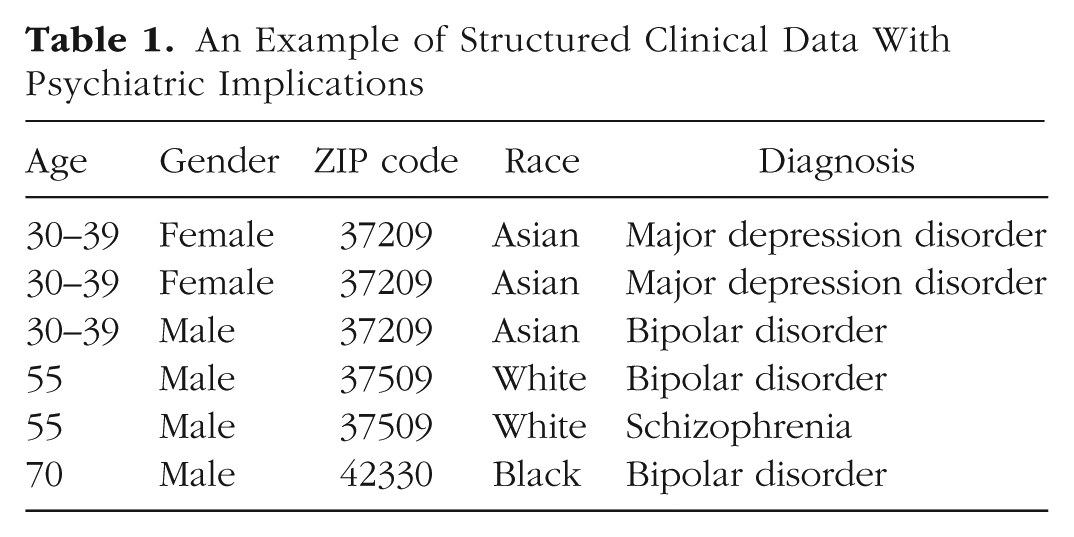
Data suppression, data generalization, and addition of noise are de-identification mechanisms that have been used for decades. Data suppression refers to removing certain records or values from the data—usually the ones with the highest probability of being leveraged for re-identification. For example, in Table 1, there is only one record for a 70-year-old Black man whose lives in ZIP code 42330. If the data recipient happens to know that a particular person with this description is represented in the data set, then the recipient may be able to re-identify this record uniquely. To prevent such exposure, the data sharer could drop this record from the data set. Though simple to apply, suppression can result in a substantial loss of information and, thus, influence the results of subsequent scientific investigations.
By contrast, generalization refers to replacing specific values in the data with coarser values. For example, in Table 1, the age values in the first three records have been replaced with a 10-year interval. Notably, generalization maintains truthfulness in the data, but it also influences the statistics of the data set (albeit in a more controlled manner than suppression). Generalization and suppression are often combined to form the optimal de-identification solution, which minimizes the modification of the original data while ensuring that re-identification risk is below some accepted threshold.
Addition of noise (e.g., via randomization) is another de-identification mechanism that has been adopted in various settings (Adam & Wortman, 1989; Willenborg & de Waal, 2001). This mechanism involves perturbing values by injecting noise using some known function, such as a Gaussian or Laplacian distribution. For example, a date could be changed to another date randomly selected from a fixed period of time. Although the addition of noise, when performed appropriately, maintains the values of certain summary statistics, it reduces the truthfulness of the data (Samarati, 2001). Thus, this mechanism is not appropriate when the truthfulness of the data is of vital importance. Furthermore, if the noise introduced to the data is optimized for a predefined analysis, the use of the data will be highly limited for subsequent analyses (El Emam et al., 2009).
New alternatives to these conventional mechanisms are being developed to generate simulated clinical data that retain statistical properties of relationships between the variables in the original data set. These new mechanisms often retain first-order relationships in the data, such as the means and variances of certain descriptive statistics. Most recently, it has been shown that generative adversarial networks, a variation of deep neural networks, can be learned from real-world electronic medical records (Choi et al., 2017). This is notable because these networks can capture highly complex nonlinear relationships between the variables. A generative adversarial network can subsequently be applied to simulate records. Sharing the simulated records produced by such neural networks may thwart re-identification, attribute disclosure, and even membership disclosure through the application of some controlled noise mechanism (e.g., differential privacy; Beaulieu-Jones et al., 2017). As for the utility of the simulated data, it has been demonstrated that they can be used in place of the real data in machine-learning models predicting various physical-health concerns (e.g., cardiovascular problems). Such methods may be applicable to psychological and psychiatric investigations as well. However, if the uses to which the shared data will be put are unknown (or not supported by the simulated data), the data sharer will still need to resort to conventional methods to de-identify the originally identified data.
De-identification of unstructured clinical data
Whereas structured data provide clear information about a patient, psychiatric notes are generally in plain text, or natural language. Natural language provides extremely rich analytic data, but de-identifying such text is arduous and fraught with complications compared with de-identifying structured data. It benefits from significant effort by researchers, health-care professionals, and information-systems teams. One of the core challenges remains correctly detecting and removing information that may identify an individual (e.g., a date, Social Security number, or name) in the unstructured text without incorrectly detecting and removing relevant clinical information about a patient’s mental-health status.
There are two types of approaches for detecting identifiers in such data: those based on rules and those based on artificial intelligence (AI). Regardless of approach, the process often begins with designing a two-class classifier that distinguishes between identifiers and nonidentifiers among all the terms in the text. There are three competing goals that need to be balanced in building a classifier that de-identifies data: high recall, high precision, and low cost. Recall (i.e., the proportion of real identifiers that are detected) and precision (i.e., the proportion of detected instances that are in fact real identifiers) reflect the risk of identity disclosure and the utility of the data, respectively. If recall is high, then the risk of identity disclosure is likely to be low. If precision is high, then the utility of the data is likely to be high. The cost of building a classifier includes, for example, the monetary cost of collecting and preprocessing the data and training the classifier with the resulting data. Higher precision and recall can sometimes be reached by collecting a larger data set (when using the AI approach) or defining a richer set of rules, but this effort can often be limited by budget and time requirements (Carrell, Cronkite, Malin, Aberdeen, & Hirschman, 2016). Therefore, the cost of building de-identification algorithms must be balanced with the utility of the data and the re-identification risk (M. Li et al., 2016).
Rules-based approaches tend to use a combination of pattern matching and dictionaries of terms (e.g., patients’ names) that should be discovered and transformed (Douglass et al., 2005). Although rules-based approaches can be tuned to achieve high fidelity in specific circumstances and the computational cost is low, they do not always lead to solutions that are scalable or generalizable, which is critical if psychological data are to be collected and studied in a big-data setting.
AI-based approaches invoke the assistance of modern AI technology to detect and transform identifiers. Application of an AI approach to de-identification was first demonstrated in 1996 (L. Sweeney, 1996). Since then, a variety of techniques (Meystre, Friedlin, South, Shen, & Samore, 2010) based on machine learning have been introduced; these range from decision trees (Szarvas, Farkas, & Busa-Fekete, 2007) to grammar parsers (e.g., conditional random fields; Wellner et al., 2007), and, most recently, deep learning methodologies, which have proven successful in a growing array of medical classification problems. These techniques have been shown to hold merit detecting identifiers as well (Liu et al., 2017). Some of these approaches have been deployed in open-source software (Aberdeen et al., 2010; Gardner & Xiong, 2009).
With respect to de-identification of psychiatric intake notes, various methodologies have been developed. These methodologies were on display in a recent public competition that used a corpus of 1,000 psychiatric intake notes (Stubbs, Filannino, & Uzuner, 2017). In comparison with previous sets of medical records used for de-identification challenges (Uzuner, Luo, & Szolovits, 2007), these data contained (a) a greater quantity of personal and social details about the patients (e.g., professions, hobbies, residences, and employers) and (b) more instances of personal identifiers but in a much larger quantity of narrative text (i.e., much lower identifier density per unit of text). The results of the competition showed that even the best existing out-of-the-box de-identification approach performed poorly (the balanced average of precision and recall achieved, called the F-measure, was .799, and recall was .733). Even when the participating teams were provided with gold-standard data for training their de-identification models, the highest performance scores were reasonable but still short of what one would want to see in practice (an F-measure of .914 and recall of .889). In particular, instances of professions, locations, and names were among the most difficult to detect, especially when contextual cues were limited. Still, the competition showed that the highest performance can be achieved when ensemble (or combination) approaches, rather than individual machine-learning methods, are applied.
Hiding in plain sight
Despite advances in processing of natural language, no system is likely to achieve 100% detection of all identifiers. Therefore, detection and redaction methods must be partnered with additional risk-mitigation strategies that address the problem of residual identifiers (i.e., identifiers that a human or computer failed to detect and are thus left in place). One such method is hiding in plain sight (HIPS; Carrell et al., 2013), which is illustrated in Figure 2.
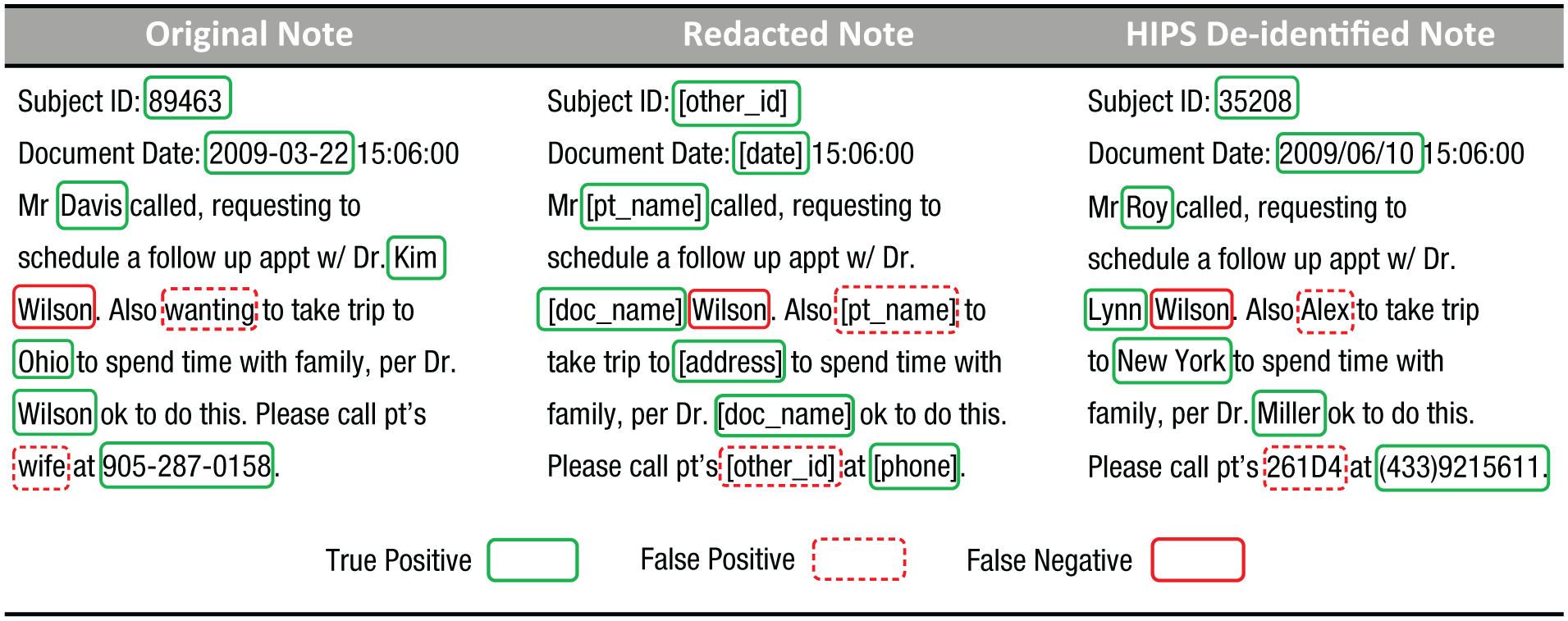

An example of an original clinical note (left), its redacted version (middle), and the note after a hiding-in-plain-sight (HIPS) resynthesis (right). True positives are real identifiers that were correctly detected as identifiers, false positives are nonidentifiers that were incorrectly detected as identifiers, and false negatives are real identifiers that were not detected as identifiers.
According to the theory behind HIPS, all detected identifiers should be replaced by fictitious surrogates that are similar semantically to what they are replacing (e.g., replace the name “Tom” with “Bob” and the date “7/30/2017” with the randomly generated date “5/28/2014”). As a result of these substitutions, the residual real identifiers will look similar to the surrogates, which allows them to hide in plain sight. Experiments with human readers suggest that HIPS can substantially reduce the possibility that data recipients will be able to detect residual identifiers. Still, care must be taken, as recent evidence suggests that if HIPS is insufficiently applied, a data recipient might be able to differentiate the signal from the noise (B. Li, Vorobeychik, Li, & Malin, 2017; M. Li et al., 2016).
Practical Examples: Vanderbilt University Medical Center’s Synthetic Derivative
To a certain extent, psychiatric data are already being shared through large research repositories for de-identified data. One example of such a resource, which has facilitated our own efforts to predict nonfatal suicide attempts (Walsh, Ribeiro, & Franklin, 2017), is the Synthetic Derivative (SD) at VUMC. The SD is a clinical-data repository with more than two decades’ worth of de-identified electronic medical records, including psychiatric notes, for more than 2 million patients (Roden et al., 2008). It has adopted a multilayered approach to privacy, by combining many of the protections described earlier in this article. First, it applies the safe-harbor model of de-identification to all of the structured clinical data (e.g., patients’ demographics). Second, it applies natural-language processing to detect and redact safe-harbor-designated identifiers in clinical notes. The SD was created before HIPS was invented, so to minimize privacy risks, the system was tuned to be conservative. Specifically, the de-identification engine was oriented to overredact when in doubt about the status of a term, in order to ensure that the rate of leaked identifiers would be sufficiently small. Given that there is residual risk in the semantics of the natural language itself (e.g., a clinical record might read, “the mayor’s wife” or “the quarterback of the university football team”), all users of the system must enter into a data-use agreement that explicitly prohibits attempts at re-identification. Although the agreement itself does not prevent re-identification, it does permit the institution to assign accountability and pursue penalties as deemed necessary. Finally, use of the data was originally restricted to Vanderbilt University or VUMC employees and students. As the system became more popular and outside investigators gained an interest in studying the data, they were allowed to do so provided that they paired with a local investigator who assumed responsibility for appropriate use and interpretation of the data.
Suicide research is a notable domain to consider in examining open-data concerns because of recent scientific advances enabled by the research infrastructure for clinical data (Barak-Corren et al., 2017; Kessler et al., 2015; Pestian et al., 2017; Walsh et al., 2017). In the SD setting, our colleagues and experts in suicide research were able to review patients’ records, including de-identified clinical text, to validate whether structured diagnostic codes truly described self-injury with suicidal intent. We discovered that in our health system, diagnostic codes for self-injury alone were often not associated with suicidal intent and were therefore inaccurate predictors of such intent nearly 42% of the time. The algorithms developed from this validation benefited from this discovery, which would not have been possible with a relatively small investment of time and money in the absence of the infrastructure and data-sharing capabilities of the SD.
On a broader scale, VUMC participates in various clinical-data research networks (CDRNs), such as the Mid-South CDRN (Rosenbloom et al., 2014), which itself is part of the broader Patient Centered Outcomes Research Network (PCORnet; Collins, Hudson, Briggs, & Lauer, 2014). Large CDRNs aimed at accelerating discovery in psychology and psychiatry are continuing to form around the United States. The Mental Health Research Network (MHRN), the Observational Healthcare Data and Informatics Network (OHDSI), and other large conglomerates within PCORnet are examples of successful research collaboration that translates discovery into practice (Ahmedani et al., 2017; Fleurence et al., 2014; Hripcsak et al., 2015).
All of these networks require data sharing in some form, whether they use a centralized model (i.e., data are deposited at one coordinating site) or a federated model (i.e., data remain at partner sites that adhere to a shared format, such as the Common Data Model). Most active CDRNs emphasize sharing of de-identified structured data in a federated model.
Conclusion and a Look Toward the Future
As the clinical psychology and psychiatry communities move toward open-science initiatives, they must take care to erect infrastructure that supports data sharing without inducing high privacy risks to patients. Risk in this setting is proportional to the potential harm that can transpire if clinical data are misused or abused in some fashion. As a result, it is up to the organizations managing clinical data to assess risks given the context in which the data will be made accessible. Although privacy regulations define de-identification, they do not preclude clinical psychologists and psychiatrists from adopting flexible mechanisms for making data available. As we illustrated in our discussion of VUMC’s SD, a strategy for mitigating privacy risk can combine a belt-and-suspenders approach in which de-identification is applied with contractual agreements that ensure accountability.
Still, clinical data may be shared in a tiered manner that takes into account the perceived risk associated with different contexts. If clinical data and the psychological or psychiatric status of a patient are to be shared in a restricted setting with only trusted users, then less obfuscation can be introduced, which would make the data more useful for research studies. However, if the data are shared in an unrestricted setting (i.e., deposited in a public repository that can be accessed by anyone in the world), then risk will be higher and a more conservative de-identification strategy may be more prudent. The challenges that face the clinical psychology and psychiatry communities include (a) characterizing plausible threats to privacy in their data, (b) determining the appropriate risk thresholds for different settings and recipients of the data, (c) defining how many tiers of access can be managed feasibly, (d) assessing how different de-identification mechanisms (e.g., generalization vs. randomization vs. generation of simulated data) influence the utility of clinical psychological and psychiatric data, and (e) harmonizing de-identification strategies across repositories so that individuals’ identities are protected.
If a tiered approach to access is applied, the fundamental tension between openness and privacy should be recognized. Specifically, as the tiers become more open, the risk of an attack on or misuse of clinical data grows. As a result, to ensure that the risk remains sufficiently low, it will be necessary to de-identify data to a greater extent, which will limit its reuse for validation of published findings and discovery of new associations.
Finally, it is critical to recognize that de-identification does not mean the data are devoid of re-identification risks—it means only that the re-identification risk is considered to be small.
Footnotes
Action Editor
Daniel J. Simons served as action editor for this article.
Author Contributions
C. G. Walsh and B. A. Malin jointly developed the idea for this manuscript. C. G. Walsh, W. Xia, M. Li, J. C. Denny, P. A. Harris, and B. A. Malin all contributed to writing and revising the manuscript and approved the final submitted version.
Declaration of Conflicting Interests
The author(s) declared that there were no conflicts of interest with respect to the authorship or the publication of this article.
Open Practices
The authors support open practices. This article does not report on data, methodology, or a preregistered study and therefore does not qualify for any of the badges.
