Abstract
Ionizing radiation-induced formation of genomic DNA damage can be modulated by nearby chemical species such as heavy metal ions, which can lead to non-linear dose response. To investigate this phenomenon, we studied cell survival and formation of 8-hydroxyguanine (8-OHG) base modifications and double strand breaks (DSB) caused by combined action of cadmium (Cd) and gamma radiation in cultured medaka fish (Oryzias latipes) fibroblast cells. Our data show that the introduction of Cd leads to a significant decrease in the fraction of surviving cells and to increased sensitivity of cells to ionizing radiation (IR). Cd also appears to cause non-linear increases in radiation-induced yields of 8-OHG and DSB as dose-yield plots of these lesions exhibit non-linear S-shaped curves with a sharp increase in the yields of lesions in the 10–20 μM range of Cd concentrations. The combined action of ionizing radiation and Cd leads to increased DNA damage formation compared to the effects of the individual stressors. These results are consistent with a hypothesis that the presence of Cd modulates the efficiency of DNA repair systems thus causing increases in radiation-induced DNA damage formation and decreases in cell survival.
INTRODUCTION
Relatively few mechanistic studies have been carried out on the interactions of radiation and chemicals in tissues or whole organisms. In part, this is due to the complexity of the changes in cellular processes perturbed by combinations of ionizing radiation and chemicals. How can we approach the design of mechanistic studies on combined exposures in model systems to help elucidate molecular mechanisms underlying a possible variety of additive, synergistic or antagonistic responses? One approach is to select a chemical agent that acts in cells similarly to ionizing radiation. In the present work we selected combinations of cadmium, a soluble metal ion found in numerous waste sites around the United States, and gamma radiation from Cesium-137, a radioactive isotope found in a number of the same waste sites.
Cd acts via a number of mechanisms, which have also been associated with the actions of ionizing radiation. For example, Cd can activate oncogenes (Fang et al. 2002), induce apoptotic cell death (Szuster-Ciesielska et al. 2000; Watjen et al. 2002; Pulido and Parrish 2003; Jimi et al. 2004), induce reactive oxygen radical species (Szuster-Ciesielska et al. 2000; McMurray and Tainer 2003; Waisberg et al. 2003; Bertin and Averbeck 2006), inhibit DNA repair (Yang et al. 1996; Hartwig et al. 2002a, b; Hartwig and Schwerdtle 2002; Waalkes 2003; Waisberg et al. 2003; Durham and Snow 2006; Giaginis et al. 2006), and induce metallothionein (MT) formation in tissues (Chan and Cherian 1992). The diverse mechanisms whereby Cd can induce damage, as well as alter biological responses caused by other contaminants, suggest that Cd can potentially affect all stages of the carcinogenic process, including initiation, promotion and progression (Waalkes 2003) just as has been shown for ionizing radiation.
Genotoxic and genomic studies on combined exposure to ionizing radiation and Cd
Very little is known about the combined effects of ionizing radiation and metals, such as Cd, on DNA damage and gene expression in mammalian cells or tissues. A recent study (Hornhardt et al. 2006) on human lymphoblastoid cells included exposures to combined levels of Cd and Cs gamma-rays. Their data on the formation of micronuclei as a marker suggest that a combination of Cd and radiation had an additive effect on the frequency of micronuclei formation. It was also shown that exposure to 10 μM Cd led to a significant increase in γ-radiation induced 8-OHG formation in human fibroblast GM00637 and HeLa S3 cells by inhibiting the expression of an 8-oxoguanine DNA glycosylase (Youn et al. 2005). A synergistic increase in enzyme activities of lactate dehydrogenase, alkaline phosphatase, and acid phosphatase in Wistar rats under combined action of Cd and γ-radiation was also reported (Salovsky et al. 1993). It has been suggested that Cd inhibits the repair of potentially lethal damage caused by UV–irradiation in cultured human and simian cells (Nocentini 1987). In combination with gamma irradiation, Cd has been shown to increase the level of metallothioneins in the bone marrow and liver of mice (Koterov et al. 1997). Increases in the levels of lipid peroxidation in tissues, accumulation of its end products in biosubstrates, and activation of catalase and peroxidase oxidoreductases in blood of test animals have also been shown (Honcharuk et al. 1995).
Cd fed to rats as a dietary supplement slightly enhanced the cytogenetic damage caused by chronic and acute γ-radiation and decreased the cytogenetic adaptive response induced by chronic γ-radiation. (Zaichkina et al. 2001). An antagonistic effect of the combined action of Cd and γ-radiation also was reported as mice treated with Cd 48 h prior to irradiation exhibited decreased levels of DNA damage in peripheral blood lymphocytes and splenocytes in comparison with untreated mice. (Privezentsev et al. 1996). In addition, the pre-irradiation administration of Cd to mice is reported to have accelerated repair processes in the bone marrow and spleen as well as increased the number of leukocytes and thrombocytes in the peripheral blood (Mackova et al. 1996).
It is possible that such broad and high Cd activity, associated with the ability of this metal to bind to proteins and DNA and to change the conformation of these macromolecules (Predki and Sarkar 1994; Asmuss et al. 2000a; Hartwig 2001; Kopera et al. 2004; Witkiewicz-Kucharczyk and Bal 2006), can lead to nonlinear or even hormetic dose-response relationships in combination with ionizing radiation.
In this paper we present the results of some of our initial studies on the combined effects of Cd and ionizing radiation exposures on DNA damage in cultured medaka fish fibroblast cells.
MATERIALS AND METHODS
Cell culture treatments
CAB fibroblast medaka fish (Oryzias latipes) cell line was provided by Dr. Wendy Kuhne (Colorado State University) with kind permission of Dr. Hiroshi Mitani (The University of Tokyo). Cells were incubated in T75 (Corning, NY) flasks in cell culture medium (L15 Leibovitz Medium (HyClone, Logan, UT), 20% Fetal Bovine Serum (HyClone, Logan, UT), 0.5 U/ml Penicillin – Streptomycin (HyClone, Logan, UT)) with 0 −50 μM CdCl2 at 30° C for 24 hrs. After a Cd treatment, they were γ-irradiated at the Colorado State University 137Cs gamma radiation facility at a dose-rate of 2.5 Gy/min with doses from 0 to 5 Gy for each Cd concentration. After irradiation, cells were immediately placed on ice. For Cd-only treatments cells were incubated with CdCl2 for 48 and 72 hours and used immediately. For the colony formation assay approximately 300 cells were plated in each 16mm well in 24-well tissue culture plates (Falcon; Becton Dickinson & Co, Oxnard, CA) and incubated with 0 −50 μM CdCl2 at 30° C for 24 hrs. Immediately following this treatment cells were γ-irradiated with doses from 0 to 15 Gy at a dose-rate of 2.5 Gy/min.
DNA isolation
DNA from the cells was extracted by the chaotropic NaI method (Helbock et al. 1998). Approximately 7.5 105 cell fractions were used for preparation of agarose plugs (Wang et al. 1995).
Colony formation assay
After treatment, cells were incubated for 14 days to allow for the growth of clonogenic colonies. Colonies were fixed with methanol and treated with Crystal Violet stain. Colonies with 50 or more cells were counted (Herskind et al. 2000; Potts et al. 2003; Akudugu et al. 2004; Nair et al. 2004; Youn et al. 2005; Franken et al. 2006).
Determination of 8-hydroxy-2′-deoxyguanosine (8-OHdG) levels in isolated DNA
The quantitation of 8-OHdG levels in the isolated DNA samples was carried out via high-performance liquid chromatography with electrochemical detection (HPLC-ECD) (Nicotera and Bardin 1998; Pouget et al. 2000). The assay performance was tested by quantitation of 8-OHdG formation in γ-irradiated aqueous solutions of salmon sperm DNA (Sigma, St. Louis). Aqueous solutions of 1μg/μl DNA in 10mM NaClO4 were γ-irradiated with 5 doses from one to 100 Gy at a dose rate of 30 Gy/hour. After enzymatic digestion of DNA to nucleosides (Helbock et al. 1998) HPLC analysis was done with a Waters 2690 Alliance System equipped with an Atlantis™ dC18 reversed-phase column and a Waters 2487 dual λ UV absorbance detector. An ESA Model 5600A CoulArray electrochemical detector with 8 electrochemical cells received the eluent from the UV detector and provided the ECD spectra used in the analysis. For each sample, the amount of DNA injected onto the column was estimated using the UV signal of unmodified nucleosides after appropriate calibration. The amount of 8-OHdG was determined using the ECD signal after appropriate calibration with a commercially available standard (Berry & Associates Inc., Dexter, MI). The radiation yield of 8-OHdG obtained from the yield – dose dependence was 1.1 ± 0.05 8-OHdG/105 dG Gy−1 and is in good agreement with values found in the literature (Dizdaroglu 1985; Fuciarelli et al. 1990; Douki et al. 1997; Svoboda and Harms-Ringdahl 1999).
Quantitation of double-strand breaks in isolated DNA
A CHEF Mapper XA Pulsed Field Electrophoresis System (Bio-Rad, Hercules, CA) was used for measurements of double strand break (DSB) formation. DNA fragments in the size region of 200 kb −6 Mb were separated according to a previously published protocol (Elia and Nichols 1993). DNA fragments in the 200 bp −200 kbp region were resolved by a modified method (Carle et al. 1986). Quantation of double-strand breaks was done by number average length analysis (Sutherland et al. 2003a, b; Gradzka and Iwanenko 2005).
Detection of apoptosis
Cells were incubated with several Cd concentrations in the 10–50 μM range at 30°C for 24 hrs. After incubation, cells were harvested and DNA was extracted with an Apoptotic DNA Ladder Extraction kit (BioVision, Mountain View, CA) and analyzed by electrophoresis on a 1.2% agarose gel (Kang et al. 2006). DNA extracted from medaka cells treated with Camptothecin (CAM) was used as positive control for apoptotic DNA fragmentation (Tonini et al. 2004).
Statistical analysis. All data displayed graphically are expressed as mean ± SD values obtained from 3–5 repeats. Linear regression analysis and non linear curve fitting were done with Microcal Origin software (OriginLab, Northampton, MS).
RESULTS AND DISCUSSION
Effect of Cadmium on Medaka Cell Survival and DNA Damage Formation
We have performed studies of Cd and combined Cd and γ-irradiation effects on medaka CAB (Oryzias latipes) fibroblasts. Endpoints measured consisted of cell survival, induction of apoptosis and quantitation of DNA damage markers (8OHdG and double strand break [DSB] formation).
Cell survival data resulting from Cd exposures of medaka fibroblast cells are presented in Figure 1. These data indicate that acute (24 hour) exposures of the cells to Cd concentrations up to 10 μM did not produce statistically significant changes in the survival fraction (SF). However, Cd concentrations in the range of 20 to 50 μM resulted in decreases in SF up to a factor of 30. (Fig. 1 [1]). The data were analyzed by use of a non-linear curve fitting analysis based on a quadratic model (Chadwick and Leenhouts 1973; Hethcote et al. 1976). The quadratic model provided an accurate (reduced chi-square < 0.005) fit to the experimental data.
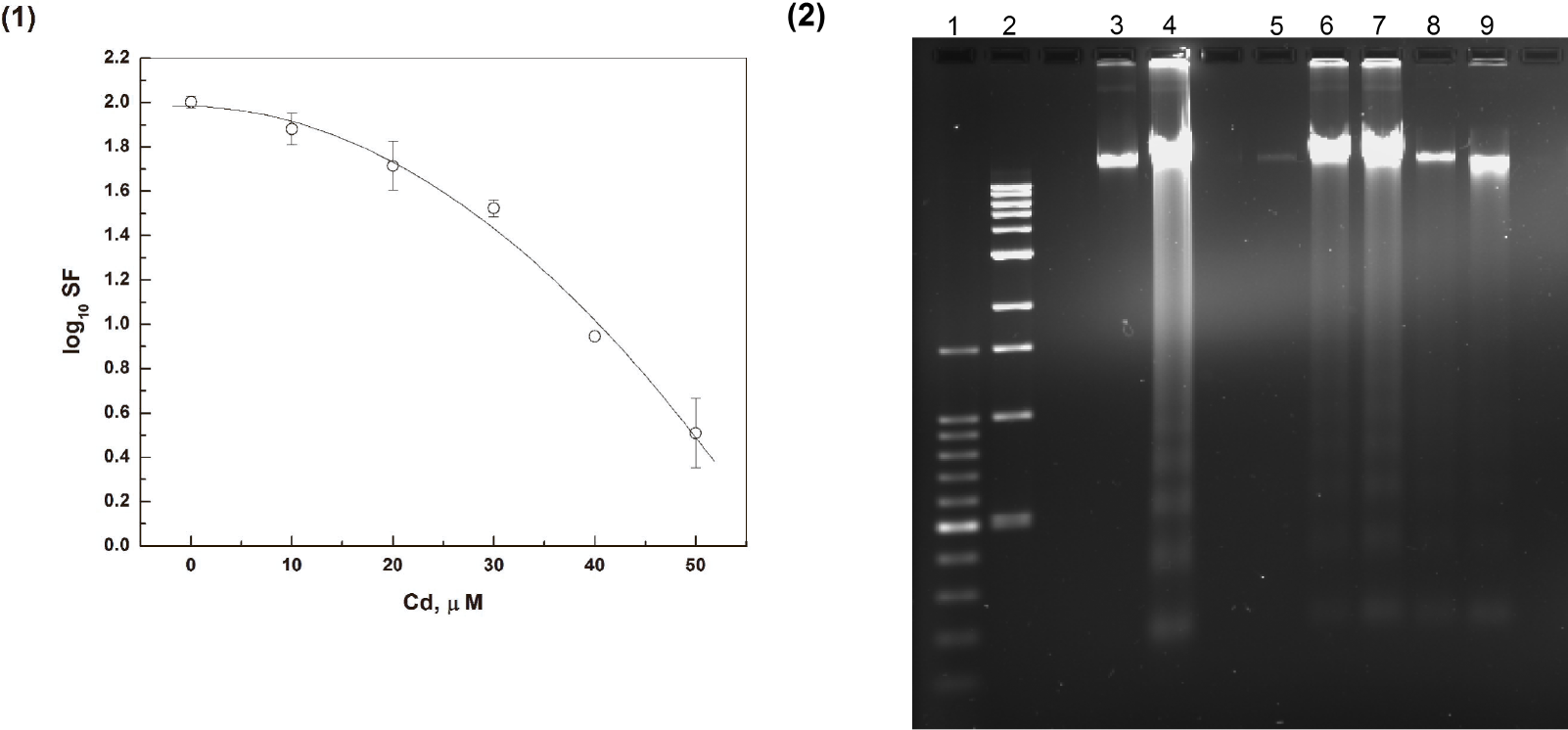
The effect of Cd on the survival of medaka cells: (1) The dependence of cell survival fraction (SF) on Cd concentration. The cells were incubated for 24 hours with various Cd concentrations; The data are fit very well by a quadratic model (Chadwick and Leenhouts 1973; Hethcote et al. 1976), reduced chi-square < 0.005; (2) Apoptotic medaka cell DNA fragmentation patterns caused by Cd exposures: Lane (1) −1 kb DNA Ladder; lane (2) −100 bp DNA Ladder; lane (3) - Control DNA from the medaka cells without any treatment; lane (4) - DNA from the medaka cells incubated with CAM; lanes (5) – (9) - DNA from the medaka cells incubated with 10, 20, 30, 40 and 50 μM of Cd correspondingly.
Gel electrophoresis results (Fig. 1 [2]) show apoptotic DNA fragmentation caused by incubation of the cells with Cd. The fragmentation takes place only if Cd concentrations are higher then 10 μM. DNA extracted from medaka cells treated with Camptothecin (CAM) was used as a positive control for apoptotic DNA fragmentation (Tonini et al. 2004).
Incubation of cells with Cd also leads to increases in levels of DNA damage markers (8-hydroxyguanine and DSB). This effect depended not only on concentration but also on exposure time. (Fig. 2).

The effect of Cd on the yield of DNA damage markers in treated medaka cells: (1) The dependence of the yield of 8OHdG on time of exposure to Cd (lesions per μM); (2) The dependence of the yield of DSB on time of exposure to Cd (lesions per μM).
Quantitation of DNA damage markers −8-OHG (Fig. 2 [1]) and DSB (Fig. 2 [2]) - in medaka cells indicated that the yields of DNA lesions per Cd concentration unit (μM) strongly depended on the exposure time.
Similar cytotoxic and genotoxic effects of Cd have been widely described in the literature (Latinwo et al. 1997; Mikhailova et al. 1997; McMurray and Tainer 2003; Filipic and Hei 2004; Bertin and Averbeck 2006; Martelli et al. 2006; Badisa et al. 2007). However, the underlying mechanisms that could be responsible for such effects are still not clearly understood. The toxicity produced by Cd exposures could be due to several simultaneous processes such as induction of apoptosis (Szuster-Ciesielska et al. 2000; Watjen et al. 2002; Pulido and Parrish 2003; Jimi et al. 2004), indirect generation of reactive oxygen species (Szuster-Ciesielska et al. 2000; Galan et al. 2001; McMurray and Tainer 2003; Waisberg et al. 2003; Watanabe et al. 2003), and interference with DNA repair (Yang et al. 1996; Asmuss et al. 2000b; Hartwig et al. 2002b; Hartwig and Schwerdtle 2002; Jin et al. 2003; Potts and Pasqualotto 2003; Potts et al. 2003; Waalkes 2003; Waisberg et al. 2003; Youn et al. 2005; Durham and Snow 2006; Giaginis et al. 2006). Moreover, Cd exposure can affect cell cycle progression, proliferation, differentiation, and DNA replication (Dong et al. 2001; Fang et al. 2002; Yang et al. 2004; Oh and Lim 2006).
The data presented in Fig. 1 indicate that Cd exposures induced apoptosis and reduction in SF in medaka fibroblast cells in a concentration-dependent manner. The induction of apoptotic cell death by Cd has also been demonstrated in some other mammalian and fish cell lines. (Achanzar et al. 2000; Szuster-Ciesielska et al. 2000; Lag et al. 2002; Watjen et al. 2002; Pulido and Parrish 2003; Jimi et al. 2004; Risso-de Faverney et al. 2004; Gonzalez et al. 2006). Such induction can occur through two main pathways: extrinsic and intrinsic. The extrinsic pathway is triggered by ligand-induced activation of the death receptors and the intrinsic pathway is induced by cellular stress signals (Watjen et al. 2002).
The presence of a shoulder on the survival-dose curve (Fig. 1 [1]) and the absence of detectable apoptosis (Fig. 1 [2]) at Cd concentrations up to 10 μM may be consistent with the involvement of metallothioneins. Metallothioneins – cysteine-rich heavy metal-binding proteins – can play a protective role against Cd toxicity (Andrews 2000). Cd-induced overexpression of metallothioneins has been detected in mammalian cells (Li et al. 2005) as well as in fish tissues (Woo et al. 2006). Overexpression of metallothioneins in the cells was shown to reduce the effects of Cd at lower (up to 10 μM) but not at higher (20–50 μM) concentrations (Li et al. 2005). Further, low concentrations of Cd can produce stimulation and hormesis effects under certain conditions (Weis and Weis 1986; von Zglinicki et al. 1992; Damelin et al. 2000; Gaddipati et al. 2003; Fulladosa et al. 2005).
The dose-dependent accumulation of unrepaired DNA damage and the concomitant increases in the damage yields per μM of Cd in the Cd-exposed cells with time (Fig. 2) appear to be consistent with a mechanism involving increases in reactive oxygen species (ROS) production and/or suppression of DNA repair by Cd similarly to previously reported results (Dally and Hartwig 1997; Mikhailova et al. 1997; Filipic and Hei 2004; Pan and Zhang 2006; Emmanouil et al. 2007).
Although Cd does not generate free radials directly (Lloyd et al. 1998; Valko et al. 2005), it can do so indirectly by facilitating increases in lipid peroxidation (Filipic and Hei 2004), decreases in intracellular glutathione content, decreases in activities of cellular antioxidant enzymes including superoxide dismutase, peroxidase and catalase, (Hussain et al. 1987; Waisberg et al. 2003), and increases in iron and copper concentrations (Koizumi et al. 1992; Casalino et al. 1997), all of which can lead to increasing levels of ROS.
On the other hand, Cd has been shown to inhibit DNA repair processes such as base-excision repair (BER) (Dally and Hartwig 1997; Potts et al. 2001; Potts et al. 2003; Youn et al. 2005), nucleotide-excision repair (NER), and mismatch repair (MMR) (Jin et al. 2003). Additionally, Cd inhibits repair of oxidative DNA damage by inhibiting the expression of 8-oxoguanine DNA glycosylase (Potts and Pasqualotto 2003; Potts et al. 2003; Youn et al. 2005). The data presented in the next section illustrate the interference of Cd with DNA repair systems.
Combined Effects of Cadmium and Ionizing Radiation on Cell Survival and Formation of DNA Damage
Cell survival data obtained from combined Cd and γ-irradiation exposures of medaka fibroblast cells are presented in Figure 3. It can be seen that a prior 24 hour exposure to Cd significantly increases the sensitivity of cells to γ-radiation (Fig. 3 [1]). The data were plotted by use of a nonlinear curve fitting analysis based on a quadratic model (Chadwick and Leenhouts 1973; Hethcote et al. 1976). The quadratic model provided an accurate (reduced chi-square < 0.07) fit to the experimental data. As Cd exposure concentrations are increased up to 30 μM, the median lethal radiation dose (LD50) decreases by a factor of 3 (Fig. 3 [2]). Further increases in the concentration of Cd from 30 to 50 μM no longer affect the LD50.

The effect of combined action of Cd and γ-radiation on the survival of medaka cells: (1) The dependence of the cell survival fraction (SF) on radiation dose after 24 hours incubation with various Cd concentrations. The data are fit very well by a quadratic model (Chadwick and Leenhouts 1973; Hethcote et al. 1976), reduced chi-square < 0.07; (2) The effect of Cd concentration on radiation Lethal Dose (LD50).
Medaka cells treated with combined Cd and γ-irradiation exhibit increases in levels of DNA damage markers compared with the levels observed after exposures to single contaminants. These increases can be seen in the bar graphs presented in Figures 4(1) and 4(2) as well as in the three-dimensional graphs shown in Figures 5 (1) and 5 (2).
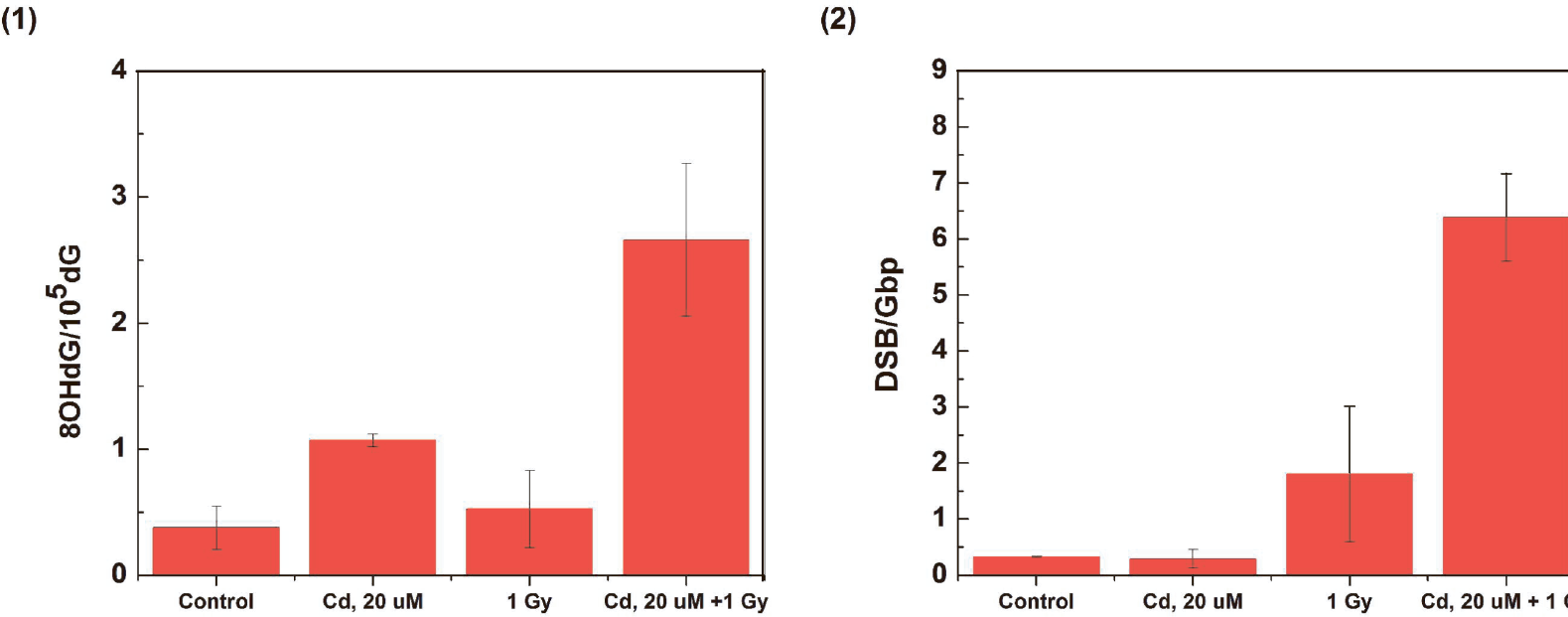
The effect of combined Cd and γ-radiation exposures on the levels of DNA damage markers in medaka cells: (1) Quantitation of 8OHdG lesions in medaka cells produced in control cells incubated for 24 hours; cells incubated for 24 hours with Cd (20 μM); cells irradiated with γ-rays (1 Gy); and cells treated with a combination of Cd and γ-rays; (2) Quantitation of DSB lesions produced in cells treated as shown on graph. The control is the background level of DSB frequency in the medaka cell DNA.

The effect of Cd exposures on the levels of DNA damage markers caused by ionizing radiation in treated medaka cells. The cells were incubated for 24 hours with various Cd concentrations prior to irradiation. (1) The dependence of the 8OHdG/10−5dG level on radiation dose and Cd concentration: X-axis - Cd concentration in μM; Y-Axis - radiation dose in Gy; Z-axis −8OHdG/10−5dG level. (2) The dependence of the DSB/Mbp on radiation dose and Cd concentration: X-axis - Cd concentration in μM; Y-Axis - radiation dose in Gy; Z-axis - DSB/Mbp.
In the Graphs shown in Figure 5, it can be seen that the slopes of the dose-response curves for the formation of both 8OHdG and DSB increase with increasing Cd concentrations.
The radiation-induced yields of 8OHdG and DSB in medaka cell DNA were determined by linear regression analyses from the slopes of the linear portions of the dose–response graphs for various Cd concentrations (Fig. 6). These dose-response graphs show that the Cd concentration dependence has a nonlinear character with curves for both 8OHdG and DSB yields being similar in shape. Increases in Cd concentration up to 10 μM do not produce changes in the yields of DNA damage markers. In the concentration range from 10 to 20 μM, the yields of markers increase sharply by a maximum factor of approximately two. No further significant changes are found for Cd concentrations above 20 μM.
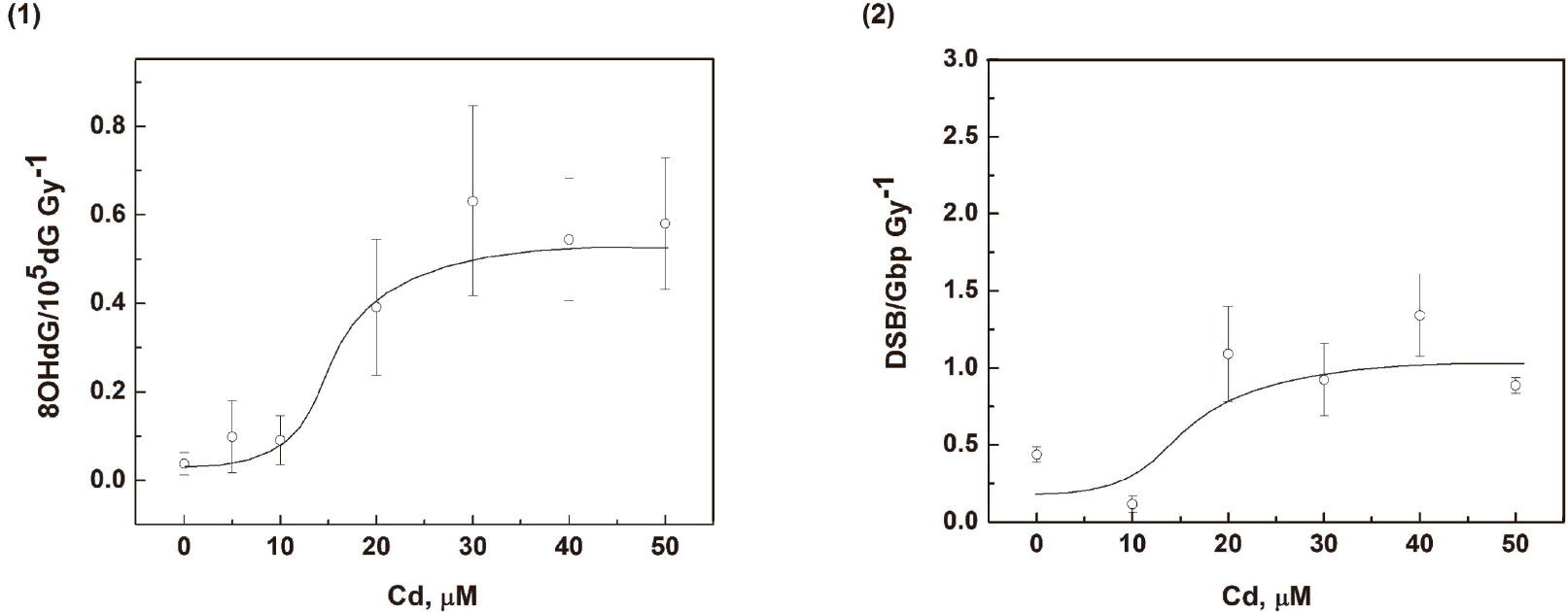
The effect of Cd exposure on the yield of DNA damage markers in treated medaka cells. The cells were incubated for 24 hours with various Cd concentrations prior to irradiation (1) The dependence of the radiation yield of 8OHdG (lesions per Gy) on Cd concentration.; (2) The dependence of the DSB radiation yield (lesions per Gy) on Cd concentration.
As mentioned above, Cd can impair the efficiency of DNA repair systems, which will lead to increases in the sensitivity of cells to ionizing radiation and in the yields of radiation-induced DNA damage as the concentration of Cd is increased. (Fig. 3 [2]; Fig. 5). Higher yields of DNA damage observed after a combined exposure of cells to Cd and ionizing radiation compared with the yields observed after exposures to the single agents may be consistent with the action of two parallel processes - the indirect formation of additional ROS by Cd along with the direct radiation-induced formation of ROS and a greater reduction in efficiency of repair of DNA damage caused by both radiation and Cd-generated ROS.
It has been shown that Cd can reduce or even inhibit DNA binding of a number of proteins involved in DNA repair processes (Bertin and Averbeck 2006) such as xeroderma pigmentosum A (XPA) protein required for recognition of DNA lesions in NER system (Hartmann and Hartwig 1998; Hartwig 1998; Hartwig et al. 2002a; Hartwig and Schwerdtle 2002); formamidopyrimidine glycosylase (Fpg) that recognizes and repairs 8-oxo-7,8-dihydroguanine base modifications in the Base Excision Repair (BER) system in Escherichia coli (O'Connor et al. 1993), a mammalian homologue of Fpg (Potts et al. 2003) 8-oxoguanine-DNA glycosylase-1 (OGG1), and specificity protein-1 (Sp1) transcription factor (Watkin et al. 2003). The mechanism underlying the ability of Cd to inhibit binding of DNA repair proteins is consistent with displacement of zinc (Zn) by Cd in the Zn finger structure of these proteins (Predki and Sarkar 1994; Asmuss et al. 2000a; Hartwig 2001; Kopera et al. 2004; Witkiewicz-Kucharczyk and Bal 2006). Also, Bertin and Averbeck (Bertin and Averbeck 2006) have noted that proteins thought to be involved in homologous recombination (HR) and non-homologous end joining (NHEJ) such as BRCA1 (Miki et al. 1994), ATR (Villard et al. 1997), and DNA ligase III alpha and beta (Martin and MacNeill 2002) can be inhibited by Cd. It has not yet been demonstrated (Bertin and Averbeck 2006) that Cd can inhibit the repair of DNA DSBs in mammalian cells or tissues, although such inhibition has been observed in the repair of single-strand breaks (SSB) induced by external agents (Littlefield et al. 1994; Lynn et al. 1997; Pruski and Dixon 2002; Fatur et al. 2003; Hengstler et al. 2003). In contrast, inhibition of DNA SSB repair in HeLa cells by Cd was not detected (Dally and Hartwig 1997).
We have demonstrated in this study that Cd not only causes the formation of DNA DSB in medaka cells (Fig. 4 [2]) but also affects DSB repair (Fig. 6 [2]). In the latter figures, the dose-response curves for radiation-induced yields versus Cd concentration are non-linear for both 8OHdG and DSB yields up to 10 μM of Cd followed by sharp increases in yields in the narrow concentration range from 10 μM to 20 μM and then followed by saturation of the yields in the 20–50 μM interval. The finding that both dose-response relationships have similar character suggests that the underlying mechanisms can be similar or the same. For example, one such mechanism could involve the competitive displacement (Scatchard 1949) of Zn by Cd in the Zn finger structures. As stated previously, overexpression of metallothioneins can reduce the effects of Cd at relatively low concentrations up to 10 μM Cd (Li et al. 2005) and can be responsible for the 0–10 μM Cd threshold observed in Fig. 6.
CONCLUSIONS
Summarizing the data presented above we conclude that Cd exposures significantly reduce the survival of medaka fish fibroblast cells and cause concentration-dependent increases in DNA damage markers such as 8OHG and DSB. Further, Cd decreases the observed LD50 of γ-irradiated cells and increases the radiation yields of the DNA damage markers. The dose-response curves for these effects are nonlinear. The mechanisms underlying these dose-responses likely involve several processes including the interaction of Cd with metallothioneins and competitive binding of Cd to proteins involved in DNA repair systems.
Footnotes
ACKNOWLEDGMENTS
This work was supported by NIH/NCI T32 Training Grant CA-09236-29
