Abstract
Arginase-1, an enzyme that catalyzes the reaction of L-arginine to L-ornithine, is implicated in the tumor immune response and represents an interesting therapeutic target in immuno-oncology. Initiating arginase drug discovery efforts remains a challenge due to a lack of suitable high-throughput assay methodologies. This report describes the combination of self-assembled monolayers and matrix-assisted laser desorption ionization mass spectrometry to enable the first label-free and high-throughput assay for arginase activity. The assay was optimized for kinetically balanced conditions and miniaturized, while achieving a robust assay (Z-factor > 0.8) and a significant assay window [signal-to-background ratio > 20] relative to fluorescent approaches. To validate the assay, the inhibition of the reference compound nor-NOHA (Nω-hydroxy-nor-L-arginine) was evaluated, and the IC50 measured to be in line with reported results (IC50 = 180 nM). The assay was then used to complete a screen of 175,000 compounds, demonstrating the high-throughput capacity of the approach. The label-free format also eliminates opportunities for false-positive results due to interference from library compounds and optical readouts. The assay methodology described here enables new opportunities for drug discovery for arginase and, due to the assay flexibility, can be more broadly applicable for measuring other amino acid–metabolizing enzymes.
Introduction
Arginase-1, an enzyme responsible for catalyzing the conversion of L-arginine to L-ornithine and urea, is implicated in T-cell activity, offering promising avenues for immunotherapy.1,2 Arginase-1 is significantly upregulated and promotes tumor cell growth in select solid tumors, such as breast cancer3,4 and colorectal cancer. 5 Since L-ornithine is a biosynthetic precursor of polyamines that facilitate tumor cell growth and proliferation, arginase inhibition disrupts the L-ornithine supply for polyamine biosynthesis, thereby inhibiting cell proliferation.4,6,7 Indeed, arginase inhibition was shown to block the growth of lung carcinoma in a murine model. 8 Arginase-1 can also be upregulated by M2 macrophage cells in response to external stimuli, such as the Th2 cytokines interleukin-4 (IL4) and IL13, and is functionally associated with the M2 alternative activation pathway of macrophages. 9 Inhibiting Arginase-1 may therefore drive macrophages toward their tumor-killing response function.10,11 Drug discovery efforts focused on arginase are limited, largely due to a lack of suitable high-throughput screening (HTS) assays. Historically, arginase assays have relied on functionalization of the urea side product to generate a colorimetric signal.12,13 This indirect methodology requires harsh reagents such as sulfuric acid that are hazardous and not always compatible with automated platforms used in HTS. Recently, a fluorescent probe termed Arginase Gold was developed that exhibits a decrease in fluorescence due to quenching on conversion of arginine to ornithine. 14 While the probe enables a homogeneous, high-throughput arginase assay capable of analysis in kinetic mode, the robustness (Z-factor = 0.62) and small assay window in particular [signal-to-background (S/B) ratio = 1.52] 14 challenge the distinction between a hit and non-hit, and fluorescent assays are susceptible to false-positive and -negative results.
As an alternative to labeled assays, mass spectrometry (MS) has emerged as a powerful approach to analyze biochemical reactions. MS reports on the mass of the substrates and products, providing an information-rich and quantitative readout. Conventional MS methodologies, such as electrospray ionization (ESI), offer high sensitivity that is well suited for analyzing biochemical reactions. Reactions, however, require processing (often by solid-phase extraction) prior to analysis, limiting their use in large-scale screening campaigns. In one example, the RapidFire MS system has been used to screen a small library of 18,000 compounds against arginase-2. 15 A recent study showcased a novel RapidFire method called BLAZE-mode that eliminates the sample washing and enrichment steps, enabling a fourfold increase in throughput. 16 This approach may be attractive for distinct targets, provided the reactions do not require detergents or high salt concentrations that induce ion suppression in MS instruments or dilute samples that fall lower than the limit of detection. Alternatively, high-throughput MS applications, including matrix-assisted laser desorption ionization (MALDI)17,18 and acoustic dispense, 19 offer an accelerated option for HTS campaigns with a throughput of 100,000 compounds per day. The increased throughput is attributed to the elimination of the column chromatography washing and enrichment steps of the liquid chromatography (LC) system. With minimal or no sample preparation, assay development must focus on not only the optimization of the target reaction but also the MS instrument.
This study reports the development of the first label-free, high-throughput assay for measuring arginase-1 activity. The approach combines self-assembled monolayers of alkanethiolates on gold with matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF-MS). This methodology, termed SAMDI-MS (self-assembled monolayer desorption ionization mass spectrometry), overcomes the limitations of traditional MALDI-TOF-MS and other MS approaches. The specific immobilization of the analyte(s) of interest to the monolayer, while all other components are washed away, enables the analysis of virtually any enzyme activity that results in a mass shift. Moreover, the platform is amenable to any buffer system, including detergents, high salts, carrier proteins, organic additives, and even cell lysates.20–22 SAMDI-MS has been reported to measure biochemical activities on peptide,23–26 DNA, 27 and RNA substrates. 28 This is the first report to apply SAMDI-MS to a large-scale screen for inhibitors of an enzyme acting on a small-molecule substrate. The assay was biochemically characterized to ensure conditions were in a kinetically meaningful range, including assessing linear enzyme activity and determining substrate KM values. Furthermore, the assay was validated using the reference arginase inhibitors nor-NOHA (Nω-hydroxy-nor-L-arginine) 29 and ABH [2-(S)-amino-6-boronohexanoic acid]. 30 Finally, the SAMDI-MS assay was used to screen 175,000 compounds in <16 h, which represents, to the best of our knowledge, the largest screen reported for arginase. The advantages of the SAMDI-MS assay over traditional arginase assays and implications for improved drug discovery efforts are discussed.
Materials and Methods
Proteins and Compounds
Arginase-1 was purchased from BPS Bioscience (cat. no. 71658, San Diego, CA). L-arginine and L-ornithine were purchased from Sigma-Aldrich (St. Louis, MO). Nor-NOHA was purchased from Cayman Chemical (cat. no. 10006861, Ann Arbor, MI). The N-hydroxysuccinimide (NHS) ester probe was synthesized from BioPeptide (>95% purity; San Diego, CA). All other chemicals were purchased from Sigma-Aldrich.
Arginase-1 Mass Spectrometry Assay
Arginase-1 assays were performed in 6 µL volume in 384-well low-volume polypropylene microtiter plates (cat. no 784201, Greiner Bio-One, Kremsmünster, Austria) at ambient temperature. The optimized assay buffer was 50 mM Hepes HCl pH 7.5, 0.005% Tween-20, and 1 mM MnCl2. For compound screening, arginase-1 (final concentration = 10 nM) was added using a Multidrop Combi (Thermo Fisher Scientific, Waltham, MA) and pre-incubated for 30 min with small molecules to allow for slow on-rates. Reactions were initiated by the addition of the L-arginine (final concentration = 5 mM) and incubated for 120 min. Reactions were quenched by the addition of 0.5% formic acid (final) with subsequent neutralization using 1% sodium bicarbonate (final). A biotinylated probe featuring an NHS ester was added (final concentration = 2 µM) in water to react with the alpha amino group of L-arginine and L-ornithine. For SAMDI-MS analysis, 2 µL of each reaction mixture was transferred using a 384-channel automated liquid handler to SAMDI biochip arrays functionalized with a neutravidin-presenting self-assembled monolayer, as previously reported. 28 The SAMDI arrays were incubated for 30 min in a humidified chamber to allow the specific immobilization of the biotinylated probe–arginine and –ornithine conjugates. The samples were purified by washing the SAMDI arrays with deionized ultrafiltered water (50 µL/spot) and dried with compressed air. A matrix comprising 20 mg/ml alpha-cyano hydroxycinnamic acid (CHCA) in 80:20 of acetonitrile–dihydroxy ammonium citrate (DHAC; 10 mg/mL final in water) was applied in an automated format by dispensing 50 nL to each spot in the array. SAMDI-MS was performed using the reflector-positive mode on an AB Sciex TOF-TOF 5800 System (AB Sciex, Framingham, MA) with a laser intensity of 3000 using 324 shots/spectrum in a random raster sampling (18 shots/subspectrum with 18 subspectra pass acceptance), 400 Hz laser frequency, a bin size of 1 ns, and a detector voltage multiplier of 0.52. A mass window of m/z 1100 to m/z 1800 was used, and a mass threshold of 0.5 Da applied. Areas under the curve (AUCs) were generated using the OEM Applied Biosystems Series Explorer software (Thermo Fisher Scientific), and the amount of product generated was calculated using the ratio of product AUC divided by the sum of the AUC of the substrate and product. The third-degree polynomial (cubic) normalization equation, determined through the standard curve, is y = 0.000103x3 − 0.00322x2 + 0.359x − 0.702. Assay robustness was determined by Z-factor, calculated using the following equation: Z-factor = 1 – 3(σ+ + σ−) / (µ+ − µ−), where σ is the standard deviation, and µ is the average inhibition of positive (+) and negative (−) controls.
High-Throughput Screening
The 175,000 compounds included in this screen (SAMDI Tech collection) are derived from a diverse compound library synthesized in 2020 and dissolved in DMSO, with each compound achieving an average purity of >95%. Compounds were screened in pools of eight (12.5 µM final concentration each compound) by stamping 60 nL of 1.25 mM compounds into 384-well plates using a Mosquito HTS Automated Liquid Handler (SPT Labtech, Melbourn, UK). The assay was performed as described above. A selection of compounds was reassessed by assaying each compound individually in the arginase assay. The 100% inhibition controls using 0.5% formic acid (final) were placed in 32 wells, and 0% inhibition controls of 60 nL DMSO were added to another 32 wells. Assay quality was monitored using the Z-factor and S/B ratio for each 384-well plate.
Results
Development of a Mass Spectrometry Assay of Arginase-1 Activity
To develop a SAMDI-MS-based enzymatic assay for arginase-1 activity, we designed a probe featuring three components: a biotin group, a linker with a basic region and tetra(ethylene glycol) spacer, and an NHS ester that reacts with the amino group of arginine and ornithine post reaction. The biotin group allows for the specific immobilization onto neutravidin-presenting self-assembled monolayers for SAMDI-MS analysis, and the linker improves ionization efficiency during MALDI analysis ( Fig. 1A ). On arginase activity, the ornithine product is distinctly resolved in the SAMDI-MS spectrum from the arginine substrate by a shift of 42 Da ( Fig. 1B ), corresponding to the loss of the guanidinium group. To quantitatively assess product yield in an MS assay, it is important to consider ionization efficiency of the substrate and product. To test ionization efficiency, eight distinct ratios of arginine and ornithine were prepared and analyzed by SAMDI-MS. The spectra revealed the substrate peak at m/z 1487.0 and the product at m/z 1445.0. The amount of product observed is calculated by dividing the AUC of the product by the sum of the AUC of substrate and product. The amount of ornithine is then compared to the amount observed to generate a normalization factor that enables a linear relationship between actual and observed product ( Fig. 1C ), supporting the use of SAMDI-MS to quantitatively report on arginase activity.
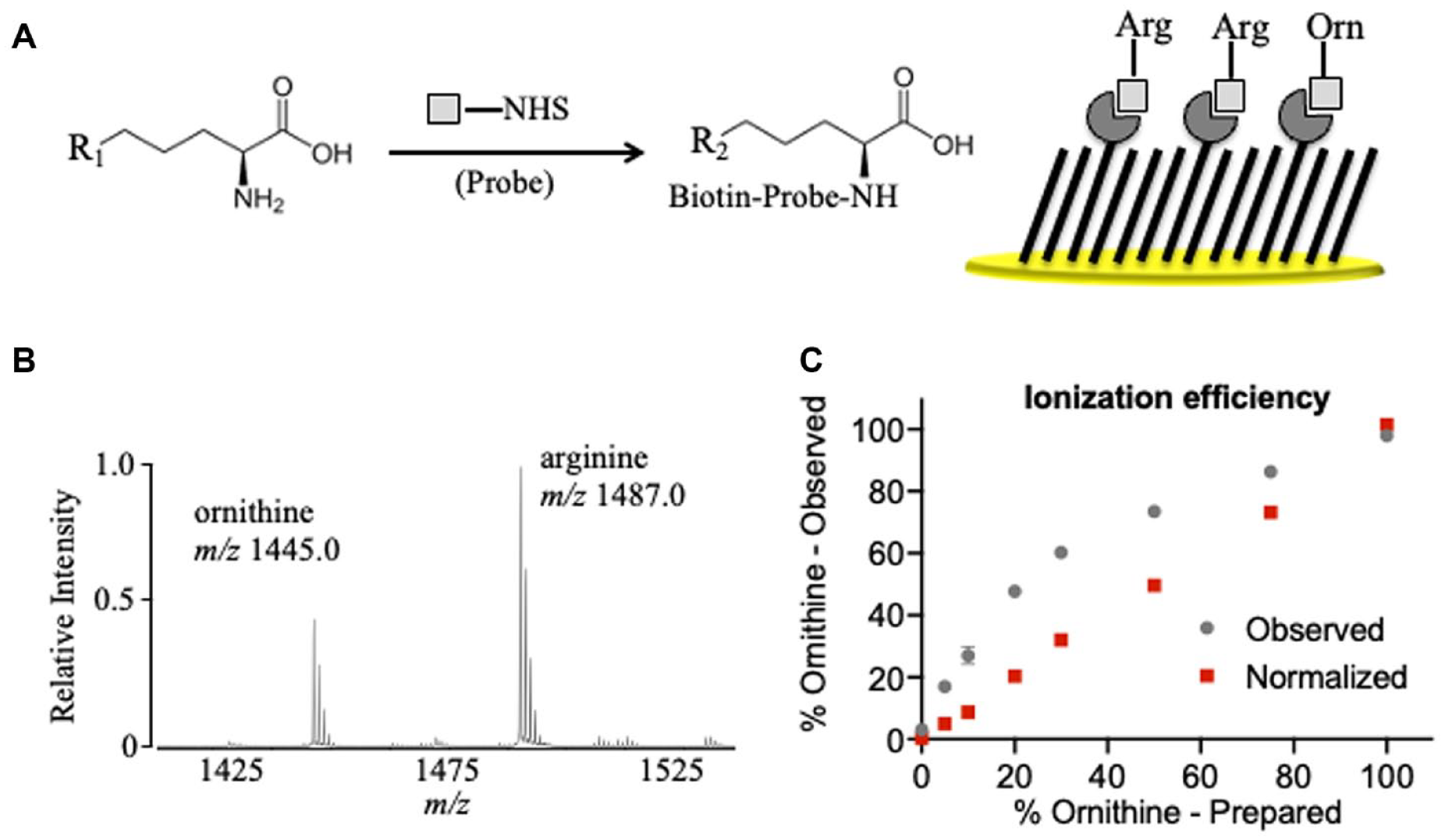
Self-assembled monolayer desorption ionization mass spectrometry (SAMDI-MS) assay of arginase-1 activity. (
Optimization of Assay Conditions and Determination of Kinetic Constants
Assay development is a critical process in drug discovery to optimize conditions suitable for evaluating small-molecule modulators. SAMDI-MS assay development started with an enzyme titration at 5 mM arginine to determine enzyme linearity in a buffer containing 50 mM Hepes pH 7.5, 0.005% Tween-20, and 1 mM MnCl2. Product formation increased with enzyme concentration over the range of 1.56 to 100 nM (
Fig. 2A
). Next, the KM and kcat were measured using SAMDI-MS. The arginine concentration was titrated between 200 µM and 25 mM, and the product was monitored throughout time using 10 nM enzyme. Given the alkaline nature of arginine, higher concentrations of arginine can increase the pH of the reaction, which has been shown to increase arginase activity. Therefore, the pH of the 25 mM arginine reaction was confirmed to be pH 7.5. Plotting the initial velocity (V0) of the linear portion versus the substrate concentration revealed a Michaelis–Menten curve with an apparent KM of 5 mM and a kcat of 93 s−1 (
Fig. 2B
). The kinetic values are consistent with several reports using alternative assay methodologies under similar assay conditions (pH 7.4–7.5: KM 1.5 mM, kcat 190 s−1;
31
KM 2.3 mM, kcat 300 s−1;
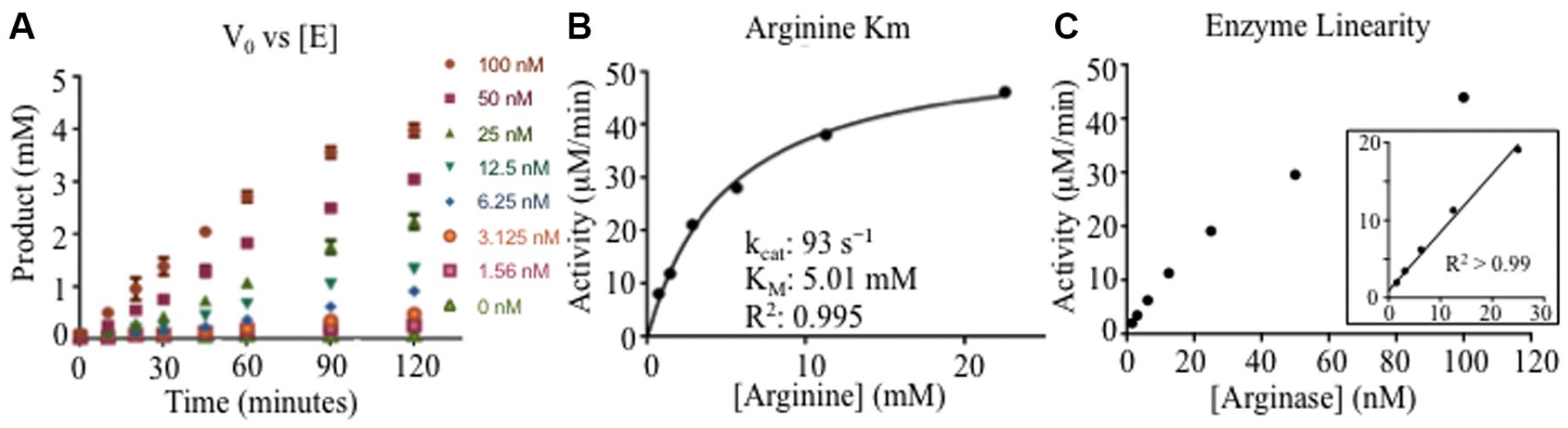
Arginase-1 assay development using self-assembled monolayer desorption ionization mass spectrometry (SAMDI-MS). (
Assay Performance and Validation with Positive Controls of Arginase-1 Inhibition
The robustness of the SAMDI-MS assay was assessed in 6 µL volumes in 384-plate format using the optimized conditions. Columns 1 and 2 of each plate included 100% inhibition controls by pre-quenching with 0.5% formic acid (final). The presence of positive and negative controls permitted calculation of a Z-factor (a measure of robustness), which considers the mean and standard deviation for positive and negative controls. The Z-factors among three plates on three different days were determined to be 0.810, 0.838, and 0.858 along with a S/B ratio of 24.0, 26.7, and 32.3, thus supporting a robust and reproducible assay ( Fig. 3A ). This significant S/B ratio measured by SAMDI-MS is attributed to the distinct mass of the probe–ornithine conjugate product relative to the probe–arginine conjugate substrate ( Fig. 1 ). Finally, the SAMDI-MS assay was used to determine the potency of the reference arginase inhibitors nor-NOHA and ABH.29,30 The SAMDI-MS assay used the optimized conditions and assessed the inhibition in the presence of nor-NOHA or ABH in a 10-point dose response in triplicate. The data point to an IC50 of 181 nM (132–247 nM) for nor-NOHA and 581 nM (504–664 nM) for ABH ( Fig. 3B ), which are in line with the reported IC50s of 119 nM and 554 nM, respectively, 14 using similar assay conditions, supporting the biochemical relevance of the SAMDI-MS assay.
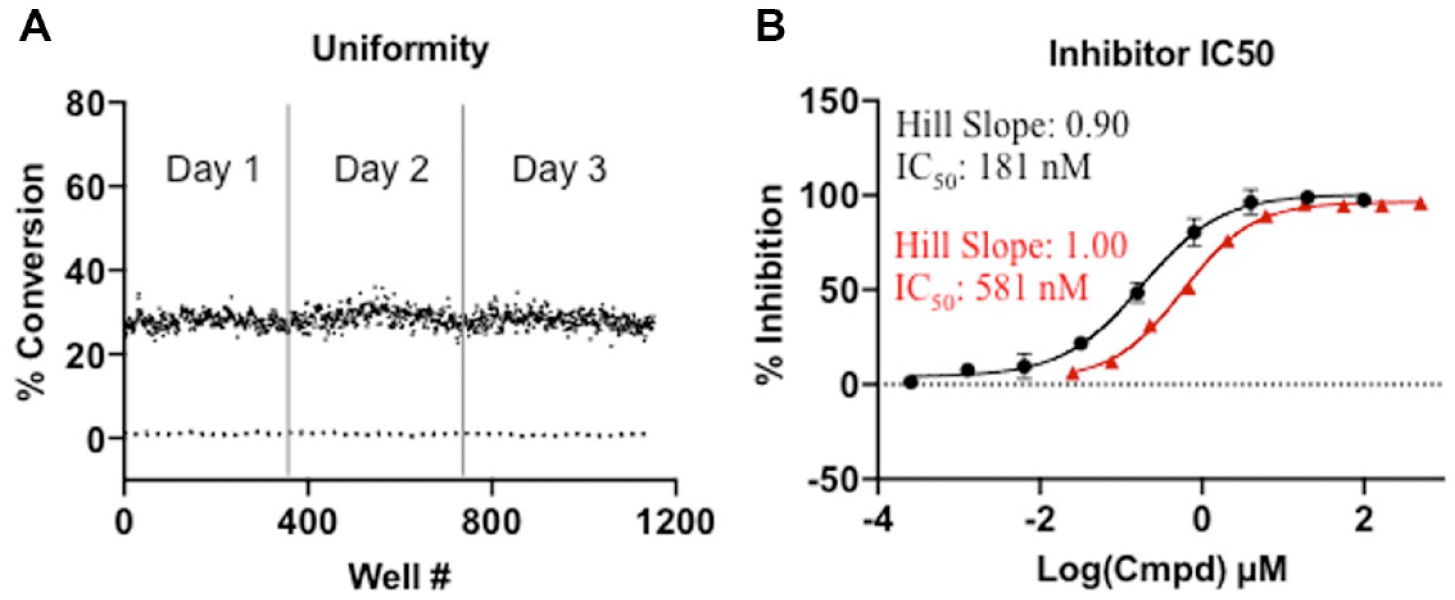
Self-assembled monolayer desorption ionization mass spectrometry (SAMDI-MS) assay robustness and validation. (
High-Throughput Screen of Arginase-1 Activity
To demonstrate the screening capability of the SAMDI-MS arginase assay, a chemically diverse 175,000-compound library was screened for inhibitors of arginase. This diverse set was synthesized in 2020 and features an average purity of >95%. Each compound is dissolved in DMSO, and the library was pooled with eight compounds per well prior to analysis, in which each compound was tested at a final concentration of 12.5 µM and a final DMSO concentration of 1%. The screen of 175,000 compounds was complete in <16 h, showing consistent activity across the screen (
Fig. 4A
), with an average Z-factor of 0.8 and an average S/B ratio of 22.0 (
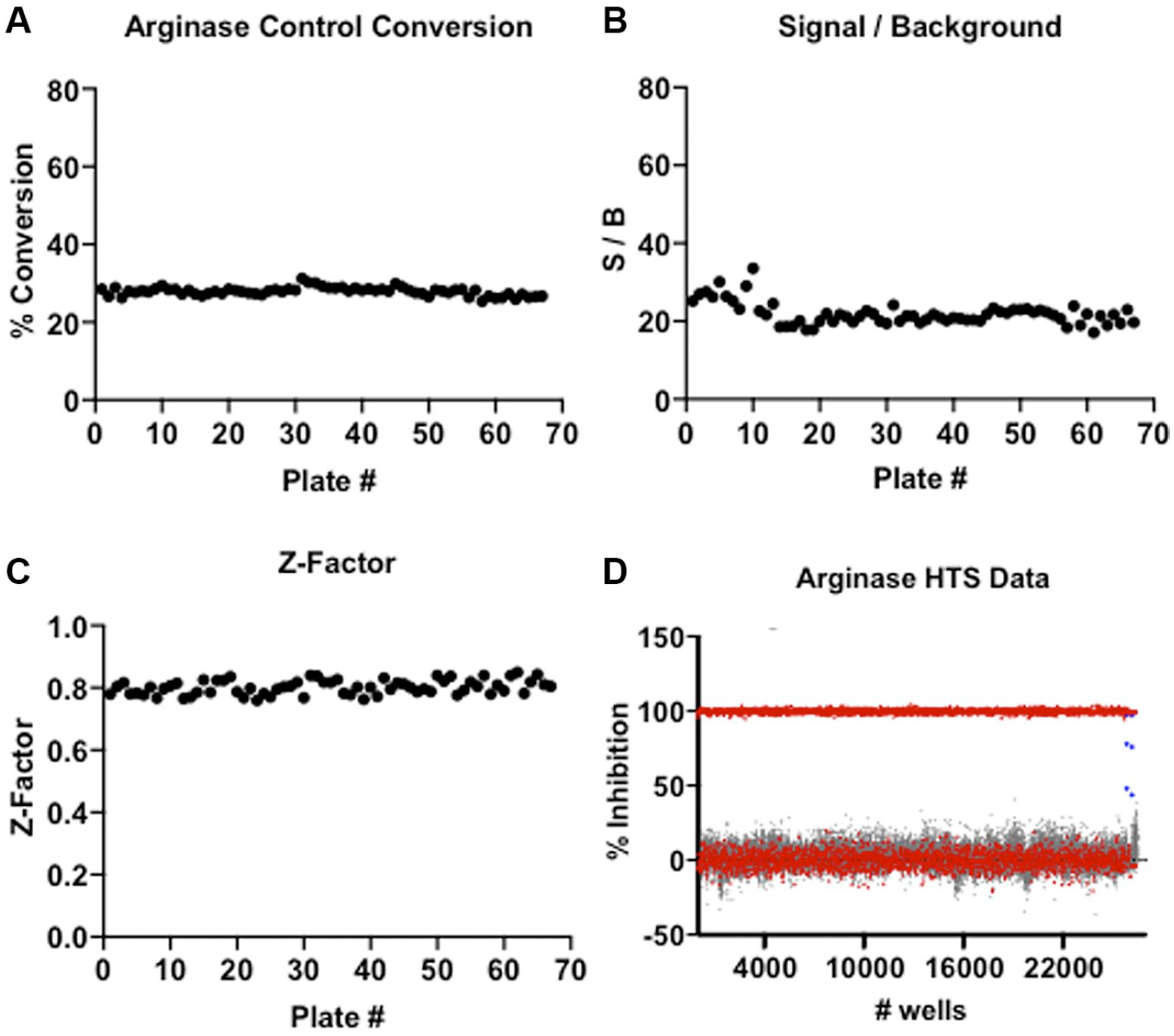
High-throughput screening data. (
Discussion
This study reports the first label-free and high-throughput assay to measure arginase-1 activity and opens avenues for improved drug discovery efforts to find small-molecule inhibitors. SAMDI-MS technology has been reported to characterize dozens of biochemical activities, including posttranslational modifying enzymes,25,34–38 RNA-modifying proteins, 28 proteases, 26 and enzymes that metabolize small molecules,39–41 demonstrating the assay flexibility. SAMDI-MS offers several solutions over other MS approaches, including traditional MALDI. Analyzing small molecules (<200 Da) by MALDI is challenging due to matrix interference, making this assay intractable by MALDI. By immobilizing the analytes onto self-assembled monolayers, the SAMDI-MS approach benefits from shifting the mass of small analytes to eliminate matrix interference effects. Moreover, while salts and detergents can induce ion suppression in traditional MALDI (as well as all traditional MS methods), the SAMDI-MS technology is amenable to any buffer component due to the specific immobilization of the analyte and robustness of the self-assembled monolayers. This capability enables the development of assay conditions that are optimized according to the target without having to compromise for the detection methodology. SAMDI-MS, however, retains the benefits of MALDI analysis, including detection of a singly charged, nonfragmented molecule for simplified data interpretation; the diverse analytes it can detect (e.g., small molecules, lipids, peptides, proteins, and oligonucleotides); the high-throughput readout; mass accuracy; and the robustness of the instrument throughout the screen. While many SAMDI-MS assays require incorporating a chemical handle onto the analyte of interest, such as a thiol for immobilization,23,42 introducing such a handle on arginine would likely affect its recognition by the enzyme. Therefore, this assay developed a probe featuring an NHS ester that reacts with the primary amino groups of arginine and ornithine post reaction. The probe also contains a biotin group to immobilize the arginine– and ornithine–probe conjugates to a neutravidin-presenting monolayer, and it features a linker with a basic region that improves the ionization efficiency of the substrate and product and a tetra(ethylene glycol) spacer for improved solubility. The assay conditions are designed such that the probe will react with the alpha amino group of arginine and ornithine (pKa 9.0) before the side-chain amino group of ornithine (pKa > 10). Notably, the probe has the potential to interact with other primary amines, including compounds such as nor-NOHA and ABH. This challenge is overcome by the fact that the concentration of the compounds is significantly lower than the arginine and ornithine substrate and product, and the probe–molecule conjugate has a distinct mass and therefore does not interfere with distinguishing arginine and ornithine. The probe approach enables the use of the high-throughput SAMDI-MS capability to evaluate enzyme activities on challenging substrates and has recently proven to be a powerful tool to measure cellular metabolites, 38 offering an orders-of-magnitude increase in speed and scale compared to conventional LC-MS systems.
The SAMDI-MS assay offers distinct advantages over label-dependent approaches that use colorimetric or fluorescent quenching to report on arginase activity. First, SAMDI-MS eliminates the requirement for an expensive fluorescent probe, saving on assay costs, while the MS readout provides a superior robustness (Z-factor = 0.8, relative to 0.62 for fluorescent approaches 14 ) and more than a 10-fold increase in assay window (S/B > 20, relative to 1.52 using fluorescent approaches and similar assay conditions 14 ) ( Fig. 4 ). Second, SAMDI-MS directly reports on the distinct masses of the arginine substrate and ornithine product, eliminating the additional step of harsh reagents needed to functionalize urea and produce a colorimetric signal. Third, SAMDI-MS eliminates any compound autofluorescence or fluorescence masking that occurs in most labeled assay approaches, 26 thereby allowing for screening in a compressed format of eight compounds per well. The label-free format of the SAMDI-MS assay eliminates the high rates of false-positive and -negative results common to optical readouts. 26 The approximate 65% false-positive and -negative hit rate due to compound autofluorescence or interference with the fluorescent probe exemplifies the caveat of optical readouts. 14 Characterizing small molecules with label-free SAMDI-MS ensures that only the most promising compounds progress toward the clinic, saving time and resources to drive drug discovery.
Screening 175,000 compounds and identifying minimally active compounds support the challenges with modulating arginase-1 with small molecules. Arginine derivatives represent most of the reported active compounds (nor-NOHA, CB-1158, and ABH).2,29,30 Future work will continue to explore a larger chemical space to identify novel chemical matter for this challenging target. Importantly, the SAMDI-MS assay will offer a high-throughput label-free methodology to test diverse chemical matter, including fragments and compounds arising from virtual screens or rational design methodologies.
In conclusion, the SAMDI-MS technology not only represents the first label-free and high-throughput assay for screening and characterizing inhibitors of arginase, but also is characterized by advantages in sensitivity, selectively, and robustness. Furthermore, the approach is flexible and can be applied to other amino acid–metabolizing enzymes, opening avenues for drug discovery around challenging metabolic targets.
Footnotes
Acknowledgements
The authors wish to thank Sandra Dib for assistance in preparing library compound pools and assay-ready plates, and the SAMDI Tech team for critical discussions.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
