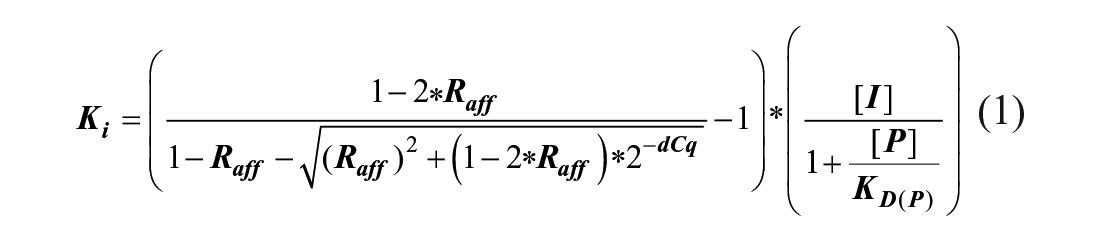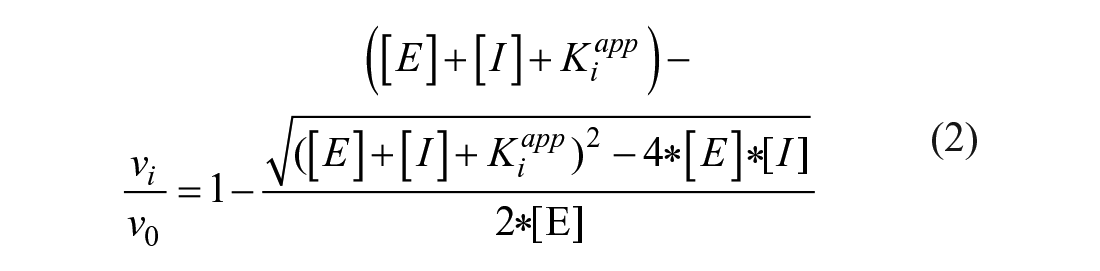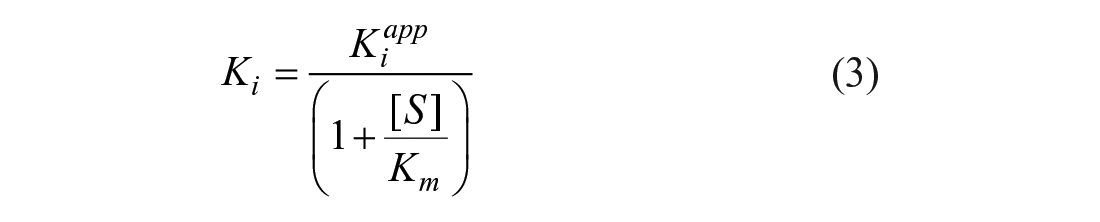Abstract
The DNA-linked inhibitor antibody assay (DIANA) has been recently validated for ultrasensitive enzyme detection and for quantitative evaluation of enzyme inhibitor potency. Here we present its adaptation for high-throughput screening of human carbonic anhydrase IX (CAIX), a promising drug and diagnostic target. We tested DIANA’s performance by screening a unique compound collection of 2816 compounds consisting of lead-like small molecules synthesized at the Institute of Organic Chemistry and Biochemistry (IOCB) Prague (“IOCB library”). Additionally, to test the robustness of the assay and its potential for upscaling, we screened a pooled version of the IOCB library. The results from the pooled screening were in agreement with the initial nonpooled screen with no lost hits and no false positives, which shows DIANA’s potential to screen more than 100,000 compounds per day.
All DIANA screens showed a high signal-to-noise ratio with a Z′ factor of >0.89. The DIANA screen identified 13 compounds with Ki values equal to or better than 10 µM. All retested hits were active also in an orthogonal enzymatic assay showing zero false positives. However, further biophysical validation of identified hits revealed that the inhibition activity of several hits was caused by a single highly potent CAIX inhibitor, being present as a minor impurity. This finding eventually led us to the identification of three novel CAIX inhibitors from the screen. We confirmed the validity of these compounds by elucidating their mode of binding into the CAIX active site by x-ray crystallography.
Keywords
Introduction
Carbonic anhydrase IX (CAIX), 1 of 12 catalytically active human carbonic anhydrase (CA) isoforms, is a promising cancer target. 1 It is an integral type I transmembrane protein with a catalytically active domain facing to the extracellular space, which catalyzes the hydration of carbon dioxide to carbonate and thus helps cells to maintain their acid–base balance. 2 CAIX is upregulated by hypoxia and is thus overexpressed in a variety of solid tumors.3,4 Through its enzymatic activity, CAIX is responsible for acidification of the extracellular space of tumors, which is associated with their aggressiveness. Simultaneously, it counteracts acidification of intracellular space, which is particularly important in tumors due to their metabolic acidification.5,6 Therefore, it is not surprising that inhibition of CAIX has been shown to impair the growth of CAIX-expressing tumors in both in vitro and in vivo models. 7 Furthermore, due to its rather specific expression in several solid tumors, CAIX is investigated as an address for specific delivery of cytotoxic drugs.8,9
Novel CAIX inhibitors are needed to enable clinical exploitation of CAIX, but current assays for activity/inhibition testing of CAs are laborious and time- and enzyme-consuming. 8 Currently, all inhibitors of CAs investigated for potential clinical use are derivatives of sulfonamides. Use of sulfonamide derivatives for clinical CAIX targeting is hindered by several obstacles. First, they have unfavorable pharmacokinetic properties, mainly due to high nonspecific adsorption to serum proteins. 10 Second, it has been estimated that sulfonamide administration leads to allergic reactions in approximately 3%–6% of the population, which can evolve in a life-threatening state.11,12 Finally, sulfonamides act nonselectively by binding to zinc ions contained in the active site of all human CAs and therefore show poor selectivity across the CA family. 13 Given these issues, the identification of a new structural class of inhibitors could represent a breakthrough in the design of selective CAIX inhibitors and enable significant advances in both primary and translational research employing CAIX inhibitors as an anticancer therapy target. Even though methods like isothermal titration calorimetry (ITC) and fluorescent thermal shift assay (FTSA) were used for the assessment of large numbers of CAs inhibitors, they lack sufficient throughput and are quite demanding on sample consumption to represent effective platform for high-throughput screening (HTS) of CAIX inhibitors. 14
The DNA-linked inhibitor antibody assay (DIANA) has been recently developed for ultrasensitive enzyme detection and inhibitor screening. 15 In DIANA, the target protein is captured by an immobilized antibody and subsequently detected by a fully synthetic detection probe, which comprises a small-molecule ligand (i.e., competitive inhibitor) of a target enzyme covalently linked to a DNA oligonucleotide.
On top of the sensitive detection of protein targets, DIANA can also be used for the screening of enzyme inhibitors. The inhibition potency of the test compounds is determined by their ability to outcompete binding of the probe to the enzyme. DIANA has several unique advantages over state-of-the-art screening technologies, such as (1) a wide dynamic range enabling the determination of the inhibition potency (Ki value) from a single well measurement, (2) an extremely high signal-to-noise ratio, and (3) the ability to screen inhibitors with minuscule amounts of unpurified enzymes. The robustness of DIANA can be proven by Z′ values, which are commonly in the range of 0.85–0.90. 15 The initially reported DIANA protocol was performed manually in a 96-well plate format and was not suitable for high throughput. However, adapting DIANA to automated 384-well format would represent significant improvement of its HTS capability. An assay such as DIANA would address several main drawbacks of current inhibitor screening assays, such as low accuracy, narrow dynamic range, and only qualitative output.
The IOCB library represents a unique proprietary well-curated collection of small molecules, which were synthetized at the IOCB of the Czech Academy of Sciences. It contains various unique chemical scaffolds typically not present in commercial libraries (i.e., natural compounds, steroids, peptides and peptidomimetics, and nucleosides and nucleotides), with the main emphasis on nucleoside and nucleotide analogs. Importantly, most compounds present within the library were prepared as a result of medicinal chemistry efforts to develop potential drugs against various pharmaceutically relevant targets, which ensures their drug-likeness. Indeed, there are already several FDA-approved drugs originating from this library, such as tenofovir, one of the most widely used inhibitors of HIV reverse transcriptase; adefovir, a potent virostatic used in the treatment of hepatitis B infection; and cidofovir, an effective anticytomegalovirus compound. 16 The library currently contains more than 3000 compounds and it is still growing, as more compounds are continuously prepared at IOCB. Considering all the abovementioned properties, the IOCB library represents a very useful compound collection for the screening of pharmaceutically relevant enzymes such as CAIX.
Here, we present and describe the identification of novel CAIX inhibitors using DIANA adapted to HTS format combining several strategies for an increased throughput: (1) reformatting the assay from 96- to 384-well plate format, (2) substituting all manual pipetting steps by liquid handling platforms, and (3) pooling of test compounds for the screen. We used DIANA for screening more than 2800 compounds from the IOCB library, and thoroughly validated the obtained hits from the assay in terms of their purity, inhibition activity, and mode of binding toward CAIX using x-ray crystallography.
Materials and Methods
HTS of IOCB Library by DIANA against CAIX
The general DIANA protocol for CAIX HTS was performed as previously described with minor modifications. 15 Briefly, 25 ng of M75 antibody was coated on a FrameStar 384 Well Skirted PCR Plate (4ti-0381; 4titude Ltd., Surrey, UK) in Tris-buffered saline (TBS) followed by addition of casein blocking agent diluted fivefold in TBS (CBC1; SDT, Baesweiler, Germany). After washing, 5 µg of HT-29 cell lysate, which endogenously expresses CAIX protein, was added to the wells, incubated, and washed away. The bivalent detection probe CAIX probe 2 (structure published previously 15 ) was added to the wells together with the test compounds. If not stated otherwise, the final probe concentration was 500 pM and the concentration of test compound was 10 µM (during pooled testing each compound in the well was 9.1 µM so the final concentration of all test compounds within one well was 100 µM). After additional washes, the quantitative PCR (qPCR) mixture was added to the wells and the signal was detected using a LightCycler 480 II qPCR instrument (Roche, Basel, Switzerland). Threshold cycles (Cq) were obtained from the measured fluorescence curves using the method of maxima of the second derivative in the LightCycler 480 II Software (Roche). The acquired data were then analyzed and Ki values were calculated based on the previously described equations. 15
For the washes between all the steps TBS + 0.05% Tween-20 using BlueWasher (BlueCatBio, Neudrossenfeld, Germany) was used. During all experiments using the 384-well plate format, the last two columns of the 384-well plate were used for control samples that contained dilution series (3–10,000 nM) of the known potent CAIX inhibitor acetazolamide (AZA) in duplicates and wells without any inhibitor in at least octaplicate. These controls served as intraplate and interplate controls throughout the screens and enabled the determination of the assay window and calculation of Z′ factors.
For all DIANA measurements, the Ki values of test compounds were calculated using the equation for bivalent probe:
where Raff is defined as a ratio of dissociation constants of bivalent to monovalent probes toward CAIX, which is equal to 0.022; dCq is the difference between Cq values of wells with and without test compound; [I] is the working concentration of the test compound (variable); and [P] and KD(P) are the working concentration (500 pM) and dissociation constant (1.5 nM) of the bivalent probe, respectively. Detailed derivation of the equation is described in the supplementary information of Navrátil et al., 15 where it is listed as eq 15.
Ki Determination of Screen Hits by DIANA
To validate results from HTS, the Ki values for identified hits were determined again by DIANA in the 96-well plate format. The protocol was identical to HTS, except each test compound was measured in four wells with different concentrations of test compound ranging from 50 µM to 50 nM. The acquired data were then analyzed and Ki values were calculated based on the previously described equations. 15
CA Inhibition Assay
A stop-flow instrument (Applied Photophysics, Surrey, UK) was used to measure the CA-catalyzed CO2 hydration activity in the presence of inhibitors. 17 The assay was performed as described previously with minor modification. 18 The CA concentrations in the reaction mixture ranged from 5 to 19 nM. The apparent inhibition constants were obtained by nonlinear least-squares methods in GraphPad Prism version 8.0.0 for Windows (GraphPad Software, San Diego, CA; www.graphpad.com), which explore nonlinear fitting of the Williams–Morrison equation: 19
The Ki values were derived from the apparent Ki using the Cheng–Prusoff equation: 19
FBSA Quantifi cation by LC-MS
All hits from the DIANA screen (compounds
Results
Adapting DIANA into HTS Format
To enable screening of large compound libraries via DIANA, we first needed to adapt the assay to the 384-well format and automate all liquid handling steps. A detailed scheme of the final DIANA protocol is described in the Materials and Methods section and summarized in Supplemental Table S1 . Briefly, the developed protocol consists of several incubation and washing steps, and its total duration is less than 3 h, when starting with precoated and blocked plates. The rate-limiting step in the protocol is qPCR, which takes less than 30 min. As a result, 16 plates per 8 h can be run on a single qPCR instrument resulting in a throughput of 5632 compounds per day having 352 compounds per plate.
Screen of the IOCB Library against CAIX Using DIANA
To evaluate DIANA screening capabilities, we performed a screen of 2816 compounds from the IOCB library against CAIX using DIANA, that is, eight 384-well plates with each plate comprising 352 wells with test compounds and 32 wells with controls. The screen revealed 13 hits and high signal-to-noise ratios expressed by the Z′ factor ranging between 0.91 and 0.95 for each of the eight plates. The results from the screen are summarized in Figure 1A .
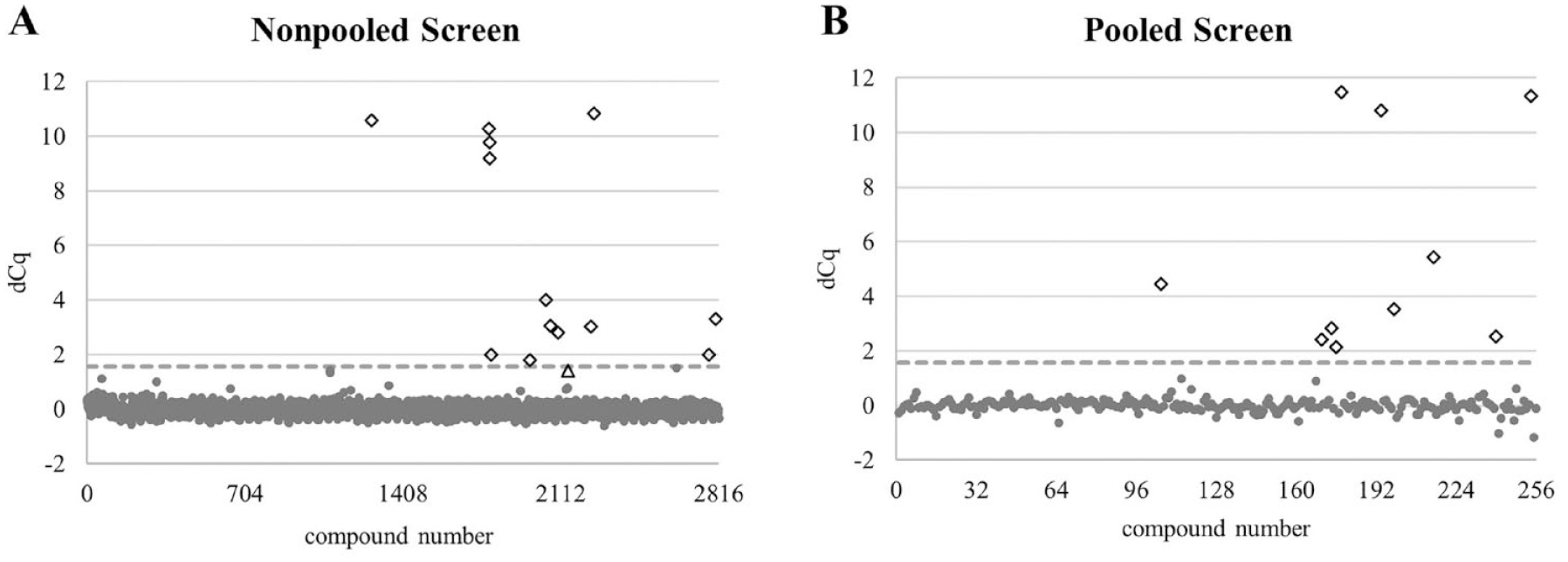
Results of nonpooled and pooled DIANA screen of IOCB library against CAIX. Each test compound is represented by a dot and the shown dCq values are calculated as Cq of the well without any test compound subtracted from Cq of the well with compound (Cqcpd – Cqblank). The empirical threshold (gray dashed line) shows the dCq value corresponding to 43% inhibition, that is, 10 µM Ki value (dCq = 1.57). Compounds with dCq higher than 1.57 (Ki < 10 µM) are shown as diamonds, and compounds below this threshold are shown as gray circles. The compound shown as a triangle (dCq = 1.42) represents a compound that was identified as a hit in the pooled screen but not in the nonpooled screen. DIANA was performed as described in Materials and Methods. Briefly, CAIX was captured on a 384-well plate via the M75 antibody. Subsequently, each test compound (
We set the cutoff threshold to dCq 1.57, which corresponds to approximately 43% inhibition of CAIX under the assay conditions. In this way, we identified 13 positive hits (diamonds in
Given the very low chance of obtaining false-positive hits in DIANA (see Discussion for more information), we also investigated the ability of DIANA to screen pooled libraries. We pooled 11 compounds from the IOCB library per well and thus limited the number of tested wells from 2816 to 256. As shown in Figure 1B , we identified 10 positive wells from the screen using identical cutoff criteria as for the nonpooled screen. The Z′ factor for the pooled screen was 0.89. The general characteristics of both the nonpooled and pooled DIANA screens are summarized in Table 1 .
General Characteristics of DIANA CAIX Screen.
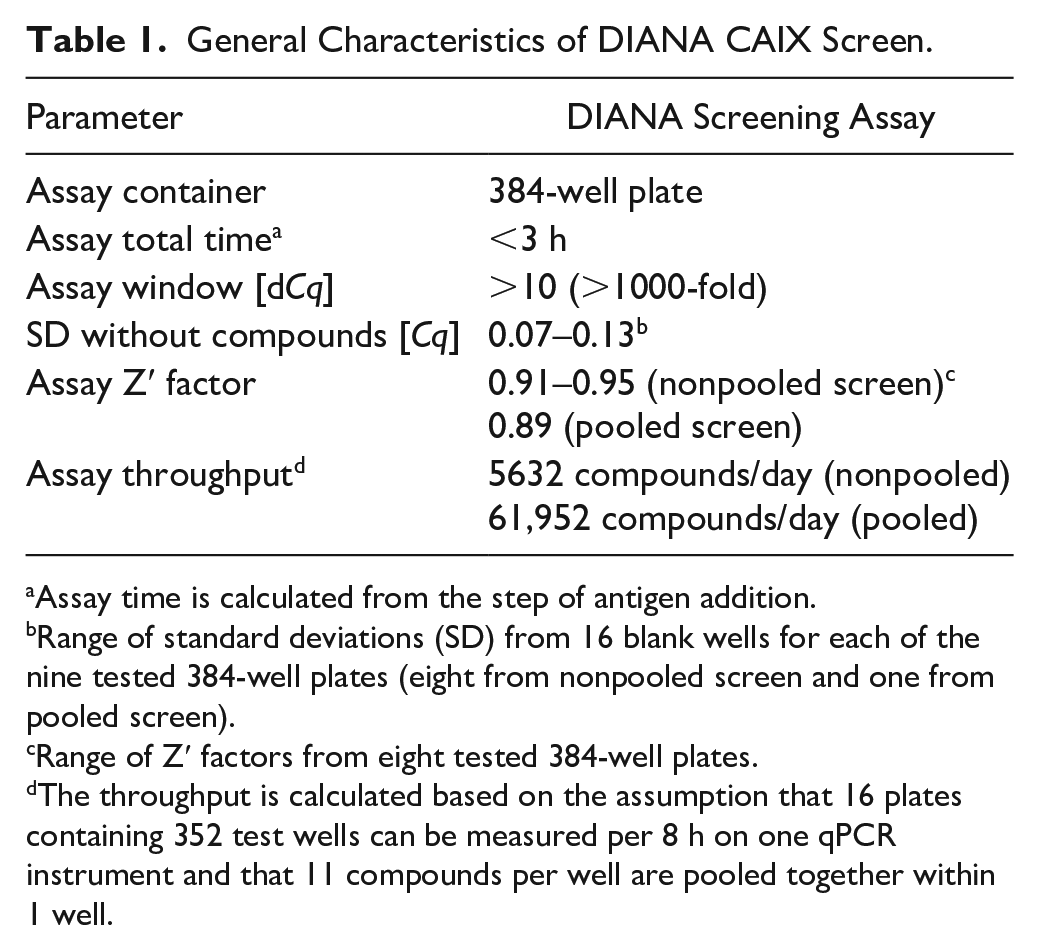
Assay time is calculated from the step of antigen addition.
Range of standard deviations (SD) from 16 blank wells for each of the nine tested 384-well plates (eight from nonpooled screen and one from pooled screen).
Range of Z′ factors from eight tested 384-well plates.
The throughput is calculated based on the assumption that 16 plates containing 352 test wells can be measured per 8 h on one qPCR instrument and that 11 compounds per well are pooled together within 1 well.
Comparison of Nonpooled and Pooled DIANA Screens
A closer look at the identified hits from both nonpooled and pooled DIANA screens revealed that all 13 hits identified during the nonpooled screen were also identified by the pooled screen with two pool wells (nos. 194 and 215) containing more than one hit. The determined dCq values for each hit are summarized in
Table 2
, with each row comprising one well from the pooled screen and all identified hits from the nonpooled screen within that well. Additionally, one positive well in the pooled screen (no. 170) was found that did not contain a positive hit from the nonpooled screen. However, examining the signal of 11 compounds present within the pooled well (no. 170), there was one compound (no. 2143) that provided a signal very close to the cutoff threshold (highlighted as triangle in
Summary of Hits Identified during Nonpooled and Pooled DIANA Screen against CAIX.
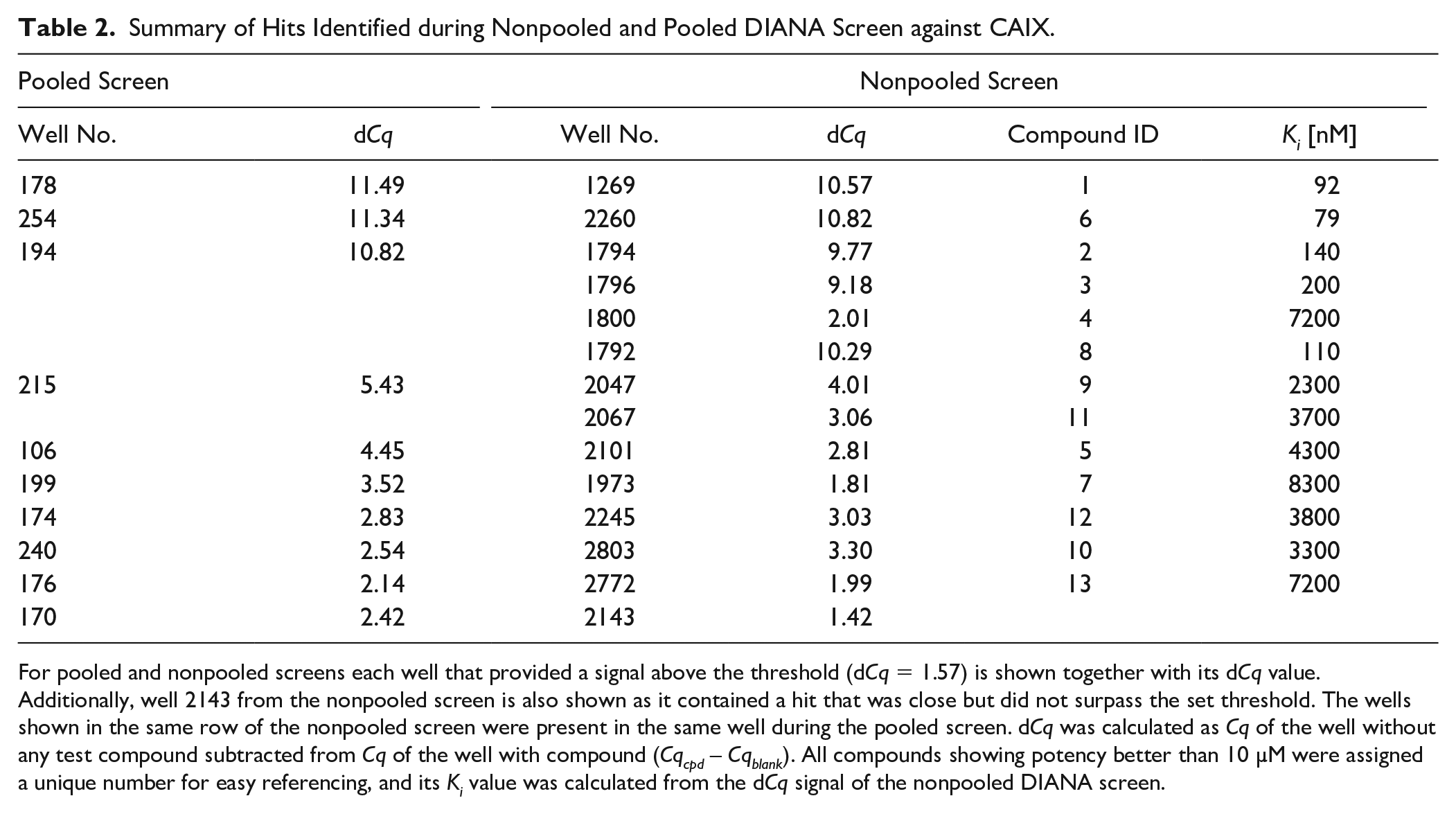
For pooled and nonpooled screens each well that provided a signal above the threshold (dCq = 1.57) is shown together with its dCq value. Additionally, well 2143 from the nonpooled screen is also shown as it contained a hit that was close but did not surpass the set threshold. The wells shown in the same row of the nonpooled screen were present in the same well during the pooled screen. dCq was calculated as Cq of the well without any test compound subtracted from Cq of the well with compound (Cqcpd – Cqblank). All compounds showing potency better than 10 µM were assigned a unique number for easy referencing, and its Ki value was calculated from the dCq signal of the nonpooled DIANA screen.
Overall, the comparison of these two datasets confirmed that the pooled DIANA screen maintains its robustness (high Z′ factor) and that it has identified the identical set of hits as the nonpooled screen.
Determination of Ki Value from Single Well Measurement
One of the key features of DIANA is the fact that it is a quantitative method; that is, it enables determination of hit Ki values from a single well measurement (see Table 2 ). In this particular case, in which 10 µM compounds were tested, the assay window enabled precise determination of Ki values in the range of two orders of magnitude between 100 nM and 10 µM; this range depends on several factors, such as assay window, valency of the used probe, and test compound working concentration. 15 Given these assay features for the CAIX DIANA, all Ki values close to 100 nM could be in theory underestimated and the actual inhibition potency of these compounds may have been better.
To confirm this assumption, we performed DIANA at four different concentrations of compounds
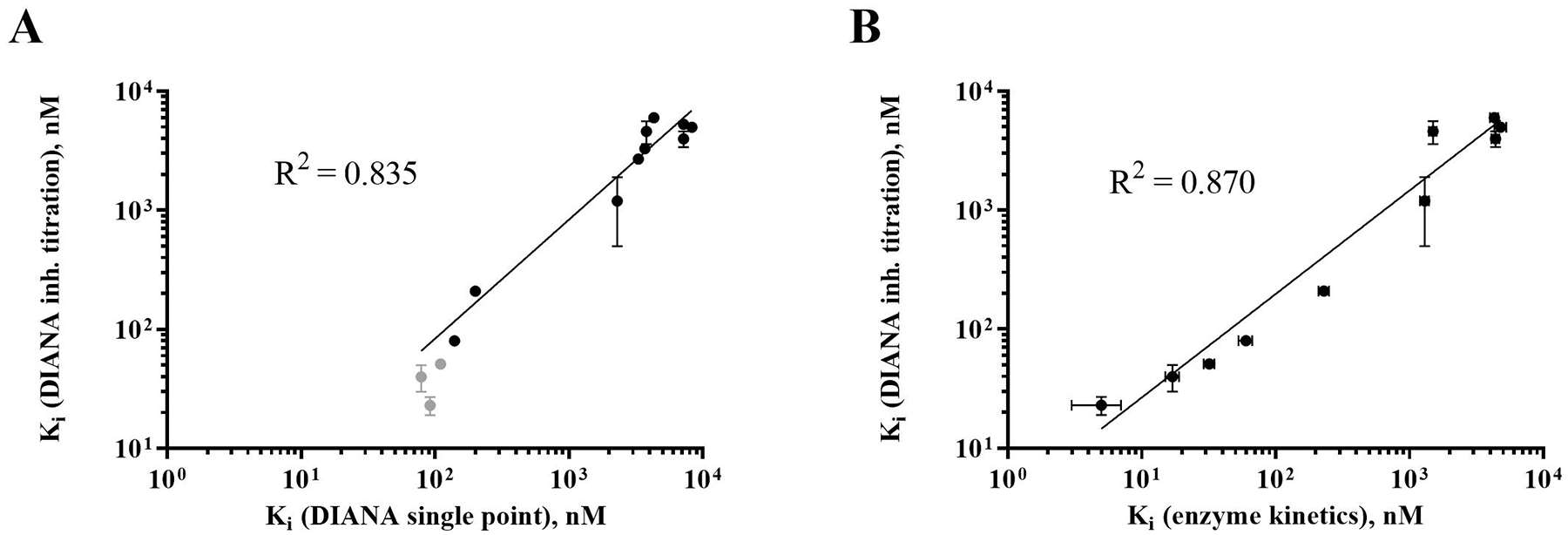
Comparison of determined Ki values of identified inhibitors against CAIX. (
To further validate the precision of Ki value determination of DIANA, we determined the Ki values of the 10 most potent hits by an established orthogonal enzymatic inhibition assay, which is based on the measurement of pH changes of CA-catalyzed CO2 hydration using a spectrophotometric readout. As shown in Figure 2B , we observed a good correlation of Ki values obtained by DIANA (DIANA inh. titration) and enzymatic inhibition assay (enzyme kinetics; R2 = 0.870), confirming the robustness and specificity of DIANA (see raw data in Suppl. Table S2 ).
Validation of the Hits from DIANA Screen against CAIX
After identification of the 13 hits by DIANA screen, we performed a thorough analysis of these compounds by liquid chromatography–mass spectrometry (LC-MS) to confirm their chemical purity and homogeneity. The LC-MS analysis provided several important findings. First, we found that compounds
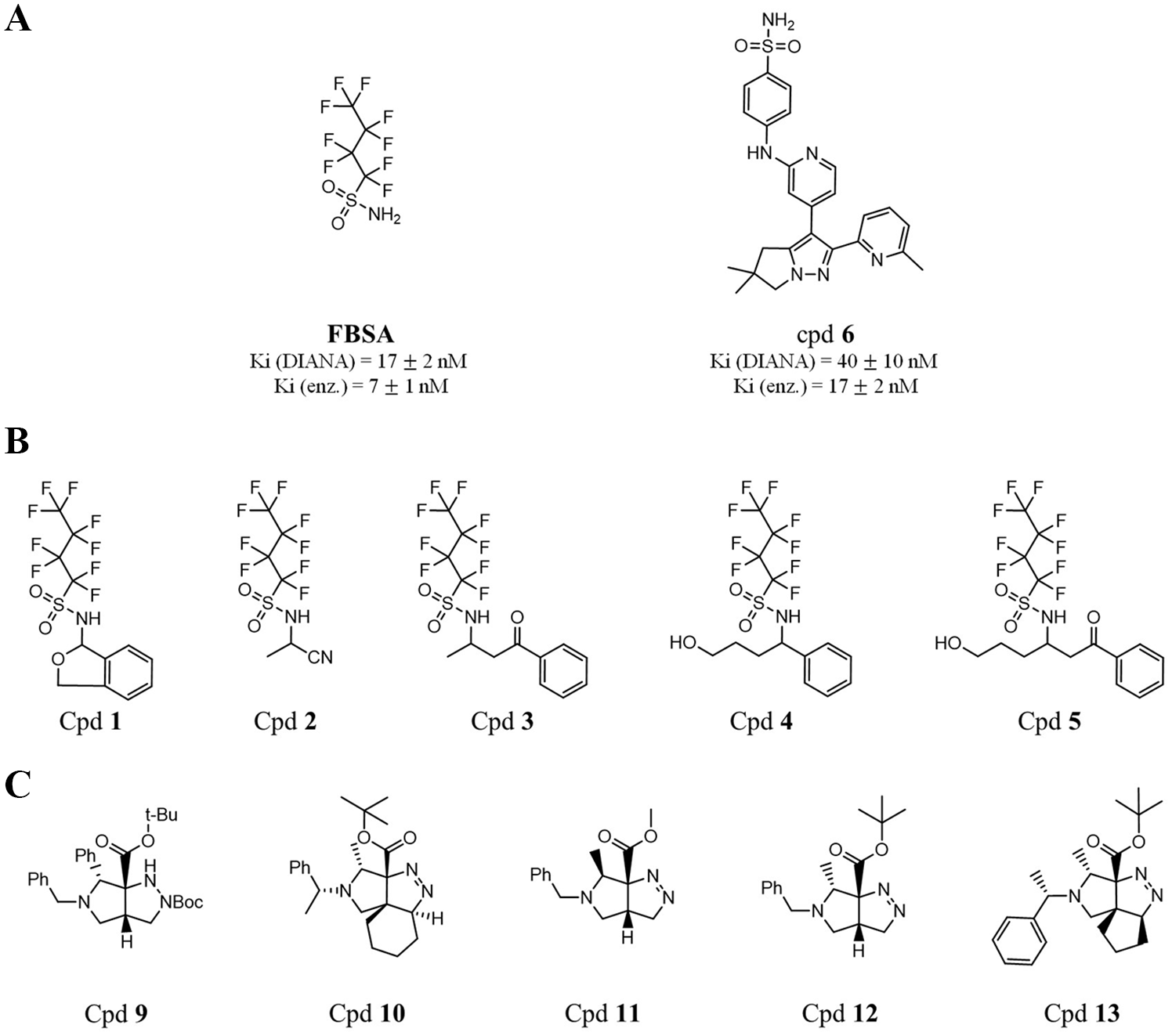
Chemical structures of identified hits against CAIX by DIANA. (
To confirm the presence of FBSA in the analyzed compounds and to find out whether the observed inhibition activity against CAIX is attributed to the actual compounds or FBSA, we synthesized FBSA, analyzed it by LC-MS, and determined its inhibition potency against CAIX. Indeed, the chromatographic and mass spectral data of FBSA confirmed its presence in compounds
To conclude, the number of hits identified via the DIANA screen was upon validation reduced from 13 to 3 unique chemical entities, namely, FBSA, compound
Binding Modes of Novel CAIX Inhibitors
To elucidate the binding mode of the identified compounds, we co-crystallized FBSA and compound
The high quality of electron density maps allowed for unambiguous modeling of compounds in the CAIX active site (see
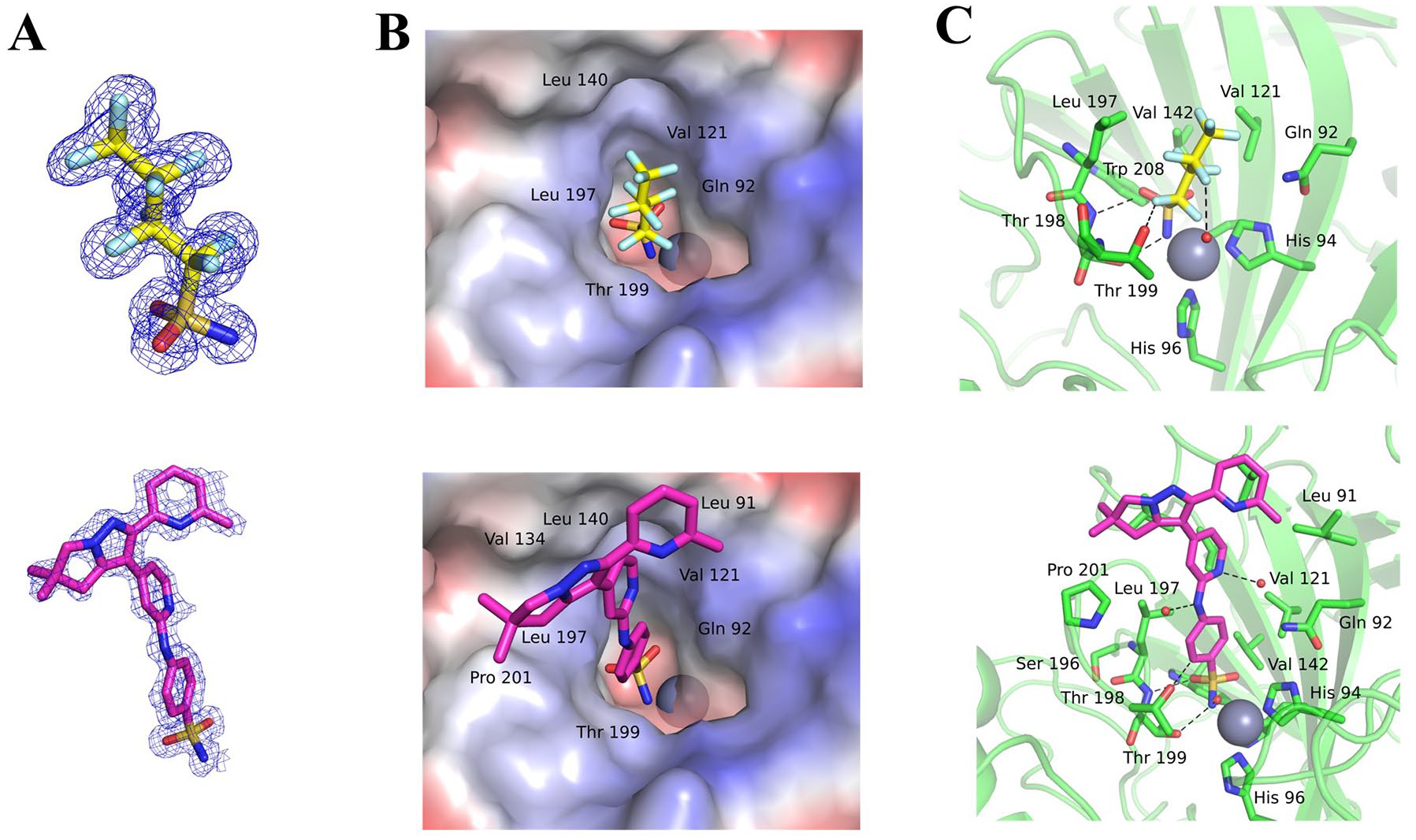
Structures of FBSA and compound
Furthermore, FBSA forms an additional 29 interatomic contacts with Gln92, Val121, Leu140, Val142, Leu197, Thr198, Thr199, and Trp208, thus filling the bottom of the active site cavity. In contrast, much bulkier compound
Discussion
In this work, we present the development of a high-throughput screen of CAIX inhibitors from a compound library and structural characterization of their binding mode to the enzyme. For the screening, we adapted the recently published DIANA method and used the proprietary in-house compound library prepared at IOCB Prague.
DIANA Development and Validation
First, we analyzed if the HTS format does not compromise DIANA’s advantageous features. Compared with the original DIANA, the current assay setup was miniaturized to the automated 384-well plate format while it maintained an identical assay window and consumption of target enzyme (approximately 400 pg of CAIX present in 5 µg of the HT-29 cell lysate), thus confirming that the assay keeps its minimal requirements on target amount and purity. Furthermore, implementation of the liquid handling platforms minimizes human errors and increases assay reproducibility. This fact together with the large assay window contributed to excellent Z′ factors (>0.89) for the assay in all conducted experiments, demonstrating its overall robustness.
Additional advantageous features of DIANA are its low false-positive and false-negative rates. While the determination of false positives of the assay is straightforward, using orthogonal inhibition assays after the initial screen, the determination of the false negatives is challenging since these compounds are not identified during the initial screen as hits. However, during our validation of the pooled DIANA screen, we collected two independent datasets (from pooled and nonpooled screen) that enabled us to at least estimate the false-negative rates of the assay to be very low. As summarized in
Table 2
, we identified all the hits from the nonpooled screen that were also in the pooled screen. Moreover, we identified one extra positive well in the pooled screen that did not contain a hit from the individual screen. However, this pooled well contained one compound that provided a dCq signal of 1.42, which is barely below the set threshold (dCq = 1.57). Regarding the false-positive rate of the assay, we did not identify a single compound that would be positive in DIANA and not show the inhibition activity in the orthogonal inhibition assay (
As stated in the Results section, we intentionally set the cutoff threshold in the DIANA screen to a dCq value of 1.57, which corresponds to the Ki value of the compound concentration in the screen (10 µM). Given the very low standard deviation (SD) of DIANA and the large assay window, our cutoff threshold was much more stringent (eight times SD above blank wells without inhibitors) than the generally accepted statistical threshold (three to five times SD above blank). This assay feature ensures a very low probability of obtaining false-positive hits; in theory, the statistical chance of false-positive hit occurrence assuming normal data distribution would be lower than 10–14. If the threshold would be calculated as a signal corresponding to blank measurement with five times of its SD added (dCq = 0.92), it would correspond to only 28% inhibition of CAIX under the assay condition and still provide a stochastic false-positive rate lower than 0.001%. Such a threshold would enable us to identify compounds with a Ki value lower than 20 µM while using only the 10 µM compound concentration during the screen.
Yet another beneficial feature of DIANA is its ability to calculate the compound inhibition potency from a single well measurement and thus rank and quantify the inhibition potency of the hits right after the initial screen. This is enabled by the sensitivity of the detection and by the possibility to accurately determine inhibition in a wide range. Since qPCR has a dynamic range of up to nine orders of magnitude and it is a logarithmic readout measuring the fold change rather than absolute change, and at the same time DIANA is detecting the amount of free enzyme unoccupied by the inhibitor, virtually any inhibition above 25% can be measured with high precision. Specifically, the relative coefficient of variation of determined inhibition is usually lower than 10% of the remaining free enzyme; for example, 90% inhibition is determined with an error of less than 1%, while 99.9% inhibition has an error of less than 0.01%. These properties translate into the ability of DIANA to determine Ki values from a single well measurement.
To confirm that the results obtained during the DIANA screen at one compound concentration can be used for Ki determination, we compared the Ki values from the nonpooled screen with Ki values from both DIANA run at four different compound concentrations and an established orthogonal CAIX enzymatic inhibition assay. As shown in Figure 2 , data from both assays show very good correlation with data obtained from the DIANA screen apart from compounds that had high inhibition potencies (Ki < 100 nM). This observation can be explained by the fact that under the given assay conditions (such as the assay window, the concentration of test compound, and usage of bivalent probe), the range of Ki values that can be reliably determined in the presented CAIX DIANA screen at a 10 µM concentration of the test compounds is 100 nM to 10 µM. The compounds with a Ki lower than 100 nM are able to outcompete almost all DIANA probes, and thus all of them appear as 100 nM inhibitors, while the compounds with Ki higher than 10 µM are unable to outcompete a sufficient amount of probe to provide a reliable signal change compared with the signal from the noninhibited reaction. This phenomenon can be demonstrated on the known potent CAIX inhibitor AZA that was used as a control for all DIANA experiments. From the DIANA screens the Ki value of AZA calculated from the 10 µM concentration was 99 ± 10 nM (average of 10 measurements with SD; data not shown), while if calculated from serial dilutions within the DIANA range the Ki value was 7 ± 1 nM (the value is an average from eight measurements with SD; data not shown), a value that is in agreement with previously reported data. 13 Nevertheless, this feature does not represent a significant disadvantage of the assay, since all highly potent compounds will still be ranked as the most potent hits within the screen and there is no danger that they would be omitted from further analysis. The only limitation will lie in the fact that within this group of potent hits the ranking will be less precise.
The range of inhibitory potencies that can be accurately determined in DIANA is very broad thanks to the possibility to calculate a Ki value from a single point measurement. The lowest detectable potency (highest Ki) approximately equals the highest tested inhibitor concentration. The highest accurately measurable potency (lowest Ki) is limited by the assay window and by the amount of enzyme used for screening, as the inhibitor should not be assayed below the enzyme concentration. In this assay using endogenous CAIX, it is possible to calculate Ki values in the range of 0.01-fold to 1-fold of the assayed inhibitor concentration; that is, the lowest measurable Ki is ~100-fold below the used enzyme concentration. Specifically, CAIX concentration in the assay is approximately 2 nM, which translates into the lowest precisely measurable Ki of 20 pM, while inhibitors with lower Ki would appear as 20 pM as well. A higher assay window, which can be gained by using recombinant and not endogenous CAIX, would enable precise measurement of even lower Ki values. As shown for prostate-specific membrane antigen, the DIANA window can be as high as six orders of magnitude, resulting in the lowest measurable Ki value of 1 fM. 15
In DIANA, any compound capable of outcompeting the probe out of the active site is identified as a hit, and thus any competitive inhibitor should be identified. Even though the probe binds to the active site, the inhibitor does not necessarily need to bind to the same site. For example, allosteric inhibitors bind to the exosites and cause structural changes within the enzyme active site and thus prevent the substrate/probe binding. Indeed, such an allosteric inhibitor has been identified during an internal DIANA screen against another enzyme (prostate-specific membrane antigen) at our institute and its allosteric mode of action was confirmed by x-ray crystallography (unpublished data). However, if the enzyme contained two completely independent binding sites, such as cofactor and peptide binding site, and inhibitors of both sites were of interest, then the screening could be run with two probes, each targeting different binding sites. Even though this is associated with extra effort to develop two probes, the extra time and cost associated with such a screen could be minimized by multiplex qPCR, and thus screening both probes in the same well. Use of multiple probes would bring the possibility to decipher the binding site of the hits.
Even though only a small compound library (~3000 compounds) was used for validation of DIANA, the performed analysis confirms that the assay is highly robust with very low false-positive and false-negative rates. Notably, the presented pooling approach significantly reduces the time of the screen as it first tests mixtures of compounds in the initial pooled screen, followed by deconvolution testing of only positive pooled wells. The number of compounds pooled together must be appropriately chosen taking into the account the expected positive hit rate; for the majority of random libraries the expected positive hit rate ranges from 0.2% to 0.5 %. Additionally, it is important to have a reasonable estimate of the false-positive rate of the used screening assay; since we did not identify a single false-positive hit, this rate will be <0.04% for DIANA. Based on the presented data, we deduce that the ideal number of pooled compounds will be in the range of 10–20 compounds per well. In line with these findings, we have pooled 11 compounds per well. Considering the expected ultra-low false-positive rate of DIANA, this number could be even bigger for difficult targets with low hit rates.
Finally, it should be noted that DIANA’s throughput is in the range of 50,000 to 100,000 test compounds per day on one qPCR instrument when 10–20 compounds are pooled per well. Since the current bottleneck of the assay is the qPCR readout, the utilization of more than one qPCR instrument enables further upscaling of the method to significantly more than 100,000 test compounds per day.
Hit Validation and Characterization
From the initial screen we identified 13 hits with inhibition potency below 10 µM. During assay validation we repeatedly determined the inhibition potency of the hits not only from solutions stored within the library but also from freshly prepared solutions of the stored powders. During this analysis we identified an unexpected discrepancy for compounds
This finding would suggest that FBSA is responsible for the inhibition activity of compounds
We identified three novel inhibitors (FBSA and compounds
Since sulfonamide derivatives are generally nonspecific CA inhibitors, it can be expected that the identified inhibitors will also inhibit other CA isoforms. Considering the extensive interatomic interaction of compound
In conclusion, we present data from a screen of the IOCB medicinal compound library against CAIX using the novel HTS method DIANA. We showed that DIANA represents a very robust and efficient HTS platform for the screening of enzyme inhibitors that requires a minimal amount of nonpurified target enzyme, has negligible false-positive and false-negative rates, and enables screening in compound mixtures. Moreover, DIANA can be easily adapted to various enzymatic and nonenzymatic (e.g., protein–protein interactions) targets. From the screen, we identified three novel inhibitors of CAIX within the IOCB library. They are all sulfonamide-based compounds, and thus we have not identified any structurally unexplored scaffolds within the IOCB library. Nevertheless, the presented DIANA is the first assay suitable for HTS of CAIX inhibitors, and its use on larger libraries may lead to the identification of completely novel scaffolds in the future.
Supplemental Material
DISC918836-Supplement – Supplemental material for Identification of Novel Carbonic Anhydrase IX Inhibitors Using High-Throughput Screening of Pooled Compound Libraries by DNA-Linked Inhibitor Antibody Assay (DIANA)
Supplemental material, DISC918836-Supplement for Identification of Novel Carbonic Anhydrase IX Inhibitors Using High-Throughput Screening of Pooled Compound Libraries by DNA-Linked Inhibitor Antibody Assay (DIANA) by Jan Tykvart, Václav Navrátil, Michael Kugler, Pavel Šácha, Jiří Schimer, Anna Hlaváčková, Lukáš Tenora, Jitka Zemanová, Milan Dejmek, Vlastimil Král, Milan Potáček, Pavel Majer, Ullrich Jahn, Jiří Brynda, Pavlína Řezáčová and Jan Konvalinka in SLAS Discovery
Footnotes
Acknowledgements
The use of the beamline MX14.1 operated by the Helmholtz-Zentrum Berlin at the BESSY II electron storage ring (Berlin-Adlershof, Germany) to collect diffraction data is gratefully acknowledged. We would also like to thank Milan Fábry for preparation of expression plasmid for the CAIX mimic and Radko Souček for his excellent assistance with the quantification of FBSA by LC-MS.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Patent applications describing the DIANA technological platform have been filed. V.N., J.S., J.K., and P.Š. are co-owners of a company commercializing the technology.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by the Grant Agency of the Czech Republic, projects 19-10280S and 17-14510S; in part by the European Regional Development Fund, OP RDE (no. CZ.02.1.01/0.0/0.0/16_019/0000729); and in part by the Ministry of Education of the Czech Republic (program “NPU I”), project LO1304.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.