Abstract
Background:
In orthopaedic foot and ankle procedures, bone debris generated during osteotomies is typically discarded. However, this autograft bone debris has good handling properties and can be amenable for use as a stimulus to bone healing at the site of the osteotomy.
Methods:
In the present study, discarded bone debris was harvested intraoperatively during minimally invasive chevron Akin (MICA) hallux valgus corrections, isolated Akin osteotomies of the proximal phalanx of the great toe, cheilectomies, and calcaneal osteotomies from 9 participants.
Results:
Multiplex protein arrays of 40 cytokines and 41 growth factors identified 76 immunomodulatory and reparative proteins within the bone debris. Fifteen key growth factors and cytokines (VEGF, PDGF-BB, M-CSF, EGF-R, HGF, ICAM1, GCSF, TIMP-2, sTNFRII, MCP-1, GM-CSF, IL-6sR, IL-10, MCP-2, RANTES) were prominent, suggesting that bone debris proteins may have potential effects on immunomodulation and bone regeneration including cytokine-cytokine receptor interaction, interleukins, MAPK, PI3K-Akt, Wnt, BMP, and TGFβ signaling pathways.
Mesenchymal stem/stromal cells (MSCs) were isolated from bone debris of the osteotomies. MSCs expressed genes involved in bone and cartilage formation, homeostasis, angiogenesis, and immunomodulation. Fourteen genes were associated with maintaining cell stemness, whereas seventeen genes were linked to osteochondral development and spatial organization. Additionally, the study identified eight genes promoting angiogenesis and ten genes regulating immune responses in the mesenchymal stem cell environment.
Conclusion:
This repurposed surgical waste contains a concentrated array of growth factors, antiinflammatory mediators, and viable MSC that might enhance bone healing when reintroduced to surgical sites.
Clinical Relevance:
The results could serve as a foundation for repurposing previously discarded bone debris as autologous bone grafts for reimplantation in minimally invasive orthopaedic procedures to potentially enhance bone tissue healing. To confirm clinical relevance, further well-controlled trials are required to establish whether these findings improve bone healing rates and related patient-reported outcomes.
Keywords
Introduction
Bone graft substitutes and augmentation strategies represent a critical frontier in orthopaedic surgery, with the global bone graft and substitutes market projected to exceed $4.6 billion by 2030. 1 Despite advances in synthetic and allogeneic materials, autologous bone grafting remains the gold standard because of its osteogenic, osteoinductive, and osteoconductive properties.2,3 Traditional autograft harvesting extracts cancellous or corticocancellous bone from the calcaneus, tibia, or iliac crest. 4 However, this method requires removal of additional bone, and iliac crest harvests carry significant morbidity, including chronic pain, infection, and nerve injury in up to 25% of patients.5,6 This clinical challenge has driven the search for alternative autologous bone sources that minimize patient morbidity while maximizing therapeutic efficacy.
In orthopaedic foot and ankle surgery, procedures such as the minimally invasive chevron and Akin (MICA) procedure for hallux valgus correction, 7 cheilectomies for hallux rigidus,8,9 and sliding calcaneal osteotomies for hindfoot deformities 10 generate substantial quantities of bone debris through high-speed burr osteotomies. Remarkably, this potentially valuable autologous material is routinely discarded, a missed therapeutic opportunity. Previous studies have shown that mesenchymal stem/stromal cells (MSC) isolated from surgical waste, such as reamer-irrigator-aspirator (RIA) aspirates, retain native molecular signatures and express genes associated with multipotentiality, osteogenesis, adipogenesis, and angiogenic support. 11 Unlike bone marrow aspirates, which require separate harvesting procedures and contain limited solid matrix components,12,13 or cortical bone grafts that necessitate additional surgical sites, 14 surgical bone debris represents an immediately available, zero-morbidity autologous resource generated as an inherent byproduct of the primary procedure.
Despite theoretical advantages, bone debris’s bioactive profile in foot and ankle surgery is uncharacterized. Current literature addresses marrow- or cortical-derived therapeutics,15,16 leaving a gap for mechanically processed debris. Moreover, no previous studies have employed comprehensive multiplex protein arrays to profile the immunomodulatory and reparative protein content within surgical bone debris, limiting our understanding of its therapeutic mechanisms.
The clinical implications of successfully harnessing surgical bone debris are profound. Delayed union and nonunion rates in foot and ankle surgery range from 5% to 15% depending on the procedure,17,18 often requiring revision surgeries, prolonged nonweightbearing periods, and significant health care costs. 19 Enhanced bone healing through immediate autologous augmentation could dramatically reduce these complications while eliminating the need for synthetic substitutes or secondary harvest procedures.
This study represents the first comprehensive characterization of surgical bone debris generated during minimally invasive foot and ankle procedures, employing advanced multiplex protein arrays to quantify more than 40 key regenerative and immunomodulatory proteins. It is also the first study to extract, isolate, and harvest MSC from this source, targeting the unique profile of mechanically processed debris, an unexplored autologous resource. We hypothesize that this repurposed surgical waste contains a concentrated array of growth factors, antiinflammatory mediators, and viable MSC that could significantly enhance bone healing when reintroduced to surgical sites.
Materials and Methods
A translational laboratory study was performed using human tissue samples, with level III evidence.
Bone Debris Collection
Bony debris samples were collected from 9 patients, 5 males and 4 females, ranging in age from 40 to 63 years and body mass index from 20 to 40. The surgeries performed included 1 cheilectomy, 1 MICA, 3 lateral calcaneal slide osteotomies, 1 Cotton and medial slide osteotomy, 2 Lapidus/Akin procedure, and 1 first toe proximal phalanx Akin osteotomy. The indications for surgery included 4 halluces valgi, 1 halluces rigidi, 2 varus hindfoot deformities, and 2 pes planovalgus deformities. All collected bone debris was pooled, and analysis was not divided by site of origin.
Bone Debris Preparation and Protein Analysis
Bone debris was rinsed with saline and then frozen at –80 °C freezer. On experiment day, the bone debris was thawed, rinsed at least 3 times with 1X Dulbecco’s phosphate-buffered saline (DPBS; Sigma Aldrich), and lysed with RIPA lysis buffer (Gibco; Thermo Fisher Scientific) to release cellular proteins.
Multiplex protein arrays of 40 cytokines and 41 growth factors (RayBio C-Series; RayBiotech; Supplementary Table S1) were used to determine proteins present in 7 bone debris samples. Specifically, 1 mL of bone debris lysate was used for each assay following the manufacturer’s instructions. Data shown represent 30-second exposure in FluorChem E chemiluminescence imaging system (ProteinSimple). Results were quantified using ImageJ Fiji software (Fiji/ImageJ; National Institutes of Health). Results generated by the software were introduced into Microsoft Excel–based analysis software tools for the automatic computation of the extracted numeric data obtained from the array image (https://www.raybiotech.com/tools/array-analysis-tool, accessed March 10, 2024). All signal intensities were normalized with the background signal of each array as well as normalized to positive and negative signal to reduce batch variability. Finally, the quantified signal intensity for each protein spot was proportional to the relative concentration of the antigen in the sample determined by BCA protein assay (Thermo Fisher Scientific).
Isolation and Culture of Bone-Derived MSC
Bone debris was collected as described via the aforementioned surgical procedures, washed with saline to remove blood, and then immediately transferred to the laboratory in a sterile specimen container (Figure 1). The washed bone fragments were then cultured in complete growth medium (PromoCell). Cultures were maintained at 37 °C in a humidified atmosphere containing 5% CO₂. After 10 days, when outgrowth of adherent cells was observed, bone fragments were removed, and cells were allowed to expand to 70% to 80% confluence. Cell morphology was assessed using confocal microscopy (Dragonfly; Andor) (Figure 1D).

Bone extraction and MSC harvesting technique from minimally invasive orthopaedic procedures. (A) Saline-cooled burr used to drill into bone for intended surgery, while preventing thermal damage and maintaining MSC viability. (B) Bone debris rinsed with saline then transferred to sterile specimen container. (C) Bone debris cultured in growth media for 10 days. (D) Microscopic view of the isolated MSC, highlighting their density and characteristic morphology.
RNA Isolation and Gene Expression Analysis
Total RNA was extracted from cells using RNeasy Mini Kit (Qiagen) according to the manufacturer’s instructions. RNA was measured, prepared as 1-μg reactions using SuperScript VILO cDNA Synthesis Kit (Invitrogen), and transcribed to cDNA using a thermocycler. Subsequently, real-time quantitative polymerase chain reactions (qPCRs) were performed using RT² Profiler Arrays for Human Mesenchymal Stem Cells (Qiagen). Briefly, master-mixes for each MSC sample was prepared with SYBR Green Supermix, 100 µL cDNA (100 μg), and ultrapure water according to the manufacturer’s instructions for RT² profiler arrays, and 20 µL master-mix was added into each well of the array plates. Ct values were obtained and oriented into Excel spreadsheets for importing into Qiagen’s Data Analysis Center where ΔCt was obtained using beta-2-microglobulin housekeeping gene (B2M). Expression levels were calculated using the 2−ΔCt method.
Results
Seventy-six immunomodulatory and reparative proteins were identified in the harvested bone debris. These proteins enhance various biologic processes including angiogenesis, chondrogenesis, osteogenesis, immune function, regulation of cell proliferation, response to stimuli, cell migration, signal transduction, signal receptor activity, and protein phosphorylation.
Growth Factors in Viable Bone Debris
A diverse array of growth factors was detected in the bone debris isolated from foot and ankle osteotomies (Figure 2A). These include angiogenic factors (VEGF, VEGF R2, VEGF R3, VEGF D, and PIGF), fibroblast and epithelial growth factors (bFGF, FGF-4, FGF-6, FGF-7, HB-EGF), as well as TGF-α and TGF-β family members. Additionally, hematopoietic and immune-regulating factors (GM-CSF, G-CSF, M-CSF) and insulinlike growth factor binding proteins (IGFBP-1 to -6) were present.
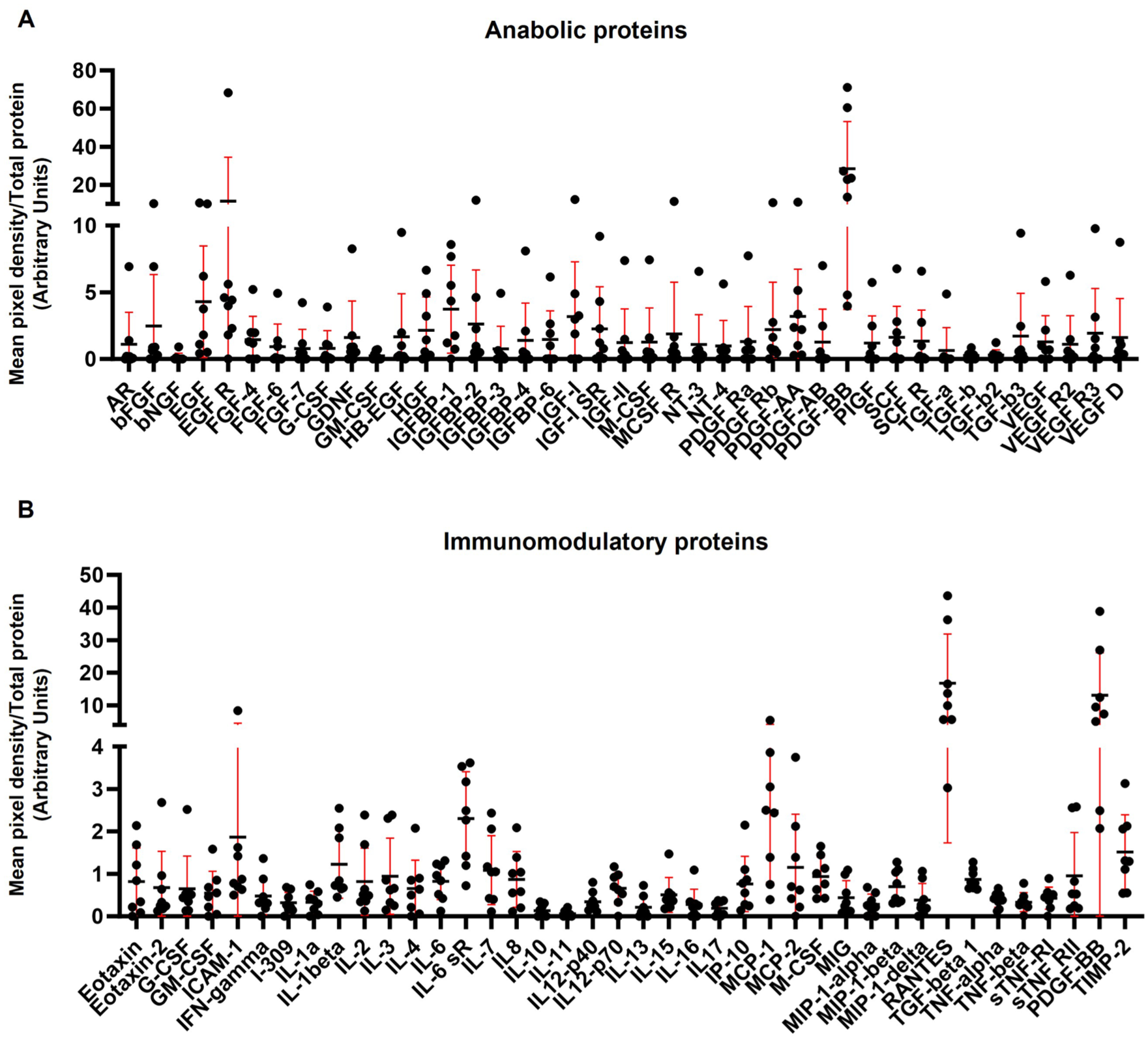
Protein profile of bone debris collected from minimally invasive foot and ankle surgeries reveals a diverse array of anabolic and immunomodulatory proteins. (A) Anabolic proteins present within bone debris include angiogenic factors (VEGF family and PIGF), fibroblast and epithelial growth factors (βFGF, FGF-4/6/7, HB-EGF), hematopoietic factors (GM-CSF, G-CSF, M-CSF), and TGF family proteins. (B) Identified cytokines encompass proinflammatory (IL-1α, IL-1β, IL-6, TNF-α, TNF-β, IFN-γ, IL-17) and antiinflammatory (IL-4, IL-10, IL-11, IL-13) types, along with MIP family members (MIP-1α, MIP-1β, MIP-1δ) and tissue inhibitor of metalloproteinases-2 (TIMP-2).
Immunomodulatory Proteins in Viable Bone Debris
Multiple immunomodulatory proteins were detected in the viable bone debris (Figure 2B). These include proinflammatory (IL-1α, IL-1β, IL-6, TNF-α, TNF-β, IFN-γ, IL-17) and antiinflammatory (IL-4, IL-10, IL-11, IL-13) cytokines. Several chemokines were also identified, including Eotaxin, Eotaxin-2, MCP-2, RANTES, IP-10, MIG, and MIP family members (MIP-1α, MIP-1β, MIP-1δ), which support immune cell recruitment. TIMP-2, a key regulator of extracellular matrix remodeling, was also identified.
Anabolic and Immunomodulatory Effects of Viable Bone Debris Proteins
Pathway analysis revealed that the detected proteins are involved in key biologic processes including cytokine-cytokine receptor interaction (46.9%), MAPK signaling pathway (34.6%), PI3K-Akt signaling pathway (38.3%), Ras signaling pathway (28.4%), Rap1 signaling pathway (23.5%), Jak-STAT signaling pathway (23.5%), signaling by interleukins (42.0%), and IL-10 signaling (9.8) (Figure 3). Identified proteins were grouped into 3 K-means clustering networks with elevated protein-protein interaction (PPI) enrichment (P < 1.0e−16) and an average local clustering coefficient >0.7, indicating strong biologic connections.

Immunomodulatory and anabolic pathways affected by viable bone debris proteins. Pathway analysis revealed 8 pathways upregulated that are involved in immunomodulatory and anabolic processes. Identified proteins were involved in cytokine-cytokine receptor interaction (46.9%), MAPK signaling (34.6%), PI3K-Akt signaling (38.3%), Ras signaling (28.4%), Rap1 signaling (23.5%), Jak-STAT signaling (23.5%), interleukin signaling (42.0%), and IL-10 signaling (9.8%).
Osteoinductive and Osteoconductive Effects of Viable Bone Debris Proteins
Proteins from viable bone debris are implicated in osteoinductive and osteoconductive processes critical for bone formation (Figure 4). Eight pathways were identified to be upregulated with these biologic processes: cell population proliferation (60.5%), response to stimulus (74.0%), cell migration (37.0%), signal transduction (69.1%), signaling receptor activity (9.9%), immune process (19.8%), protein phosphorylation (50.6%), and angiogenesis (16.1%).

Osteoinductive and osteoconductive biological processes affected by viable bone debris proteins. Pathway analysis revealed 8 pathways upregulated that are involved in osteoinductive and osteoconductive processes. Upregulated pathways include cell proliferation (60.5%), response to stimuli (74.0%), cell migration (37.0%), signal transduction (69.1%), receptor activity (9.88%), immune regulation (19.8%), protein phosphorylation (50.6%), and angiogenesis (16.1%).
MSC Gene Expression Profile
Microarrays for MSCs were used to build a comprehensive gene expression analysis, which revealed a sophisticated network of molecular regulators critical to osteochondral differentiation, MSC stemness, immunomodulation, and angiogenesis.
Stemness and Differentiation Characteristics
Fourteen genes were found to regulate MSC “stemness,” namely ANPEP, CD44, THY1, ALCAM, CTNNB1, CASP3, ERBB2, JAG1, NOTCH1, LIF, POU5F1, EGF, FZD9, and FUT4 (Figure 5A). This reflects the ability of cells to remain undifferentiated with potential for bone or cartilage formation.

MSC stemness and osteochondral differentiation gene expression profile. (A) Fourteen genes regulate MSC stemness and maintenance of undifferentiated state: ANPEP, CD44, THY1, ALCAM, CTNNB1, CASP3, ERBB2, JAG1, NOTCH1, LIF, POU5F1, EGF, FZD9, and FUT4. (B) Seventeen genes are key regulators of differentiation and spatial organization during osteochondral development: COL1A1, ITGB1, ITGAV, TGFB1, PTK2, HDAC1, RUNX2, SMURF1, KAT2B, ITGA6, BGLAP, BMP4, GDF5, GDF4, BMP2, BMP6, and FGF10. Heatmap representation (color schematic explained in Supplementary Table 2) of gene expression profiling.
Seventeen key osteochondral differentiation genes were discovered which mediate MSC differentiation and spatial organization during osteochondral development. These genes include COL1A1, ITGB1, ITGAV, TGFB1, PTK2, HDAC1, RUNX2, SMURF1, KAT2B, ITGA6, BGLAP, BMP4, GDF5, GDF4, BMP2, BMP6, and FGF10 (Figure 5B).
Vascular and Immunomodulatory Gene Function
There was increased expression of eight genes involved in the formation of new blood vessels (ENG, VEGFA, PDGFRB, ACTA2, FGF2, MCAM, HGF, and KDR; Figure 6A).
The immunomodulatory gene expression profile revealed ten genes involved in regulation of the inflammatory environment (ICAM1, MMP2, SMAD4, PPARG, IL6, GDF15, CSF3, TGFB3, IL1B, and CSF2; Figure 6B).

Vascular and immunomodulatory gene expression profiles. (A) Angiogenesis-related genes showing increased expression of eight genes involved in the formation of new blood vessels: ENG, VEGFA, PDGFRB, ACTA2, FGF2, MCAM, HGF, and KDR. (B) Immunomodulatory gene expression profile revealing ten genes involved in regulation of the inflammatory environment: ICAM1, MMP2, SMAD4, PPARG, IL6, GDF15, CSF3, TGFB3, IL1B, and CSF2. Heatmap representation (color schematic explained in Supplementary Table 2) of gene expression profiling.
Discussion
Our findings demonstrate that viable bone debris from foot and ankle osteotomies contains a rich reservoir of bioactive molecules critical for bone regeneration, supporting its potential value as an autologous graft material. This approach aligns with the 4 fundamental components of bone tissue engineering: growth factors/morphogens, stem cells, biomaterials, and mechanical stimulation. 15
Protein Reparative and Immunomodulatory Functionality
This study confirms the presence of several key growth factors and signaling molecules essential for bone repair. Vascular endothelial growth factor (VEGF) plays a dual role in bone regeneration by enhancing angiogenesis and directly influencing osteoblast differentiation,20 -23 whereas platelet-derived growth factor-BB (PDGF-BB) accelerates collagen formation and endothelial cell proliferation. 24 The presence of EGF-R suggests potential for enhanced cellular proliferation, differentiation, and survival during the healing process. 25 TGF-β and FGF family members further complement this regenerative milieu by facilitating the recruitment and differentiation of osteoprogenitor cells into functional osteoblasts.26,27 The copresence of these growth factors is particularly significant, as their synergistic interactions likely amplify their individual effects, suggesting potential for accelerated bone repair.
Surgical interventions often cause significant damage to bones and surrounding tissues, triggering an immune response that, if uncontrolled, can exacerbate bone degradation. The presence of both proinflammatory (IL-1α, IL-1β, IL-6, TNF-α) and antiinflammatory cytokines (IL-4, IL-10, IL-13) suggests a capacity for self-regulation of the inflammatory microenvironment. IL-10, in particular, helps prevent excessive inflammation that could delay healing, 28 whereas sTNFRII acts as a decoy receptor to neutralize TNF-α activity, 29 TIMP-2 presence indicates protection against excessive extracellular matrix degradation, 30 ensuring appropriate tissue remodeling during regeneration. Importantly, the chemokine profile (including Eotaxin, MCP-2, and MIP family members) suggests capacity for targeted immune cell recruitment to coordinate the phagocytosis of debris and initiate tissue repair cascades without excessive inflammatory damage.31 -33
Signaling Pathways
The proteins identified in this study are central to critical signaling pathways, including MAPK, PI3K-Akt, and JAK-STAT, which regulate angiogenesis, osteogenic transcription factors, and the regenerative environment for bone repair. Specifically, the MAPK and PI3K-Akt pathways are central regulators of osteoblast proliferation and differentiation,34,35 whereas the Rap1 pathway influences cell adhesion and cytoskeletal organization critical for cellular migration and matrix production during bone formation.36 -38 Additionally, the JAK-STAT pathway enables rapid signal transduction in response to growth factors and cytokines, directly influencing osteoblast gene expression. 39 Moreover, the pronounced involvement of interleukin signaling (42.0%) and IL-10 signaling (9.8%) reflects the importance of immunomodulation in creating an environment conducive to bone regeneration. 40 Together, these interconnected pathways provide nutrients and signals to optimize the healing environment and support osteogenic production.
MSC Reparative and Immunomodulatory Functionality
The successful isolation and culture of MSC from bone debris highlights another crucial aspect of bone debris reparative capacity. The gene expression profile of MSC indicates their capacity for multilineage differentiation and tissue regeneration.
The stemness markers (including CD44, THY1, and POU5F1) confirm that MSC maintain their multipotent capacity, whereas the expression of osteochondral differentiation genes (COL1A1, ITGB1, RUNX2, SMURF2) demonstrates their propensity toward bone and cartilage formation. COL1A1 encodes the primary structural protein in bone matrix (alpha 1 chain of type I collagen). Mutations in COL1A1 are associated with osteogenesis imperfecta, leading to brittle bones. 41 ITGB1, which encodes integrin beta-1, a transmembrane receptor involved in cell adhesion and signaling, contributes to osteoclast function and bone resorption. RUNX2 is a master regulator of osteoblast differentiation. 42 Furthermore, SMURF2, a E3 ubiquitin ligase, regulates bone homeostasis by modulating the interaction between SMAD3 and the vitamin D receptor in osteoblasts, favoring osteoclastogenesis and bone remodeling. 43
Bones require oxygen, nutrients, and growth factors to heal, 44 which can all be acquired when angiogenesis is upregulated. The expression of angiogenic genes (VEGFA, ENG, and PDGFRB) supports our hypothesis that reimplantation of viable bone debris could stimulate bone repair. VEGFA encodes vascular endothelial growth factor A, coupling angiogenesis to endochondral and membranous bone formation during skeletal development and repair. 45 ENG encodes endoglin, a coreceptor for TGF-β that is essential for vascular development in bone. 46 PDGFRB encodes the platelet-derived growth factor receptor beta, which regulates MSC function and osteogenic differentiation. Moreover, PDGFRB signaling in osteoblast lineage cells controls bone resorption through upregulation of CSF1 expression, influencing osteoclastogenesis and bone remodeling. 47 Finally, ACTA2 encodes smooth muscle alpha-actin 2, which is involved in the contractile function of smooth muscle cells, coordinates nutrient delivery involved in bone formation and osseointegration. 48
The immunomodulatory gene profile further supports the potential for MSC to regulate the local inflammatory environment, creating conditions conducive to repair rather than chronic inflammation. Ten key genes (ICAM1, MMP2, SMAD4, PPARG, IL6, GDF15, CSF3, TGFB3, IL1B, and CSF2) coordinate the delicate balance between inflammation and repair. ICAM1 facilitates critical interaction with immune cells, particularly macrophages, essential for transitioning from inflammation to repair. 49 MMP2 contributes to extracellular matrix remodeling and activates latent growth factors, amplifying regenerative signaling cascades.50,51 As a critical mediator of TGF-β signaling, SMAD4 orchestrates antiinflammatory effects while supporting osteoblast differentiation and modulating T-cell responses. 52
PPARG expression enables MSC to shift macrophage polarization toward an antiinflammatory M2 phenotype, promoting tissue repair. 53 The simultaneous expression of proinflammatory (IL6, IL1B, CSF2) and antiinflammatory (TGFB3, GDF15) cytokine genes reflects capacity for context-dependent immune regulation throughout the healing process.54,55 CSF3 and CSF2 regulate myeloid cell recruitment and polarization toward phenotypes supporting bone regeneration,56,57 whereas TGFB3 exhibits potent antiinflammatory properties through induction of regulatory T cells that dampen excessive immune responses.58,59 GDF15, a divergent member of the TGF-β superfamily, reduces macrophage-mediated inflammation while promoting tissue regeneration in models of inflammatory bone loss. 60
This coordinated expression pattern enables MSC to sequentially coordinate inflammation, repair, and remodeling phases through precise immune regulation. By creating conditions that favor bone regeneration while preventing chronic inflammation, the immunomodulatory capacity of these cells complements their angiogenic and osteogenic properties, enhancing their therapeutic potential.
Limitations and Future Directions
This study provides valuable insights into the bioactive potential of viable bone debris for autologous grafting; however, limitations must be considered. The study was conducted as a proof of concept rather than a comparative analysis to traditional autografts. It was also conducted as an in vitro analysis of bone debris with a small sample size (n = 9). Larger studies with more participants would be necessary to confirm these results and ensure broader applicability across different patient populations and surgical procedures. Finally, although this study highlights promising proteomic and molecular profiles, further in vivo research, including animal models and clinical trials, is needed to assess the long-term viability, functionality, and safety of using bone debris as an autologous graft material in orthopaedic surgery.
Conclusion and Clinical Significance
This study introduces a novel and potentially transformative approach to enhancing bone healing in orthopaedic surgery by repurposing previously discarded bone debris as an autologous graft source. Our research challenges this conventional paradigm by demonstrating that this biologically active debris, particularly when produced using saline-cooled burrs, retains essential osteogenic properties, growth factors, and MSCs that can contribute to accelerated bone repair. To incorporate this material back into the patient to augment healing, we propose collecting the debris during surgical drilling and immediately reimplanting it directly into the surgical site. This method could be used either alone or in combination with established biologic stimulants such as BMPs and PDGF-BB, 61 and may offer a simple, cost-effective method to improve outcomes for fusions, osteotomies, or other bone healing procedures such as fracture repairs, nonunion treatments, and joint arthrodesis. To confirm clinical relevance, further well-controlled trials are required to establish whether these findings improve bone healing rates and related patient-reported outcomes.
Supplemental Material
sj-pdf-1-fao-10.1177_24730114251398510 – Supplemental material for Proteomic and Molecular Analysis of Viable Bone Debris From Foot and Ankle Osteotomies: Implications for Autologous Grafting
Supplemental material, sj-pdf-1-fao-10.1177_24730114251398510 for Proteomic and Molecular Analysis of Viable Bone Debris From Foot and Ankle Osteotomies: Implications for Autologous Grafting by Kirin Cromer, Kevin Liebmann, Noah Gabor, Steven Steinlauf, Thomas M. Best and Dimitrios Kouroupis in Foot & Ankle Orthopaedics
Footnotes
Appendix
Cytokines and Growth Factors Arrays.

| Cytokines Array | Growth Factors Array |
|---|---|
| Eotaxin | AR |
| Eotaxin-2 | βFGF |
| ICAM-1 | βNGF |
| IFN-γ | EGF |
| I-309 | EGF R |
| IL-1α | FGF-4 |
| IL-1β | FGF-6 |
| IL-2 | FGF-7 |
| IL-3 | G-CSF |
| IL-4 | GDNF |
| IL-6 | GM-CSF |
| IL-6 sR | HB-EGF |
| IL-7 | HGF |
| IL-8 | IGFBP-1 |
| IL-10 | IGFBP-2 |
| IL-11 | IGFBP-3 |
| IL12-p40 | IGFBP-4 |
| IL12-p70 | IGFBP-6 |
| IL-13 | IGF-I |
| IL-15 | IGF-I SR |
| IL-16 | IGF-II |
| IL-17 | M-CSF |
| IP-10 | MCSF R |
| MCP-1 | NT-3 |
| MCP-2 | NT-4 |
| MIG | PDGF Rα |
| MIP-1α | PDGF Rβ |
| MIP-1β | PDGF-AA |
| MIP-1δ | PDGF-AB |
| RANTES | PDGF-BB |
| TNF-α | PIGF |
| TNF-β | SCF |
| sTNF-RI | SCF R |
| sTNF-RII | TGF-α |
| TIMP-2 | TGF-β |
| TGF-β2 | |
| TGF-β3 | |
| VEGF | |
| VEGF R2 | |
| VEGF R3 | |
| VEGF D |
Ethical Considerations
Ethical approval for this study was waived by the University of Miami IRB as it is not human research (based on the nature of the samples as discarded tissue).
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Study supported by NIH/NIAMS grant number 5R21AR080388-0.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article. Disclosure forms for all authors are available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
