Abstract
Dysfunction of apoptosis and DNA damage response pathways often drive cancer, and so a better understanding of these pathways can contribute to new cancer therapeutic strategies. Diverse discovery approaches have identified many apoptosis regulators, DNA damage response, and DNA damage repair proteins; however, many of these approaches rely on indirect detection of DNA damage. Here, we describe a novel discovery platform based on the comet assay that leverages previous technical advances in assay precision by incorporating high-throughput robotics. The high-throughput screening (HTS) CometChip is the first high-throughput-compatible assay that can directly detect physical damage in DNA. We focused on DNA double-strand breaks (DSBs) and utilized our HTS CometChip technology to perform a first-of-its-kind screen using an shRNA library targeting 2564 cancer-relevant genes. Conditions of the assay enable detection of DNA fragmentation from both exogenous (ionizing radiation) and endogenous (apoptosis) sources. Using this approach, we identified LATS2 as a novel DNA repair factor as well as a modulator of apoptosis. We conclude that the HTS CometChip is an effective assay for HTS to identify modulators of physical DNA damage and repair.
Keywords
Introduction
Maintenance of DNA integrity is crucial for high-fidelity transmission of genetic information to progeny cells. Threats to DNA integrity can be cell-extrinsic (e.g., ionizing radiation, chemicals) and/or cell-intrinsic (e.g., metabolites, apoptosis induced). DNA damage can also interfere with critical cellular processes and lead to cancer-causing changes in the genome.1,2 DNA double-strand breaks (DSBs) in particular can have severe consequences as part of the apoptotic program, by causing genomic alterations that cause loss or dysfunction of genetic information, or by triggering cell death via DNA damage signaling. 3 DNA repair pathways safeguard the genome against the deleterious effects of DNA DSBs, and dysregulated DNA repair is often observed in pathological conditions such as cancer. 2 Likewise, apoptosis is highly regulated in normal cells, and dysregulation is a hallmark of cancer. 2 While defective DNA repair can destabilize the genome, thus promoting cancer, these same defects can render cancer cells vulnerable to DNA damage in the form of radio- or chemotherapy. Therefore, a better understanding of DNA repair and apoptosis pathways can lead to improved cancer therapy. Here, we describe an advance in the throughput of physical DNA DSB detection using a platform that is designed to be compatible with high-throughput screening (HTS) technologies. We leveraged this novel approach for gene discovery to further our understanding of DNA DSB repair and apoptotic pathways.
DSBs are repaired via two major pathways in human cells, namely, homologous recombination (HR) and nonhomologous end joining (NHEJ). 4 In the HR pathway, the broken DNA strands are end resected to create 3′ overhangs that invade homologous DNA on sister chromatids to copy genetic information for repairing the break. 5 Due to the requirement of sister chromatids as a source of template DNA for accurate repair, the HR pathway is generally most active during the S and G2 phases of the cell cycle. 6 In contrast, NHEJ does not require a homologous template, and therefore is active throughout the cell cycle. 7 For NHEJ, broken ends of DNA strands are protected from end resection by the Ku70/Ku80 heterodimer, effectively inhibiting HR. 8 DNAPKcs then binds to the Ku70/Ku80 heterodimers at the DNA DSB to form the DNAPK holoenzyme. 9 Upon activation, DNAPK is able to phosphorylate itself, as well as many downstream NHEJ factors, including the Artemis protein, 10 which is one of the major end processing endonucleases recruited to DSBs with nonligatable ends. Other end processing enzymes recruited to DSBs include PNKP, 11 which removes phosphate groups from 3′ ends and phosphorylates hydroxyl groups on 5′ ends, as well as DNA polymerases Pol µ and λ. 12 The broken DNA ends are then ligated together by a complex composed of XRCC4, XLF, PAXX, and DNA ligase IV.13,14 These key components of the HR and NHEJ pathways are critical for efficient DSB repair, and loss of function of these genes leads to persistent DSB following DNA damage.
Given that genes that modulate DNA repair could be useful targets for novel cancer therapeutics, many laboratories have screened for genes directly involved in DSB repair and for genes that indirectly impact DSB repair (e.g., genes involved in chromatin remodeling or cell signaling). For example, proximity ligation approaches coupled to mass spectrometry proteomics have revealed entire networks of proteins that associate at sites of DNA damage. 15 In addition, researchers have employed knockdown or knockout libraries to screen thousands of genes for their ability to modulate DNA damage responses. Such responses have included co-localization of proteins involved in DNA DSB responses (e.g., γH2AX, 53BP1, and Rad52), as well as synthetic lethality (e.g., sensitization of BRCA2 mutant cells). Using these methods, networks of interacting genes that modulate DNA DSB levels have been revealed, as well as specific genes that participate in DNA DSB recognition and repair.16,17 These studies have shown the utility of large-scale genomic screening to identify novel DNA repair genes that could lead to novel strategies for treating cancer. However, none of these large-scale studies exploit methods to directly measure the levels of physical DNA DSBs in cells. For example, downstream signaling events (such as phosphorylation of H2AX) or changes in protein co-localization are associated with, but do not occur exclusively in the presence of, DNA DSBs. In addition, cell survival and synthetic lethality assays rely on lethal interactions that may or may not be due to changes in DSB repair. We therefore set out to perform a large-scale RNAi screen using detection of physical DNA DSBs as an approach to identify previously unknown genes that impact DNA damage formation and repair.
The comet assay is a well-established assay for DNA DSBs 18 directly induced by DNA-damaging agents (e.g., ionizing radiation) or indirectly induced as a result of enzymatic DNA fragmentation that occurs during apoptosis.19,20 The comet assay technique, first described by Ostling and Johanson in 1984 and further developed by many others,21–24 is based on the principle that damaged DNA has greater electrophoretic mobility in an agarose gel compared with intact DNA, resulting in a comet-shaped distribution of DNA within the gel (where a nucleoid forms the head and damaged DNA forms the tail). Traditionally, the assay requires spotting a molten agarose cell suspension onto a glass slide with one condition per slide, and therefore it is extremely low throughput and tedious to perform. Users also report high variation in the results obtained from the assay, preventing its usage in large-scale screens. To address the limitations of the comet assay, Andrew Collins’s group25 and Trevigen, Inc. (Gaithersburg, MD) created slides with subregions, which greatly improved throughput and consistency, but the assay is performed using standard imaging and analysis, which can be quite slow and inconsistent among experimentalists. Wood and coworkers developed the CometChip,26–28 which also subdivides a glass slide, in this case with 96 regions in the conformation of a 96-well plate. The CometChip also exploits cell micropatterning, which maximizes the number of useful comets per unit area, prevents overlapping comets, creates a uniform focal plane (enabling the use of only one or two images to capture ~100 comets, rather than one image per comet), and enables automated imaging and analysis (increasing throughput by orders of magnitude). To create a cell array, microfabrication techniques are exploited to create a stamp that can be applied to molten agarose to create an agarose slab with an array of cell-sized microwells. When a cell suspension is placed over the agarose, cells enter the microwells by gravity and are then processed using standard comet assay conditions to create an array of comets. The bed of arrayed microwells can be made to the size of a standard multiwell plate and can be partitioned into 96 compartments by using a bottomless 96-well plate, each with ~300 comets per well. The approach effectively miniaturizes the assay, enabling conversion from one glass slide per sample (the traditional assay) to one 96-well per sample. With the CometChip, multiple samples can be prepared in parallel on the same assay substrate, and throughput is greatly increased via automated imaging and analysis using either in-house or commercially available software available through Trevigen, Inc. However, the existing hardware for performing the CometChip is not compatible with HTS robotics, because the hardware dimensions do not match a typical multiwell plate, a requirement for automated manipulation of 96-well plates. Here, we describe a novel advance in the throughput of DNA damage detection by creating hardware that conforms to the requirements of high throughput robotics and is compatible with standard automated microscopes. The HTS CometChip enables large-scale screens. We leveraged this platform to perform a large-scale shRNA screen of 2564 human genes wherein the levels of DNA DSBs were directly measured. From the screen, we identified LATS2, as a modulator of both apoptosis and DNA repair. We furthermore show that in addition to its role in regulation of cell–cell contact survival, LATS2 modulates DNA repair and radiation sensitivity. Beyond these new roles for LATS2, this work establishes the utility of the HTS CometChip as a tool for the discovery of novel genes that impact DNA damage and repair.
Materials and Methods
Cell Culture
M059K and M059J human glioblastoma cells from ATCC (Manassas, VA) were cultured in 1:1 DMEM/F12 nutrient mix (Invitrogen-Thermo-Fisher Scientific, Waltham, MA) supplemented with 10% fetal bovine serum (FBS; Atlanta Biologicals, Atlanta, GA), 1× MEM Non-Essential Amino Acids (Invitrogen), and 100 units/mL penicillin-streptomycin (Invitrogen). HeLa human cervical adenocarcinoma cells from ATCC were cultured in DMEM (Invitrogen) supplemented with 10% FBS (Atlanta Biologicals) and 100 units/mL penicillin-streptomycin (Invitrogen).
HTS CometChip Apparatus Fabrication
A 120 × 78 mm aluminum base plate was cut from a 3 mm thick aluminum sheet to fit tightly in a uniwell tray (VWR, Radnor, PA). Four aluminum posts (6 mm in diameter and 6.5 mm in height) were welded onto the base plate with their centers 28.5 mm and 7.5 mm from the short and long edges of the plate, respectively. A 2 mm wide and 5 mm deep hole was drilled parallel to the long axis of each of the posts, from the center of their flat surfaces. The outer edges of a bottomless 96-well plate (VWR) were sawed off, leaving a 110 × 74 mm grid of 96 wells. Wells A3, A10, H3, and H10 were plugged with tight-fitting polystyrene cylinders with a 2 mm wide hole drilled at their centers. HTS CometChip was assembled by sandwiching the microwell arrayed agarose between the aluminum base plate and the bottomless well plate, such that the posts on the aluminum plate meet the pegs on the bottomless plate, and fastening them together with screws.
Neutral CometChip
The standard CometChip was prepared as described previously. 27 Briefly, 12 mL of 1% molten agarose in PBS is poured over a GelBondflim (Lonza, Basel, Switzerland) placed on a uniwell tray (VWR). A reusable PDMS stamp arrayed with 40 µm wide microposts is floated on the molten agarose and the agarose is allowed to cool and solidify. Following gelation, the PDMS stamp is removed to reveal microwells for cell loading. The agarose and the GelBond film are placed on a glass plate, and a bottomless 96-well plate (VWR) is clamped onto the agarose using binder clips to assemble the standard CometChip. The HTS CometChip assembly is described in Figure 1 . The microwell array is poured on GelBond film and held between the aluminum base plate and a bottomless 96-well plate with four screws. Cell suspensions are loaded into individual wells of the bottomless 96-well plate and the apparatus is placed in a 37 °C incubator for 20 min to allow cell loading. After cell loading of either standard or HTS CometChip, excess cells are washed off with PBS, and a thin layer of 1% molten low-melting-point agarose in PBS is applied on the top surface of the CometChip. Cells within the CometChip are exposed to DNA-damaging agents (e.g., γIR) and placed in complete media for recovery. Cells are then lysed at 43 °C overnight after desired time points by submerging the CometChip into lysis buffer containing 2.5 M NaCl, 100 mM Na2 EDTA, 10 mM Tris, 1% N-lauroylsarcosine, pH 9.5, with 0.5% Triton X-100 and 10% DMSO. After cell lysis, the CometChip is washed three times with neutral electrophoresis buffer containing 90 mM Tris, 90 mM boric acid, 2 mM Na2 EDTA, pH 8.5. Electrophoresis was conducted using prechilled neutral electrophoresis buffer at 4 °C for 1 h at 0.6 V/cm and 6 mA. The CometChip is then stained with SYBR Gold (Invitrogen) according to the manufacturer’s instructions and images are taken using an upright fluorescent microscope with a 4× objective for non-high-throughput experiments (https://zenodo.org/record/2656668#.XqhZ6GhKhaQ). Comets are analyzed for the lengths of their tail region using a custom-made script written in MatLab (MathWorks, Natick, MA): the code is available from DOI: 10.5281/zenodo.2656668.
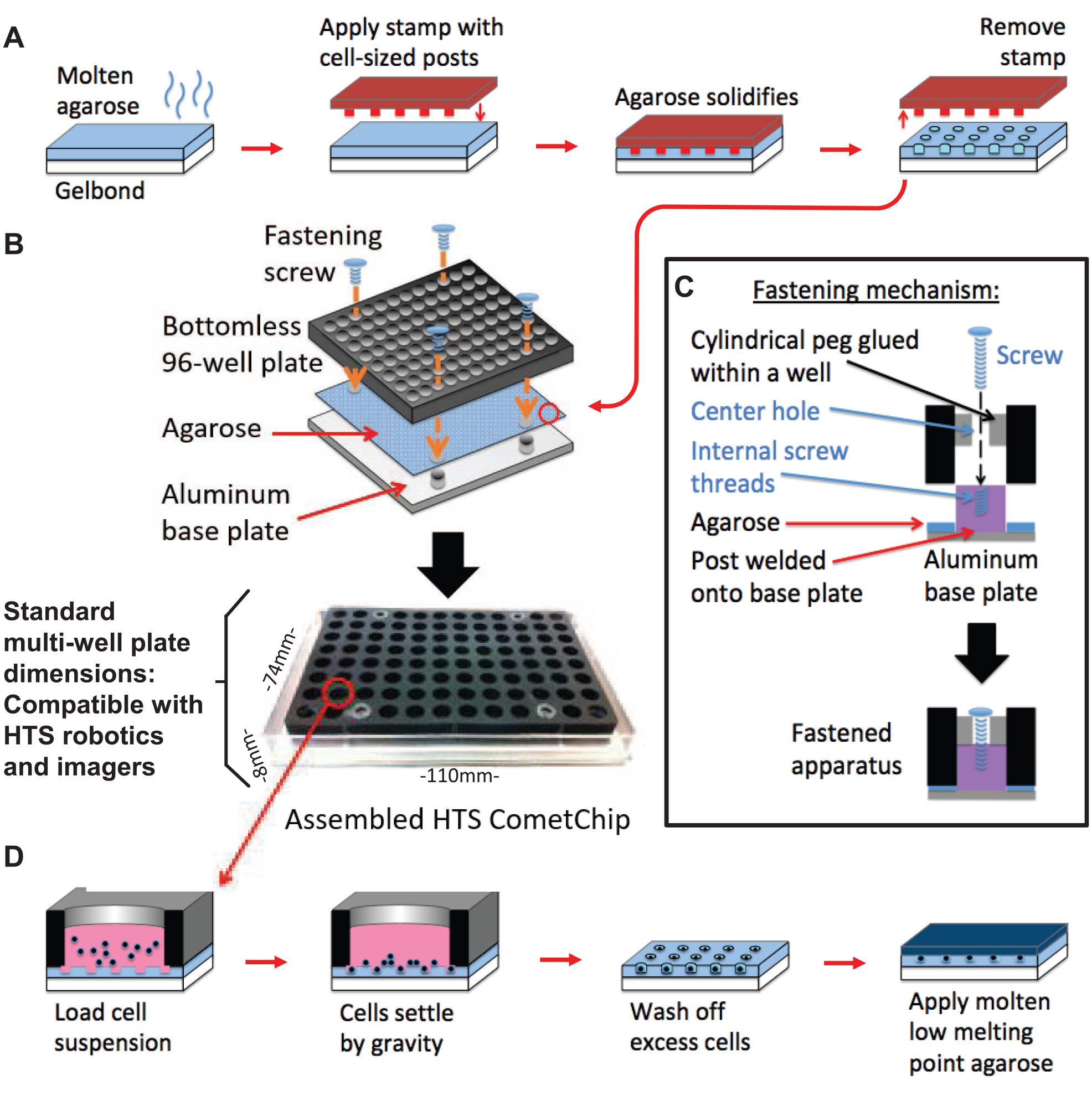
Creation of an HTS-compatible CometChip platform. (
HTS Robotics
Access to HTS robotics and equipment was obtained via MIT’s Koch Institute of Integrative Cancer Research, Swanson Biotechnology Center, High Throughput Sciences Facility. Robotic manipulation of the HTS CometChip was performed using a Evo 100 liquid handler (Tecan, Inc., Mannedorf, Switzerland), equipped with a 96-well MultiChannel Arm (MCA96) and a Robotic Manipulator Arm. The Evo 100 is also integrated with a Liconic (Woburn, MA) STX110 incubator with temperature, humidity, and CO2 control. Imaging of the HTS CometChip was performed using a Thermo Scientific Cellomics ArrayScan VTI HCS Reader with a 5× objective.
Pilot shRNA Screen
Lentiviral reagents expressing shRNA were obtained from the Genetic Perturbation Platform of the Broad Institute of MIT and Harvard (https://portals.broadinstitute.org/gpp/public/). The list of shRNA IDs and their target sequences used in the pilot screen are shown in
HTS CometChip shRNA Screen
Lentiviral reagents expressing shRNA were obtained from MIT’s Koch Institute of Integrative Cancer Research, Swanson Biotechnology Center, High Throughput Sciences Facility. This shRNA library was purchased from the Broad Institute of MIT and Harvard, The RNAi Consortium (TRC) as a subgenome library of 2564 genes, with three unique shRNAs per gene. The genes targeted by this library are included in the supplemental material available online and additional clone information is available at Broad link: https://portals.broadinstitute.org/gpp/public/.
This library was formatted as an arrayed library, with lentivirus targeting one gene in each well of a 96-well plate, across 89 plates. The screening effort was divided into alternating screening days and imaging days. Prior to screening days, M059K cells were plated on duplicate tissue culture 96-well plates at a density of 600 cells/well, with the exception of 3 wells on each plate to allow the inclusion of untransduced M059K cells (2 wells) and untransduced M059J cells (DNAPKcs null; 1 well). Cells were infected with lentiviruses at an average MOI of 8 the next day after cell plating. One well was infected with a separate preparation of DNAPKcs shRNA that performed best in the pilot screen. Transduced cells were selected with 1.5 μg/mL puromycin 48 h after infection. Puromycin was applied for 72 h and cells were allowed to recover in selection-free media for 24 h. All media changes were performed using a BioTek EL406 microplate washer under sterile conditions. On screening days, cells were transferred to HTS CometChips as described above. CometChips were then submerged in media and treated with 100 Gy of γ-rays at 1 Gy/min using a 137Cs source. Cells were allowed to repair DNA damage at 37 °C for 4 h in media before they were lysed overnight. On imaging days, the HTS CometChips were washed thrice with neutral electrophoresis buffer and electrophoresis was conducted to measure the extent of DNA DSBs. HTS CometChips were stained with SyBR Gold and imaged with a Thermo Scientific Cellomics ArrayScan VTI HCS Reader using a 5× objective.
Transfection with siRNAs
Negative control siRNA (cat. 4390843) and siRNAs targeting LATS2 (cat. 4392420, ID s25505) and DNAPKcs (cat. 4390824, ID s774) were purchased from Ambion (Thermo Scientific) and dissolved in nuclease-free water. HeLa cells (40,000) were plated on each well of a 6-well plate and transfected with siRNA reagents 24 h later as per the manufacturer’s instructions.
qRT-PCR
Total RNA was extracted from approximately 500,000 HeLa cells treated with LATS2 or negative control siRNA using RNeasy kit (Qiagen, Germantown, MD). Complementary DNA was generated using random hexamers with M-MLV reverse transcriptase (Invitrogen) according to the manufacturer’s instructions. Semiquantitative PCR was performed using an 7500 Fast Real Time PCR System machine (Applied Biosystems, Thermo-Fisher Scientific, Waltham, MA), with Fast SYBR Green master mix (Applied Biosystems) using the following primers: actin (forward)—AGA GCT ACG AGC TGC CTG AC, actin (reverse)—AGC ACT GTG TTG GCG TAC AG, LATS2 (forward)—TCA GAC AGG ACA GCA TGG AG, and LATS2 (reverse)—TAG TTT GGA GTC CCC ACC AG. Data from the experiment were analyzed using the ΔΔCT method.
Western Blotting
M059K cells were treated with lentiviral shRNA constructs targeting DNAPKcs or GFP, and whole-cell lysates were harvested after successful transduction. Whole-cell lysates were collected using RIPA buffer (Pierce, Thermo-Fisher Scientific, Waltham, MA) supplemented with HALT protease inhibitor (Thermo-Fisher Scientific) and frozen at −20 °C. Lysates were thawed on ice and spun down at maximum speed for 15 min. Lysates were mixed with Laemmli sample buffer (Bio-Rad, Hercules, CA) according to the manufacturer’s instructions and incubated at 95 °C for 5 min and then chilled on ice. Samples were loaded onto a 6% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) gel and electrophoresed at 200 V for about 90 min with Tris-glycine-SDS running buffer (Bio-Rad). Proteins were transferred onto nitrocellulose membranes (Bio-Rad) at 100 V for 2 h in Tris-glycine-methanol transfer buffer (Bio-Rad). Membranes were blocked in Odyssey blocking buffer (LI-COR, Lincoln, NE) for 2 h at room temperature with gentle shaking. DNAPKcs and Ku80 were probed with primary antibodies (Cell Signaling Technologies [Beverly, MA] cat. 4602 and cat. 2180) in blocking buffer overnight at 4 °C with gentle shaking. Secondary antibodies (LI-COR) were applied to the membrane in blocking buffer for 2 h at room temperature with gentle shaking. HeLa cells treated with LATS2 or negative control siRNA were treated with 10 Gy of ionizing radiation and allowed to incubate at 37 °C, 5% CO2 for up to 1–24 h. Whole-cell lysates were collected using whole-cell lysis buffer (0.12 M Tris-HCl, pH 6.8, 4% SDS). Lysates were sonicated at 35 amplitude for 1 min in a cup-horn sonicator. Lysates were mixed with Laemmli sample buffer and incubated at 95 °C for 5 min and separated by SDS-PAGE gels (15% for H2AX, 6% for LATS2). Proteins were transferred onto nitrocellulose membranes (Bio-Rad) at 30 V for 20 h in Tris-glycine transfer buffer. Membranes were blocked in Odyssey blocking buffer (LI-COR) for 2 h at room temperature with gentle shaking. Total H2AX and γH2AX were probed with primary antibodies (Abcam [Cambridge, MA] cat. Ab11175 and EMD Millipore [Burlington, MA] cat. 05-636) in blocking buffer overnight at 4 °C with gentle shaking. Secondary antibodies (LI-COR) were applied to the membrane in blocking buffer for 2 h at room temperature with gentle shaking. Bands were visualized using a LI-COR Odyssey scanner.
CellTiter-Glo Assay
HeLa cells treated with LATS2 or negative control siRNA were treated with up to 20 Gy of ionizing radiation in 96-well plates and allowed to recover for 3 days before performing the CellTiter-Glo assay. Fifty microliters of reconstituted CellTiter-Glo reagent (Promega, Madison, WI) was added to each well containing 50 µL of media and allowed to incubate in room temperature for 10 min with gentle shaking. Analyte was transferred to a white 96-well plate (Evergreen Scientific, Buffalo, NY) and analyzed with a Tecan M1000 plate reader, with an integration time of 1 s per well.
Annexin V Staining
HeLa cells treated with LATS2 or negative control siRNA were transferred onto HTS CometChip and analyzed immediately or after 4 h of incubation at 37 °C in media by annexin V staining (Life Technologies, Thermo-Fisher Scientific, Waltham, MA) according to the manufacturer’s instructions. DNA was stained using Vybrant DyeCycle Violet stain (Life Technologies).
Results
Fabrication of HTS CometChip Apparatus
The CometChip was developed to improve the precision and throughput of DNA damage measurements.26–28 Patterned microwells dictate the positions of cells within an array, which minimizes the footprint per sample, avoids overlapping comets, and facilitates automated imaging and image analysis (
Next, we clamped a bottomless 96-well plate onto the gel surface, so that each of the 96 wells had hundreds of microwells at its base, as was done previously for the CometChip. In order to attach a commercially available 96-well plate to the aluminum base, we sacrificed 4 wells so that they could be used as attachment points. Specifically, hollow pegs (e.g., polystyrene cylinders), each with a 2 mm hole in their centers, were fused to the inner surface of wells (
Once assembled, the HTS CometChip is placed on the work area of HTS robotic equipment. The robot pipettes cell suspensions into each well of the HTS CometChip, transfers the HTS CometChip into a 37 °C incubator for cell loading, and retrieves it after wells have been loaded with cells by gravity (
Functional Testing of HTS CometChip
To explore the possibility of using the HTS CometChip for large-scale experiments using HTS robotics, we tested the performance of our apparatus under a range of γIR doses. In order to specifically assess the efficacy of our approach in detecting DNA DSBs, we used the human glioblastoma cell lines M059K and M059J.
29
Derived from the same tumor, the M059J cell line has a truncation mutation in the DNAPKcs gene,
30
whereas the M059K cell line has intact wild-type (WT) DNAPKcs. Given the importance of the core components of the NHEK pathway for DSB repair, we reasoned that these two cell lines should give significant differences in DSB persistence following γIR exposure. Previous studies from our group using neutral CometChip also demonstrate elongated comet tails indicative of decreased DNA DSB repair capacity in cells treated with DNAPKcs chemical inhibitors prior to exposure to DNA DSB-inducing agents.
27
We therefore treated M059K and M059J cells on the HTS CometChip with increasing doses of γIR and allowed them to repair the resulting DNA damage over time. We performed the neutral HTS CometChip assay, as the neutral CometChip had previously been shown to be effective for the detection of DSBs.27,31 We observed that irradiation led to a dose-dependent increase in the comet tail lengths (
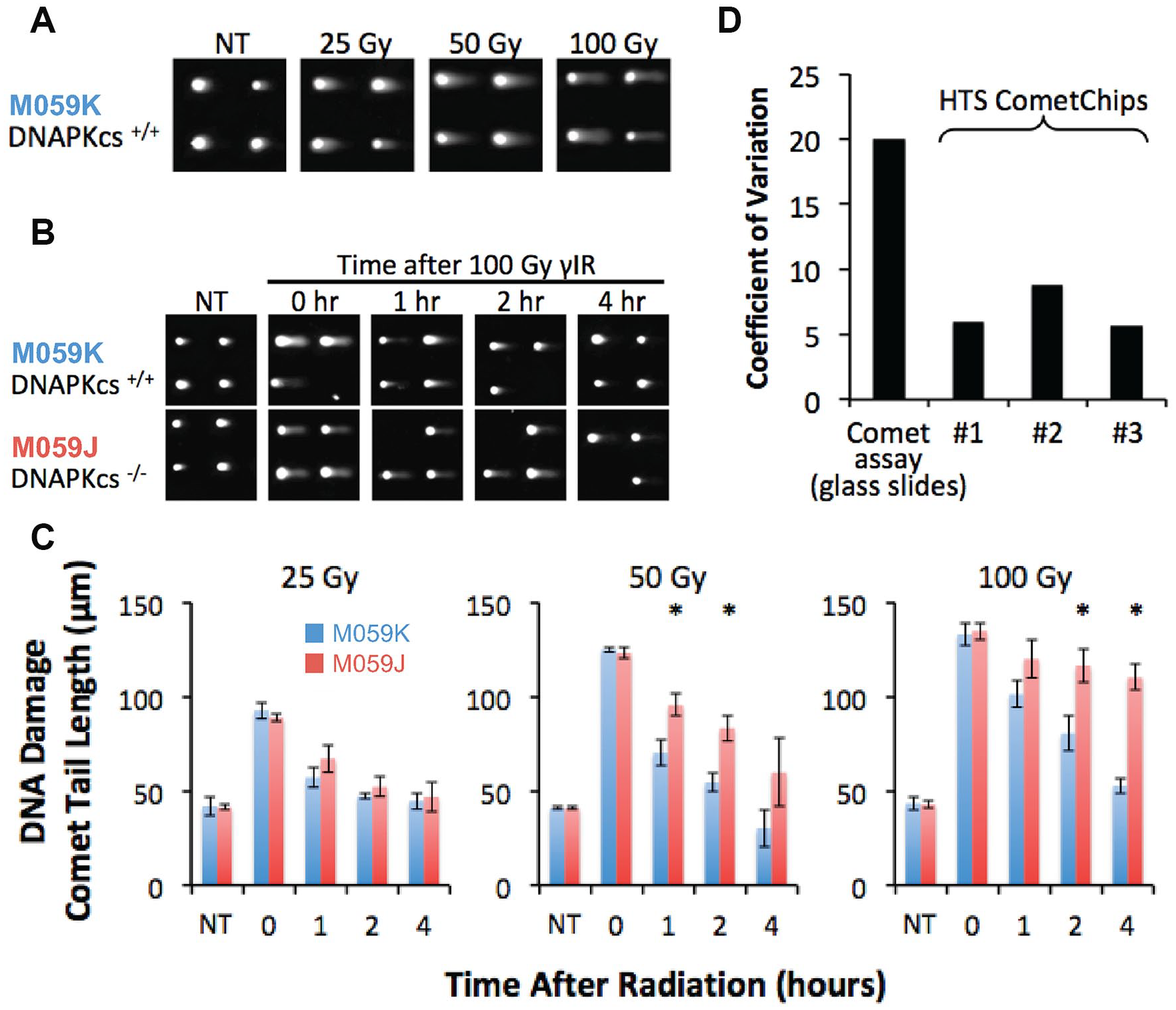
HTS CometChip assay characterization. (
Next, we tested the consistency of the data obtained among three HTS CometChips exposed to the same treatment conditions. We found that the coefficient of variation (COV) between wells on each HTS CometChip was between 5% and 10%, consistent with the original version of CometChip
27
and lower than the traditional comet assay (
Pilot Screening
Having demonstrated the efficacy of the HTS CometChip assay for measuring DSBs, we next performed a pilot shRNA screen of key NHEJ genes that should inhibit DSB repair, along with negative controls predicted to have no effect on DSB repair. We designed a custom-made 96-well plate of lentiviruses expressing shRNAs targeting canonical NHEJ genes or negative controls (
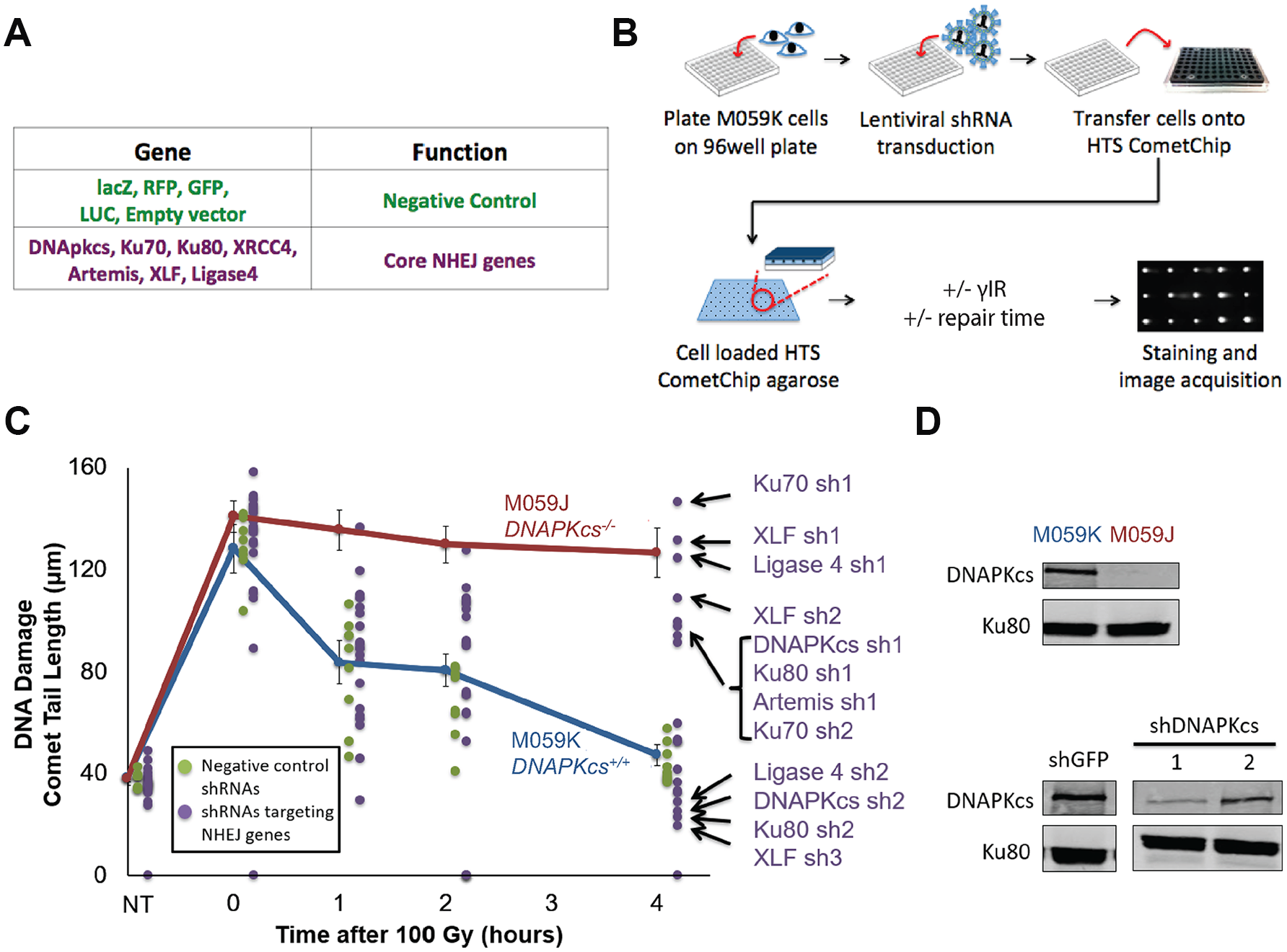
High-throughput shRNA pilot screening using the HTS CometChip. (
In our pilot screen, we assessed the performance of the HTS CometChip under screening conditions. Similar to our earlier assay performance tests, we observed that under lentiviral shRNA screening conditions, untransduced cells proficient in DSB repair (M059K) show robust DNA repair, while M059J cells show reduced repair (blue and red lines, respectively;
In some cases, we observed differing results from shRNAs targeting the same gene. We anticipated that this could be the case, given that some shRNAs can be less effective than others. We further explored this possibility by studying the knockdown efficiency for two of the shRNAs targeting DNAPKcs by Western blot. As shown in
Figure 3D
, we found that shRNA 1 had highly efficient knockdown (corresponding with high levels of persistent DNA DSBs;
Large-Scale Arrayed Lentiviral shRNA Screen for Modulators of DNA DSB Repair
We next leveraged the HTS CometChip assay to perform a large-scale arrayed shRNA library screen. The shRNA library was created by the Broad Institute as a curated subset of TRC genomic shRNA library. We specifically selected a library that harbors shRNA for oncology-related targets. This library is composed of 7692shRNAs, targeting 2564 key cancer pathway genes. This library was focused on cancer cell signaling, and therefore excluded many essential genes (such as core DNA replication machinery), whose loss would lead to cell death. The arrayed library was prepared in 89 tissue culture 96-well plates (
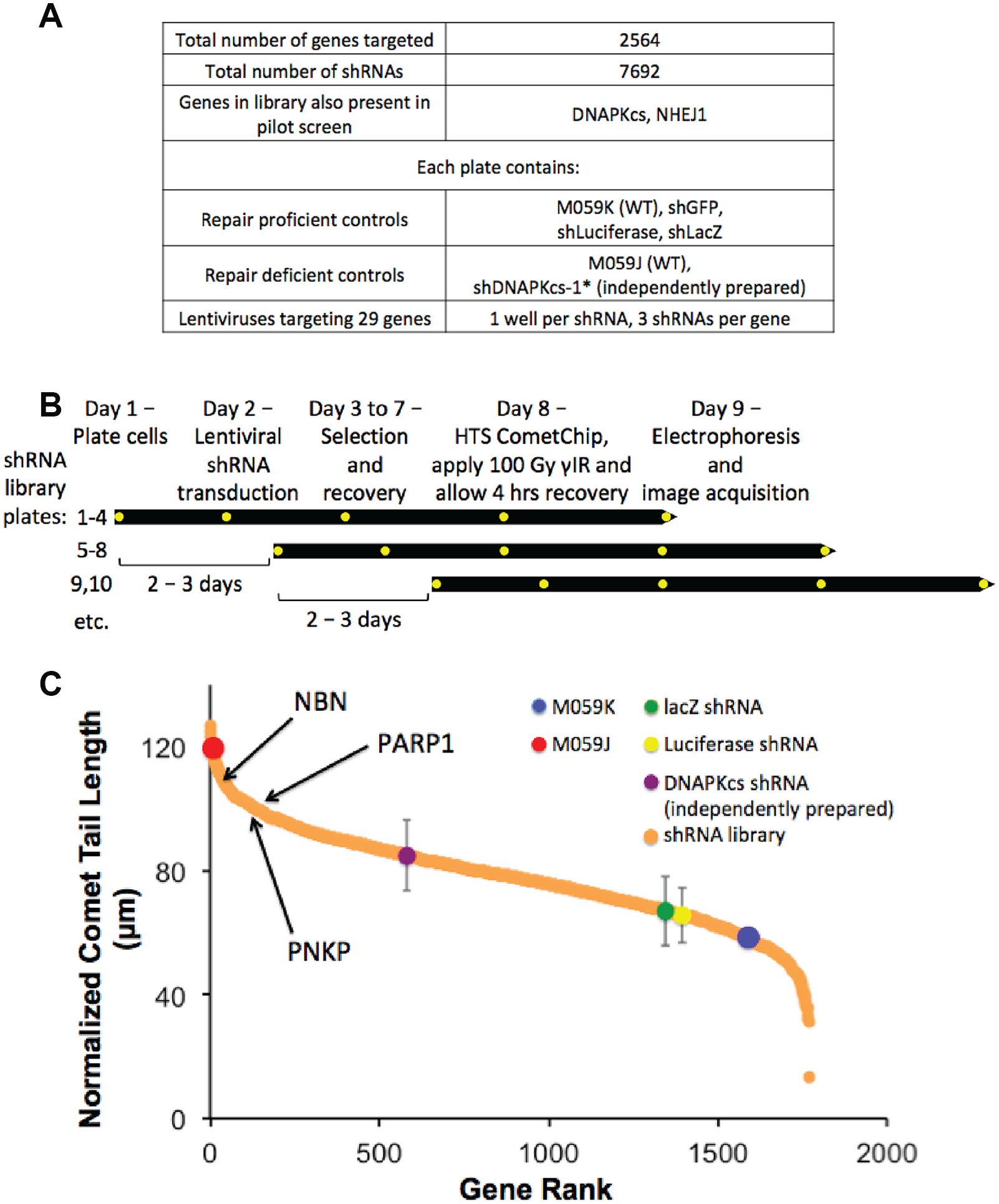
Large-scale arrayed lentiviral shRNA screen for modulators of DNA DSB repair. (
In order to efficiently assay a large number of plates in the screen, we designed a workflow that uses a staggered format (
Screen of 2564 Genes Reveals LATS2 as a Top Hit for DNA Fragmentation
We interrogated which pathways were represented in the top 150 genes from our screen by performing Ingenuity Pathway Analysis (
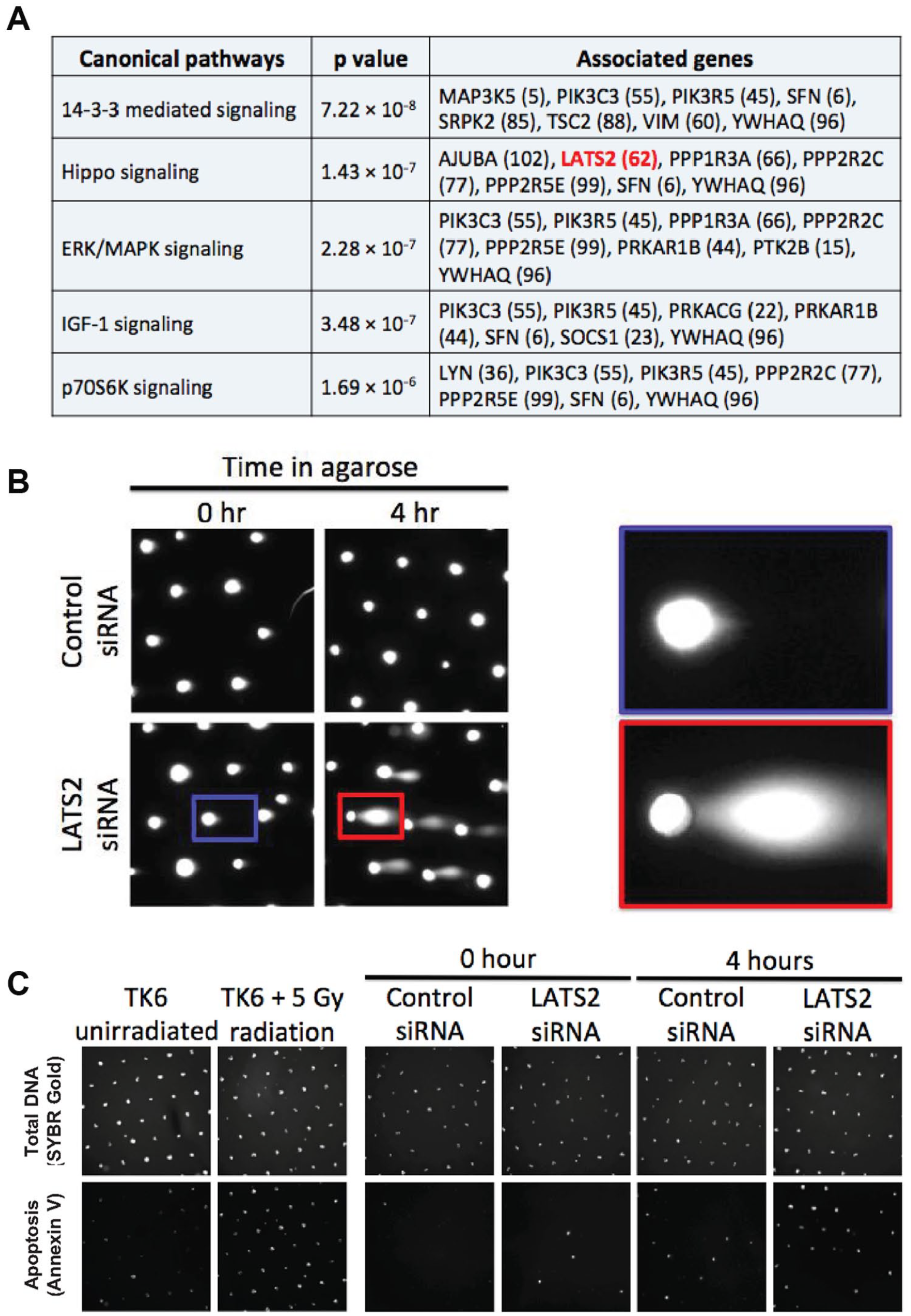
Identification of an apoptotic phenotype in LATS2-depleted cells. (
The second most significantly enriched pathway identified by Ingenuity Pathway Analysis is the Hippo signaling pathway. The Hippo signaling pathway is known to control the rates of cell growth and organ sizes in an organism. 36 In flies, inactivation of the Hippo signaling pathway leads to tumor-like growth in specific tissues, due to loss of cell–cell contact growth inhibition and overproliferation of the constituent cells. 37 Conversely, deregulation of the Hippo signaling pathway can lead to changes in the cell’s response to pro-apoptotic signals activated by chemotherapeutic agents or changes in anchorage status. Given these known roles of the Hippo signaling pathway, we reasoned that increased DNA fragmentation (longer tails) observed in cells knocked down for Hippo pathway genes could also be due to apoptosis.
To explore the possibility of apoptosis-induced comets in our screen, we analyzed the comet images from RFC5, a gene that is essential for DNA replication, as well as a prominent candidate gene belonging to the Hippo signaling pathway, LATS2. We reasoned that depleting a gene that requires DNA replication will lead to replication fork collapse during the S phase of the cell cycle, and subsequently lead to DSB formation and apoptosis. Indeed, analysis of images from RFC5 in the screen showed that many of the comets have a shape that is characteristic of apoptosis, wherein there is extensive DNA fragmentation19,20,38–40 (
To further explore the possibility that LATS2 KD cells are sensitized to apoptosis under anchorage-independent growth conditions, we used an alternative approach for assaying for apoptosis, namely, annexin V staining (
LATS2 Knockdown Cells Are Defective in DNA DSB Repair
Several proteins in the Hippo signaling pathway have also previously been shown to be activated when cells are exposed to DNA damage.
43
This led us to hypothesize that LATS2 KD cells might also be deficient in DNA DSB repair. To investigate whether or not LATS2 KD has a DNA repair defect, we treated control and LATS2 KD-irradiated cells and assayed for DNA DSBs. Specifically, we treated HeLa cells with siRNA targeting sequences in LATS2 distinct from the shRNA sequences used in the screening library, and allowed them to recover under adherent conditions on tissue culture plates, thereby maintaining cell–cell contacts, before transferring them to the CometChip to enable studies of DNA damage and repair (rather than apoptosis). In images obtained from these experiments, tail morphology was consistent with DNA damage, rather than apoptosis (
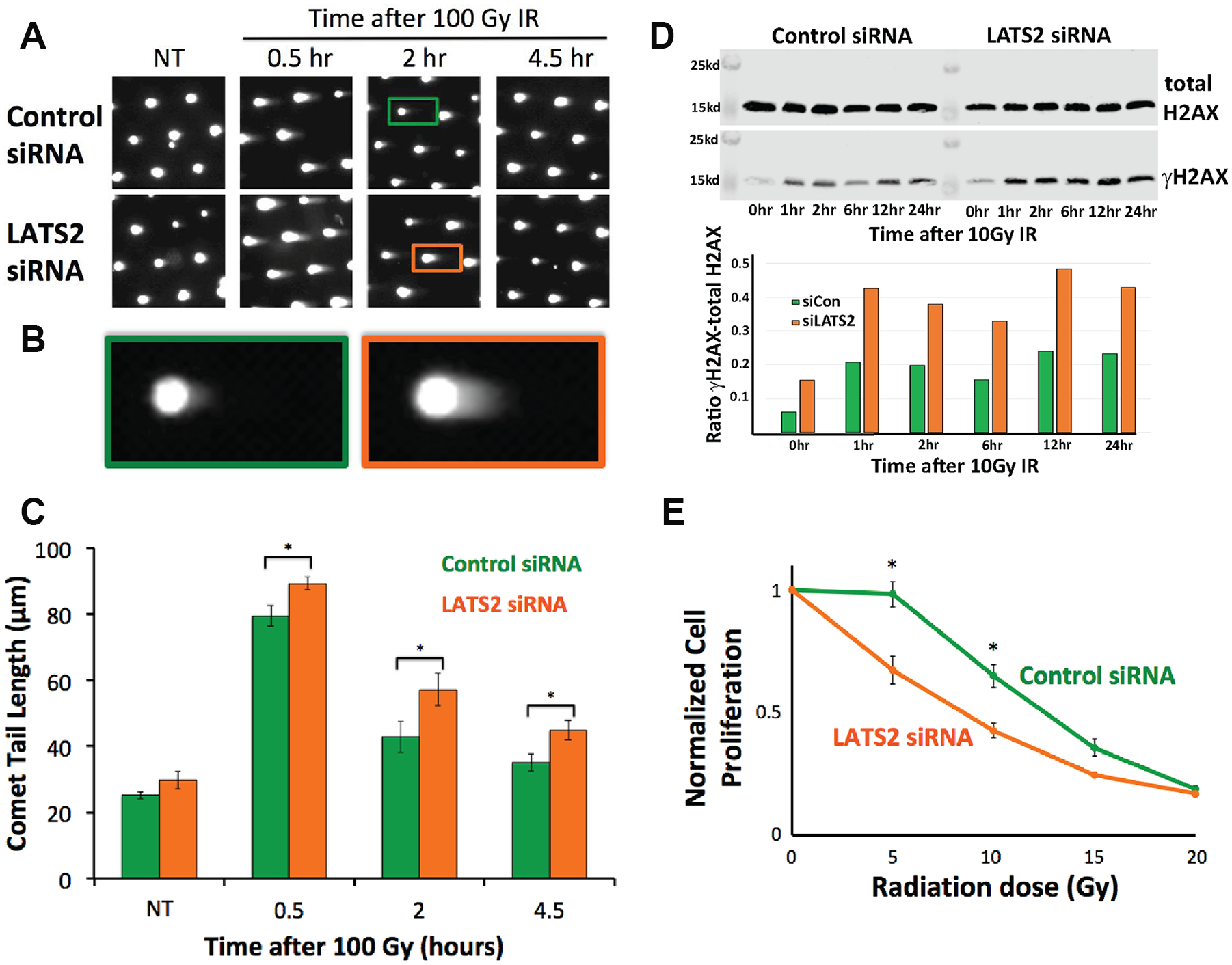
LATS2 knockdown causes DNA DSB repair defects. (
As an independent approach for evaluating the levels of DSBs, we also tested the effect of LATS2 KD on the formation of γH2AX after exposure to γIR via Western blotting. We found that LATS2 KD cells have a higher level of γH2AX after being exposed to γIR as compared with control cells, which is consistent with the presence of higher levels of DNA DSBs (
Discussion
In this study, we developed technology to perform large-scale comet assay-based experiments utilizing HTS robotics and performed a first-of-its kind DNA DSB genetic screen. The original comet assay, although well accepted by the DNA repair community, is an assay that is rarely adopted when researchers intend to perform large-scale experiments. There are two main reasons for this: (1) incompatibility with HTS robotics and (2) the high variability in data produced by the assay. In terms of sample handling, the CometChip alleviates the problems of the comet assay by maximizing the number of comets obtainable per unit area of the assay, which makes it possible to analyze 96 samples on the same assay plate. This reduces the amount of preparatory work per sample, therefore making the assay less cumbersome to perform. In terms of reproducibility, testing multiple samples on the same assay plate also reduces noise between samples, therefore reducing variability in the data produced. Furthermore, the regular array of comets produced by the patterned microwells prevents overlapping comets, enabling automated image analysis, which greatly increases the throughput and precision of the assay. While the CometChip is now well established, the HTS CometChip takes the platform a step further, enabling the use of HTS robotics. By leveraging the innovations of robotic screening and further improving precision, speed, and reproducibility, it is now possible to perform large-scale comet assay experiments.
With this platform, we queried a library of 7692 shRNAs targeting 2564 genes to identify gene targets affecting DNA repair following γIR exposure. In addition, since the comet assay detects DNA fragmentation, it was also possible to detect DNA damage from apoptosis. This comprises the first high-throughput genetic screen using physical DNA fragmentation as a readout for apoptosis. Despite the lack of shRNAs targeting canonical DNA repair genes in the screening library, we identified the 14-3-3 signaling as the top pathway from our screen. The 14-3-3 signaling pathway is intimately involved in a cell’s DNA damage response, providing evidence that our screening methodology could potentially reveal novel DNA repair genes and pathways. This screen identified LATS2 as an IR-induced DNA damage repair factor. Interestingly, LATS2 also scored in our assay for apoptotic DNA damage. As such, the platform can be used both for studies of apoptosis and for studies of DNA damage and repair.
In our study, we identified the Hippo pathway protein kinase LATS2 as a novel DNA DSB repair modulator. The Hippo pathway was also identified as the second most significantly enriched pathway in the top 150 genes from our screen (7 out of 150 genes). The Hippo pathway is known to control cell growth rates and organs.36,37,44 The members of the Hippo pathway were first identified in Drosophila, where mutant Hippo pathway genes led to massive overgrowth in the organs that harbored the mutated cells. The Hippo pathway consists of four kinases, MST1, MST2, LATS1, and LATS2, which receive input signals from environmental cues such as cell adhesion and cell–cell contact. 45 The end result of the kinase cascade activation in this pathway is the phosphorylation of the YAP/TAZ transcriptional co-activators, leading to their degradation. When the kinases are inactive, YAP and TAZ can translocate into the cell nucleus and promote the activity of TEADs and SMADs, which in turn promotes cell growth and tumorigenesis. 46 In the context of these cell–environment signaling pathways, LATS2 and the Hippo pathway were connected to DNA damage via Abl kinase. 47
Our data demonstrate that the loss of LATS2 in cells led to sustained levels of DNA DSBs after exposure to γIR, as well as to increased levels of γH2AX signaling. We also observed a radiation-induced growth defect in LATS2-depleted cells. This, coupled with the DSB repair defect we observed, suggests that increased radiation sensitivity could be due to unrepaired DSBs following LATS2 loss of function.
We also observed that LATS2 depletion led to increased comet tail length when the cells were dispersed and transferred from tissue culture plates to agarose. This phenotype was observed even when cells were not exposed to γIR, suggesting that these comet tails could be caused by DNA fragmentation from apoptotic signaling. Since the Hippo pathway responds to changes in cell–cell contacts and to changes in the extracellular matrix environment, we hypothesize that transferring the cells from a monolayer on a tissue culture plate to a low-adherence single-cell 3D environment encapsulated by agarose may change the activity of the pathway, and that loss of LATS2 in this context may lead to activation of apoptosis. We confirmed this by demonstrating annexin V staining in LATS2-depleted cells treated with the same nonirradiated conditions. Therefore, our identification of LATS2 demonstrates the ability of the HTS CometChip to detect modifiers of both radiation- and apoptosis-induced DSBs.
In terms of efficacy for use in a large screen, it is helpful to compare HTS CometChip throughput to what is feasible with the traditional assay. Our screening library was distributed across eighty-nine 96-well plates, with each shRNA occupying a single well. We collected 9 images from each well, capturing an average of 50 comets per shRNA. Including the wells that were assigned for controls on each plate, we collected ~130,000 images, containing more than 750,000 comets. To put the scale of the screen in perspective, given a liberal estimate of being able to complete 100 traditional comet assay slides per week, it would take a typical researcher at least 150 weeks to collect the same amount of data as we did in about 10 weeks. Thus, the HTS CometChip can be used to screen thousands of genes for their potential impact on DNA fragmentation.
Although beyond the scope of this particular study, it is possible to further increase the rate of data collection when using the HTS CometChip in a genomic screen. In our proof-of-concept screen, we fabricated and loaded 8 HTS CometChips at a time each day, taking about 1.5 h to complete. This can be easily quadrupled, to yield data from approximately 2600 samples (32 HTS CometChips × 84 sample wells) per day or 13,000 samples per 5-day workweek. A screen using a typical genomic shRNA library that targets approximately 20,000 genes using three distinct shRNAs per gene will take as little as 10 weeks to complete, with duplicate data for each shRNA. The possibility of performing a genome-wide screen for factors that modulate DNA DSBs by measuring physical DNA damage can now be fully realized with the advent of HTS CometChip as an enabling innovation.
Our study focused on the application of HTS CometChip to detect physical DNA DSBs, but the HTS CometChip can also be used to detect other forms of DNA damage. To discover genes that are involved in DNA DSB repair, we performed the neutral comet assay; however, different conditions can detect other classes of DNA damage. For example, base damage induced by oxidizing or alkylating agents can be detected when it is converted to strand breaks either enzymatically (e.g., during base excision repair) or by incubation under alkaline conditions. 48 Alternatively, it is also possible to detect subclasses of damage such as oxidized purines. These lesions can be detected by incubation with specific endonucleases after cell lysis, which converts undetectable base lesions into detectable abasic sites and/or single-strand breaks. 49 The ability to measure single-strand breaks that are formed during excision repair further expands the utility of the assay by enabling analysis of the DNA repair capacity for damaged bases. Furthermore, the HTS CometChip can also measure epigenetic marks such as DNA methylation. Specifically, the recently reported EpiComet-Chip 50 exploits recognition of 5MeC by the McrBC restriction enzyme, and converts sites of CpG methylation into detectable single-strand breaks. These can then be detected readily by the HTS CometChip. Therefore, we propose that the HTS CometChip is useful for a broad range of applications. Finally, it was recently shown that the microwell platform used for these studies can also be cross-purposed to monitor cell growth, thus facilitating parallel analysis of genotoxicity and cytotoxicity. 51
All of the proposed variants of the HTS CometChip experiments (e.g., measuring DSBs, alkali-labile sites, base damage, and 5MeC) can also be adapted for small-molecule screens as part of a drug discovery process. Small-molecule libraries are frequently arranged in an arrayed format (each drug/dose condition in the library occupies one well in a multiwell plate), and this format is highly compatible with the arrayed nature of the HTS CometChip. A small-molecule library can also be used in tandem with a DNA damage inducer (such as γIR) to identify DNA repair chemical inhibitors. These inhibitors can form the basis of drug development to produce adjuvant therapies that can enhance the effectiveness of DNA-damaging cancer treatments, leading to improved outcomes.
In conclusion, we have developed the HTS CometChip to render the powerful CometChip technology compatible with automated robotic systems found in HTS facilities. We used this assay to query thousands of genes for their ability to modulate DNA DSB levels by measuring levels of physical DNA DSBs in cells exposed to radiation. From a screen of more than 7500 shRNAs, we identified LATS2 as a gene that, when depleted, leads to inefficient DNA DSB repair and radiosensitivity. We also showed that LATS2-depleted cells undergo apoptosis more readily when they were encapsulated in agarose, indicating that the HTS CometChip can also be used for studies of apoptosis. Given the myriad permutations to the comet assay for detecting different classes of damage, excision repair capacity, and even global methylation levels, the HTS CometChip is an effective tool for performing a broad range of high-throughput screens with applications in cancer biology.
Supplemental Material
20200227Supplemental – Supplemental material for HTS-Compatible CometChip Enables Genetic Screening for Modulators of Apoptosis and DNA Double-Strand Break Repair
Supplemental material, 20200227Supplemental for HTS-Compatible CometChip Enables Genetic Screening for Modulators of Apoptosis and DNA Double-Strand Break Repair by Ian J. Tay, James J. H. Park, Anna L. Price, Bevin P. Engelward and Scott R. Floyd in SLAS Discovery
Supplemental Material
Supplemental_Table_1 – Supplemental material for HTS-Compatible CometChip Enables Genetic Screening for Modulators of Apoptosis and DNA Double-Strand Break Repair
Supplemental material, Supplemental_Table_1 for HTS-Compatible CometChip Enables Genetic Screening for Modulators of Apoptosis and DNA Double-Strand Break Repair by Ian J. Tay, James J. H. Park, Anna L. Price, Bevin P. Engelward and Scott R. Floyd in SLAS Discovery
Supplemental Material
Supplemental_Table_2 – Supplemental material for HTS-Compatible CometChip Enables Genetic Screening for Modulators of Apoptosis and DNA Double-Strand Break Repair
Supplemental material, Supplemental_Table_2 for HTS-Compatible CometChip Enables Genetic Screening for Modulators of Apoptosis and DNA Double-Strand Break Repair by Ian J. Tay, James J. H. Park, Anna L. Price, Bevin P. Engelward and Scott R. Floyd in SLAS Discovery
Footnotes
Acknowledgements
This work was made possible with the assistance and expertise of the MIT Central Machine Shop for the fabrication of the HTS CometChip apparatus. We thank the Koch Institute’s Robert A. Swanson (1969) Biotechnology Center for technical support, specifically, the High Throughput Sciences Facility.
Supplemental material is available online with this article.
Authors Note
Scott R. Floyd is currently affiliated with Department of Radiation Oncology, Duke University School of Medicine, Durham, NC, USA.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Joint Center for Radiation Therapy foundation (JCRT), the Koch Institute Clinical Investigator Award, and the Burroughs Wellcome Career Award for Medical Scientists (awarded to S.R.F.). I.J.T. was supported by the Agency for Science Technology and Research (A*STAR) National Science Scholarship (NSS-PhD), with support from P01CA26731 and R44ES024698 (awarded to B.P.E.). This work was also supported by the MIT Centre for Environmental Health Sciences (P30ES002109), and by the Koch Institute Support (core) Grant P30-CA14051 from the National Cancer Institute.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
