Abstract
The COVID-19 pandemic has clearly brought the healthcare systems worldwide to a breaking point, along with devastating socioeconomic consequences. The SARS-CoV-2 virus, which causes the disease, uses RNA capping to evade the human immune system. Nonstructural protein (nsp) 14 is one of the 16 nsps in SARS-CoV-2 and catalyzes the methylation of the viral RNA at N7-guanosine in the cap formation process. To discover small-molecule inhibitors of nsp14 methyltransferase (MTase) activity, we developed and employed a radiometric MTase assay to screen a library of 161 in-house synthesized S-adenosylmethionine (SAM) competitive MTase inhibitors and SAM analogs. Among six identified screening hits, SS148 inhibited nsp14 MTase activity with an IC50 value of 70 ± 6 nM and was selective against 20 human protein lysine MTases, indicating significant differences in SAM binding sites. Interestingly, DS0464 with an IC50 value of 1.1 ± 0.2 µM showed a bisubstrate competitive inhibitor mechanism of action. DS0464 was also selective against 28 out of 33 RNA, DNA, and protein MTases. The structure–activity relationship provided by these compounds should guide the optimization of selective bisubstrate nsp14 inhibitors and may provide a path toward a novel class of antivirals against COVID-19, and possibly other coronaviruses.
Introduction
COVID-19, a severe acute respiratory syndrome in humans, is caused by SARS-CoV-2. It first surfaced in December 2019 1 and soon became a pandemic with more than 140 million confirmed cases and more than 3.1 million deaths reported worldwide at the time of this writing (https://www.who.int/emergencies/diseases/novel-coronavirus-2019). SARS-CoV-2 belongs to the Coronaviridae family of viruses,2,3 which also includes the severe acute respiratory syndrome coronavirus (SARS-CoV) and Middle Eastern respiratory syndrome coronavirus (MERS-CoV), which caused the SARS and MERS epidemics in 2002 and 2012, respectively.4,5 SARS-CoV-2 contains a nonsegmented, positive-sense 30 kb RNA that consists of 14 open reading frames (ORFs), 6 encoding 16 nonstructural proteins (nsps) and four main structural and accessory proteins. 7 The 16 nsps (referred to as nsp1 to nsp16) are more conserved among coronaviruses compared with the structural and accessory proteins. 8 These nsps in coronaviruses form a replicase–transcriptase complex and are essential for the transcription and replication of the virus. 9 Among these, nsp14 and nsp16 are RNA methyltransferases (MTases) involved in RNA capping. 10
Nsp14 is a bifunctional protein with a C-terminal MTase domain catalyzing N7- guanosine methylation and an N-terminal exoribonuclease domain (
While complex formation between nsp10 and nsp14 is required for enhanced exoribonuclease activity, the MT activity of nsp14 is independent of nsp10-nsp14 complex formation.15,16 The nsp14 S-adenosylmethionine (SAM)-dependent MTase activity is essential for viral mRNA capping.16,17 The cap1 structure at the 5′-end of viral RNA helps in masking the virus from the host immune system.18,19 The cap (GpppN) structure in nascent RNA of coronaviruses is formed by nsp1320,21 and a guanylyltransferase (GTase). Nsp14 methylates this cap structure at the N7 position of the guanosine, forming a cap-O (N7mGpppN). 17 Nsp16 further 2′-O-methylates the product of the nsp14 MTase activity, completing the capping process (N7mGpppNm).16,22
Nsp14 is conserved among the seven coronaviruses known to infect humans to date (
Material and Methods
Reagents
S-Adenosylhomocysteine (SAH) and sinefungin were purchased from Sigma-Aldrich (St. Louis, MO). S-adenosyl-
All compounds tested in vitro were ≥95% pure, except JL27-56A1, which was 86% pure at the time of screening. Purity was determined by analytical high-performance liquid chromatography (HPLC) on an Agilent 1100 series instrument equipped with a Phenomenex KINETEX column (50.0 mm × 4.6 mm, C18, 2.6 μM) at 25 °C. A linear gradient starting from 5% acetonitrile and 95% water to 95% acetonitrile and 5% water over 4 min, followed by elution at 95% acetonitrile and 5% water, was employed. Formic acid (0.1%) was added to all solvents.
Protein Expression and Purification
The expression and purification of SARS-CoV-2 nsp14 are provided as supplemental data.
Radiometric Assay Development
Assay Optimization
The MTase activity of nsp14 was measured using a radiometric assay. The transfer of the 3H-methyl group from 3H-SAM to the RNA substrate (5′ GpppACCCCCCCCC-Biotin 3′) was monitored using a scintillation proximity assay (SPA). Unless stated otherwise, all reactions were carried out in a 20 µL final volume and in triplicate at room temperature. For assay optimization, several concentrations of nsp14 (0.5 nM to 1 µM) were tested by mixing 1 µM 3H-SAM and 1 µM RNA in 20 µL of buffer. The reaction was stopped after 30 min by adding 20 µL of 7.5 M guanidium chloride and 20 µL of 20 mM Tris HCl, pH 8. The overall assay mixture was then transferred to a 384-well FlashPlate (SPA plate) coated with streptavidin/scintillant. After 3 h, the amount of methylated RNA formed was quantified using a TopCount (counts per minute [CPM]). To test additives and buffers, nsp14 (1.5 nM) was mixed with 50 nM RNA and 250 nM 3H-SAM and the reaction was stopped after 30 min. Each buffer (Tris, HEPES, and BTP) was tested at 20 mM at pH 7.5. The effect of pH was evaluated in Tris, pH 6.5–9.5. The effect of additives was monitored individually by titrating the reaction mixture with varying concentrations of DTT (from 0.1 mM to 100 mM), MgCl2 (from 0.1 mM to 100 mM), Triton X-100 (from 0.01% to 10%), and DMSO (from 1.25% to 10%) and comparing their relative activity to the samples with no additive.
Kinetic Characterization
The MTase activity of nsp14 was determined using the optimum buffer conditions identified (20 mM Tris HCl, pH 7.5, 250 µM MgCl2, 5 mM DTT, and 0.01% Triton X-100). The kinetic parameters (Kmapp and kcatapp) of nsp14 were determined using a series of reactions containing nsp14 at a saturating concentration of one substrate (1 µM RNA or 1 µM 3H-SAM) and varying concentrations of the others (from 15.6 nM to 2000 nM for 3H-SAM and from 7.8 nM to 1000 nM for RNA). Reactions were stopped at various time points (2, 5, 10, 15, 20, and 30 min). Initial velocities of the reaction were calculated from the linear portion of the reaction curves (see
IC50 Determination
Compounds were tested at various concentrations from 12 nM to 200 µM final concentration to determine their half-maximal inhibitory concentration (IC50) values. Potent compounds (SS148, SAH, and WZ16) were tested from 3 nM to 50 µM. The final DMSO concentration was 2%. The final reaction mixture consisted of 1.5 nM nsp14, 250 nM 3H-SAM, 50 nM RNA in 20 mM Tris HCl, pH 7.5, 250 µM MgCl2, 5 mM DTT, and 0.01% Triton X-100. The reaction time was 20 min. Data were fitted to the four-parameter logistic equation using GraphPad Prism 8.
Z′-Factor Determination
To evaluate the effectiveness of the nsp14 assay for screening purposes, the Z′ factor was determined by running 96 different reactions in the presence or absence of 1 µM sinefungin, a known MTase inhibitor. The final reaction consisted of 1.5 nM nsp14, 250 nM 3H-SAM, 50 nM RNA in 20 mM Tris HCl, pH 7.5, 250 µM MgCl2, 5 mM DTT, and 0.01% Triton X-100. Reactions were stopped after 20 min. The Z′ factor was calculated as described by Zhang et al. 24
Screening
Nsp14 was screened against the in-house library of 161 compounds at 50 µM in 1% DMSO. Compounds with inhibition of more than 75% were selected as screening hits for further analysis. The hits were tested for assay signal quenching at 50 µM. The signal was generated using 0.1 µM 3H-biotin in a 384-well FlashPlate that is coated with streptavidin/scintillant. Compounds that did not quench the signal were tested by dose-response against nsp14 at various concentrations from 12 nM to 200 µM final concentration to determine their IC50 values.
Mechanism of Action
The mechanism of action (MOA) of DS0464 was investigated by determining the IC50 values of the compound for nsp14 at the saturating concentration of one substrate (250 nM RNA or 1.25 µM 3H-SAM) and varying concentrations of the others (from 62.5 nM to 1.25 µM for 3H-SAM and from 12.5 nM to 250 nM for RNA).
Selectivity Assays
Selectivity assays were performed as previously described. 25 Briefly, compounds were tested at 50 µM in duplicate using radiometric assays. IC50 values were determined for compounds with higher than a 50% inhibitory effect, as described above.
Surface Plasmon Resonance
KD values for initial screening hits for nsp14 were determined by surface plasmon resonance (SPR) using a Biacore T200 from GE Healthcare (Chicago, IL). N-terminally biotinylated nsp14 (amino acids 1–527) and C-terminally biotinylated SETD3 (amino acids 1−605, as control) were coupled on a CM5 SPR Sensor chip (GE Healthcare). Compounds were injected into the sensitized chip at five concentrations (0.6, 1.9, 5.6, 16.7, and 50 µM) plus 0.5% DMSO at 50 µL/min, using HBS-EP Plus buffer (10 mM HEPES, pH 7.4, 150 mM NaCl, 2 mM EDTA, and 0.005% Tween-20) and 0.5% DMSO. Compounds with lower IC50 values (SS148, SAH, WZ16, and JL27-56A1) were injected at the following concentrations: 0.06, 0.19, 0.56, 1.67, and 5 µM. The contact time was 60 s and the disassociation time was 120 s. Buffer alone (plus 0.5% DMSO) was used for blank injections, and buffers containing 0.4%–0.6% DMSO were used for buffer corrections.
Modeling
DS0464 was docked into the MTase active site of SARS-CoV nsp14 (Protein Data Bank [PDB] code 5C8S) with ICM (Molsoft, San Diego, CA). PDB coordinates were loaded, side chains that were missing due to poor electron density were built and energy minimized with a biased-probability Monte Carlo simulation, hydrogens were added, and rotameric states of hydroxyl groups, terminal amides, and histidine side chains were optimized in the internal coordinate space. Ligand docking was conducted with fully flexible ligand and a grid representation of the protein with a Monte Carlo-based energy minimization. 26
Results
Assay Development and Optimization for High-Throughput Screening
Developing therapeutics for COVID-19 and other coronaviruses requires reliable high-throughput screening assays. Radiometric assays have been widely used for developing potent substrate and SAM competitive inhibitors for human MTases within the last decade.
25
Using biotinylated RNA substrate, we developed a radiometric nsp14 MTase activity assay with 3H-SAM as a methyl donor (

Kinetic characterization and screening of nsp14 MTase activity. The optimized radiometric MTase assay was used to determine the Kmapp and kcatapp values for ( ) and presence (
) and presence ( ) of 1 µM sinefungin. (
) of 1 µM sinefungin. ( ) were selected for follow-up experiments. Values in
) were selected for follow-up experiments. Values in
Kinetic Parameters of SARS CoV-2 Nsp14 MTase Activity
Using optimized assay conditions, we assessed the linearity of initial velocities (activity vs time) at various concentrations of RNA at a fixed SAM concentration (1 µM) (
Assay Amenability to High-Throughput Screening
In small-molecule screening campaigns, typically the assays are performed at Km of the substrates to allow potential inhibitors to compete with the substrates and allow their binding to be detected. However, the activity of the enzyme should be linear during the assay period. As the radiometric MTase assays are endpoint assays, lack of linearity may mask inhibition of some compounds. Testing the activity of nsp14 at 50 nM RNA and 250 nM SAM indicated that the assay can be run for at least 20 min while maintaining the linearity (
Screening SARS CoV-2 nsp14 against a Collection of Potential SAM Competitive Inhibitors
An in-house library of 161 SAM competitive MTase inhibitors and SAM analogs was screened against nsp14 at 50 µM, and 19 compounds were identified that inhibited nsp14 MTase activity more than 75% (
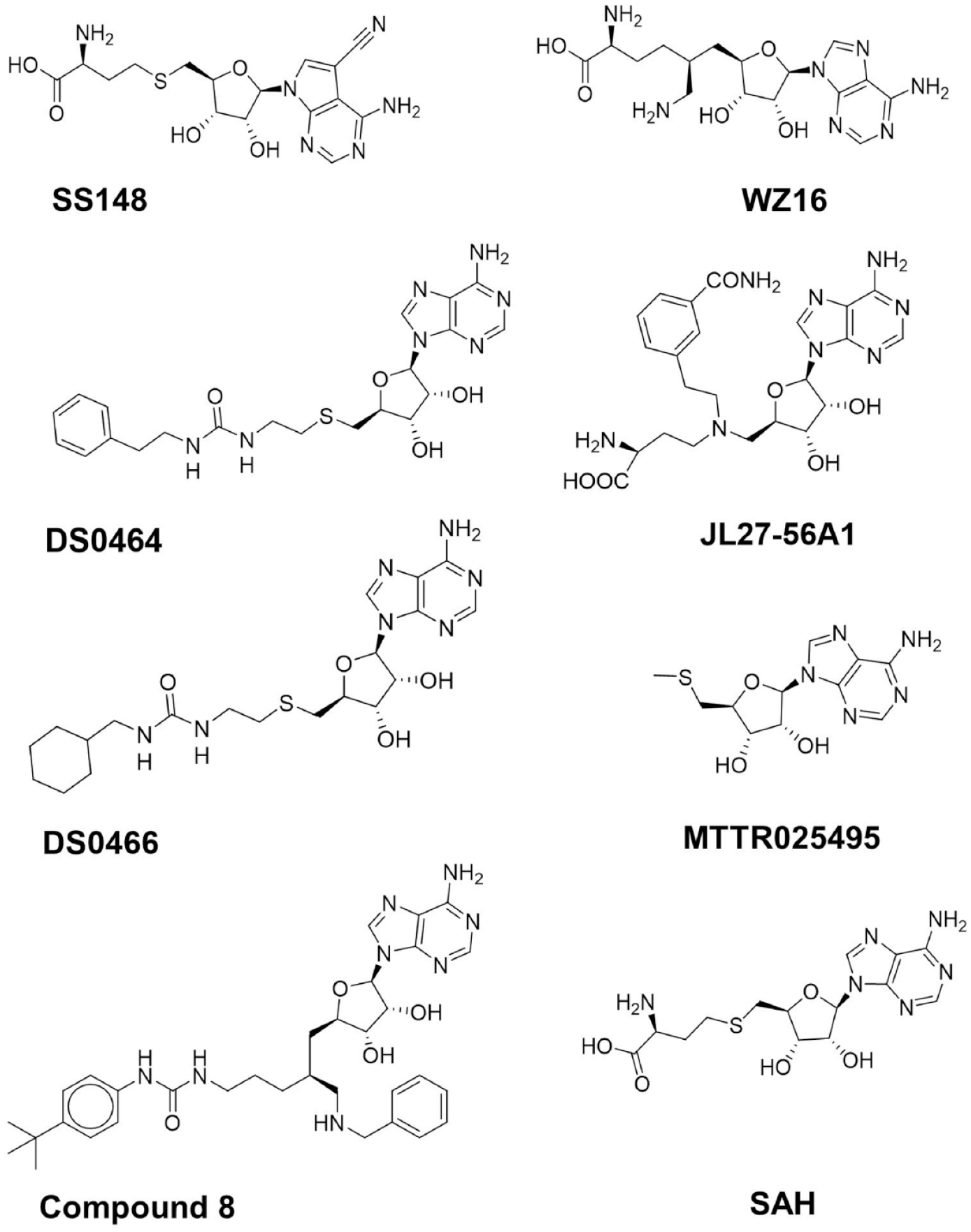
Structures of nsp14 screening hits.
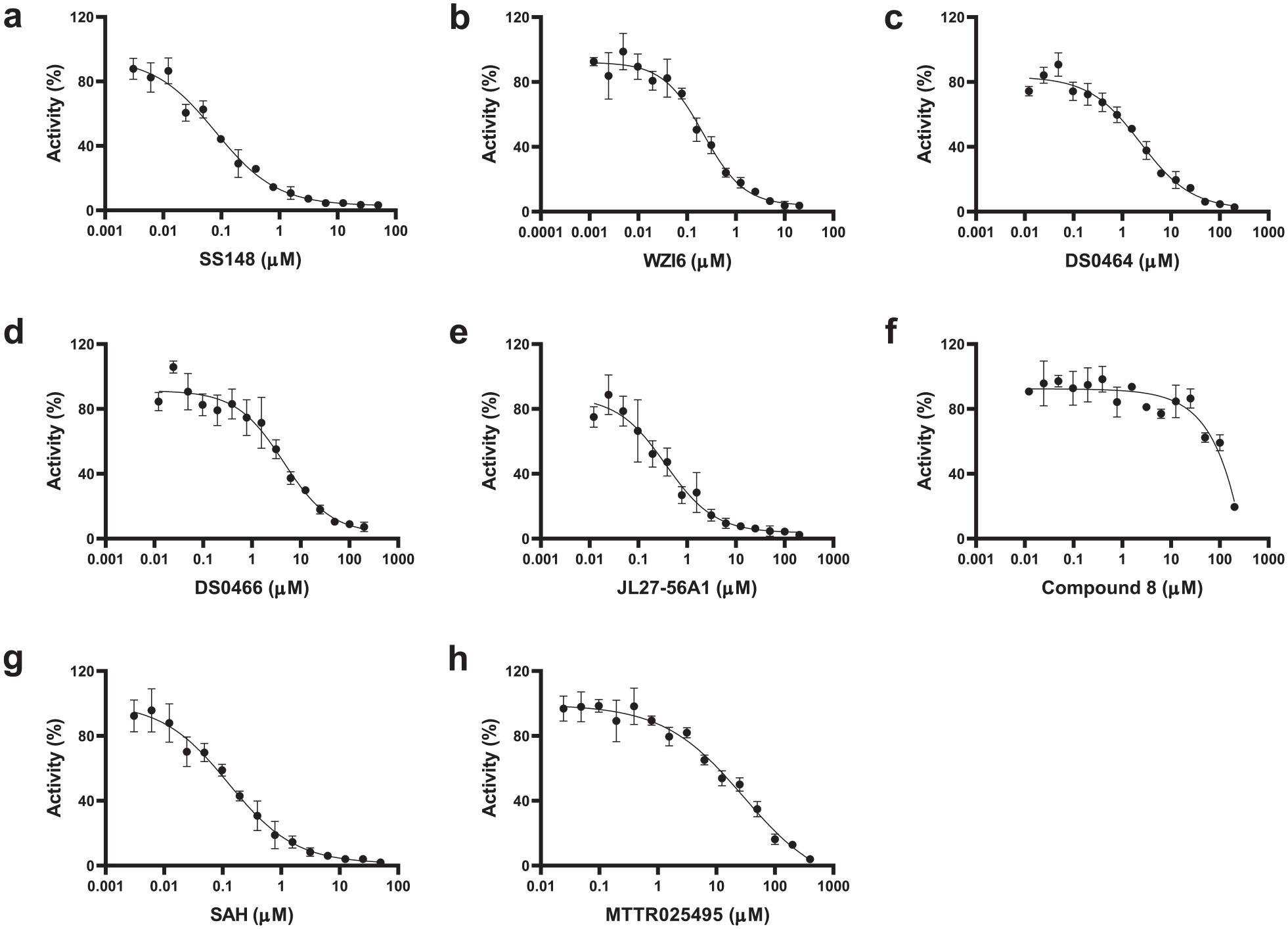
Dose–response curves of nsp14 screening hits. The IC50 values were determined for (
Confirmation and Selectivity of nsp14 Screening Hits.
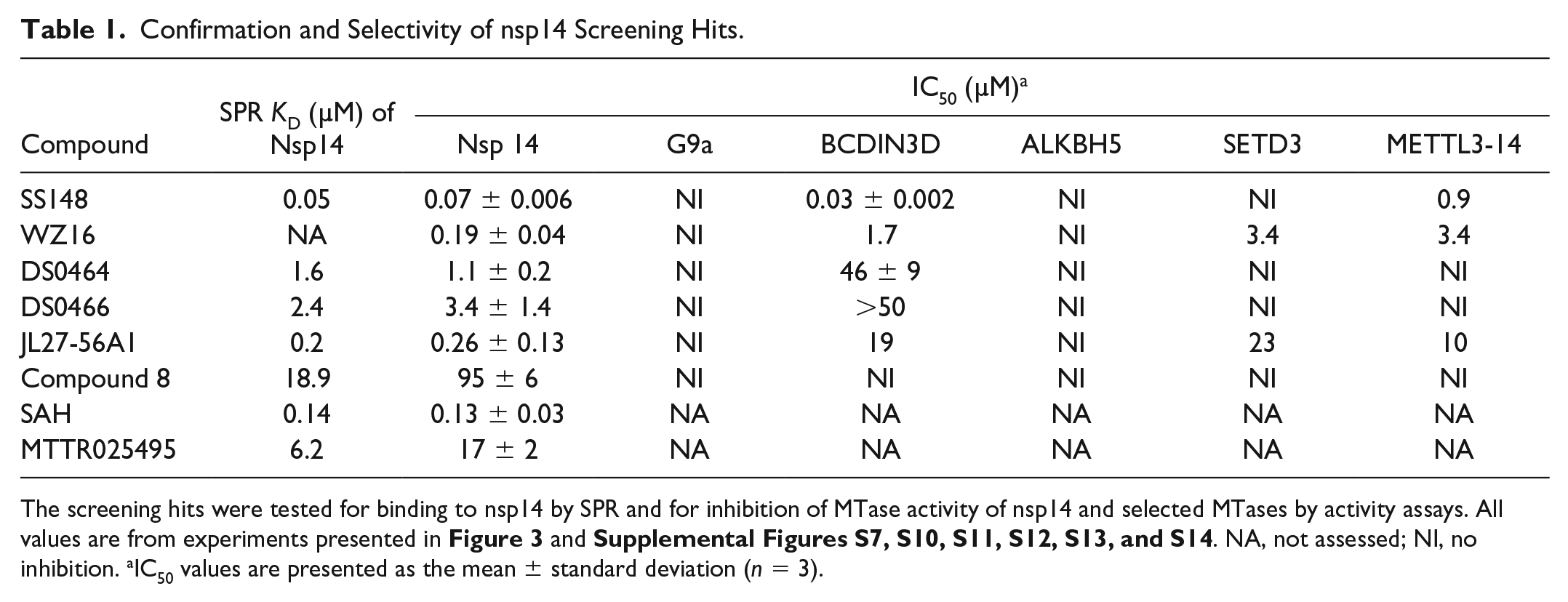
The screening hits were tested for binding to nsp14 by SPR and for inhibition of MTase activity of nsp14 and selected MTases by activity assays. All values are from experiments presented in Figure 3 and Supplemental Figures S7, S10, S11, S12, S13, and S14. NA, not assessed; NI, no inhibition. aIC50 values are presented as the mean ± standard deviation (n = 3).
Docking and Modeling
This limited and focused screening exercise provided important insights on the structural chemistry of SARS CoV-2 nsp14 inhibition (
Table 1
). First, we noted that the demethylated cofactor S-adenosyl-
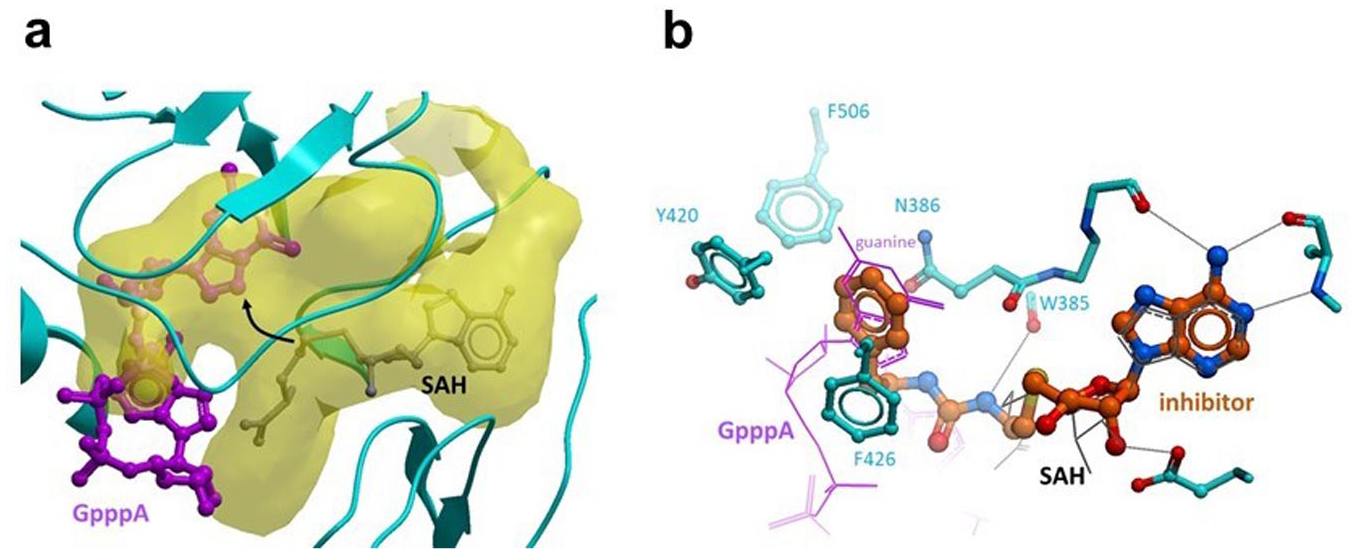
Docking model of DS0464. (
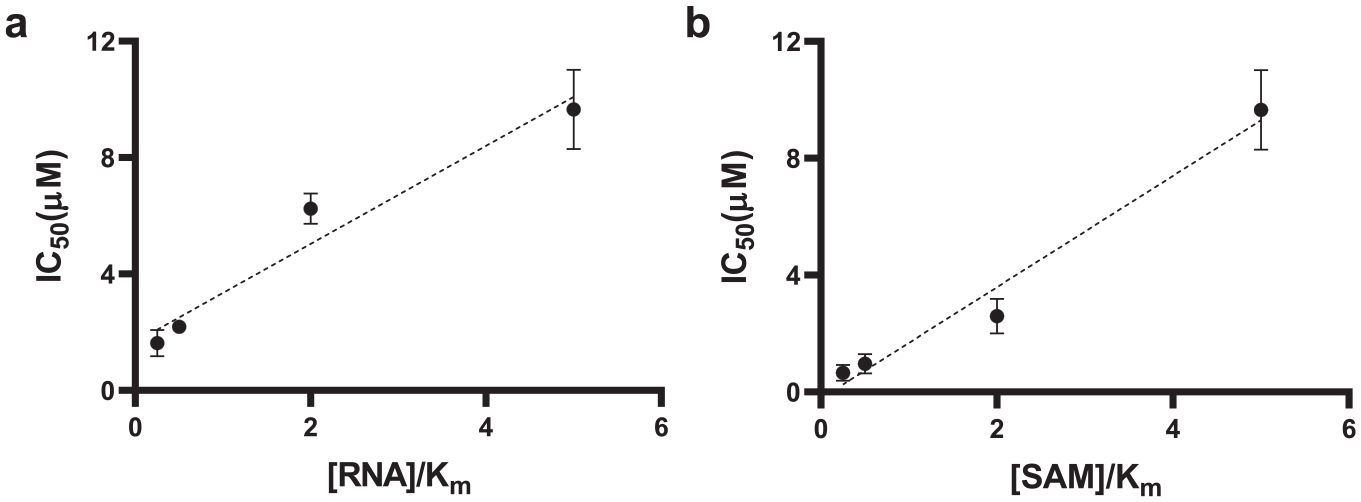
MOA of DS0464. IC50 values were determined for DS0464 at (
Selectivity of Screening Hits
The selectivity of all six compounds was tested against the human RNA MTase BCDIN3D and the METTL3-METTL14 complex (METTL3-14), the RNA demethylase ALKBH5, and protein lysine MTases G9a and SETD3 (
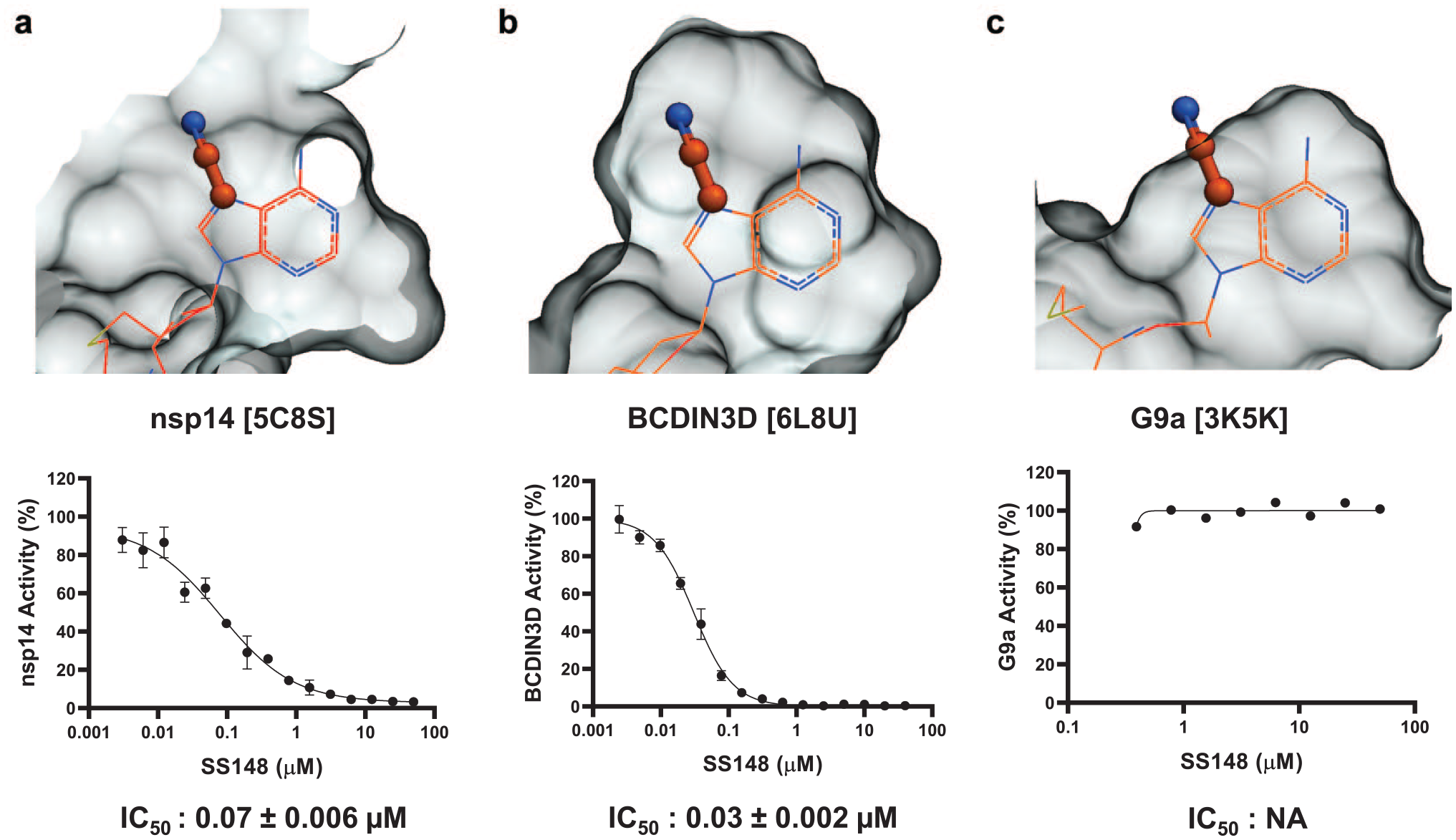
Structural determinant for SS148 selectivity. A nitrile group (stick) at position 7 of the cofactor adenine ring is compatible with the crystal structure of SAH (wire) bound to (
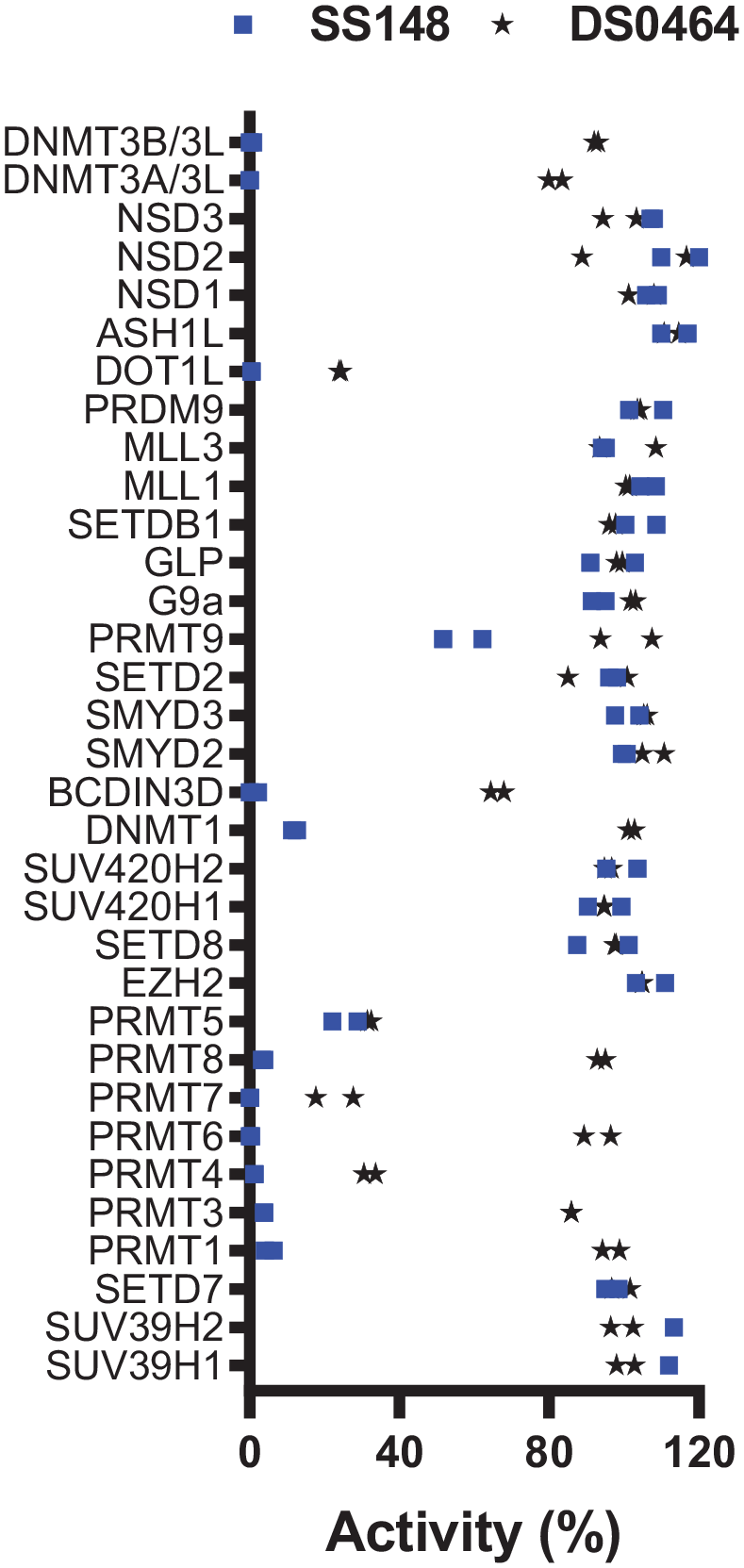

Selectivity of SS148 and DS0464. Both SS148 and DS0464 were screened for selectivity against 33 human RNA, DNA, and protein MTases at 50 µM. All experiments were performed in duplicate. When significant inhibition was observed, IC50 values were determined and are presented in Table 2.
Selectivity of SS148 and DS0464.
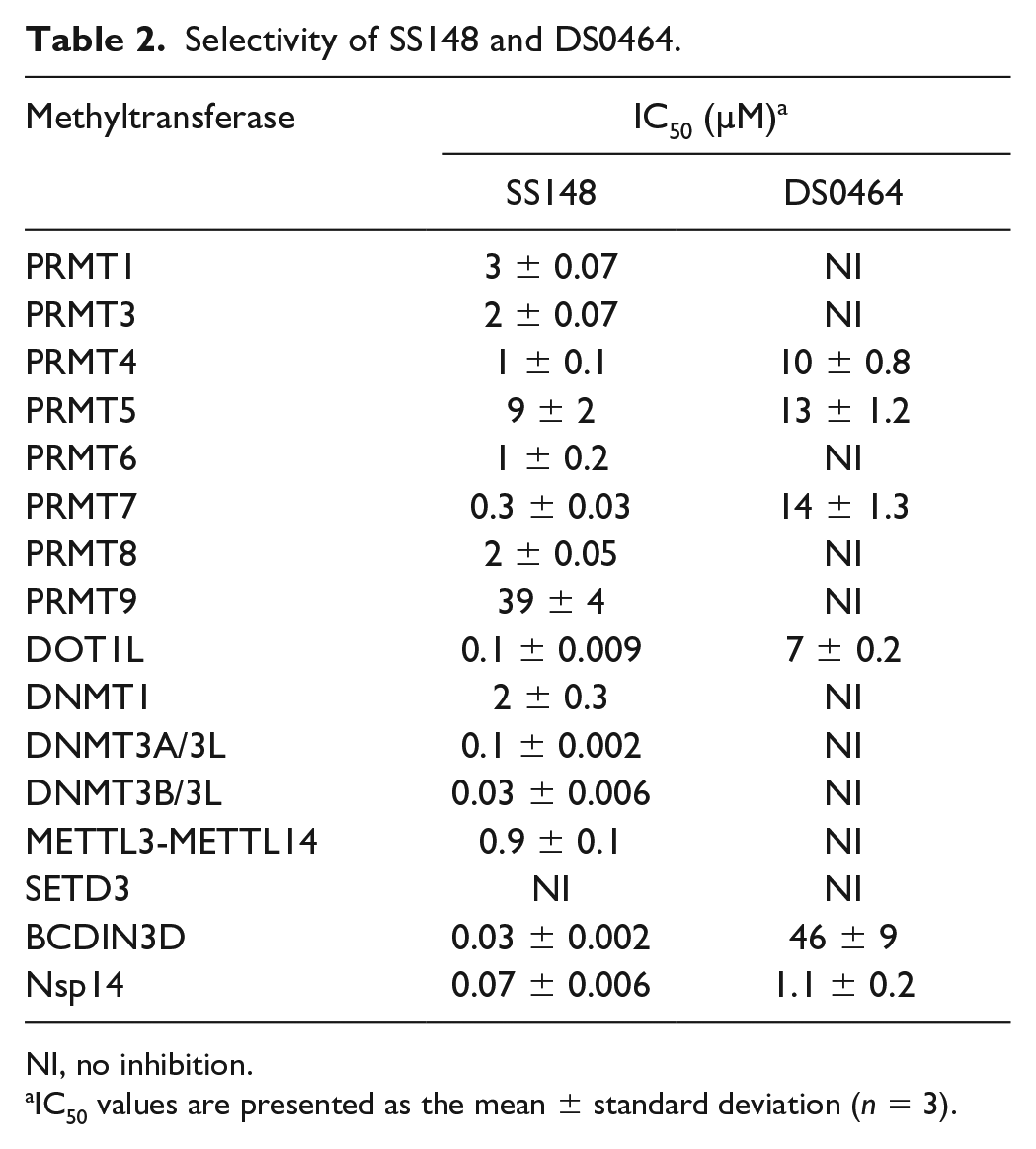
NI, no inhibition.
IC50 values are presented as the mean ± standard deviation (n = 3).
Discussion
The frequent emergence in the last two decades of novel coronaviruses as human pathogens, highlighted by the current COVID-19 pandemic, urgently needs to be addressed, preferably with pan coronavirus drugs. Nsp14 is an essential MTase in RNA cap formation that is required for protecting viral RNA and proper replication of coronaviruses. Therefore, targeting nsp14 MTase activity would be a viable option toward developing antiviral therapeutics. 28 MTases are druggable. 29 In the last decade, a significant number of selective and cell-active small molecules (chemical probes) have been discovered for human MTases,25,30,31 and some are in clinical trials for various cancers.30,31
Key to a successful discovery campaign of such chemical probes is the availability of reliable screening methods that could enable medium- to high-throughput screening with low false-positive and false-negative rates. Various assays including mass spectrometry,32–34 fluorescence, 35 and radiometric assays 25 have been used for screening libraries of compounds. Mass spectrometry-based assays require more expensive instrumentation and expertise. Fluorescence assays can be performed in any lab; however, many fluorescent compounds in chemical libraries may increase the background and lead to high numbers of false positives to triage following screening large libraries. Radiometric assays are typically more reliable and have fewer false positives, leading to identifying more reliable screening hits. 25 In this study, we have developed a radiometric assay for nsp14 and employed this assay for screening a small library of selected SAM competitive inhibitors and analogs.
Targeting the SAM binding site has successfully led to the discovery of chemical probes for human MTases such as DOT1L,36,37 EZH2/EZH1,
38
and SMYD2.
39
Synthesis of adenine dinucleoside SAM analogs as specific inhibitors of SARS-CoV (SARS) nsp14 RNA cap guanine-N7-MTase activity has also been recently reported.
40
SS148 (nsp14 IC50 = 70 ± 6 nM) was reported as a DOT1L inhibitor with a nitrile as a nontraditional replacement for heavy halogen atoms.
41
This is consistent with the selectivity of SS148 against all other protein lysine methyltransferases (PKMTs) due to narrower active sites that could not fit the added nitrile group (
Supplemental Material
sj-pdf-1-jbx-10.1177_24725552211026261 – Supplemental material for Probing the SAM Binding Site of SARS-CoV-2 Nsp14 In Vitro Using SAM Competitive Inhibitors Guides Developing Selective Bisubstrate Inhibitors
Supplemental material, sj-pdf-1-jbx-10.1177_24725552211026261 for Probing the SAM Binding Site of SARS-CoV-2 Nsp14 In Vitro Using SAM Competitive Inhibitors Guides Developing Selective Bisubstrate Inhibitors by Kanchan Devkota, Matthieu Schapira, Sumera Perveen, Aliakbar Khalili Yazdi, Fengling Li, Irene Chau, Pegah Ghiabi, Taraneh Hajian, Peter Loppnau, Albina Bolotokova, Karla J. F. Satchell, Ke Wang, Deyao Li, Jing Liu, David Smil, Minkui Luo, Jian Jin, Paul V. Fish, Peter J. Brown and Masoud Vedadi in SLAS Discovery
Footnotes
Acknowledgements
We thank Dr. Aled Edwards and Dr. Cheryl Arrowsmith for continued support, and Brian Wilson for performing compound quality control.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by University of Toronto COVID-19 Action Initiative-2020 and COVID-19 Mitacs Accelerate postdoctoral awards to A.K.Y. and S.P.; U.S. National Institutes of Health grant R35GM131858 to M.L.; and NIH/NIAID contract HHSN272201700060C to K.J.F.S. The Structural Genomics Consortium is a registered charity (no. 1097737) that receives funds from AbbVie, Bayer AG, Boehringer Ingelheim, Canada Foundation for Innovation, Genentech, Genome Canada through Ontario Genomics Institute (OGI-196), EU/EFPIA/OICR/McGill/KTH/Diamond Innovative Medicines Initiative 2 Joint Undertaking (EUbOPEN grant 875510), Janssen, Merck KGaA (aka EMD in Canada and the United States), Pfizer, Takeda, and Wellcome (106169/ZZ14/Z).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
