Abstract
Oxidative stress induced by excessive levels of reactive oxygen species (ROS) underlies several diseases. Therapeutic strategies to combat oxidative damage are, therefore, a subject of intense scientific investigation to prevent and treat such diseases, with the use of phytochemical antioxidants, especially polyphenols, being a major part. Polyphenols, however, exhibit structural diversity that determines different mechanisms of antioxidant action, such as hydrogen atom transfer (HAT) and single-electron transfer (SET). They also suffer from inadequate in vivo bioavailability, with their antioxidant bioactivity governed by permeability, gut-wall and first-pass metabolism, and HAT-based ROS trapping. Unfortunately, no current antioxidant assay captures these multiple dimensions to be sufficiently “biorelevant,” because the assays tend to be unidimensional, whereas biorelevance requires integration of several inputs. Finding a method to reliably evaluate the antioxidant capacity of these phytochemicals, therefore, remains an unmet need. To address this deficiency, we propose using artificial intelligence (AI)-based machine learning (ML) to relate a polyphenol’s antioxidant action as the output variable to molecular descriptors (factors governing in vivo antioxidant activity) as input variables, in the context of a biomarker selectively produced by lipid peroxidation (a consequence of oxidative stress), for example F2-isoprostanes. Support vector machines, artificial neural networks, and Bayesian probabilistic learning are some key algorithms that could be deployed. Such a model will represent a robust predictive tool in assessing biorelevant antioxidant capacity of polyphenols, and thus facilitate the identification or design of antioxidant molecules. The approach will also help to fulfill the principles of the 3Rs (replacement, reduction, and refinement) in using animals in biomedical research.
Introduction
The production of reactive oxygen species (ROS) as by-products of cellular metabolism constitutes the mechanistic basis of oxidative stress in living organisms. Oxidative stress caused by excessive levels of ROS is long established as a contributing or causal factor in the pathophysiology of many chronic and degenerative diseases, because it engenders oxidative damage to the mitochondrial membrane, other cellular membranes, and cellular components such as proteins and nucleic acids. 1 Therapeutic strategies to protect against oxidative damage are, therefore, a subject of active and intense scientific investigation to date, including the use of antioxidants.
The potential of structurally diverse phytochemicals found in seeds, fruits, and vegetables to afford disease prevention (chemoprevention) and health promotion has been linked to their antioxidant action. Certain classes of phytochemicals that have been shown to exhibit antioxidant action include polyphenols (e.g., flavonoids and bioflavonoids), phenolic acids, carotenoids, trienols, and so on. 2 In addition to antioxidant action, however, many of these phytochemicals exhibit other health-promoting benefits through a variety of non-antioxidant mechanisms. 3 It would, therefore, require novel and more encompassing assays to profile the health benefits of phytochemicals reliably and fully. Distinct pharmacological actions elicited by phytochemicals that exhibit polypharmacology require uniquely distinct chemistries as a basis for validated assay methodology.
The structural diversity of phytochemical antioxidants also warrants that bespoke assay methods be developed to profile the antioxidant potential of various classes of antioxidant compounds. 4 Different chemistries would usually be optimal for different pharmacophores because the structural details often predict the most likely underlying mechanism(s) of action. The proposition that the antioxidant action of polyphenols is mediated through hydrogen atom transfer (HAT), while carotenoids act through single-electron transfer (SET), represents a form of structure–function relationship. The hydrophilic-lipophilic balance (HLB) of the compounds is also a reliable predictor of the most likely site(s) of action in vivo. The HLB of compounds relative to the details of the assay protocol (aqueous or non-aqueous environment) could indicate the biorelevance of methods. 5 A review of antioxidant chemical assay literature, therefore, concluded that there was yet no official method for an antioxidant assay (and perhaps, still is none), due to a variety of problems associated with the methodologies, which question the biorelevance of assay results. 6
It was reported about 20 years ago that the biorelevance of antioxidant chemical assays is questionable because most of the assays are one-dimensional. 7 The bioactivity of drugs and other xenobiotics administered orally depends on the bioavailability of the drugs/xenobiotics and subsequent drug action at the tissue or cellular level. Bioavailability is the rate and extent of absorption and is influenced by permeation through gut wall, a process that could be influenced by gut-wall metabolism and first-pass metabolism during hepatic circulation.
Three distinct dimensions can be identified in the process of antioxidant action exhibited by, for instance, polyphenols in living organisms:
Permeability, which depends on the physicochemical properties of the compound, for example its lipophilicity and ionization constant 8
Metabolism of the compound, either by gut-wall or first-pass metabolism during hepatic circulation. Metabolism typically affects the fraction of the compound that exists as intact molecule and thus affects the fraction of the dose that is absorbed. 9
HAT to trap ROS through phenolic bond cleavage of polyphenols 1
These three dimensions are illustrated in Figure 1 .
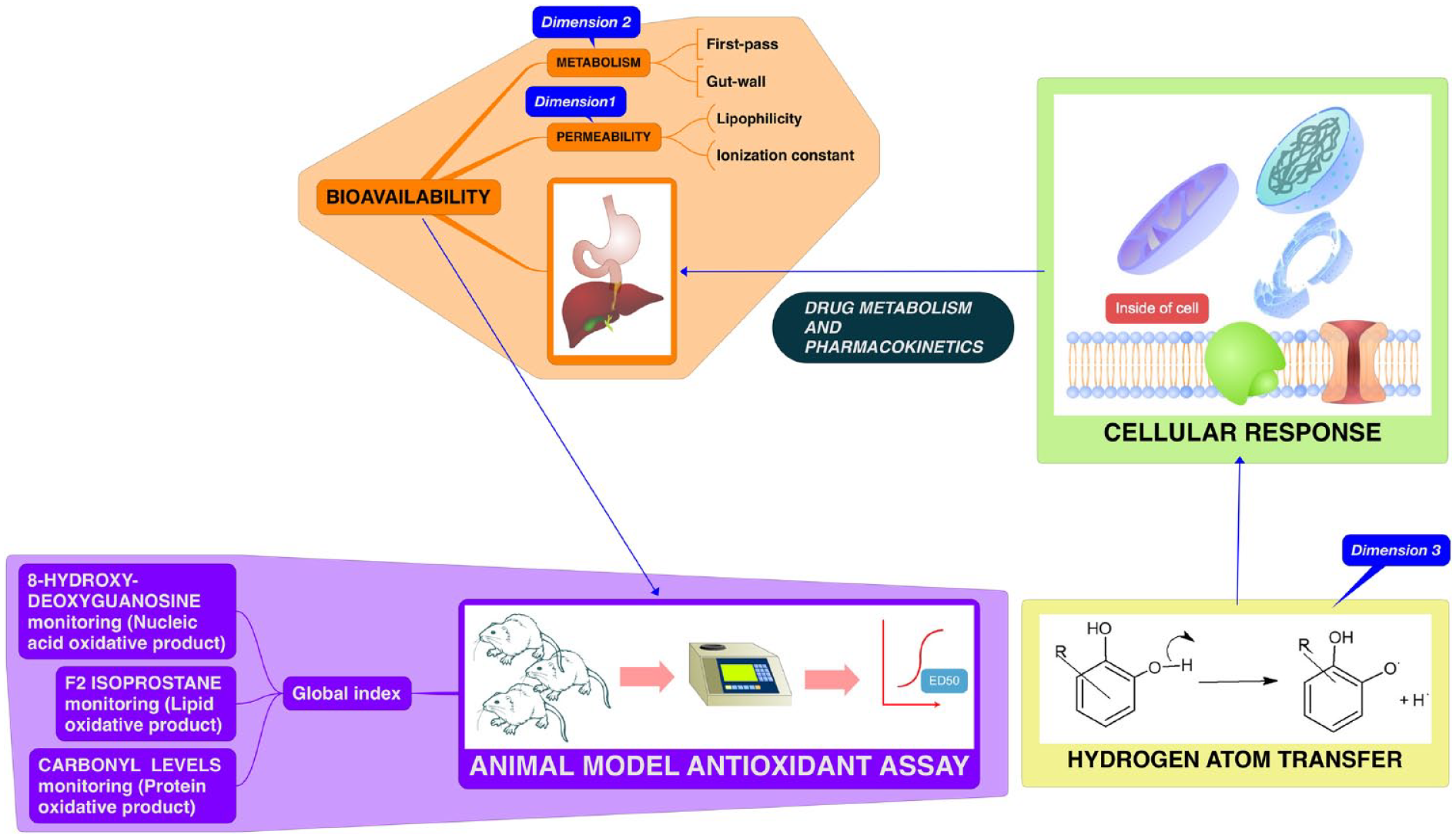
Schematic illustration showing interdependence of physicochemical properties and molecular behavior as regulators of antioxidant action in a biological system, and hence required input variables in a machine learning (ML) model for predicting biorelevant antioxidant capacity.
In vitro antioxidant assays based on chemical reactions or cell-based methodologies are usually deficient in one or more of the dimensions described above, leading to biologically irrelevant assay results. There is, therefore, a critical need for novel methodological platforms to overcome these deficiencies, which, among other benefits, could improve the translatability of preclinical drug discovery findings.
Application of Computational Systems Biology for Improving Biorelevance of Antioxidant Assays: Artificial Intelligence (AI) and Machine Learning (ML)
Kitano described computational systems biology as a combination of experimental and computational approaches to system-level understanding of biology. 10 This strategy was proposed to be capable of producing biological insights and predictions, the type required to obtain biorelevant measurement of antioxidant capacity.
In recent times, machine learning (ML), a subfield of artificial intelligence (AI), has been demonstrated to afford a useful strategy of relating a dependent variable (e.g., antioxidant action of phytochemicals in a biological system) to several independent variables. A learned algorithm that emerges from rule discovery based on several observations can then become a predictive tool for other usage environments involving newer but related members. Such an ML model was recently adopted to distinguish between different genders, ethnicities, and ages of volunteers, through mass spectrometry and imaging of latent fingerprint that was largely composed of lipids in sweat. The ML algorithm sorted a very large amount of chemical information provided by tandem mass spectrometry to pinpoint specific molecules that are different from one person to another. 11
A Global Index as a Reliable Measure of Oxidative Damage (Output Variable)
In an antioxidant assay context, a proposed ML workflow is displayed in Figure 2 . The goal is to relate the output of an animal model antioxidant assay to molecular descriptors, either experimental or calculated, that are relevant to the three dimensions of permeability, metabolism, and HAT. Several biomarkers have been investigated as indicators of oxidative damage in biological systems, which typically apply to lipids, proteins, and nucleic acids.
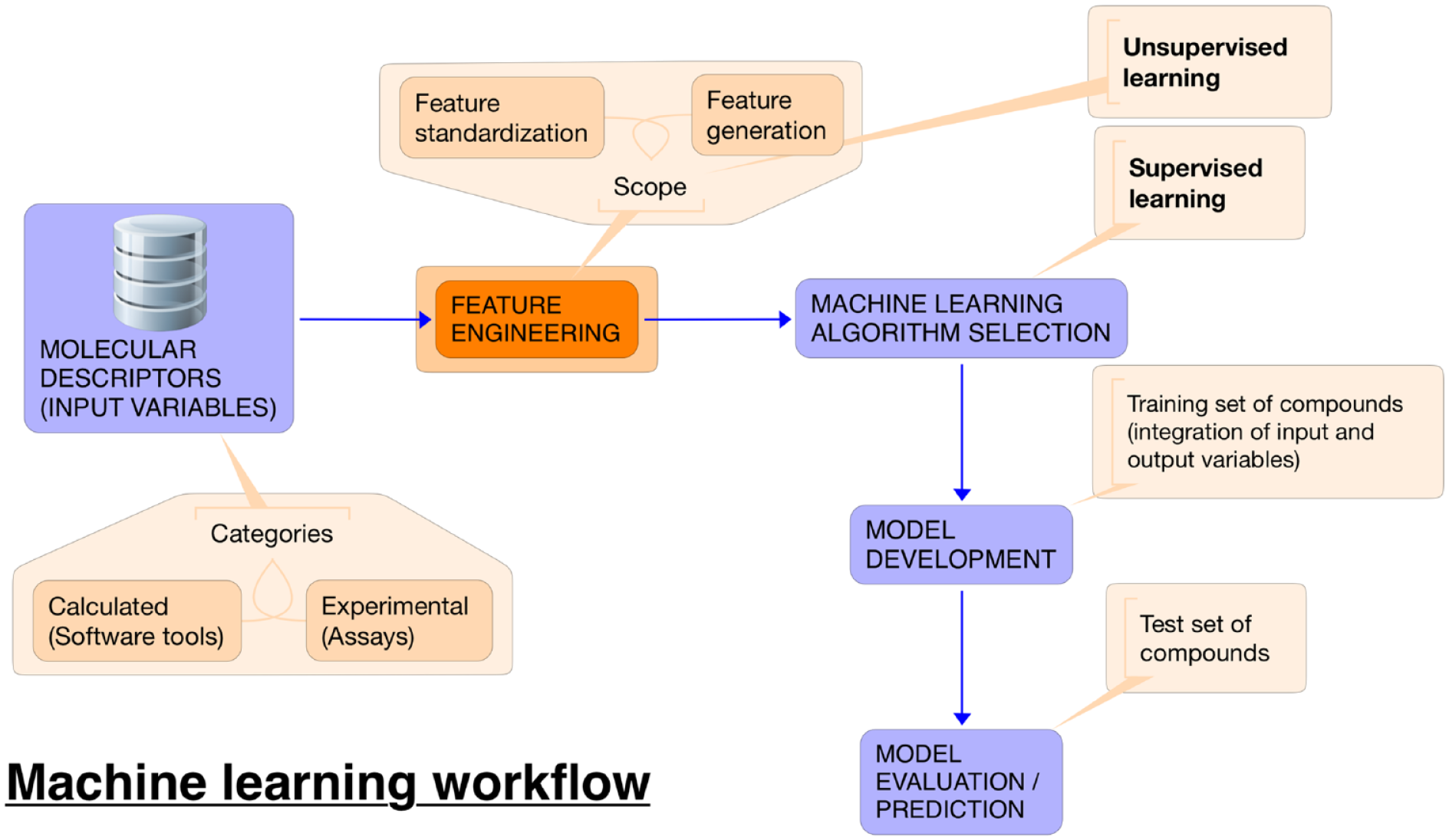
Overview of machine learning workflow for prediction of biorelevant antioxidant capacity.
Measurement of Lipid Peroxidation
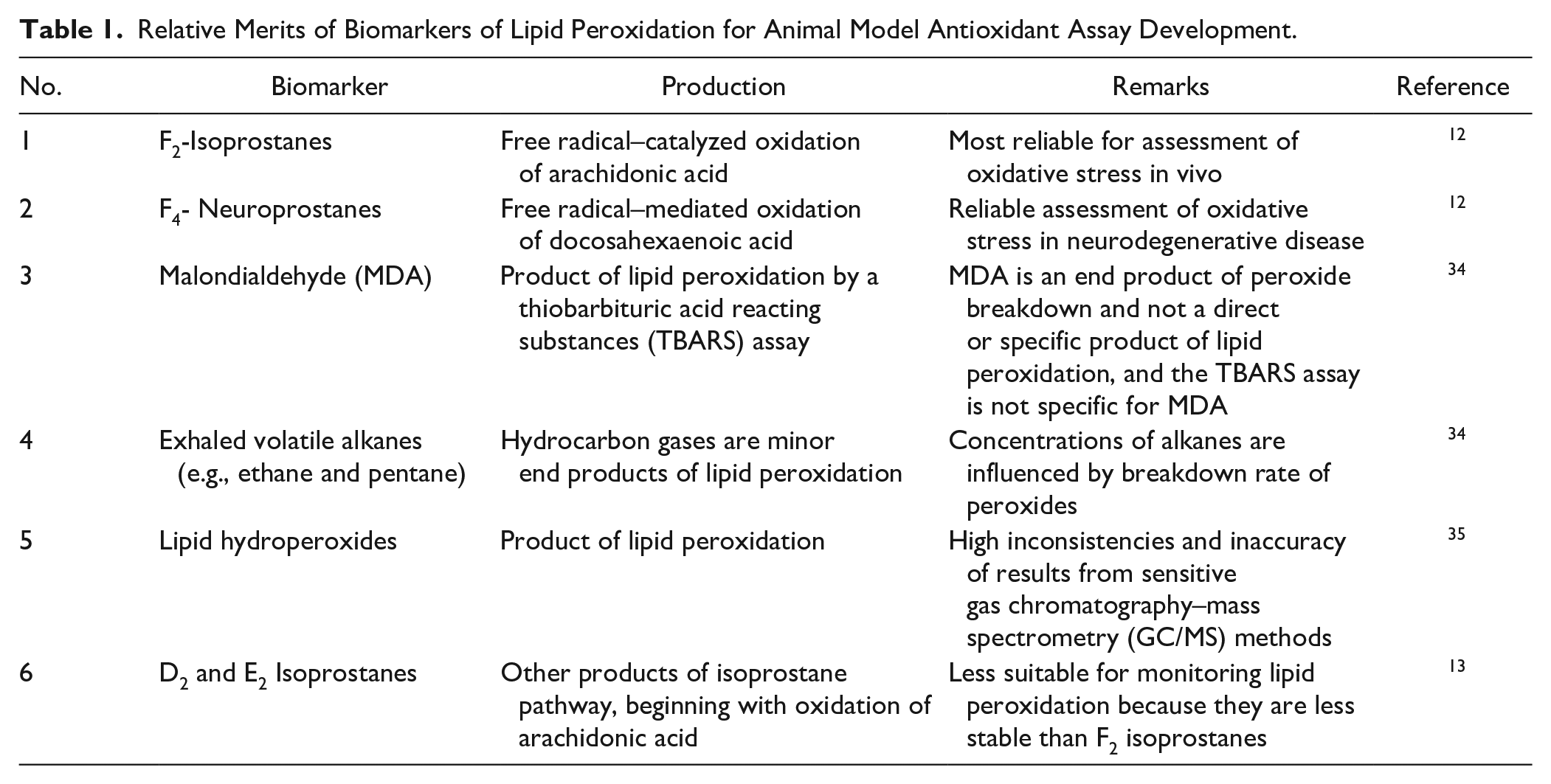
Several biomarkers of lipid peroxidation have been investigated to monitor the status of oxidative stress in vivo, but they vary widely in their reliability. The relative merits of some key biomarkers are displayed in Table 1 . F2-isoprostanes have emerged as one of the most reliable biomarkers for assessment of oxidative stress in vivo, and their measurement has implicated oxidative stress in the pathophysiology of several diseases, as shown in Figure 3 . 12 Their superiority as a biomarker is due to certain distinctive and favorable characteristics: 13 (1) They are chemically stable compounds; (2) they are a specific product of lipid peroxidation; (3) they are produced in vivo; (4) they are found in detectable amounts in all normal tissues and biological fluids, which in turn permits definition of a normal range; (5) in animal models of oxidant injury, a substantial increase in their levels is obtained; (6) they are unaffected by lipid content of diet; and (7) they might provide a sensitive biochemical basis for studies aimed at dose optimization for antioxidants.
Relative Merits of Biomarkers of Lipid Peroxidation for Animal Model Antioxidant Assay Development.
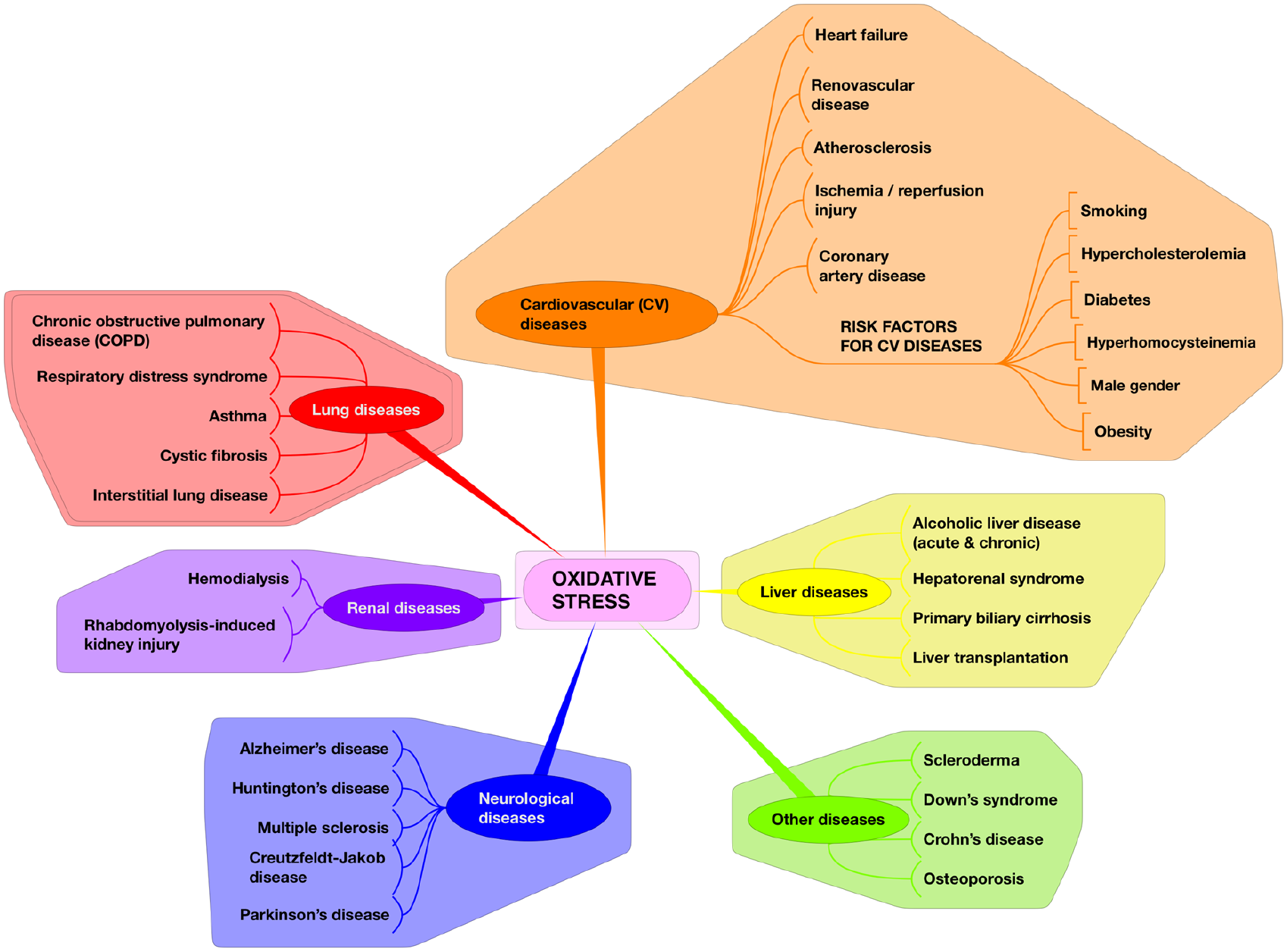
Oxidative stress as a key mechanism in the pathophysiology of several diseases. Measurement of F2-isoprostanes has implicated oxidative stress in the pathophysiology of several diseases affecting many target organs. This underscores the potential of antioxidant chemical entities as adjuncts to standard treatments, chemopreventive agents for disease prevention and health promotion, and candidates for novel therapeutics (see Ref. 12 and references cited therein).
F2-isoprostanes are produced by non-enzymatic, free radical–catalyzed peroxidation of arachidonic acid. The measurement of F2-isoprostanes is, therefore, recognized to be a reliable tool to assess the status of oxidative stress in vivo 12 and could be adapted for an antioxidant capacity assay. This will provide the response variable for ML.
Despite the attractiveness of isoprostanes as biomarkers for antioxidant assay development, their formation is impaired at elevated oxygen tensions, because of the mechanism of formation of F2-isoprostanes. The outcome is that their measurement as a marker of lipid peroxidation is insensitive in settings of elevated oxygen tension, such as hyperoxia-induced lung injury. Formation of isofurans are favored in such situations and are considered a better measure of lipid peroxidation. 12 This fact illustrates the limitation of using a single biomarker to measure oxidative stress.
Measurement of Oxidative Damage to Proteins
Protein oxidation results in an increase in protein carbonyl levels and glutamine synthetase activity. 14 The protein carbonyls can be assayed by stable hydrazone derivatives formed when they react with 2,4-dinitrophenylhydrazine. 15
Measurement of Oxidative Damage to Nucleic Acids
Oxidative damage to DNA is caused by modification in bases by ROS. This leads to genetic defects if the modifications are not repaired. The DNA base guanine is sensitive to oxidation, which makes 8-hydroxydeoxyguanosine (8-OHdG) a suitable biomarker for oxidative injury. 16
A global index integrating measures of oxidative injury to lipids, proteins, and nucleic acids will give a more robust monitoring of oxidative damage in biological systems. It will also afford a better metric for evaluating the efficacy of phenolic antioxidants as output variables for building ML models. The value of a global index of oxidative damage was proposed in evaluating the clinical significance of biomarkers of oxidative stress. 17
ML Workflow
Molecular Descriptors (Input Variables)
The numerical indices that constitute molecular descriptors are either experimental or calculated metrics that define molecular behavior with respect to permeability, metabolism, and HAT. Possible descriptors are displayed in a fishbone diagram in Figure 4 .
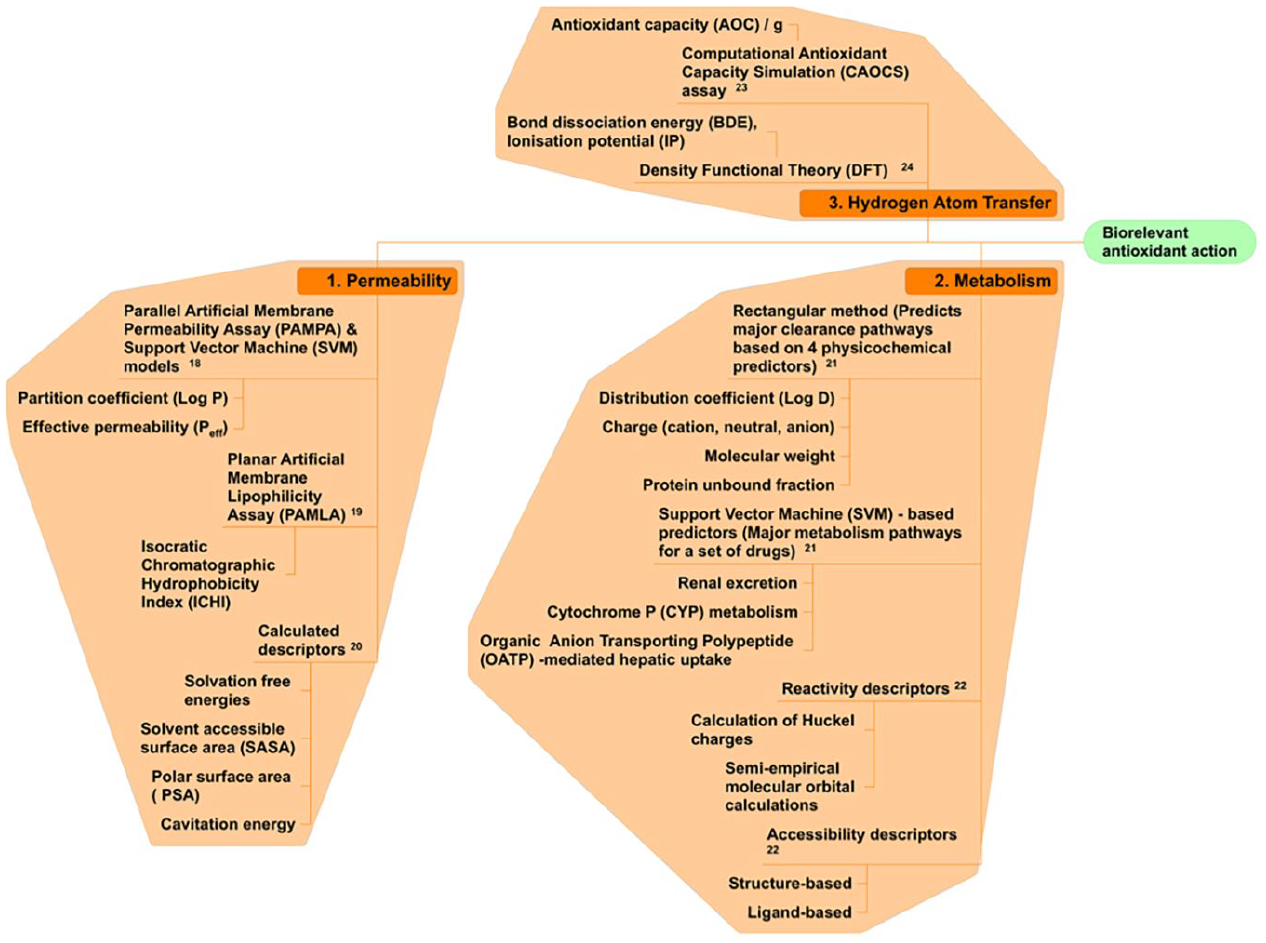
Schematic illustration of parameters (molecular descriptors) associated with each of the three dimensions governing in vivo antioxidant activity (biorelevant antioxidant action)—permeability, metabolism, and hydrogen atom transfer—with potential for use as input variables in developing a machine learning (ML)-based antioxidant capacity (AOC) assay.
Permeability of drug molecules can be measured by the planar artificial membrane permeability assay (PAMPA), a popular experimental technique with measures that are related to partition coefficient (Log P) and effective permeability (Peff). 18 An alternative descriptor obtainable from a planar artificial membrane lipophilicity assay was described as the isocratic chromatographic hydrophobicity index (ICHI). 19 In addition, some computational molecular descriptors have been shown to predict permeability. These include solvation-free energies, solvent-accessible surface area (SASA), polar surface area (PSA), and cavitation energies. 20
The major clearance pathway of drugs is predicted by a rectangular method that is based on four physicochemical predictors: distribution coefficient (Log D), charge (cation, neutral, and anion), molecular weight, and protein unbound fraction. A support vector machine-based predictor of metabolism was developed to be based on renal excretion, cytochrome P (CYP) metabolism, and organic anion transporting polypeptide (OATP). 21 In addition to these, computational descriptors have been developed for predicting metabolism of drugs. These include reactivity descriptors (e.g., the calculation of Huckel charges and semi-empirical molecular orbital calculations) that approximate the hydrogen and electron abstraction processes fundamental to the CYP catalytic cycle. In addition, there are accessibility predictors, which could be structure-based (using docking approaches to insert the molecule into the CYP binding pocket) or ligand-based (using molecular fingerprints to approximate steric hindrance and orientational concerns), designed to complement reactivity metrics. 22
The kinetics of HAT is an important rate process at the cellular level that distinguishes phenolic antioxidants from each other. Effective antioxidants can transfer hydrogen atoms much faster than ineffective compounds, and the pattern is a function of their chemical structure. Informatics tools that relate the chemical structure of compounds to their effectiveness as antioxidants are, therefore, potentially useful in ML model building for antioxidant assays. A bespoke assay for the antioxidant action of phenolics adapted proton transfer kinetics modeling as a surrogate rate process for measuring HAT. The metric, described as antioxidant capacity, integrates the proton transfer reaction constant and the concentration of standard test solutions in a manner that captures the structure–function relationship. This is a reasonable descriptor of kinetics of phenolic bond cleavage. 23
Density functional theory (DFT) was also applied to compute bond dissociation energy (BDE) and ionization potential (IP) as computational molecular descriptors of cleavage of the phenolic O–H bond, 24 which could be adapted to ML model building.
Feature Engineering
The most challenging part of the ML workflow is, arguably, knowing what data to feed into the models. The main question to be answered is “How can we find the most relevant data to train the models to solve the specific problem?” Three key considerations to identify the most relevant data are: (1) the right data attributes from which to infer realistic patterns and rules, (2) enough samples to train the model, and (3) engineering the data the right way (e.g., aggregate, integrate, transform, etc., the data) before feeding into the models. Feature engineering comprises feature generation and feature standardization. Feature generation entails looking at the attributes of the available data and, second, integrating and testing a new data source. Given the large number of possible molecular descriptors that represent potential input variables, it is a dataset without structure. Unsupervised learning is often used to develop a structure from a large dataset. Feature standardization refers to tools that make data “more appealing” to the model (e.g., imputation, normalization, and scaling). 25
ML Algorithm Selection
A well-trained and validated ML model based on a large library of polyphenols could provide a predictive tool for evaluating the potential of newer polyphenols for effective antioxidant action in biological systems. ML models that are applied in quantitative structure–activity relationship (QSAR) studies and potentially useful for developing AI-based antioxidant assays are classified into (a) supervised learning, and (b) unsupervised learning. Examples of techniques under each of the two groups are displayed in Figure 5 .
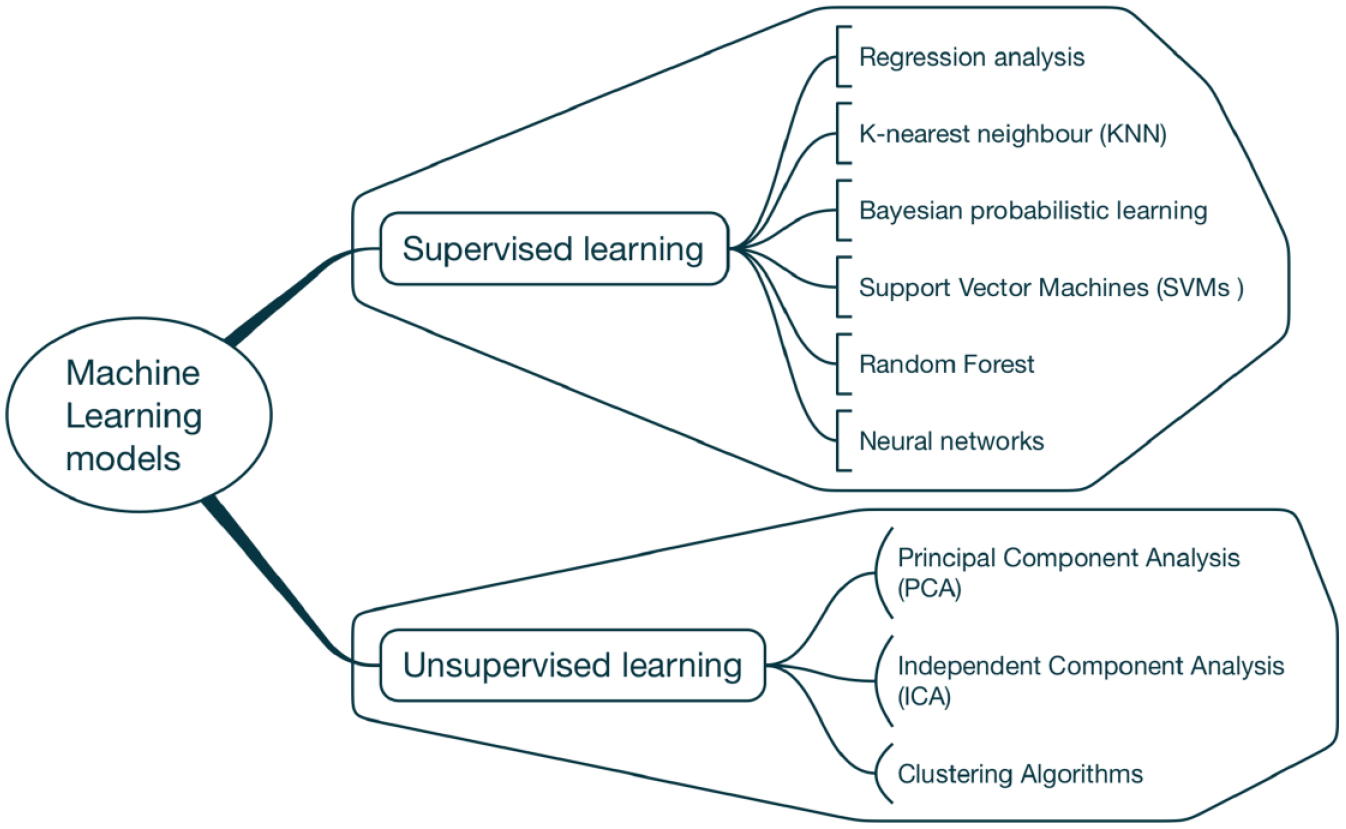
Supervised and unsupervised machine learning (ML) models. ML models that are applicable in quantitative structure–activity relationship (QSAR) studies consist of an array of techniques broadly classified as supervised and unsupervised learning.
In supervised learning, labels are assigned to training data, and once trained, the model can predict labels for given data inputs; whereas, in unsupervised learning, the techniques learn underlying pattern of molecular features directly from unlabeled data, such as dimensionality reduction techniques (principal component analysis and independent component analysis) and clustering algorithms. Clustering algorithms invariably involve (1) dividing a dataset by predefined distance metrics in high-dimensional space, and (2) assigning labels based on a number of observed categories. 26 While unsupervised learning is particularly important for feature generation, supervised learning is more important for selection of ML algorithms.
Artificial Neural Networks (ANNs) and Deep Learning (DL)
The autonomous knowledge acquisition from the molecular properties of compounds requires the use of ML techniques such as k-nearest neighbor (k-NN) and artificial neural networks (ANNs). An ANN is derived from the biological concept of neurones possessing dendrites, soma, axon, and synaptic dendrites. An ANN behaves as a neurone by working on three layers. The input layer takes input (like dendrites), the hidden layer processes the input (soma and axon), while the output layer sends the calculated output. The variants of this technique are supervised, unsupervised, and reinforced neural networks. 27 Deep learning, a branch of ANN that has found successful application in medicinal chemistry, can be described as a class of representation-learning techniques that are able to discover, from the raw data, multiple-level representations of increasing complexity by composing nonlinear models. 28
k-NN Algorithm
The k-NN algorithm assumes that similar things exist in close proximity and can be used to solve both classification and regression problems. The algorithm works by finding distances between a query and all the examples in a data, selecting the specified number examples (k) closest to the query, and then voting for the most frequent label (i.e., mode) in the case of classification. In contrast, it votes for the average of the labels (i.e., mean) in the case of regression. It has a major drawback of becoming slow as the size of the data in use grows. 29
Support Vector Machine
A support vector machine (SVM) works on the principle of margin calculation. Basically, margins are drawn between the classes in such a manner that the distance between the margin and the classes is maximum, hence minimizing the classification error. 27
SVMs have gained some popularity in recent times for developing in silico models based on large datasets to facilitate the task of lead optimization in drug discovery science. For example, support vector regression and classification models were developed based on oral permeability data obtained from PAMPA measurement of more than 4000 compounds in the same laboratory. 18 Likewise, an in silico model for prediction of major clearance pathways based on CYP enzymes, OATP-mediated hepatic uptake, and renal excretion was developed and optimized, using variants of SVM models and more than 800 descriptors, and the model with feature-selected descriptors was found optimal with respect to predictive power. 21
Bayesian Probabilistic Learning
This learning algorithm is mainly used for clustering and classification purposes. The underlying architecture of naïve Bayes depends on conditional probability. It creates trees, known as Bayesian networks, based on their probability of happening. 27
Random Forest
Decision trees are the types of trees that group attributes by sorting them based on their values. They are used mainly for classification purposes. Each tree is made up of nodes and branches. Each node represents attributes in a group that is to be classified, and each branch a value that the node can take. 27 Random forest (an example of ensemble methods) builds multiple decision trees and merges them to get a more accurate and stable prediction. An important advantage of random forest over simple decision trees is that it can be used for both regression and classification problems. 30
Model Development
Input and output variables from a dataset generated from a training set of antioxidant polyphenols will be fed into the selected algorithm to construct the AI model. The predictive power of the model depends on the quality of feature engineering, the number of compounds in the training set, and the performance of the selected algorithm.
Model Evaluation and Prediction
To validate the usefulness of the predictive model, a test set of compounds in the same chemical/biological class (e.g., phenolic antioxidants), which were not part of the training of the model, is assembled, and the predicted output is compared with experimental output. The training of the model might go through several iterations of feature generation to improve the performance of the predictive model and hence the reliability of prediction made by AI.
Merits and Demerits of ML in Predicting Biological Response
The merits of machine learning consist in the array of powerful techniques to explore the nonlinearity that characterizes structure–activity relationships (SARs). The palpable fact is that biorelevant antioxidant capacity measurement is multidimensional, and a dynamic nonlinear interaction between the dimensions is expected in biological systems. This characteristic is uniquely suited for ML exploration because it affords a robust integration of several parameters associated with the various dimensions, in a nonlinear way, in defining a single output variable.
Deep learning has outperformed traditional techniques in some drug design–related applications, principally due to some peculiar characteristics of biological and chemical data, such as complexity, uncertainty, diversity, and high dimensionality. 28
Some drawbacks of applying ML techniques include the problem of overfitting of data, which is minimized by different strategies for different techniques. An ensemble of classifiers was proposed to minimize the inherent weaknesses of each ML technique; however, this approach generated other weaknesses like decreased comprehensibility of the model, increased storage, and increased computation requirement. 31 Other drawbacks of applying deep learning include the limited number of data in certain areas of study and difficult interpretation of the chemical and biological mechanisms involved in deep learning models. 28
Part of what is outstanding in current antioxidant research is to optimize the choice and dose of antioxidants in investigating the impact of antioxidant therapy on amelioration of specific diseases. In addition to creating a global index, because antioxidant efficacy depends on the dose of the antioxidant administered, one way to incorporate this variable into the ML workflow (input) is to ensure the assays that monitor the respective biomarkers incorporate the dose–response curve of a well-known oxidant. The assay output will thus have the quality of being able to detect antioxidant or pro-oxidant effects of a fixed dose of the antioxidant. The ED50 for the model oxidant used as positive control will be compared with determined ED50 after pre-treatment/treatment with the antioxidant under study (i.e., pre-treatment with antioxidant, followed by treatment with the same antioxidant in the presence of the oxidant). The fact that polyphenols exhibit concentration-dependent cellular response, with their antioxidant effect profile typically found at relatively low concentrations (usually in the low micromolar range), warrants the dose optimization exercise for polyphenol antioxidants. 23
A multidimensional assay as proposed in this perspective will facilitate our understanding of the clinical pharmacology of antioxidants ( Fig. 6 ) by integrating the bioavailability and biodisposition of candidate antioxidants in evaluating their antioxidant potential. Such a model could facilitate molecular design tailored to optimize bioactivity and used to design candidate molecules that could be useful as powerful adjuncts in the treatment of diseases such as cancers and degenerative conditions, 32 which are linked to oxidative stress. 33
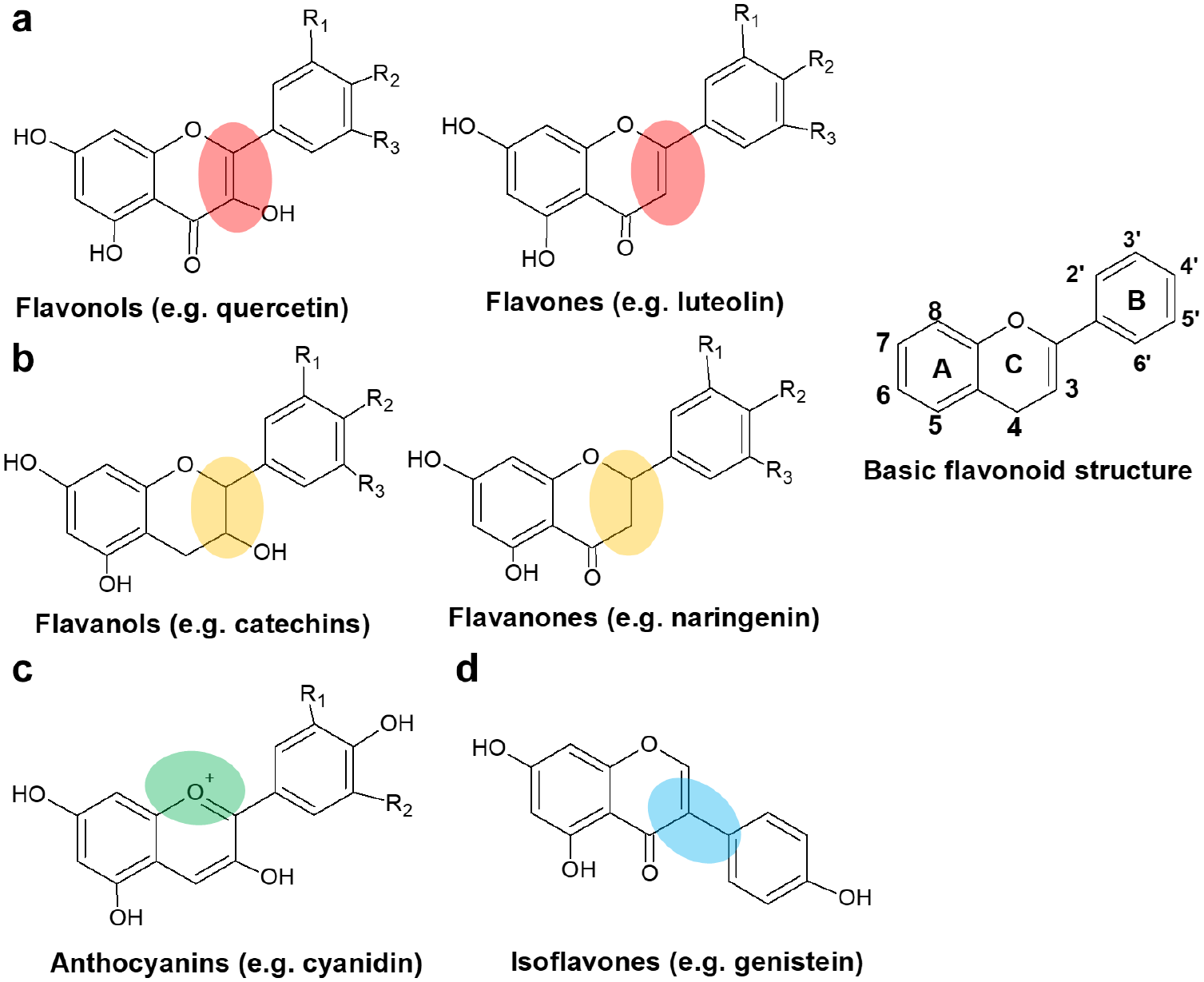
Chemical structures of different groups of antioxidant flavonoids (polyphenols) showing the basic structure and six major subclasses. (
Conclusion
Artificial intelligence–based approaches such as machine learning hold significant promise for improving the biorelevance of in vitro biological assays, including antioxidant assays, because machine learning models will have exceptionally better inherent accuracy in predicting in vivo outcomes than current in vitro assay models. Their deployment will also facilitate and accelerate the realization of the principles of the 3Rs (replacement, reduction, and refinement) with respect to the use of animals in research.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
