Abstract
Systems biology methods, such as transcriptomics and metabolomics, require large numbers of small model organisms, such as zebrafish embryos. Manual separation of mutant embryos from wild-type embryos is a tedious and time-consuming task that is prone to errors, especially if there are variable phenotypes of a mutant. Here we describe a zebrafish embryo sorting system with two cameras and image processing based on template-matching algorithms. In order to evaluate the system, zebrafish rx3 mutants that lack eyes due to a patterning defect in brain development were separated from their wild-type siblings. These mutants show glucocorticoid deficiency due to pituitary defects and serve as a model for human secondary adrenal insufficiencies. We show that the variable phenotypes of the mutant embryos can be safely distinguished from phenotypic wild-type zebrafish embryos and sorted from one petri dish into another petri dish or into a 96-well microtiter plate. On average, classification of a zebrafish embryo takes approximately 1 s, with a sensitivity and specificity of 87% to 95%, respectively. Other morphological phenotypes may be classified and sorted using similar techniques.
Introduction
Zebrafish are increasingly popular model organisms in biological research. This is largely due to their transparent embryos, which are produced in large numbers and allow for detailed in vivo microscopy, as well as high-throughput drug screening approaches. 1 Originally used mainly for developmental biology, zebrafish have recently been used to address questions in a variety of biological disciplines, ranging from neurobiology and behavior to cancer research and toxicology.2–5 The endocrinology of the zebrafish equally shows many similarities to that of mammals, even at larval stages. 6 This includes the glucocorticoid hormone system. For example, larvae homozygous for a strong mutant allele of the rx3 gene exhibit a glucocorticoid deficiency resembling human secondary adrenal insufficiencies due to defects in pituitary formation. 7 Originally, these mutants were isolated in a screen for eye development, 8 and their lack of eyes is a salient morphological feature that allows for easy identification of homozygous mutant animals. Recent research has focused on systems biology studies, including transcriptomics and metabolomics using this model, revealing a strong impact of glucocorticoids on daily patterns of transcription and metabolism. 9 Due to the small size of the larvae, large quantities of embryos need to be sorted for such time-course experiments. Manual separation of mutant embryos from wild-type embryos is a tedious and time-consuming task that is prone to errors. Additionally, separation of embryos is difficult at early stages of development, and especially if there are variable phenotypes of a mutant. Automation of mutant phenotype recognition and separation is therefore highly desirable. Two systems for sorting zebrafish embryos into a 96-well plate have been published.10,11 However, sorting embryos by features is not possible yet with these systems. For this reason, a new fish embryo sorting system was designed that is a further development with significant enhanced functionality of a previously presented system. 12 There are several requirements and challenges for sorting phenotypically different zebrafish embryos, such as rx3 mutants, at an early stage of development (i.e., prior to hatching) from phenotypically wild-type embryos:
The embryo may be orientated in any three-dimensional position within the chorion ( Fig. 1A–D ).
Some wild-type embryos may have weakly pigmented eyes due to pigmentation variants in the genetic background ( Fig. 1C ).
Dirt, empty chorions, coagulated eggs, algae, hatched larvae, or fish scales may be present ( Fig. 1E–G ).
Blurring artifacts may arise due to movements of the embryo or misaligned focus.
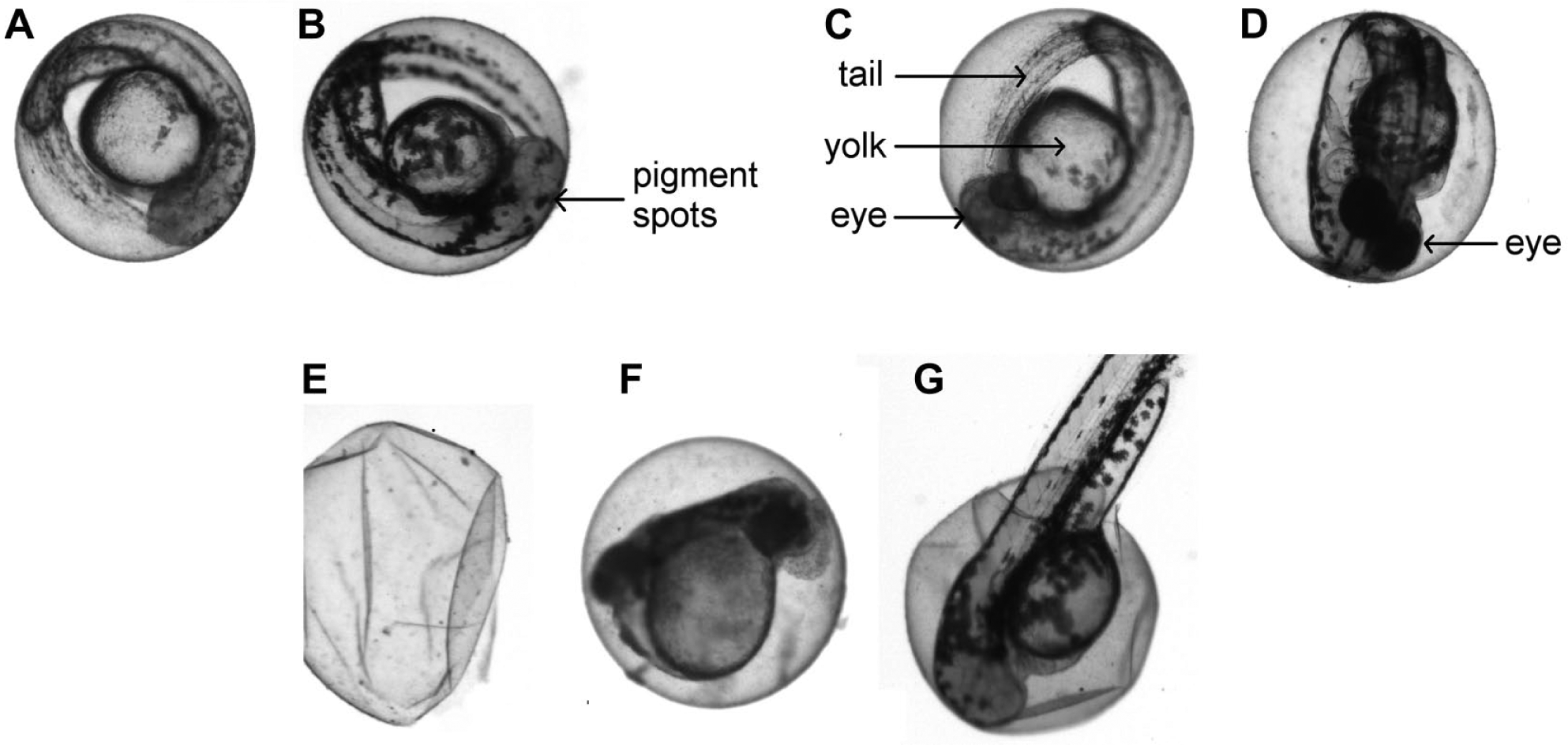
Examples of rx3 embryos without eyes that have to be sorted (
So far, different algorithms have been described in the literature for automated segmentation and for phenotypic differentiation, and newer methods continue to be investigated.1,13–15
Concept and Methods
The new sorting system consists of two cameras and a suction hose that are positioned by a three-axis linear robot, a mounting plate for the petri dish and the well plate, and below the mounting plate, a LED panel for bright-field illumination ( Fig. 2 ). The robot platform and pipetting module are further developments of a zebrafish handling system displayed in more detail in a former publication. 12 First, a standard petri dish with a diameter of 90 mm has to be fed with 100–150 zebrafish embryos. 16 After that, an overview camera, C1 (IDS UI-5240SE-C, IDS, Obersulm, Germany), in combination with a lens with a focal length of 10.27 mm (CMFA 2020ND, Lensation, Karlsruhe, Germany), is positioned by the robot over the petri dish with the embryos to be analyzed and sorted. With a resulting camera resolution of 29 µm and image processing methods, most of the dark eyes in zebrafish embryos with normal pigmentation can be detected ( Fig. 1D ). This allows for identifying and excluding the majority of embryos that do not comply with the phenotype of rx3 mutants. Additionally, the precise positions of all embryos within the petri dish are cached and all remaining embryos that potentially match the phenotype of a rx3 mutant ( Fig. 1A–C ) are inspected by the second camera, C2. The same camera model is used as for the overview process. However, a more detailed view of a single zebrafish embryo is achieved by using a telecentric lens with a magnification of 4× (TC2040, Lensation). This enables better distinction between the rx3 mutant phenotype and variable wild-type phenotypes.
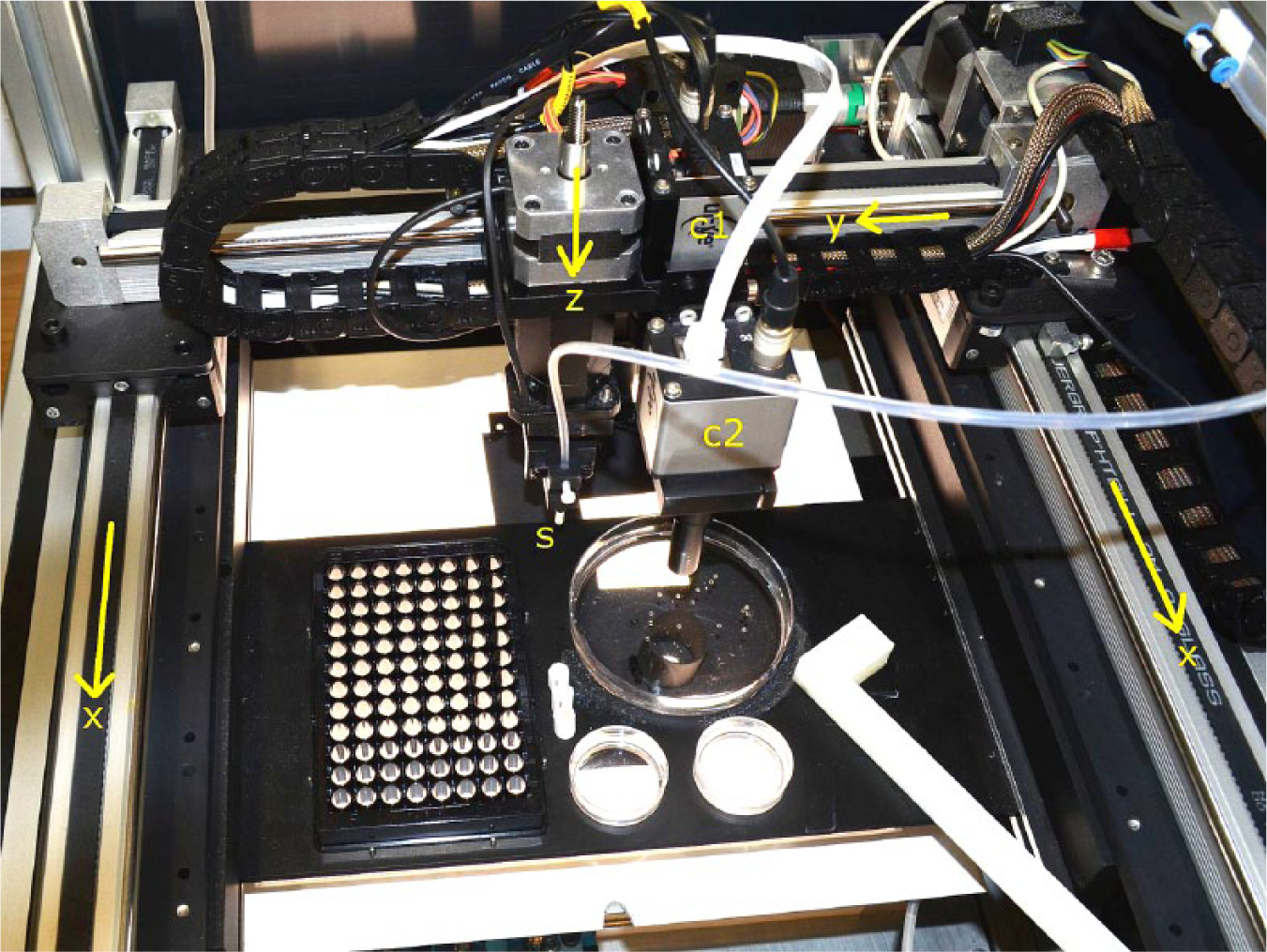

The sorter comprises three linear axes (x, y, and z axes) for moving the sucking tips and the two cameras, the overview camera (C1) and the camera for detailed inspection (C2), above the illuminating LED panel.
Image preprocessing is a basic prerequisite to classify whether an embryo is either a mutant with phenotypic deviations or a wild-type embryo or something different. Four preprocessing operators for filtering the images from both cameras were used within the IMAQ Vision for LabVIEW software (National Instruments, Austin, TX):
Threshold filter: Isolates the objects from the background.
Masking: Isolates parts of an image for further processing. This technique saves memory and processing time by limiting the image analysis to only the part of the image containing relevant information.
Particle area: Measures the areas of all the large circular particles of the image and is used to filter out all objects that are significantly smaller or bigger than the average zebrafish egg.
Particle circularity: Calculates the perimeter of an object divided by the circumference of a circle with the same area. The closer this circularity ratio is to 1, corresponding with the shape of a disk, the more likely the object is an unhatched zebrafish embryo. Dirt, hatched larvae, torn chorions, and all embryos that are not sufficiently round can be excluded ( Fig. 1D–F ).
All remaining objects after these preprocessing filtering steps are classified by applying template-matching algorithms. Template matching is a technique for finding parts of an image that match a template image by applying normalized cross-correlation or sum of absolute differences.17,18 Two-dimensional rotation of the embryos compared with the templates was also checked by the template-matching algorithm.
In a first experiment, we used templates of zebrafish without eyes in order to directly detect rx3 mutants. However, this attempt was not successful. By contrast, using templates of zebrafish with eyes and with less pigmented eyes to detect all phenotypic wild-type embryos gave better results, and they were applied subsequently.
Templates may be defined using a special editor of NIs Vision Assistant that allows for specifying the important features within a picture. In this setting, the prevalence of one or two eyes and parts of the head was crucial. For both detection phases, several templates were defined ( Fig. 3 ) and modified, and either the required minimum or least score was adjusted by testing against testing sets of rx3 mutant embryos and phenotypic wild-type embryos. By means of parallel processing of the template-matching algorithms, the speed of feature detection was doubled.
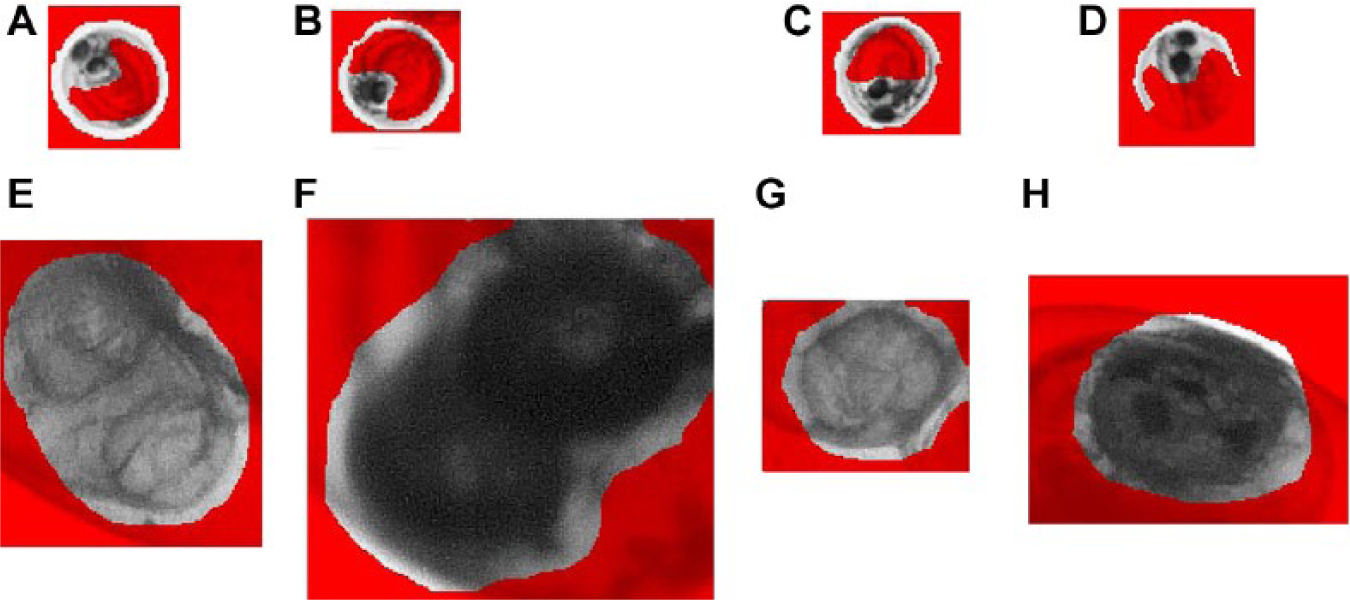
Examples of templates. Templates for the phase 1 classification (
Finally, the suction hose of the sorting robot is positioned accurately over the selected zebrafish embryo and transports the embryo with 30 µL of water to another petri dish or multiwell plate.
Results
Different sets of images were used for training and testing. For validating the classification capabilities of the new system, image data sets of zebrafish embryos 48 h postfertilization (hpf) with a premixed ratio of low and high pigmentation were used. The detection of phenotypic wild-type embryos in the first phase of image analysis (images from the overview camera, C1) differed significantly with the number of templates used. On average, 36% of all embryos present in the overview were identified as being phenotypic wild-type embryos. Thus, 64% of embryos remained for further inspection in the second phase, with eight templates applied for feature detection. When 14 templates were used, the detection rate of phenotypic wild-type embryos increased to 54% and 46% remained for further examination. An even higher number of templates did not sufficiently increase the number of embryos to be classified. Examples of templates used for classification are given in Figure 4 . The second phase of image analysis (images from the detailed-view camera, C2) was evaluated with a total of 3804 images of rx3 embryos and non-rx3 embryos (images of phenotypically wild-type embryos, empty chorions, and coagulated eggs). The images were derived from nine different batches and were labeled according to the two categories by the authors. The sensitivity (true-positive rate or percentage of rx3 embryos that are correctly identified as such) was 90% when 30 templates were used for classification and 89% for 117 templates ( Table 1 ). However, the specificity (true-negative rate or percentage of images that are correctly identified as non-rx3 embryos) was 87% and 95% when 30 and 117 templates were used for classification, respectively. In a subsequent experiment with another 948 images, the sensitivities and specificities were even higher when images showing debris, empty chorions, and coagulated eggs instead of healthy embryos were removed prior to classification. On average, classification of an embryo took less than 1 s on a computer with a core i7 CPU, and sorting into another petri dish took another 14 se. Examples of classified objects from the camera with a detailed view, C2, are given in Figure 5 .
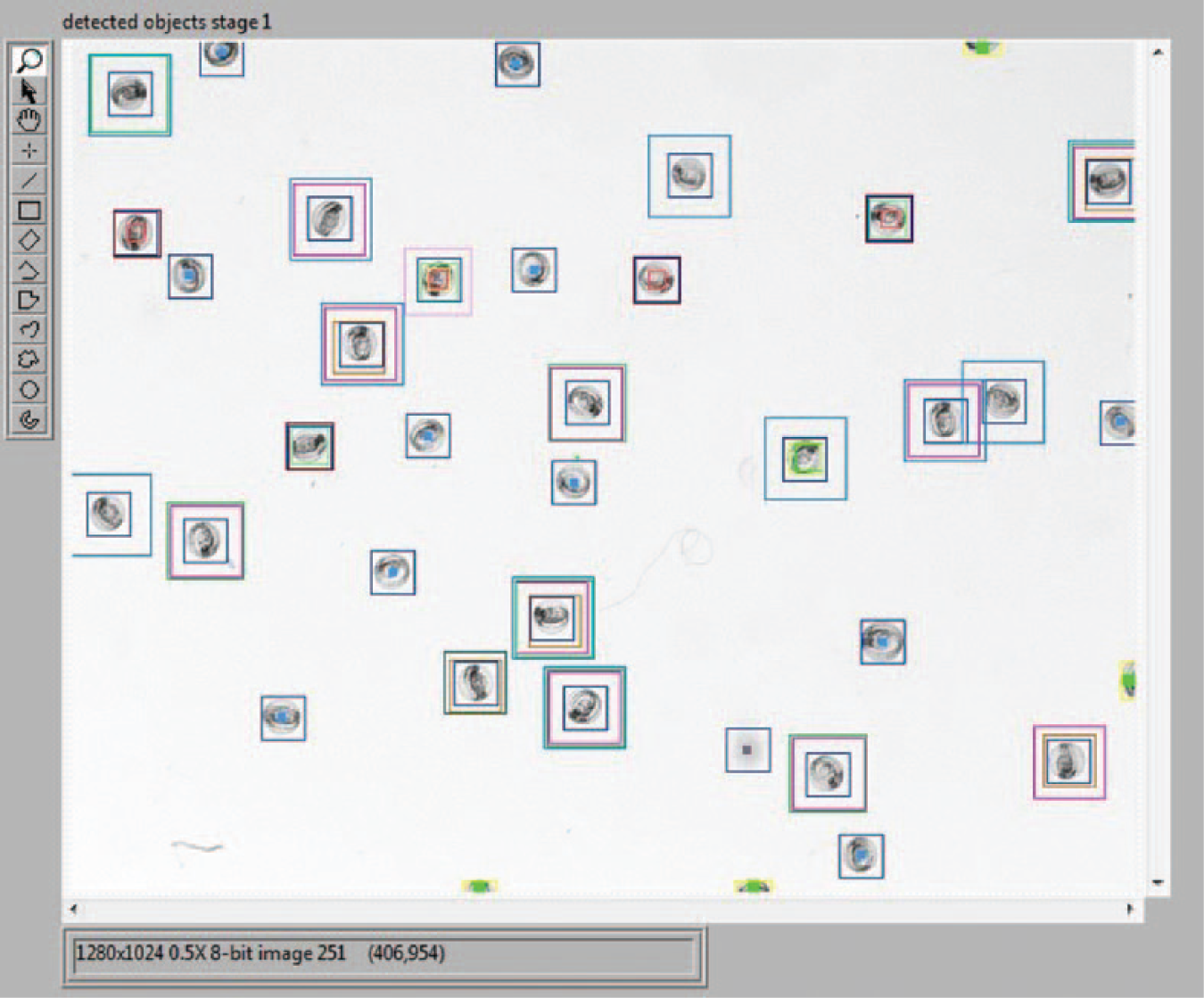
Example for classified results of phase 1. All embryos classified as rx3 are marked with a blue square in the center; all other objects are marked with multiple squares. A gray square indicates an empty chorion, and green or yellow squares indicate debris or coagulated eggs.
Classification Results of Two Data Sets of Images from the Detailed-View Camera (Phase 2).
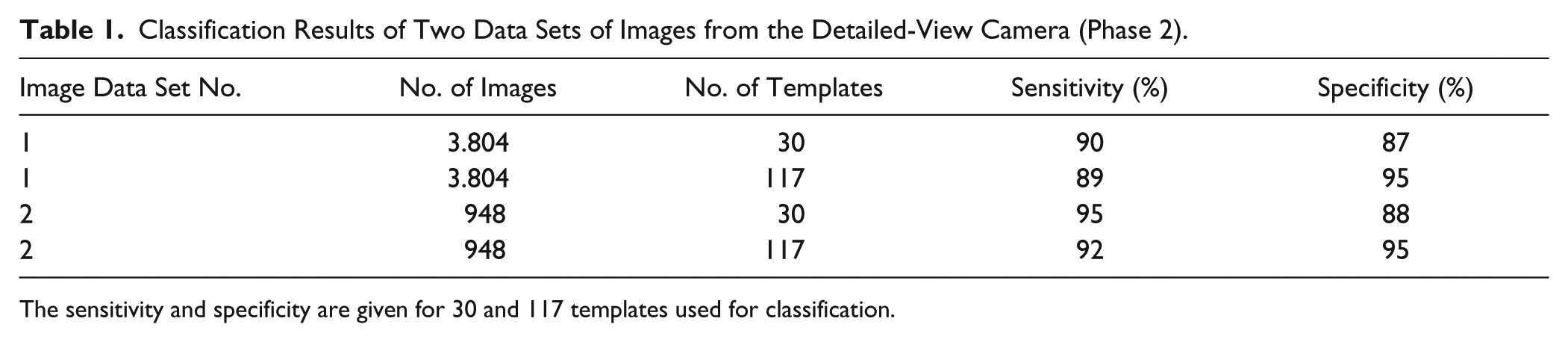
The sensitivity and specificity are given for 30 and 117 templates used for classification.
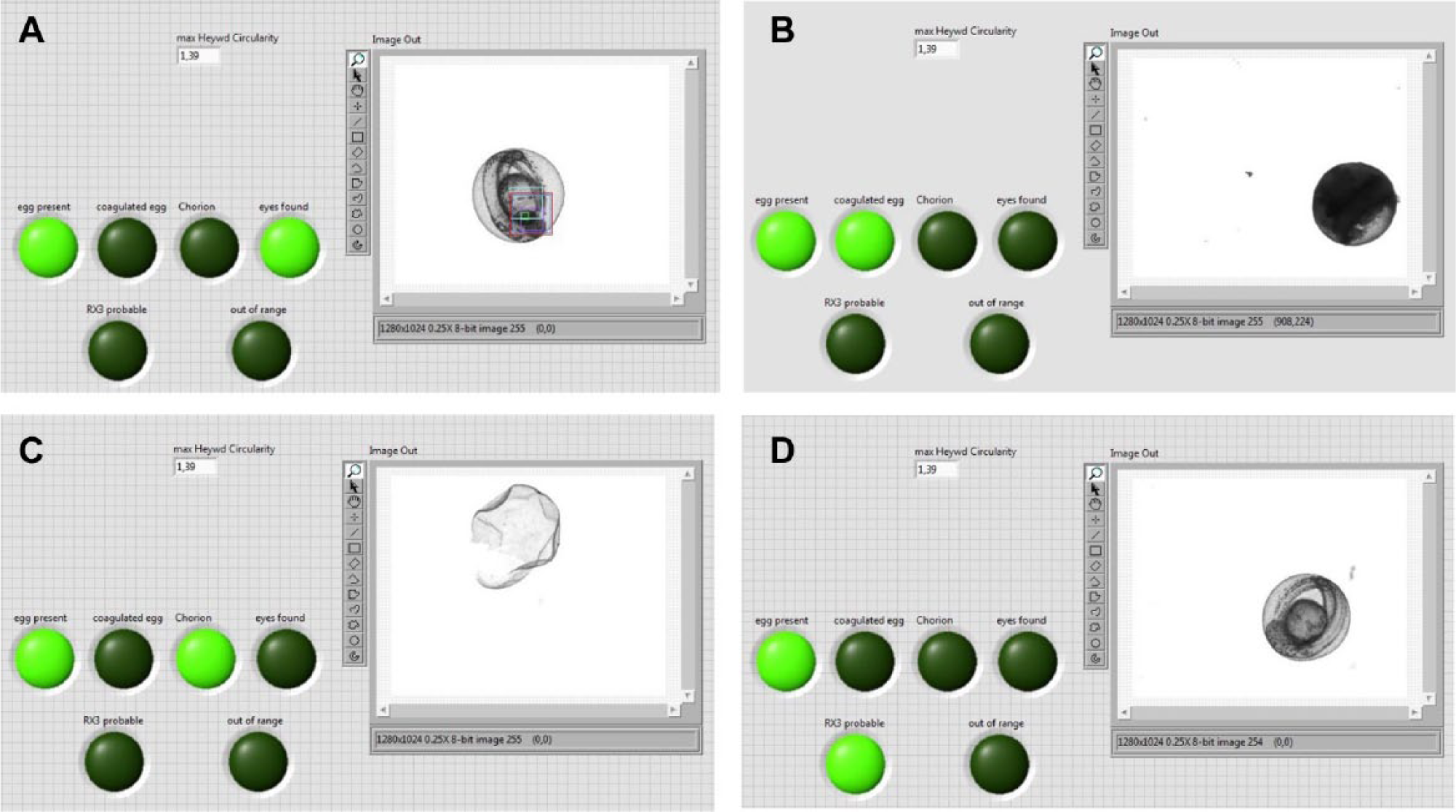

Examples for classified results of phase 2. (
Discussion
Automated sorting of zebrafish embryos with a specific phenotype requires approximately twice as long as manual sorting. However, this was no problem for all users who have tested the system and saved them time for doing other laboratory work in the meantime. Furthermore, manual sorting of larger numbers of embryos under the microscope is a tedious task and prone to errors. Additionally, the manual after-sorting of 10 incorrect candidates, if necessary, was rated as being acceptable for the researchers.
In order to accelerate the classification, the less effective but faster low-discrepancy sampling (LDS) algorithm was used instead of the grayscale value pyramid-matching (GSVP) algorithm offered by the NI IMAQ library.17,18 If the objective of sorting phenotypes is to achieve a higher specificity, the GSVP template-matching algorithm should be used instead. However, this will also increase the classification time by a factor of 6. Identification of rx3 mutants was also accelerated significantly by classifying one-third to half of the objects on images from the overview camera, C1, instead of inspecting every single object with the detailed-view camera, C2. Thus, classification in a two-stage procedure accelerates the sorting procedure significantly. By choosing a higher number of templates, the specificity of classification can be increased. Nevertheless, the number of templates had only a minor impact on the sorting time since the classification time was less than 1 s, compared with 14 s of picking and placing the embryo by the robot.
Repeated evaluation of the system showed that approximately 90% of all rx3 embryos in the reservoir petri dish were identified and sorted into another petri dish. The false-positive rate was caused mainly by embryos with less pigmented eyes that were misclassified as rx3 embryos. It has to be mentioned that sorting of wild-type embryos with less pigmented eyes ( Fig. 1C ) from rx3 embryos ( Fig. 1A,B ) that look very similar is a difficult task for a lab assistant too. The classification result was in the same range as that of a radial basis function support vector machine that was used before to discriminate hatched embryos, 15 but it has not been tested with unhatched embryos yet. Even higher classification rates might be achieved by methods of machine learning, but large numbers of image data are required to train the algorithm. 19 The system can easily be extended to sort phenotypes with other morphological features by using templates of these phenotypes. 20
Footnotes
Acknowledgements
The authors thank Nadine Borel and her team at the KIT EZRC for providing us with zebrafish embryos. We also thank Dr. Bernd Köhler for his technical support.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research received funding provided by the Helmholtz Association program “BioInterfaces: Molecular and Cellular Interactions at Functional Interfaces.”
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
