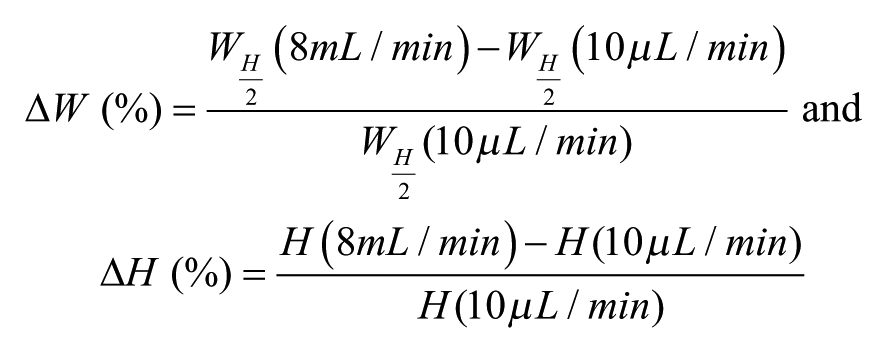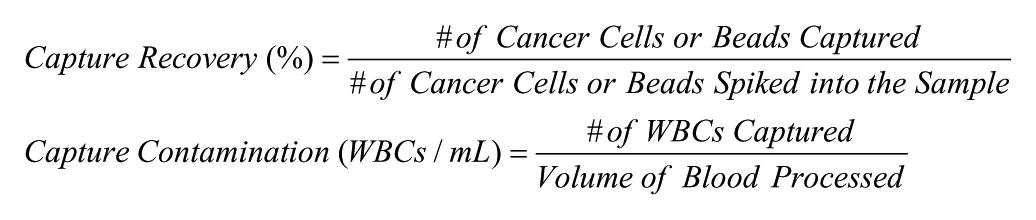Abstract
Tumor tissue biopsies are invasive, costly, and collect a limited cell population not completely reflective of patient cancer cell diversity. Circulating tumor cells (CTCs) can be isolated from a simple blood draw and may be representative of the diverse biology from multiple tumor sites. The VTX-1 Liquid Biopsy System was designed to automate the isolation of clinically relevant CTC populations, making the CTCs available for easy analysis. We present here the transition from a cutting-edge microfluidic innovation in the lab to a commercial, automated system for isolating CTCs directly from whole blood. As the technology evolved into a commercial system, flexible polydimethylsiloxane microfluidic chips were replaced by rigid poly(methyl methacrylate) chips for a 2.2-fold increase in cell recovery. Automating the fluidic processing with the VTX-1 further improved cancer cell recovery by nearly 1.4-fold, with a 2.8-fold decrease in contaminating white blood cells and overall improved reproducibility. Two isolation protocols were optimized that favor either the cancer cell recovery (up to 71.6% recovery) or sample purity (≤100 white blood cells/mL). The VTX-1’s performance was further tested with three different spiked breast or lung cancer cell lines, with 69.0% to 79.5% cell recovery. Finally, several cancer research applications are presented using the commercial VTX-1 system.
Introduction
Cancer is the second leading cause of death in the United States, with 1,688,780 new cases expected in 2017 1 (https://www.cdc.gov/nchs/fastats/deaths.htm). Cancer research is still needed to continuously address critical challenges, such as cancer screening for earlier diagnosis and better prognosis, or developing targeted therapies for more personalized and efficacious treatment. Procuring tissue or cell samples from the tumor site is a prerequisite to making a definitive diagnosis and gaining biological information on a patient’s cancer. Tissue biopsies, the current standard of care, are invasive and can be risky and costly procedures that can be limited by small specimen size and/or tumor accessibility. Moreover, single-site tumor biopsies may not reveal intra- or intertumor heterogeneity and may fail to reflect the genetic diversity of the disease. 2 These limitations can be overcome with a liquid biopsy. Liquid biopsies are noninvasive blood tests that aim to isolate and analyze circulating biomarkers, such as circulating tumor cells (CTCs), cell-free DNA (cfDNA), and exosomes, which are shed into the bloodstream by both primary and metastatic lesions.3–5 As a minimally invasive blood draw, liquid biopsies are compatible with serial sampling and could provide real-time and representative information about tumor evolution, treatment effectiveness, and cancer metastatic risks. Ultimately, liquid biopsies may allow for earlier cancer detection, relapse detection, and more personalized treatment of cancer. There are many studies aiming to examine whether CTCs or cfDNA will “win the war in the era of precision oncology.” 6 As discussed by Ignatiadis et al., 3 plasma cfDNA assays may prove more useful for monitoring disease burden and performing limited molecular profiling, but CTCs can provide a more comprehensive characterization of tumor DNA, RNA, and protein levels of residual malignant cells throughout disease course, and they may be useful for functional studies and drug sensitivity evaluations as well. Current evidence seems to indicate both CTCs and cfDNA provide important insights into a patient’s cancer biology, and each will likely play a role in precision cancer care. For genomic profiling specifically, studies have identified mutations present in CTCs but not detectable in cfDNA and vice versa, emphasizing the complementarity of both biopsies to better represent the heterogeneity of the primary tumor and metastases.3,7,8
CTCs are extremely rare cancer cells shed from primary tumor or metastases into the peripheral blood of patients with solid tumors. CTC counts have been clinically correlated with the prognosis of patients with breast, colon, and prostate cancers, and their molecular characterization could provide additional insights into their functional roles in cancer progression, thereby helping to guide treatment toward personalized therapies.3,9,10 Isolating rare CTCs among a background of millions of white blood cells (WBCs) and billions of red blood cells (RBCs) while preserving their utility for downstream characterization remains a technical challenge limiting their broader use in research and clinical laboratories.
The CellSearch system, recently acquired by Menarini-Silicon Biosystems, Inc., was the first instrument available for CTC isolation and remains the only CTC capture platform approved by the Food and Drug Administration (FDA) for prognosis. It relies on a positive selection using epithelial markers such as epithelial cell adhesion molecule (EpCAM) to immunomagnetically capture CTCs.10,11 This approach requires that CTCs express the markers of interest to be captured and consequently would miss the cells lacking these markers. 12 Beyond this dependence on expressed markers, CTC isolation using CellSearch is a multistep process originally designed for enrichment and enumeration. More recent technologies such as the CTC-iChip, 12 the MagSweeper,13,14 Fluxion, 15 Adnagen, 16 RareCyte, 17 and EPIC, 18 among others, 19 allow the isolation and analysis of CTCs beyond their enumeration, but some still require preexisting knowledge of the molecular biomarkers expressed or require an upstream sample preparation that may lead to cell loss and/or impact on CTC integrity for downstream assays. To address these limitations and the need for preexisting knowledge of cell biomarkers, alternative label-free CTC enrichment technologies have been developed to provide researchers with a broader range of CTCs, often using their larger size compared with RBCs and WBCs, for a size-based CTC enrichment strategy. 20 For example, the ClearCell FX technology uses Dean flow and hydrodynamic focusing within a curved channel for differential collection of CTC and WBC streams. 21 Limitations of this method, however, include a requirement for lysing the RBCs prior to isolation and a final collection of the isolated CTCs into a large volume, which adds an additional centrifugation and transfer step prior to analysis. Microfilter technologies, such as ScreenCell, 22 ISET, 23 CellSieve, 24 or Parsortix, 25 involve flowing blood through pores or microfluidic steps of calibrated size to trap larger cells (i.e., CTCs) while smaller cells pass through. Such technologies or approaches have the advantages of being less complicated, sometimes rapid, and needing minimal equipment. However, some of these approaches may be prone to clogging and can make releasing the CTCs into suspension for further analysis after their physical trapping challenging, often resulting in higher numbers of contaminating WBCs.
Thus, there is still an unmet need for a fast, simple, and label-free isolation of CTCs followed by their release in suspension, in a manner compatible with various downstream assays. The Vortex technology was developed to meet these needs. The Vortex technology relies on inertial microfluidics and laminar microscale vortices to isolate and concentrate CTCs from blood, based on cell size and deformability ( Fig. 1A ). The Vortex chip and the High-Throughput Vortex chip (Vortex HT), both made of polydimethylsiloxane (PDMS) and operating with a manual platform using two syringe pumps, were previously introduced, characterized, and validated for CTC isolation from blood samples of metastatic cancer patients with breast, colon, lung, or prostate metastatic cancer.8,26–29
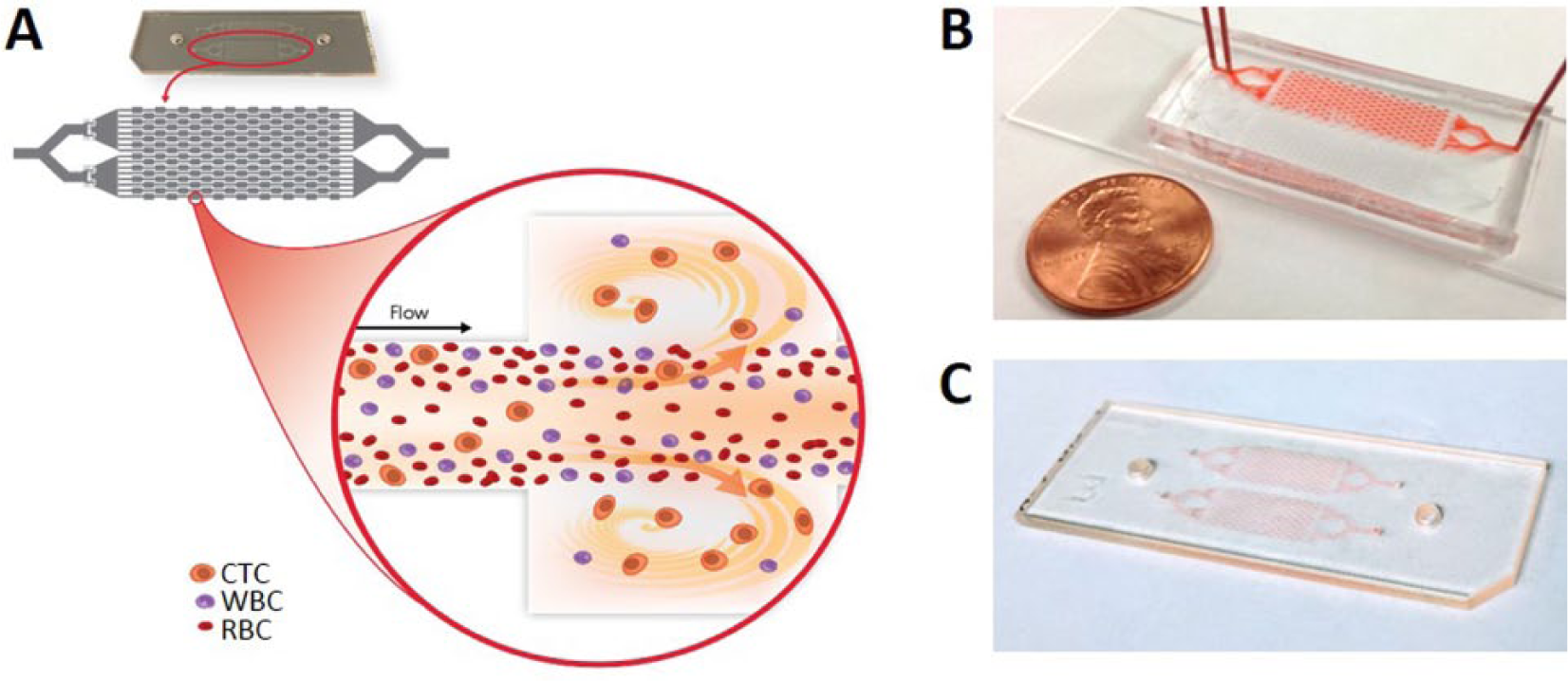
Vortex technology and microfluidic chip. (
Here, we present the commercialization efforts and challenges of transferring this technology from a research platform to a commercial, reliable, and fully-automated fluidic instrument. First, we discuss the issues of PDMS as a material for the microfluidic chip, its deformability, and the inherent impact on capture performance compared with a rigid plastic, poly(methyl methacrylate) (PMMA; Fig. 1B, C ). Second, we compare side by side the manual research platform using two syringe pumps to the automated fluidic processing using the VTX-1 Liquid Biopsy System. In a third section, the VTX-1 system and workflow are further characterized and validated for various cancer research applications, including processing of different cancer cell lines, compatibility with live cell downstream assays, and the isolation of CTCs from patient-derived xenograft blood samples from mice.
Methods
Vortex Microfluidic Chip Design, Fabrication, and Operation
As previously introduced, 27 Vortex HT is a 70 µm deep PDMS microfluidic chip comprising a parallelized array of 16 straight 40 µm wide channels, leading to a series of 12 rectangular trapping reservoirs (480 × 720 µm). Conventional PDMS fabrication processes were used to create Vortex HT PDMS chips ( Fig. 1B ) with a 1:10 PDMS mix.27,30 Plastic chips were fabricated using a standard lithography process and PMMA as the material ( Fig. 1C ). A second generation of plastic chip was composed of 16 channels in parallel, with a series of nine reservoirs. To operate the PDMS chip, flow was driven through two inlets using two syringe pumps (Harvard Apparatus), one for the sample (beads or cells) and one for the wash buffer (phosphate-buffered saline [PBS]).26,27 After a priming step to fill in the chip with PBS at 8 mL/min for 30 s, a bead suspension (19 µm polystyrene particles, Thermo Scientific, coefficient of variation [CV] ≤ 16%, 10% w/w) or a cancer cell sample was infused at 7 mL/min together with wash buffer at 1 mL/min to achieve bead/cell capture in vortices created in the chip reservoirs. The number of beads/cells varied depending on the experiment. To wash contaminating cells or debris from the beads/cells trapped in the vortices, the buffer flow rate was increased to 8 mL/min, while infusion of the sample was stopped. Captured beads/cells were then released from the vortices by stopping the flow of the wash solution. To operate the plastic chip manually, a similar method was used, with a plastic manifold and connections (Upchurch) to connect the tubing (Tefzel [ETFE] Tubing Natural 1/16″ OD × 0.040″ ID from IDEX) to the chip. For automated processing, the same plastic chips were integrated into a disposable plastic cartridge (Vortex Biosciences) and then inserted into the VTX-1 instrument (Vortex Biosciences) for fluidic processing.
Chip Deformation
The PDMS chip wall thickness is about 5 mm. The glass side sealed to the PDMS channel being rigid (E ~63 GPa for Pyrex versus 2.5 MPa for PDMS 1:10), this imposes a zero displacement between the glass and PDMS. The deformation pattern can be approximated, as illustrated in Supplementary Figure S1 , with mainly a vertical deformation of the roof and a constrained lateral deformation.31–33 To visualize microfluidic channel deformation, 33 fluorescein was injected at different flow rates (10 µL/min and 8 mL/min). Fluorescent images were recorded at the same locations for both flow rates, using a Photometrics CoolSnap HQ2 camera, a Zeiss Axio Observer microscope, and Zeiss 2012 Blue edition software. The intensity profile was plotted for both flow rates at a constant exposure time for cross sections normal to the flow direction. The intensity profile obtained at 10 µL/min is considered as a baseline with no deformation. The deformation at 8 mL/min (in percentage) represents the relative changes in depth (ΔH) and width (ΔW) between 8 mL/min and 10 µL/min ( Suppl. Fig. S1 ), that is, the normalized expansions in both axis y and z at 8 mL/min using 10 µL/min as the baseline, as defined in the following equations:
The width is measured at the middle height. In such cases, for example, a “20% deformation” means a 20% expansion observed at 8 mL/min relative to the dimension measured at 10 µL/min.
Chip Capacity
To determine the capacity of a microfluidic chip to capture particles, different amounts of 19 µm beads (500, 1000, and 3000) were mixed with water and 0.01% Triton-X and injected into the chip according to the protocol described above. During the wash step, the chips were imaged (Phantom Miro EX2 high-speed camera from Vision Research; Phantom Camera Control software) to determine the number of beads trapped in each reservoir.
Cancer Cell Lines Culture and Harvesting
All cell lines used for characterization were grown to 30% to 60% confluence at 37 °C in a humid atmosphere of 5% CO2. Cell lines were evaluated for mycoplasma contamination by PCR (Venor GeM Mycoplasma Detection Kit) and authenticated by IDEXX BioResearch. MCF7 (breast carcinoma, ATCC) cells were cultured in RPMI medium supplemented with 10% fetal bovine serum (FBS) previously inactivated at 56 °C for 30 min, 1% penicillin/streptomycin (Invitrogen), and 0.01 mg/mL human recombinant insulin (Life Technologies). MDA-MB-231 (breast carcinoma, ATCC) cells were grown in DMEM medium supplemented with 10% inactivated FBS and 1% penicillin/streptomycin. HCC827 (lung carcinoma, ATCC) cells and H1975 (lung carcinoma, ATCC) cells were grown in RPMI medium supplemented with 10% inactivated FBS and 1% penicillin/streptomycin. Adherent cells were dissociated with TrypLE express cell dissociation reagent (Gibco) and resuspended in complete media. Their viability and average diameter were measured using acridine orange, propidium iodide solution on a Cellometer K2 (Nexcelom). Cells (from 50 to 500 cells depending on the experiment) were then spiked into 0.5 to 4.0 mL healthy human blood diluted 10× in PBS pH 7.2, prior to injection through the chip.
Donor Recruitment and Blood Collection
Blood donors were recruited and informed consent obtained according to protocols approved by Institutional Review Boards (Stanford IRB protocol 5630) from the Stanford University School of Medicine or from the Stanford Blood Bank. Blood samples were drawn from healthy volunteers with no known illness or fever at the time of draw. Peripheral blood was collected into 10 mL EDTA-coated tubes (BD Vacutainer), transported, and processed at room temperature within 12 h of the blood collection.
Cell Immunostaining and Enumeration
After Vortex processing, cells were collected in a cell culture-treated 96-well plate or a PolySorp-coated eight-well strip (Nunc), fixed with 4% paraformaldehyde (Electron Microscopy Sciences) for 10 min, permeabilized with 0.4% v/v Triton X-100 (Sigma Aldrich) for 7 min, blocked with 10% goat serum (Invitrogen) for 30 min, and labeled for 1 h at room temperature (RT) with 4,6-diamidino-2-phenylindole (DAPI; Life Technologies), anti-CD45–phycoerythrin antibody (CD45-PE, clone HI30, BD Biosciences), and a cocktail of primary antibodies labeled with fluorescein isothiocyanate to identify cytokeratin (CK)–positive cells (clone CK3-6H5, Miltenyi Biotec, and clone CAM5.2, BD Biosciences). 28 For experiments with healthy mouse blood, a similar staining protocol was optimized but with a rat anti-mouse CD45-PE antibody (CD45-PE, clone 30-F11, BD Pharmingen). For patient-derived orthotopic xenograft (PDOX) samples, an anti-CK-AlexaFluor (AF) 488 antibody (clone AE1/AE3, eBioscience) and an anti-vimentin-AF647 antibody (clone V9, Abcam) were used in addition to the previously listed antibodies. After staining, the cells were imaged (Axio Observer Z1, Zeiss) and manually enumerated using specific classification criteria previously published. 27 Capture performance was evaluated as follows:
Clonogenic Assay
MDA-MB-231 cells, 50% confluent were harvested as described previously. One thousand cells were spiked in 40 mL of complete filtered medium (cDMEM), processed through the Vortex plastic chip in the VTX-1 instrument, and collected in a 35 mm diameter tissue-culture treated Petri dish containing 3 mL of DMEM. As a control, 350 and 500 MDA-MB-231 unprocessed cells were seeded in parallel. The Petri dishes were then imaged in bright field using a 5× objective, and cancer cells were counted. Dishes were incubated at 37 °C until control dish colonies consisted of greater than 50 cells or up to 10 d with partial media replacement every 2 d. Once colonies developed, cells were fixed and stained with crystal violet (5 mg/mL in glutaraldehyde) for 15 min at RT. Colonies consisting of more than 50 cells were counted, and plating efficiency (PE; in percentage) was determined as the number of colonies divided by the number of cells seeded. Each condition was tested in quadruplicate, and the experiment was repeated once (n = 2). Statistical differences comparing the PE of processed and control dishes were determined using a one-way analysis of variance (ANOVA; α = 0.05).
Invasion Assay
The invasion assay was performed on MDA-MB-231 cells using 96-well 3D Spheroid Cultrex BME Cell Invasion Assay (Trevigen). Briefly, MDA-MB-231 cells were cultured up to 70% confluence, harvested, and 200,000 cells were spiked into 300 µL of freshly isolated healthy mouse blood (collected via cardiac puncture). The spiked blood was diluted 20× with filtered PBS. Cells isolated via Vortex processing (using manual processing and Vortex plastic chip) were then used in a 3D Spheroid invasion assay as per the manufacturer recommendations. As a control, cells not processed by Vortex were seeded in separate wells. All samples were seeded in duplicates. Once the spheroids developed, they were placed within the invasion matrix and grown at 37 °C for up to 11 d. Images of individual spheroids in each well were taken every 24 h using a 5× objective.
Characterization/Validation of VTX-1 for Mouse Blood Samples
All animal studies were performed according to protocols reviewed and approved by the Stanford University Administrative Panels on Laboratory Animal Care (APLAC protocol 12809). MDA-MB-231 cells were cultured up to 50% confluence and harvested as described previously, for spiking of 200 cells into 600 µL of blood isolated from BALB/c mice via cardiac puncture. The spiked blood was diluted 40× in filtered PBS and processed using the Vortex plastic chip in the VTX-1 instrument (two cycles). The captured cells were released into an eight-well strip and stained for enumeration. Once validated, this workflow was applied to PDOX samples. Xenografts previously generated from a stage II patient with triple negative (ER–/PR–/HER2–) breast cancer were implanted into the fourth mammary fat pad of immunodeficient NOD scid gamma mice (NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ, Jackson Laboratory). PDOX tumors were grown for up to 3 mo, and tumor volumes were measured twice weekly. When the tumor volume reached between 1500 and 2000 mm3, the mice were euthanized, blood (~750 µL) was collected via cardiac puncture from each PDOX-bearing mouse and diluted 40× with filtered PBS. The diluted samples were processed on VTX-1, and isolated CTCs were immunostained and enumerated as previously described.
Results
Microfluidic Chip: From Deformable PDMS to Rigid Plastic
Deformation of PDMS Microfluidic Channels
The deformability of PDMS offers several advantages for microfluidic fabrication in research settings, for example, the ability to easily demold channels or to design pressure-actuated valves.30,32–34 However, the level of deformation, especially at high flow rates, can have a significant impact on the microfluidic phenomena observed. Figure 2A shows a representation of a chip, with its different features: narrow channel segments (i.e., “channel segments”) and expansion regions (i.e., “reservoirs”). Figure 2C represents the horizontal and vertical deformation of the Vortex microfluidic channels at different locations along the chip length, considering both the channel segments and the reservoirs, assuming a deformation pattern as described in Supplementary Figure S1 . A significant depth (z axis) deformation was observed for the first reservoirs, with 66% deformation for the first reservoir, 60% for the second, and still 20% deformation for the last reservoir. A lesser deformation (28%) was obtained for the narrow channel segments. Such results were expected, as the channel bottom is constrained in all directions by the presence of the glass slide, reducing the side walls’ lateral expansion, whereas the channel roof can still move vertically. Many different factors are affecting the PDMS deformation under continuous flow, including but not limited to the elastic modulus of the materials involved, the channel aspect ratio, the flow rate and pressure, as well as the feature position along the channel length as this affects in return the fluid velocity. This matter was thoroughly assessed and published by some of the authors,26,33 as well as others.32,35 Further details are available in Supplementary Figure S1 . Similar experiments were performed with rigid plastic chips and did not show any measurable deformation of the features (data not shown).
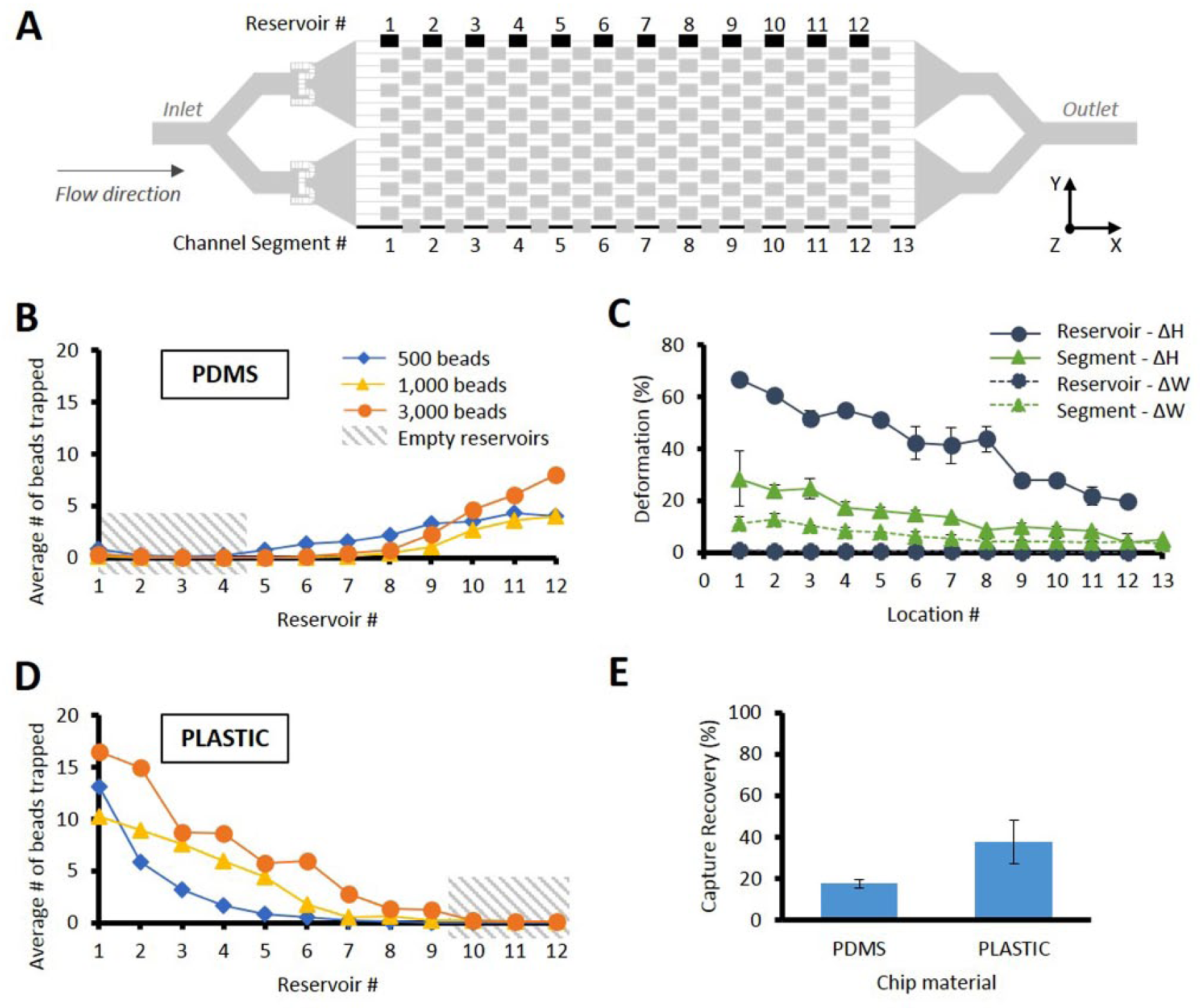
Deformation of chip microfluidic features and its effect on particle capture distribution and capture performance. (
Improving Bead Recovery by Transitioning to Less Deformable Chips
PDMS deformability had several consequences for the chip performance. Figures 2B and D show the bead distribution in the different serial reservoirs, with a side-by-side comparison between PDMS and rigid plastic microfluidic chips, for different particle numbers, at a constant flow rate of 8 mL/min. For 500 particles, in PDMS chips, particles were mainly captured toward the outlet of the chip, with an average of 72.8% of captured beads being captured in the last three reservoirs. For rigid plastic chips, the capture pattern is opposite, with the captured particles being captured near the inlets (65.7% in the first three reservoirs). For a similar flow rate, the deformation difference between the two materials has an impact on channel cross section and velocity profile along the channel length, which is expected to affect the inertial focusing positions of the different cell streamlines, as well as the Vortex flow patterns differently in each of the reservoirs. 33 The first PDMS reservoirs being more deformed, fewer beads are captured but get captured toward the end of the chip, where reservoirs are less deformed. For PMMA, the reservoirs are nondeformed, equally all along the channel length, which means the beads are mainly captured toward the inlet and further down the channel length as more cells are entering the vortices.
Figures 2B and D illustrate how the channel deformation also affects the overall capture capacity of the microfluidic chip, at the corresponding optimal flow rate ( Suppl. Fig. S2 ). As more beads were injected (500, 1000, and 3000), PDMS chip reservoirs capture recovery reached a plateau of 1000 beads ( Suppl. Fig. S3 ). Even with a higher bead concentration, particles were still only captured by the reservoirs located toward the outlet of the chips (84.5% and 81.9% of total captured beads being captured in the three reservoirs closest to the outlet, for 1000 and 3000 beads spiked, respectively). For rigid microfluidic chips, the opposite was true. The particles were first captured by the reservoirs near the inlet, and as more beads were injected, they were then captured along the channel length. More than 99% of the captured beads were captured in the first nine reservoirs. Consequently, the plastic chip was redesigned to include only nine reservoirs per channel instead of 12. This also reduced the overall pressure drop inside the chip. In addition, the difference in material deformability also affected the total number of beads captured. On average, for 3000 beads injected, the plastic chip had a capacity to capture 2.9-fold more beads in its reservoirs than its PDMS counterpart.
Improving Cell Capture Performance with Less Deformable Chips
Similarly, the cell recovery of the plastic chips was higher than the PDMS chips, as shown in Figure 2E . When 500 MCF7 cells were spiked in 0.5 mL of healthy human blood and processed through both chips side by side with the manual setup, only 17.5% were captured with the PDMS chips, against 37.7% with the plastic chips. This represents a 2.2-fold difference in cancer cell recovery.
Fluidic Processing: From Manual Syringe Pumps to the VTX-1 Liquid Biopsy System
Introducing an Automated System for Improved Sample Processing
Figure 3A illustrates the manual prototype for isolating cancer cells using the rigid plastic Vortex chips. The manual prototype relies on syringe pumps to push fluid through the chip at the appropriate flow rates and requires significant manual operations. Figure 3B outlines the fully automated VTX-1 Liquid Biopsy System. In this system, the Vortex chip is fully integrated into a cartridge ( Fig. 3C ). The user simply connects the patient blood tube to the Vortex cartridge, which is then inserted into the VTX-1 system. The VTX-1 automates the blood dilution and sample processing, collecting the isolated CTCs into a container (a microwell strip is shown in Fig. 3B ) for analysis. The isolated CTCs can be automatically collected into an Eppendorf tube, slide chamber, Petri dish, or microwell strip depending on the downstream analysis approach.
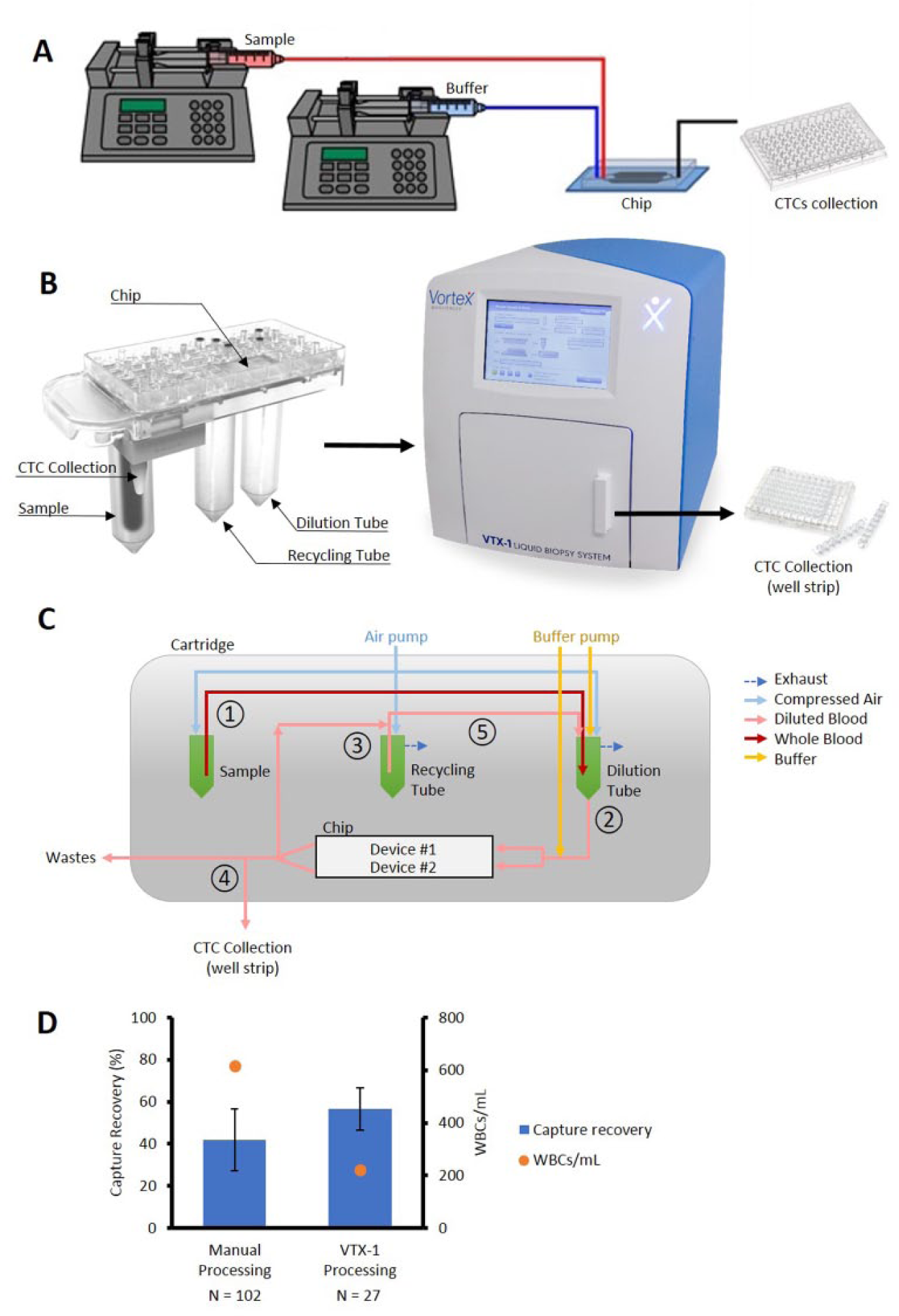
Transition from a manual setup to an automated instrument (VTX-1) and validation of this system. (
Improving Capture Performance with the Automated VTX-1 Liquid Biopsy System
Figure 3D shows the performance of both the manual and automatic setups when isolating 500 MCF7 cells spiked in healthy human blood, both using the plastic chip. The VTX-1 system captured 56.63% of the cancer cells, compared with 41.83% with the manual setup, and 2.8 times fewer contaminating WBCs were captured with the VTX-1, with 222 and 617 WBCs/mL with the VTX-1 and the manual setup, respectively. Finally, the performance obtained with the VTX-1 was more consistent: the CV was half that obtained with the manual setup (17.8% and 35.1% for the VTX-1 and the manual setup, respectively). In conclusion, the automated setup is not only easier and safer to use but also captures more cancer cells and fewer WBCs in a more reproducible way. These results indicate the superior performance obtained with the automated processing.
Optimizing VTX-1 Protocols for Cancer Cell Purity and Cancer Cell Recovery
The flexibility of the VTX-1 allowed us to target either optimal CTC recovery or optimal CTC purity depending on the intended downstream analysis. The system offers the possibility to reprocess a sample: after being processed once (one cycle), the effluent is collected on the Vortex cartridge and reinjected into the microfluidic chip for a second time (second cycle), to capture additional cancer cells missed in the first pass ( Fig. 4A ). System control protocols were optimized to use this capability to improve cancer cell recovery and minimize WBC contamination. About 50 MCF7 cells spiked in 4 mL blood were processed with up to three cycles. When processed with the high purity mode (one cycle), 53.8% of MCF7 cells were collected on average. When the same sample was processed with a high recovery mode (three cycles), 71.6% of the cells were recovered. Recycling a sample up to two times led to capturing 33.2% more MCF7 cells. In the high purity mode, only 101 contaminating WBCs/mL were collected, whereas the high recovery mode led to 350 contaminating WBCs/mL. By adjusting the number of cycles, it is possible to favor either an optimal cancer cell recovery (“high recovery mode”) or a better purity (“high purity mode”). The repeatability of the capture efficiency was also satisfactory for both protocols, with a standard deviation of 6.5% and 8.6% for the high purity mode and the high recovery mode, respectively.
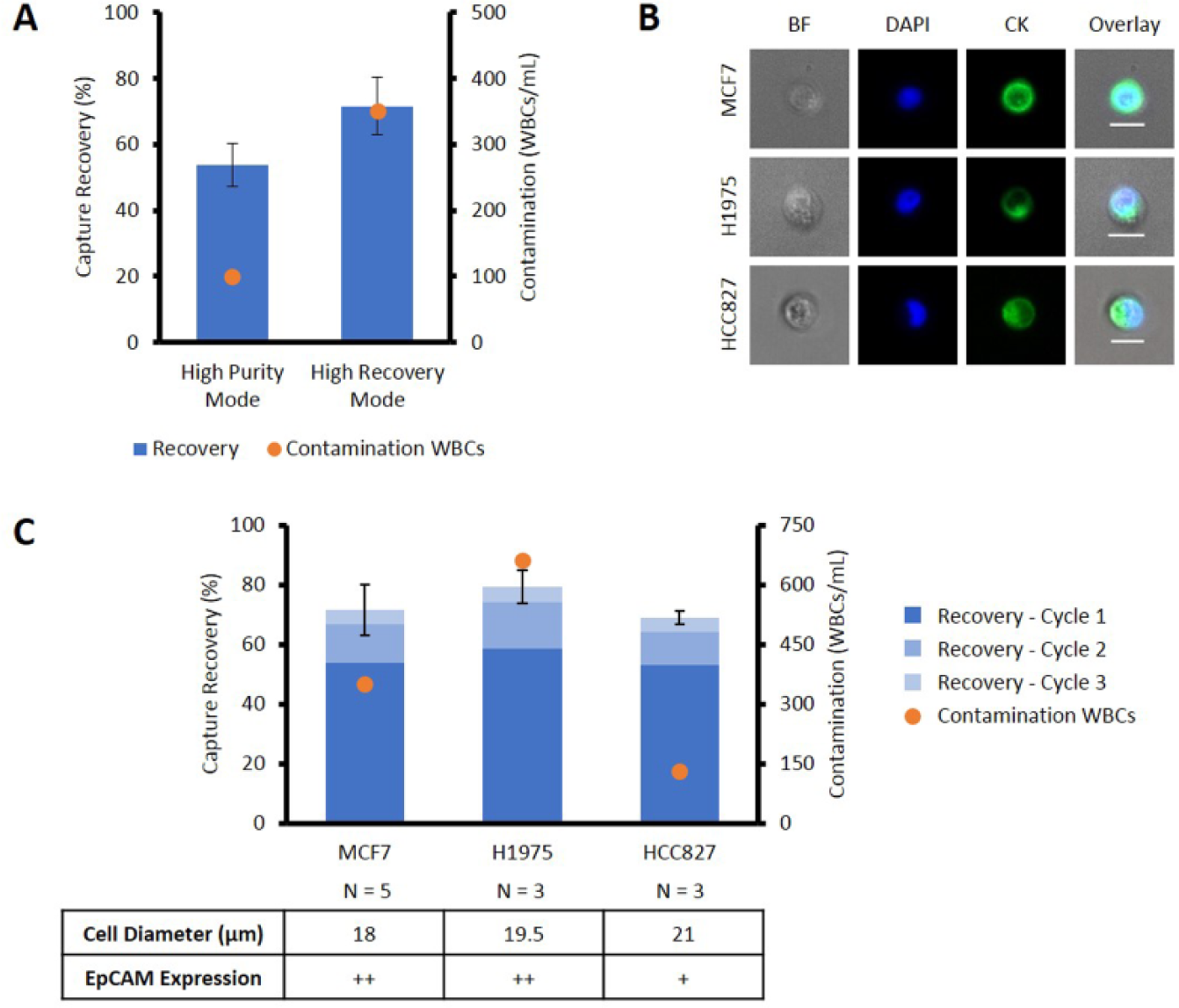
Validation of VTX-1 protocols and application to different cancer cell lines. (
VTX-1 Processing for Cell Lines with Various Epithelial Marker Expressions
The VTX-1 high recovery mode was tested on three different cells lines, with various sizes, CK expression, and EpCAM expression ( Fig. 4B, C ). High recovery rates were obtained for all of them: 74.0%, 79.5%, and 69.0% cell recovery for MCF7 cells (high EpCAM expression36,37), H1975 cells (high EpCAM expression 37 ), and HCC827 cells (medium EpCAM expression 37 ), respectively. The cell recovery profile was similar across the cell lines tested, with an average of 75.4% of the collected cancer cells captured during the first cycle. The WBC contamination over three cycles varied from 131 to 662 WBCs/mL, as the cell lines were tested with different blood donors. This shows the ability of the VTX-1 to isolate cancer cells from different cancer types and independent of their EpCAM expression levels.
Validation of VTX-1 Liquid Biopsy System for Research Assays
Invasion and Clonogenic Assays
In a previous study, we showed that cells processed through Vortex PDMS chips remained viable and could be cultured for over 7 d. 28 Using a colony formation assay, we tested whether processing through VTX-1 affects the ability of a tumor cell to form a colony. Our data showed no statistically significant differences between mean PEs of MDA-MB-231 isolated with the VTX-1 (“processed,” 26.4% ± 9.0%) and the control cells (PE: 22.4% ± 6.0% and 24.3% ± 4.0% for the 350 and 500 control cells, respectively; Fig. 5A ), demonstrating that processing on VTX-1 does not affect clonal growth potential of cells (one-way ANOVA, F(2, 21) = 0.735, p = 0.491). In a separate invasion assay, we showed that MDA-MB-231 cells maintained not only their capacity to form 3D spheroids but also their ability to invade into the extracellular matrix ( Fig. 5B ) after isolation through the Vortex microfluidic platform. Our results clearly demonstrate that Vortex processing does not affect cell viability, impair colony and tumor spheroid formation, or alter cell invasiveness.
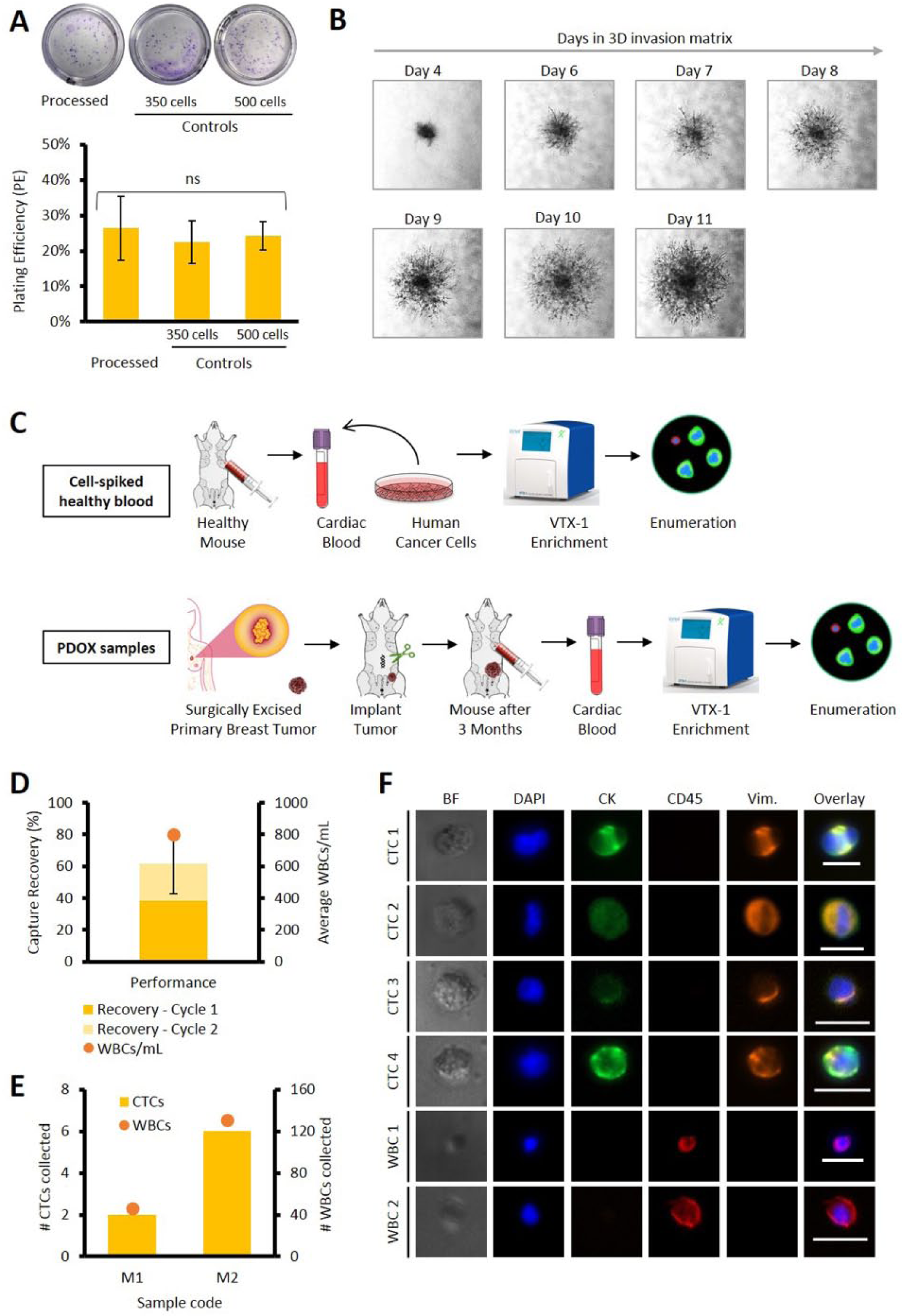
Performance of the VTX-1 instrument for various downstream applications. (
Characterization of VTX-1 for Mice Blood Samples
Blood from mouse, often less than 0.5 to 1 mL in volume, poses a big challenge for processing. This issue was overcome by diluting it 40× before process initiation. As illustrated in Figure 5C , cancer cell isolation from mice blood was first evaluated with MDA-MB-231 breast cancer cells spiked in 600 µL of healthy mouse blood, mimicking processing of mice model blood samples. The overall capture efficiency obtained with two cycles was equal to 61.30% ± 18.49% (n = 4), demonstrating the utility of the VTX-1 platform for isolating tumor cells even from small volumes of murine blood samples ( Fig. 5D ).
Validation of VTX-1 for Isolation of CTCs from Mouse PDOX Samples
Cancer cell isolation was then tested on mice implanted with the patient-derived breast tumor ( Fig. 5E, F ). Tumor volumes at the time of blood draw were 1280.97 and 1749.25 mm3 for mice 1 and 2, respectively. After cell enrichment (two cycles) and enumeration, two and six CTCs (2.67 and 8.00 CTCs/mL) were detected in the first and the second mice, respectively, with 46 and 131 WBCs (61.33 and 174.67 WBCs/mL; Fig. 5E ). The CTCs isolated stained positive for CK and vimentin, possibly indicating epithelial to mesenchymal transition ( Fig. 5F ). Our results demonstrate the ability of the VTX-1 to successfully isolate CTCs from mice bearing orthotopically implanted patient-derived xenograft of a triple negative breast tumor.
Discussion
Innovative technologies are the backbone of cancer research. New products open new approaches that can empower cancer researchers and clinicians to advance our understanding of cancer biology and improve diagnosis and treatment. Rapid prototyping can accelerate innovation at the lab scale by allowing for quick design and testing of different geometries. Multiple methods of microfabrication exist, depending on the need for rapid prototyping or industrial mass production of microfluidic devices. 33 Because of its low cost, fast turnaround, and simple fabrication, PDMS and soft embossing have been the material and technique of choice for rapid prototyping in the microfluidic research community. The low tensile modulus of PDMS offers several advantages for microfluidic applications, such as demolding microscale features without damage and ability to design pressure-actuated valves.32–34 However, among other limitations, PDMS deformability can also cause a significant deformation of fluidic features even under low pressure, as highlighted here with up to 66% vertical deformation in the first chip reservoir. This level of deformation had a negative impact on the overall capture performance of the Vortex technology. PDMS proved to be excellent for early innovation but limited our ability to transition to a robust, well-performing product. Polymers such as cyclic olefin copolymer and PMMA offer an attractive and rigid alternative to PDMS. However, introducing a new material with different properties requires not only optimization but often a redesign of the final product.
Innovative technologies are the critical first step, but the hard work of developing and optimizing a technology into a robust, well-performing product is what ultimately pushes cancer research forward. The transition from a manual research prototype to the commercial VTX-1 Liquid Biopsy System provided multiple advantages. The workflow is fully automated, with the blood collection tube being directly connected with the Vortex cartridge, then placed into the instrument (FDA class 1 medical device), where the CTCs are collected into an attached container. Experiments can be set up in 5 min with no additional sample preparation needed, and processing is completed in 1 to 2 h depending on the mode of operation. In addition to providing safer processing for the user, this provided a 1.35-fold increase in recovery and 1.97-fold increase in reproducibility, hence enhancing the performance and reliability of the instrument. The VTX-1 also enables the automated recycling of the effluent through the microfluidic chip and the cartridge. Indeed, a previous study highlighted the benefits of reprocessing the flow-through to increase overall cell recovery. 27 Recycling of the diluted blood enables the user to recover more target cells, with a 33.2% increase in recovery by adding two more cycles. However, the additional processing cycle does result in additional contaminating WBCs. By enabling automated recycling of the effluent, the system offers the flexibility to be operated in either a high purity mode (one cycle) or a high recovery mode (three cycles).
Besides its flexibility in operating modes, the system can be configured to release cancer cells into various containers attached to the cartridge by an adapter. These include an Eppendorf tube, chamber slide, Petri dish, or well strip depending on the downstream analytical assay. This means that (1) no transfer of the sample is required, resulting in less CTC loss and less variability from manual intervention, and (2) cells are not affected by the processing (i.e., in suspension). The captured cells are unaltered by labels or reagents, which provides direct access to the patient’s intact cancer biology. We demonstrated that cancer cells captured by the VTX-1 maintained their capacity to form colonies and 3D spheroids that invade into a surrounding matrix. Additional studies are now focusing on live cell assays, such as CTC-derived xenograft models, RNA sequencing, or protein assays.29,38
As highlighted with the capture of cell lines with different EpCAM expressions, the VTX-1 captures CTCs independently of the markers they express. The VTX-1 uses the physical properties of the CTCs (i.e., their size and deformability) to isolate them in flow. Isolated CTCs are thus unbiased by molecular characteristics and fully recapitulate the patient’s disease. This gives access to CTCs at different stages of the epithelial-to-mesenchymal transition, 39 but also potentially CTCs from different cancer types that do not express EpCAM markers, such as melanoma or cancers for which no specific biomarkers are known. However, as for other label-free approaches, the capture dependence on cell size opens up new questions about the expected range of CTC size. Recent studies have highlighted the clinical relevance of CTC clusters in the metastatic process.3,40,41 Numerous CTC clusters, either homogeneous (CTC-like cells only) or heterogeneous (CTCs and WBCs), have been isolated with Vortex technology in other studies. 42 However, at this stage, we have not yet fully validated our recovery of the CTC clusters present in the circulation. Studies with artificially created clusters of cell lines spiked in healthy blood could help answer this question. Further, relatively smaller CTCs may be present in patient blood, the clinical impact of which has yet to be understood, that this technology and other size-based approaches likely capture with a lower recovery rate. 43 Besides, despite the collection of various cell populations independently of their markers, downstream CTC identification is still commonly performed with immunofluorescent staining, which may underestimate the capture performance of size-based CTC enrichment.43,44 This also implies variability of cell classification between the protocols, the antibody clones, the vendors, the imaging platforms, and the trained operators. Standardized and rigorous staining and classification methods taking malignant cell structural features into consideration are still needed, with inputs from cytopathologists.45,46 Finally, other large atypical cells are circulating in the blood of cancer patients, such as circulating endothelial cells, 47 circulating cancer-associated fibroblasts, 48 or macrophage-like cells, 49 which could potentially be isolated by the VTX-1 system as well but would need specific staining for their identification.
In preliminary clinical studies performed with the PDMS chips and the manual setup,8,26,28,29,45 CTCs were isolated with high purity (from 1.4 to 92.5 WBCs/mL) from patients with metastatic breast (median 40.68 CTCs per 7.5 mL, n = 22), colorectal (median 12.23 CTCs per 7.5 mL, n = 41), non–small-cell lung (NSCLC; median 26.25 CTCs per 7.5 mL, n = 15), and prostate (median 5.63 CTCs per 7.5 mL, n = 20) cancers. Such high purity in the CTC sample increases the accuracy and sensitivity of downstream assays, such as cytology, PCR, RNA sequencing, next-generation sequencing, and Sanger sequencing. For example, using Papanicolaou staining, atypical cells were detected in 15/16 NSCLC samples, with morphological similarities observed in the corresponding primary tumor. 45 In another study, KRAS, BRAF, and PIK3CA mutations were detected by Sanger sequencing, revealing 78% concordance between CTCs and liver metastasis for colorectal cancer patients. 8 The high purity of the CTCs captured with the VTX-1 can enable CTC population-based molecular analysis that was not previously possible. Improvements made on the microfluidic chip and automated processing have significantly increased the capture recovery of our technology, with an overall 3.2-fold increase with MCF7 cells spiked in blood (with different experimental conditions). Such improvement will likely translate into even better recovery of CTCs from patient samples as well, potentially expanding the CTCs’ enrichment to earlier cancer stages. 50 New studies on CTC isolation with the VTX-1 from patient samples with cancers of different types and different stages are in progress.
In addition to the use of CTCs in the clinic for cancer patient diagnosis, prognosis, and monitoring over the course of the therapy, an emerging application is the isolation and analysis of CTCs from mouse models. Patient-derived murine cancer models provide an advanced preclinical opportunity to study human cancer and its metastatic mechanisms, as well as to design and test targeted therapies.51–53 In this study, CTCs were successfully isolated with the VTX-1 from the blood of two triple-negative breast cancer PDOX samples with good purity (<200 contaminating WBCs/sample). All CTCs were VIM+/CK+, representing an epithelial to mesenchymal transition (EMT) phenotype. This highlights that the VTX-1 platform is compatible with the study of CTCs in various stages of EMT from murine models of breast cancer and could be potentially adapted to other types of cancer xenografts (e.g., prostate, lung, colorectal cancers) or other animals (e.g., rat). Additional studies will investigate transcriptional profiling of primary and/or metastatic tumors and single CTCs in PDOX samples derived from different patients. Besides the enrichment of CTCs, such technology could be applied to the isolation of various large atypical cells as potential markers for diagnosis, such as circulating endothelial cells and fibroblasts for cancer screening47,48 and myocardial infarction, 54 or even circulating fetal nucleated cells for prenatal genetic testing. 55
The VTX-1 Liquid Biopsy System represents the transition of the innovative Vortex technology into a robust system for the efficient isolation of circulating tumor cells. In a simple, fully automated workflow, the system captures clinically relevant samples of viable CTCs with high purity and makes them available for immediate analysis and characterization.
Footnotes
Acknowledgements
We thank our clinical coordinators, the nurses, and all blood donors for their contributions toward blood and tissue sample collection.
Supplementary material is available online with this article.
Contributions
C.A.L., C.L.W., V.C.R., N.A.B., K.W.H., J.C., M.W.C., M.L.K., D.D.C., S.S.J., R.F.E., S.H., C.R., and E.S.C. designed and planned experiments. C.A.L., S.Z.L., C.L.W., V.C.R., N.A.B., K.W.H., J.C., M.W.C., M.V., and A.M.D. helped perform experiments. C.A.L., S.Z.L., C.L.W., V.C.R., N.A.B., K.W.H., J.C., M.W.C., M.V., A.M.D., C.R., and E.S.C. analyzed the data. S.S.J. helped provide blood samples. C.A.L. and E.S.C. wrote the manuscript with assistance from all authors. All authors have read and approved this manuscript.
Declaration of Conflicting Interests
The authors disclosed the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Some of the authors (C.A.L., S.Z.L., C.L.W., N.A.B., K.-W.H., J.C., M.W.C., M.V., A.M.D., D.D.C., S.C.C., R.F.E., S.H., C.R., E.S.C.) have financial interests in Vortex Biosciences and intellectual property described herein.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The Jeffrey Lab acknowledges research funding support as part of an industrial contract between Vortex Biosciences and Stanford University (V.C.R., S.S.J.). The Di Carlo Lab acknowledges research funding support from Vortex Biosciences to UCLA (D.D.C.).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.