Abstract
SARS-CoV-2, the coronavirus that causes COVID-19, evades the human immune system by capping its RNA. This process protects the viral RNA and is essential for its replication. Multiple viral proteins are involved in this RNA capping process, including the nonstructural protein 16 (nsp16), which is an S-adenosyl-
Introduction
Over the last two decades, major outbreaks of coronaviruses have been endured worldwide. These include zoonotic epidemics of severe acute respiratory syndrome coronavirus (SARS-CoV) in 2002, 1 Middle East respiratory syndrome coronavirus (MERS-CoV) in 2012,2,3 and the current pandemic of SARS-CoV-2 that started in 2019 (COVID-19). 4 With more than 56 million confirmed COVID-19 infections and more than 1.3 million deaths so far (November 20, 2020; World Health Organization; https://www.who.int/emergencies/diseases/novel-coronavirus-2019), the COVID-19 pandemic has had widespread global socioeconomic implications and overwhelmed healthcare systems worldwide.
Coronavirus is a member of the subfamily Coronavirinae within the family Coronaviridae.5,6 Many species can be infected by CoVs.
6
However, only seven coronaviruses have been documented to infect humans to date.4,7 These include SARS-CoV, MERS-CoV, and SARS-CoV-2 which could cause severe symptoms leading to higher fatalities.
4
Infection of 229E, HKU1, OC43, and NL63 has been associated with a range of relatively mild respiratory diseases.
7
SARS-CoV-2 shares 79.6% RNA sequence identity with SARS-CoV.
8
Coronaviruses are enveloped, nonsegmented positive-sense RNA viruses that have the largest genome among RNA viruses.9,10 They encode 16 nonstructural proteins (nsps) that play essential roles in RNA replication and processing of subgenomic RNA.8,11–13 SARS-CoV-2 replication involves RNA synthesis, proofreading, and capping.
14
Coronavirus mRNAs are protected at their 5′ ends by a cap structure consisting of an N7-methylated guanine linked to the first transcribed nucleotide by a 5′-5′ triphosphate bond and 2′-O-methylation (N7-meGpppN-2′-O-me) (
The capping process begins with the removal of the 5′γ-phosphate of the newly synthesized RNA chains (pppN-RNA) by nsp13.
19
Then a guanylyltransferase (GTase) catalyzes the formation of GpppN-RNA by transferring a guanosine monophosphate (GMP) molecule to the 5′-diphosphate of the RNA chains (ppN-RNA). Nsp14 methylates the cap structure at the N7 position of the guanosine, resulting in the formation of cap-0 (N7-meGpppN-RNA).
20
Ultimately, the nsp10-nsp16 complex catalyzes the addition of a methyl group on the ribose 2′-O position of the first transcribed nucleotide of the cap-0 to form a cap-1 (N7-meGpppNme-RNA) (
One of the cost-effective ways of screening for RNA substrate competitive inhibitors is to label the RNA with fluorescein and detect its binding to the target RNA methyltransferase by monitoring the change in fluorescent polarization signal. 29 Here we report the development of a fluorescence polarization (FP)-based RNA displacement assay that is suitable for high-throughput screening of the nsp10-nsp16 complex against large libraries of compounds to identify RNA competitive inhibitors of nsp16 methyltransferase activity. This assay is also a perfect tool for triaging a high number of potential inhibitors from screening by alternative methods for competing with RNA substrate and determining their mechanism of inhibition.
Materials and Methods
Reagents
S-adenosyl-
Protein Purification
His-tagged nsp10 (residues A1–Q139) and nsp16 (S1–N298) were expressed in Escherichia coli separately and purified to homogeneity using Ni-NTA resin. Purified nsp10 and nsp16 proteins were mixed at a molar ratio of 8 to 1, respectively, to prepare a stable and functional complex. The expression and purification of nsp10 and nsp16, and nsp10-nsp16 complex preparation are described in detail in the supplemental data (
FP-Based RNA Displacement Assays
All FP experiments were performed in a total assay volume of 20 µL per well in 384-well black polypropylene PCR plates (PCR-384-BK, Axygen, Tewksbury, MA) using FAM-labeled RNA (5′ N7-meGpppACCCCC-FAM 3′) and the nsp10-nsp16 complex. FP was measured after 30 min of incubation at room temperature, using a BioTek Synergy 4 (BioTek, Winooski, VT) with excitation and emission wavelengths of 485 nm and 528 nm, respectively. All experiments were performed in triplicate (n = 3), and plotted values are the average of three replicates ± standard deviation. The FP values were blank subtracted and were presented as the percentage of control (FP %). Data were visualized using GraphPad Prism software 7.04 (GraphPad, La Jolla, CA).
For assessing the binding of FAM-RNA (5′ N7-meGpppACCCCC-FAM 3′) to the nsp10-nsp16 complex, varying concentrations of the protein were incubated with 30 nM FAM-RNA in 10 mM Tris(hydroxymethyl)aminomethane (Tris) buffer, pH 7.5, containing 5 mM DTT, 0.01% Triton X-100, and 0.01% bovine serum albumin (BSA) for 30 min at room temperature in 20 µL of reaction volume. FP (485 nm/528 nm) was measured using a Biotek Synergy H1. The change in FP (mP) was plotted as a function of the nsp10-nsp16 concentration. The concentration of protein corresponding to the half-maximum FP signal (Kd) and the maximum FP signal (Bmax) was calculated using nonlinear least-squares regression to a single-site binding model in GraphPad Prism 7.04. 29
To establish the specificity of the assay, a mixture containing 0.5 µM nsp10-nsp16 complex and 30 nM FAM-RNA, in 10 mM Tris buffer, pH 7.5, containing 5 mM DTT, 0.01% Triton X-100, and 0.01% BSA, was incubated separately with varying concentrations of unlabeled RNA cap analogs: N7-meGpppG, N7-meGpppA, GpppG, and GpppA in 20 µL reaction volumes for 30 min at room temperature. The FP values were determined, and the Kdisp (K displacement) values (the concentrations required for 50% displacement of the labeled RNA) were calculated using nonlinear least-squares regression analysis to a four-parameter concentration–response curve model in GraphPad Prism 7.04.
Z′-Factor Determination
The Z′ factor was determined by incubating the nsp10-nsp16 complex (0.5 µM) with 30 nM FAM-RNA in the presence or absence of 50 µM unlabeled RNA (N7-meGpppA) in a 384-well plate for 30 min at room temperature (168 wells each). FP was measured as described above.
Screening Compounds from the Prestwick Library
The Prestwick Chemical Library was purchased from Prestwick Chemicals and screened using optimized assay conditions (10 mM Tris buffer, pH 7.5, containing 5 mM DTT, 0.01% Triton X-100, and 0.01% BSA) at a 50 µM compound concentration, with 0.5% DMSO concentration in 384-well black polypropylene PCR plates (Axygen, Tewksbury, MA, cat. PCR-384-BK) with a final reaction volume of 20 µL for 30 min at room temperature. Data were analyzed using GraphPad Prism 7.04 as described above.
Isothermal Titration Calorimetry
Binding of SAM, SAH, and sinefungin to the nsp10-nsp16 (8:1) complex was tested by Nano isothermal titration calorimetry (ITC; TA Instruments–Waters LLC, New Castle, DE) at 25 °C. The nsp10-nsp16 complex was dialyzed against 50 mM Tris-HCl buffer, pH 8.0, containing 200 mM NaCl and was loaded into the sample cell (300 µL) at a concentration of 48 µM, and a solution of 500 µM ligand (SAM, SAH, or sinefungin) was placed in the injection syringe (50 µL). Data were fitted with a one-binding-site model using TA Instruments origin software.
Results and Discussion
SARS-COV-2 nsp10-nsp16 Complex Preparation and Quality Control
The nsp10-nsp16 complex was prepared in an 8 (nsp10) to 1 (nsp16) ratio (
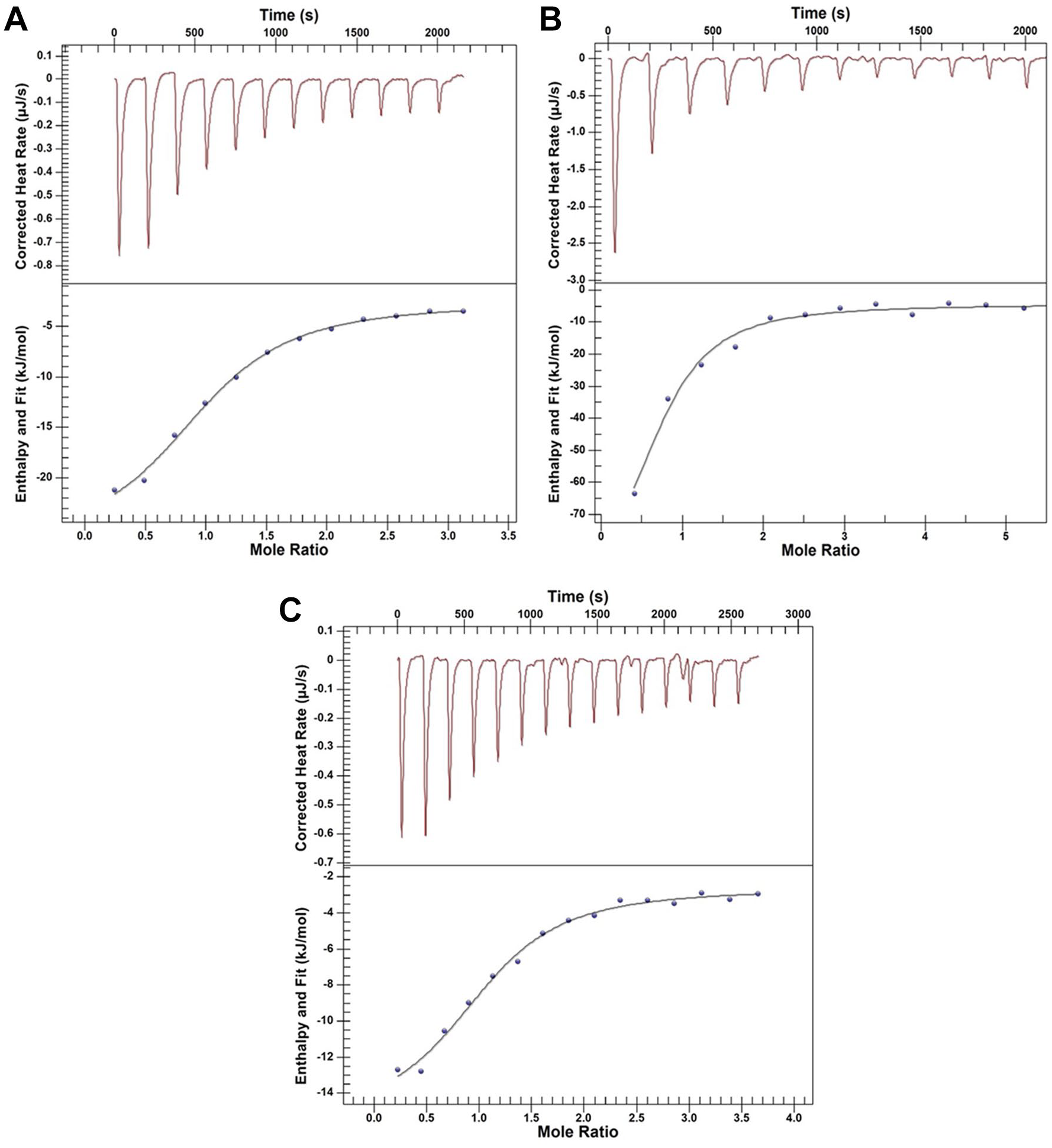
Binding assay. Binding of the nsp10-nsp16 complex to (
RNA Displacement Assay Development
The FAM-labeled RNA was used to set up the RNA displacement assay. Binding of this RNA to the nsp10-nsp16 complex was confirmed by detecting a significant increase in FP signal (
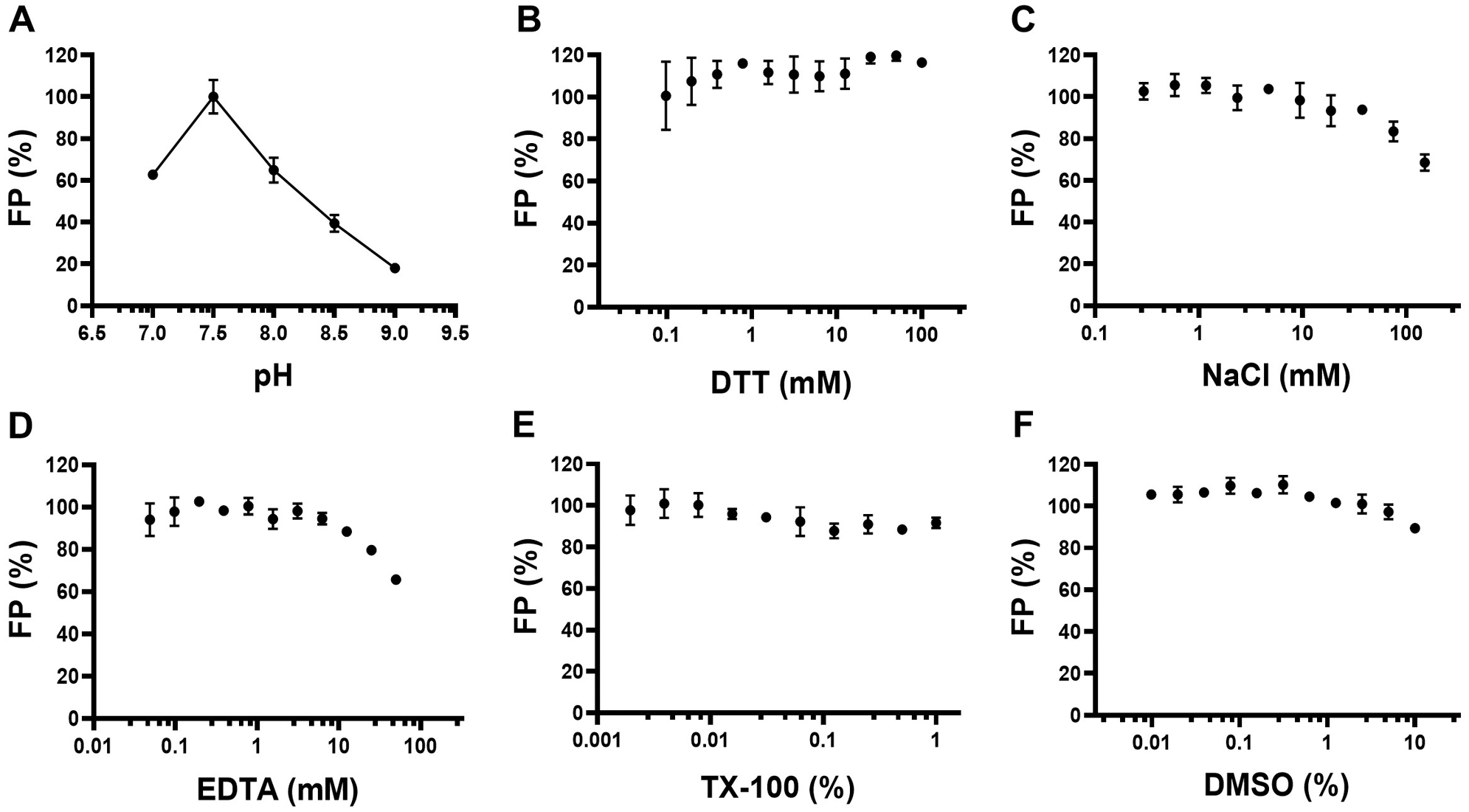
FP assay optimization for nsp10-nsp16 interaction with N7-meGpppACCCCC-FAM. The FP signal from the interaction of 0.5 µM nsp10-nsp16 complex with 30 nM N7-meGpppACCCCC-FAM was evaluated at (
Using the optimized assay conditions, the nsp10-nsp16 complex showed concentration-dependent binding to the FAM-RNA at 30 nM with an apparent Kd of 0.13 ± 0.002 µM and Bmax of 270 ± 5 mP (
RNA Displacement.
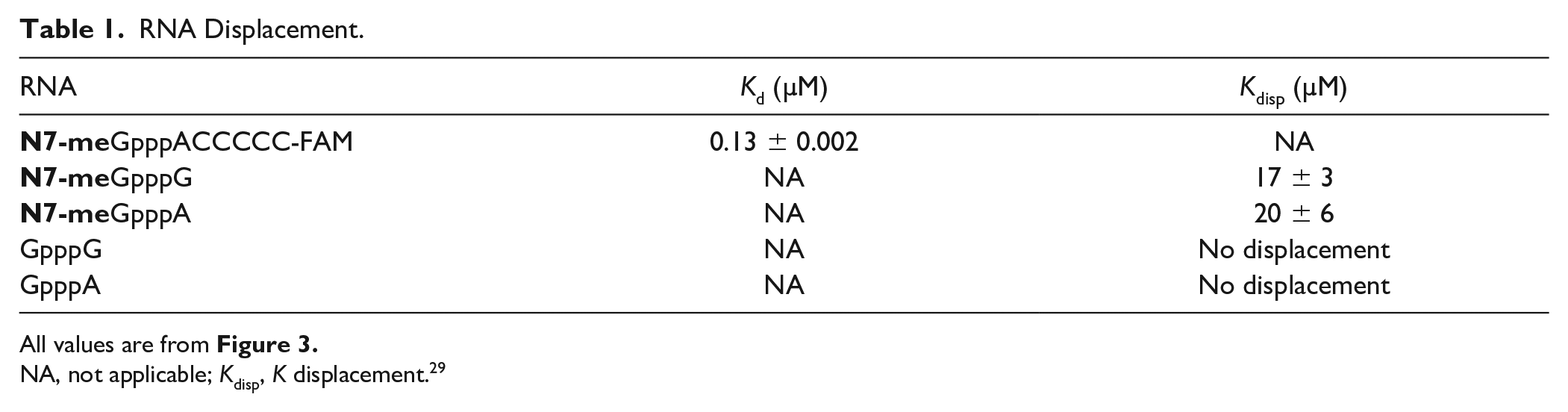
All values are from Figure 3 .
NA, not applicable; Kdisp, K displacement. 29
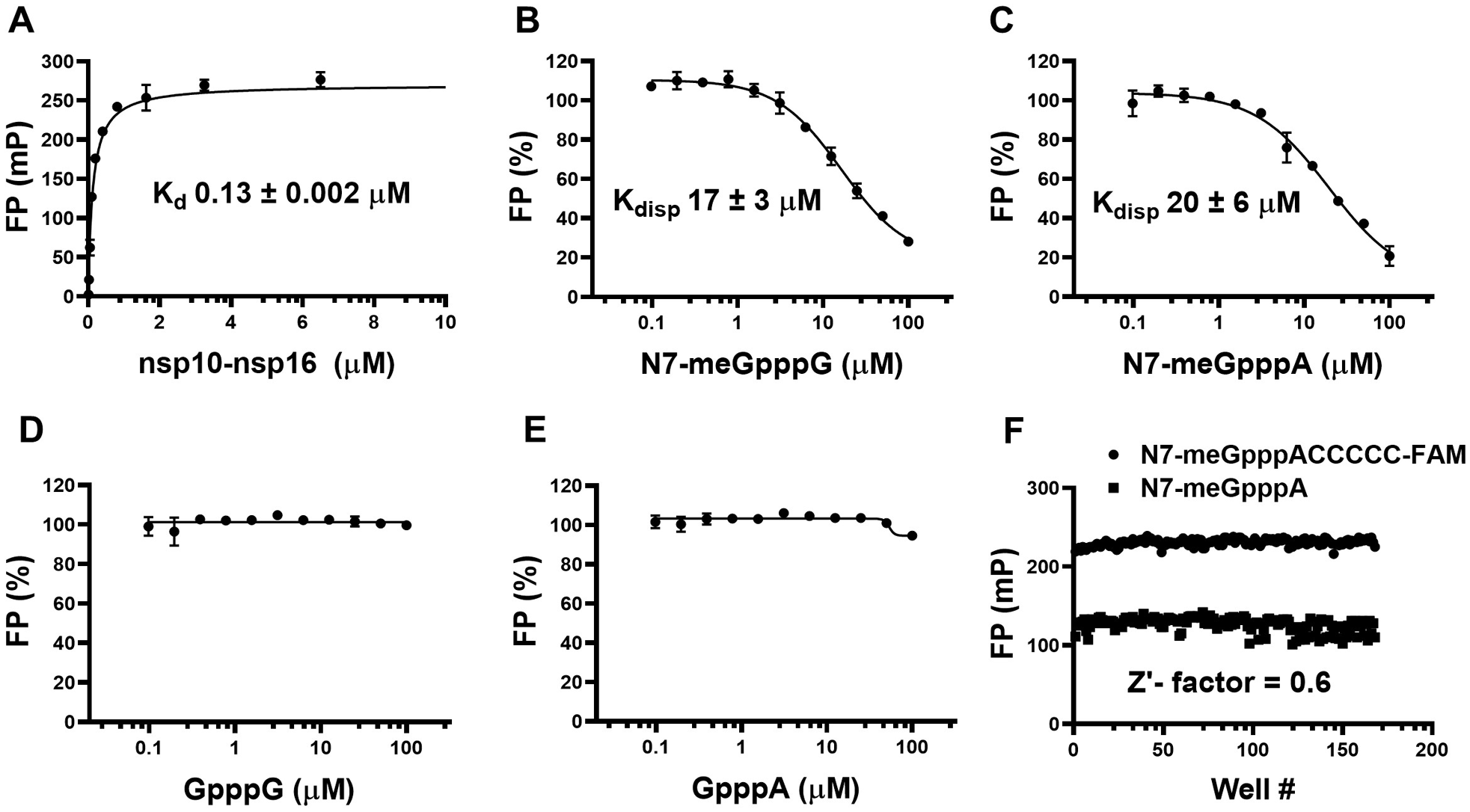
FP saturation binding and competitive displacement curves for the nsp10-nsp16 complex. (
To determine whether the RNA displacement assay at optimized conditions is amenable to high-throughput screening, the Z′ factor was determined by performing the assay in a 384-well format using FAM-RNA (5′ N7-meGpppACCCCC-FAM 3′) in the presence (168 data points) and absence (168 data points) of 50 µM unlabeled RNA (N7-meGpppA). The calculated Z′ factor was 0.6, indicating that the assay is suitable for screening for RNA competitive inhibitors (
The FAM-labeled RNA substrate in this study was designed based on the previous studies on the SARS-CoV nsp10-nsp16 complex reporting recognition of cap-0 RNA as substrate but not N7-meGpppG-capped RNA. 21 In vitro, RNA substrate for SARS-CoV nsp10-nsp16 needs to be N7-guanine methylated with adenine as the first nucleotide. 22 Similarly, no RNA displacement with the SARS-CoV-2 nsp10-nsp16 complex using the unmethylated RNA samples, such as GpppA and GpppG, was observed. These observations are also consistent with reports on similar binding assays with SARS-CoV 17 and MERS-CoV 25 proteins, where nsp16 can discriminate the cap-0 (N7-meGpppN-RNA) from an unmethylated RNA (GpppN-RNA).
Screening a Library of 1124 Compounds
A collection of 1124 drugs and drug-like compounds was screened. Initially, 15 compounds were identified as potential antagonists of nsp10-nsp16-RNA interaction based on reduction of polarization signal (>50% reduction) compared with the control at a 50 µM final compound concentration (
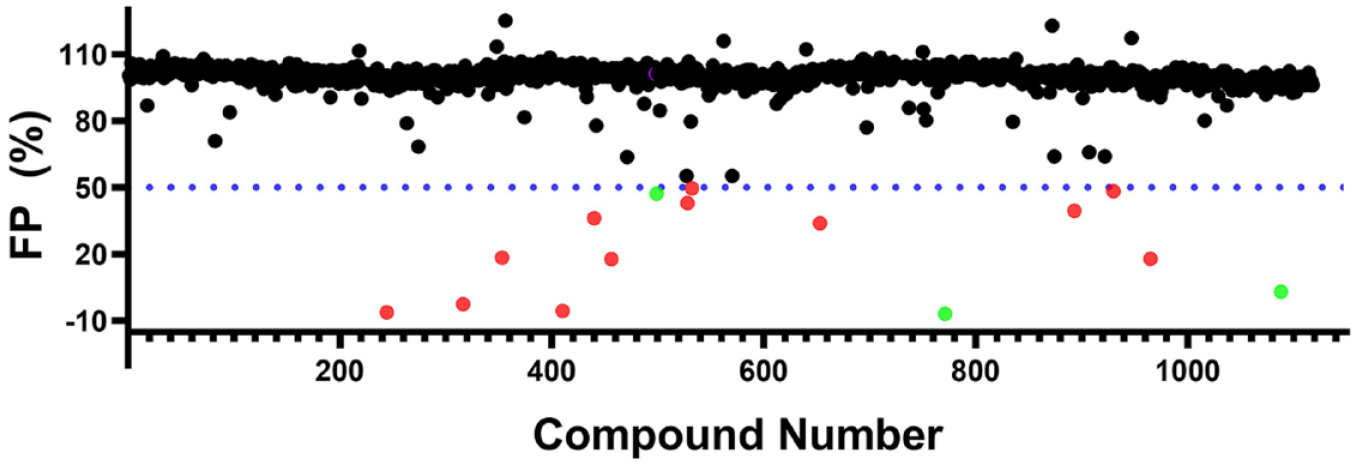
Screening Prestwick Compound Library. The distribution of the readout for 1124 compounds screened by the RNA displacement assay is shown. Compounds that significantly reduced the polarization signal (>50%) compared with the control were selected as screening hits (highlighted in red). Three compounds (highlighted in green) showed concentration-dependent inhibition. However, none of these compounds showed any binding when tested by ITC (data not shown).
All 15 compounds were tested for any possible interference with the signal readout, and 12 compounds showed a significant effect and were eliminated. Three compounds showed dose-dependent displacement with no signal interference. However, considering the high Hill slopes as a red flag, we further tested the binding of these three compounds to the nsp10-nsp16 complex by ITC, and none of the compounds were confirmed. This screening exercise confirmed the suitability of the optimized RNA displacement assay for medium- to high-throughput screening of the nsp10-nsp16 complex with a low number of false positives.
Limitation of the Assay
Methyltransferases have been shown to be druggable, 32 with more than 20 potent, selective, and cell-active small molecules (chemical probes) discovered for such enzymes in the last decade. 32 Typically, both SAM 33 and substrate binding sites of methyltransferases can be targeted for drug discovery. 32 The optimized FP-based RNA displacement assay for the nsp10-nsp16 complex is suitable for high-throughput screening to identify RNA competitive inhibitors. This assay is also suitable for triaging RNA competitive small molecules from a high number of hits identified by other methods. However, the RNA displacement assay will not detect SAM competitive inhibitors, unless they partially affect RNA binding. In addition, compounds that affect the FP signal or interact with RNA substrate could potentially increase the number of false positives. It is critical to confirm the RNA displacement-based screening hits by activity-based assays, and orthogonal methods such as ITC before the start of follow-up chemistry.
Conclusion
The COVID-19 pandemic has an unprecedented impact on the global economy and has already brought the healthcare services worldwide to a breaking point. So far, there is no effective drugs for this disease. Even vaccines developed today may not be effective on future mutated variants. 34 This necessitates the development of therapeutics that could stop the replication of the SARS-CoV-2. The RNA displacement assay in a 384-well format presented in this study provides a cost-effective method to screen the nsp10-nsp16 complex against large libraries of compounds to identify small-molecule RNA competitive inhibitors. Such molecules could lead to developing therapeutics for COVID-19 through inhibition of viral replication, and likely would be effective on other coronaviruses due to the high sequence conservation of nsp16 among this family of viruses.
Supplemental Material
sj-pdf-1-jbx-10.1177_2472555220985040 – Supplemental material for A High-Throughput RNA Displacement Assay for Screening SARS-CoV-2 nsp10-nsp16 Complex toward Developing Therapeutics for COVID-19
Supplemental material, sj-pdf-1-jbx-10.1177_2472555220985040 for A High-Throughput RNA Displacement Assay for Screening SARS-CoV-2 nsp10-nsp16 Complex toward Developing Therapeutics for COVID-19 by Sumera Perveen, Aliakbar Khalili Yazdi, Kanchan Devkota, Fengling Li, Pegah Ghiabi, Taraneh Hajian, Peter Loppnau, Albina Bolotokova and Masoud Vedadi in SLAS Discovery
Footnotes
Acknowledgements
We thank Dr. Matthieu Schapira for helpful discussions, Dr. Peter Stogios for providing reagents, Dr. Peter Brown for review of the manuscript, and Dr. Aled Edwards and Dr. Cheryl Arrowsmith for continued support.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the University of Toronto COVID-19 Action Initiative-2020, Takeda California, Inc., and COVID-19 Mitacs Accelerate postdoctoral awards to A.K.Y. and S.P. The Structural Genomics Consortium is a registered charity (no. 1097737) that receives funds from AbbVie, Bayer Pharma AG, Boehringer Ingelheim, the Canada Foundation for Innovation, the Eshelman Institute for Innovation, Genentech, the Genome Canada through Ontario Genomics Institute (OGI-196), EU/EFPIA/OICR/McGill/KTH, Diamond Innovative Medicines Initiative 2 Joint Undertaking (EUbOPEN grant no. 875510), Janssen, Merck KGaA (aka EMD in Canada and the United States), Merck & Co. (aka MSD outside Canada and the United States), Pfizer, the São Paulo Research Foundation–FAPESP, Takeda, and Wellcome (106169/ZZ14/Z).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
