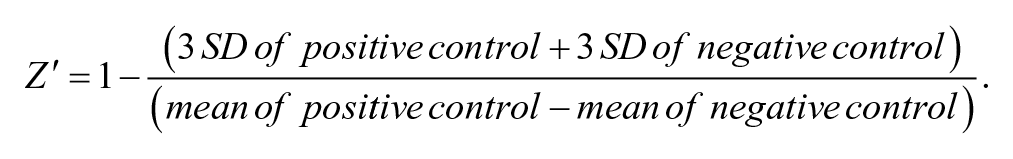Abstract
Atherosclerosis is the pathological basis of most cardiovascular diseases. Reverse cholesterol transport (RCT) is a main mechanism of cholesterol homeostasis and involves the direct transport of high-density lipoprotein (HDL) cholesteryl ester by selective cholesterol uptake. Hepatic scavenger receptor class B member 1 (SR-BI) overexpression can effectively promote RCT and reduce atherosclerosis. SR-BI may be an important target for prevention or treatment of atherosclerotic disease. In our study, we inserted human SR-BI mRNA 3′ untranslated region (3′UTR) downstream of the luciferase reporter gene, to establish a high-throughput screening model based on stably transfected HepG2 cells and to screen small-molecule compounds that can significantly enhance the mRNA stability of the SR-BI gene. Through multiple screenings of 25 755 compounds, the top five active compounds that have similar structures were obtained, with a positive rate of 0.19%. The five positive compounds could enhance the SR-BI expression and uptake of DiI-HDL in the hepatocyte HepG2. E238B-63 could also effectively extend the half-life of SR-BI mRNA and enhance the SR-BI mRNA and protein level and the uptake of DiI-HDL in hepatocytes in a time-dependent and dose-dependent manner. The structure-activity relationship analysis showed that the structure N-(3-hydroxy-2-pyridyl) carboxamide is possibly the key pharmacophore of the active compound, providing reference for acquiring candidate compounds with better activity. The positive small molecular compounds obtained in this study might become new drug candidates or lead compounds for the treatment of cardiovascular diseases and contribute to the further study of the posttranscriptional regulation mechanism of the SR-BI gene.
Keywords
Introduction
As an independent risk factor for cardiovascular disease, plasma high-density lipoprotein (HDL) cholesterol level (HDL-C) is part of the diagnostic criteria for metabolic syndromes and an important target for treatment and prevention of cardiovascular disease.1–6 The main form of anti-atherosclerotic function by HDL is through reverse cholesterol transport (RCT).6,7 RCT is an important part of cholesterol and lipoprotein metabolism. This process involves transport of excessive cholesterol from peripheral tissues (including atherosclerotic plaque) to the liver for recirculation or excretion, which can reduce lipid deposition on blood vessel walls.1–3,7 In addition to RCT, HDL has a variety of anti-atherosclerotic functions including anti-inflammation, antioxidation, inhibition of thrombosis, and improvement of endothelial cell functions.2,3,6–8 Therefore, the HDL functions and the correlation between HDL metabolism and atherosclerosis have become a research focus for the treatment of atherosclerotic cardiovascular diseases.1–3,6–8 The treatment of atherosclerotic diseases targeting HDL and its relevant receptor has received widespread attention.1–3,6–10
Human scavenger receptor class B, member 1 (SR-BI or SCARB1), also known as CD36 and lysosomal integral membrane protein II analogous 1 (CLA-1) in humans, is an important integral membrane glycoprotein of the scavenger receptor family.1–3,11 SR-BI is mainly expressed in the liver, gastrointestinal tract, and steroidogenic organs or is detected in macrophages and endothelial cells.3–5,11 Human SR-BI (CLA-1) is composed of 509 amino acid residues and shows up to 70% to 80% sequence homology across mammalian species. An extracellular domain structure containing N-glycosylation sites makes up the main part of SR-BI. In addition, two transmembrane domains and the short amino and carboxy-terminals in the cells constitute the remaining part of the molecule. 12 SR-BII is an mRNA splice variant of SR-BI with a unique carboxy-terminus.13,14 The key mechanism of HDL-C transport is the selective uptake of the cholesterol esters.1–3,6–10 As a functional HDL receptor with high expression level in the liver, SR-BI binds with HDL to mediate the selective uptake of HDL-C with high affinity and high saturation, and this process is the final rate-limiting step of RCT.1–4,6 The overexpression of hepatocyte SR-BI can indeed promote the reverse transport of HDL-C, enhance the HDL clearance rate in plasma, and reduce the morbidity and mortality of atherosclerosis and associated cardiovascular risk events.1–3,6–10 Given that SR-BI plays an important role in lipid metabolism and atherosclerosis pathogenesis, the development of new anti-atherosclerotic drugs for the treatment and prevention of cardiovascular disease using SR-BI as the target has aroused widespread attention.3,7,9,11,15
The posttranscriptional level of gene regulation is one of the main features of regulation of eukaryotic gene expression.16,17 The primary transcription product formed after transcription should be converted to mature mRNA after a series of processes, to be used as a template for protein translation.16,17 Abundance of gene expression is determined by rates of mRNA transcription and decay.17,18 The 3′ noncoding region of eukaryotic mRNA contains important regulatory sequences affecting the fate of posttranscriptional mRNA and protein biosynthesis.17,19–21 More than half of the mammalian genes produce different subtypes of mRNA due to the difference in 3′ untranslated region (3′UTR) and regulatory sequence, and a long 3′UTR shows a strong posttranscriptional level and regulatory functions at the translation level.17,20,22 3′UTR plays a crucial role in self-regulation and the highly complex regulatory systems such as competitive or synergistic microregulation of protein biosynthesis in an environment-dependent manner. 3′UTR not only is involved in the regulation of in vivo stability, the degradation rate of eukaryotic mRNA, and the control of the utilization efficiency but also can regulate the mRNA translation time, location, products, and efficiency.17,19,20,23,24 The eukaryotic gene mRNA 3′UTR has become an important target of metabolic research and drug discovery.17,20,25 By acting on stable low-density lipoprotein (LDL) receptor mRNA in the 3′UTR region, the natural medical monomer compound berberine can significantly upregulate the expression of LDLR, showing positive cholesterol-lowering activity, which is different from the mechanism of action of the lipid-lowering drugs such as statins.26,27 microRNA, which has broad regulatory functions, can directly act on the 3′UTR of target genes to perform its function. 28
In the present study, we set up a high-throughput screening (HTS) assay using stably transfected HepG2 cells with SR-BI mRNA 3′UTR in the downstream of the luciferase reporter gene (LUC), discovered and confirmed several small molecular compounds with the effect of enhancing the mRNA stability of SR-BI, and identified their roles in regulating SR-BI expression and functions.
Materials and Methods
Cell Culture
Human hepatoma HepG2 cells were cultured in Minimum Essential Medium supplemented with 10% fetal bovine serum (FBS; Hyclone, Logan, UT). Cells were grown in 5% CO2 at 37 °C in a humid atmosphere.
The stable transfected cell line CLAp-LUC HepG2 cells for luciferase reporter gene expression with human SR-BI upstream regulatory sequences (–1055 bp ~ –62 bp, with gene transcription starting site as +1) were constructed and preserved in our laboratory.29–32
Human SR-BI upstream regulatory sequences and SR-BI mRNA 3′UTR (+1 bp ~ 959 bp, with the first base after SR-BI mRNA termination codon as +1) were integrated to the upstream and downstream of LUC, respectively, to construct the stably transfected cell lines 3′UTR-LUC HepG2 cells.33,34
Analysis of Activity of Luciferase Reporter
The cells used in determining the luciferase activity were treated by the Luciferase Assay System (Promega, Madison, WI) according to the manufacturer’s instructions. The luciferase activity was detected by the multifunction microplate detector EnVision 2104-0010 (PerkinElmer, Waltham, MA).
High-Throughput Screening
Preparation of “5-in-1” compound samples
Compound samples were dissolved in 100% DMSO to prepare 10 mg/mL concentrations of stock solution. The five compound stock solutions were then mixed in equal volume, to obtain “5-in-1” compound samples (2 mg/mL of each compound in 100% DMSO). Before screening, the 5-in-1 compound samples were diluted 10-fold in distilled water to reach a final concentration for each compound of 200 µg/mL in 10% DMSO. The drug libraries of 25 755 compounds identified in this prior study were acquired from National Center for Microbial Drug Screening, Institute of Medicinal Biotechnology (Beijing, China).
Primary Screening
CLAp-LUC and 3′UTR-LUC HepG2 cells in the logarithmic growth phase were digested to single-cell suspension by trypsin. Cells were inoculated to 96-well plates at a cell density of 5 × 105 cells/mL, 100 µL per well. The complete cell culture medium was removed, and cells were rinsed once with phosphate-buffered saline (PBS) 6 h later. A total of 100 µL serum-free medium was added to each well, and then 1 µL tested compound sample was added, respectively, to get the final compound concentration of 2 µg/mL and DMSO concentration of 0.1%. At the same time, the control wells were established, and 0.1% of the DMSO was added. The luciferase activity was determined 20 h later.
First, the luciferase activity of sample wells versus control wells was defined as the upregulation rate of luciferase activity by the compounds. After deducting the upregulation activity of the compound on SR-BI promoter, the effect of the compound on the expression activity of LUC through the downstream SR-BI 3′UTR fragments of the reporter gene was obtained. Thus, the activity index of the compound was defined as the upregulation rate of luciferase activity by the compounds on 3′UTR-LUC HepG2 cells versus CLAp-LUC HepG2 cells.
According to this formula, compounds with an activity regulation rate ≥1.5 were defined as positive samples.
Secondary Screening
After the 5-in-1 positive compound samples were identified, the corresponding individual compound stock solutions of positive compound samples were reassayed, and the regulation rate of individual samples was determined according to the above method, so as to identify the active component (hit) in the 5-in-1 mixture. The effect of the positive compound on the cell growth state was also observed, and compound samples with apparent cytotoxicity were excluded from further testing.
The dose-response relationship between compound effect concentration and luciferase activity regulation rate was analyzed, and the EC50 value of the compound specimen was analyzed using Sigmaplot 12.5 (SyStat Software, San Jose, CA).
The parameters for use in evaluation and validation of HTS assays for hit identification mainly covers signal-to-background ratio
Analysis of mRNA Expression Levels
The FastStart Universal SYBR Green PCR Master kit and real-time quantitative PCR system (Roche, Mannheim, Germany) were used for detection according to the operation manual. cDNA synthesized by reverse transcription was used as template in real-time PCR. In half-life analysis GAPDH, peptidylprolyl isomerase A and beta-2-microglobulin served as reference genes. The classic and simple Livak method, the 2–∆∆Ct method, was used to analyze the relative gene expression level. 38 All of the oligonucleotide primer sequences used in the study are listed in Supplementary Table S1 and had validated efficiencies within 100% ± 5%.
Western Blot Analysis
Cells were collected and rinsed with ice-cold PBS. RIPA cell lysis buffer (25 mM Tris-HCl, 150 mM NaCl, 1% NP-40, 1% sodium deoxycholate, 0.1% sodium dodecyl sulfate, pH 7.6) was added on the ice to lyse the cells for 20 min. Protein sample loading buffer was added, and the sample was boiled for 10 min to prepare the protein specimen. Sodium dodecyl sulfate–polyacrylamide gel electrophoresis was implemented by adopting 10% separation gel. Then, the electrophoretic band was transferred to the polyvinylidene difluoride (PVDF) membrane and dipped into confining liquid and sealed for 1 h at ambient temperature. The primary antibodies (#610883, BD Biosciences, San Jose, CA) of the target protein and reference protein, respectively, were diluted with confining liquid (1:1000), incubated for 1 h at ambient temperature, or incubated overnight at 4 °C. The HRP-labeled secondary antibodies (#3700, Cell Signaling Technology, Beverly, MA) were diluted with confining liquid (1:1000) and incubated with membrane at ambient temperature for 1 h. Luminol and peroxide reagent (Millipore, Bedford, MA) were mixed with equal volume to prepare the luminescence liquid, and the luminescence liquid was dropped onto one PVDF membrane and transferred with protein, and it was exposed in the ChemiDocXRS (Bio-Rad, San Diego, CA). The band was scanned by Image Lab 4.0 (Bio-Rad) in gray scale for treatment and analyzed, and it was then standardized to the gray value of the reference protein band for quantitative measurement.
Analysis of DiI-HDL Uptake
DiI (1,1′-dioctadecyl-3,3,3′,3′-tetramethylindocarbocyanine perchlorate) is one of the most common cytomembrane fluorescent probes, presenting orange red fluorescence. After DiI was combined with a phospholipid double-layer membrane, the longest activated wavelength was 549 nm, and the longest emission wavelength was 565 nm. The cell was cultivated in 24-well plates to be detected and rinsed with PBS twice, and then 2 µg/mL DiI-HDL (Biomedical Technologies, Stoughton, MA) diluted by serum-free medium was added and then incubated at 37 °C for 4 h. The cell was rinsed with cold PBS once and treated with trypsinization. The cell was gently pipetted so that a single-cell suspension was formed. Collected cells were suspended in 0.5 mL PBS again, passed through a 300-mesh nylon net, and detected with a flow cytometer (BD Biosciences).30,31,34
Determination of mRNA Half-Life
Actinomycin D (Act D) can be implanted into the minor groove of a DNA double helix and form a heteroion with DNA, hindering transcription of RNA polymerase and restraining RNA compounds.
HepG2 single-cell suspension (1 mL, 1.0 × 105 cells/mL) was inoculated in a 12-well plate and cultivated in 10% FBS complete medium for 24 h. The culture solution was discarded, and the compound to be detected and blank reference substance were added, respectively, with a specified concentration diluted with serum-free medium and cultivation continued for 24 h. The old culture solution was discarded, and 5 μg/mL Act D diluted with serum-free medium was added and then cultivated under the conditions of 37 °C and 5% CO2. After treating with Act D to the specified time, the cells were collected and stored at −80 °C after being lysed. After all the cells were treated, total RNA was extracted and cDNA was synthesized, followed by quantitative reverse transcriptase PCR (qRT-PCR).
The formula of mRNA half-life
Statistical Analysis
Experimental data are expressed as mean ± SD, and a t test was adopted to carry out statistical analysis with pairwise comparison. One-way analysis of variance was adopted to analyze multiple comparisons. Data analyses were conducted using SPSS Statistics 19.0 (IBM), Sigma Plot 9.0, and OriginPro 8.0. Significance was set at p < 0.05.
Results
Analysis of SR-BI 3′UTR Structure and the Effect of Its 3′UTR on the mRNA Half-Life of LUC
The secondary structures of SR-BI 3′UTR predicted by RNAdraw and RNAstructure are highly similar. The predicted results showed that SR-BI 3′UTR can be folded as around 71 ring-loop structures, with the unfolded area often being where microRNA and others are apt to bind ( Fig. 1A, B ). The results of multiple species sequence alignment showed that SR-BI 3′UTR possessed a sequence conservation property at about six sections. Different truncation-length SR-BI 3′UTR were inserted into the downstream of the LUC to construct five reporter plasmids, and HepG2 cells were transiently transfected, then their influence on the expression activity of the luciferase was detected ( Fig. 1C, D ).
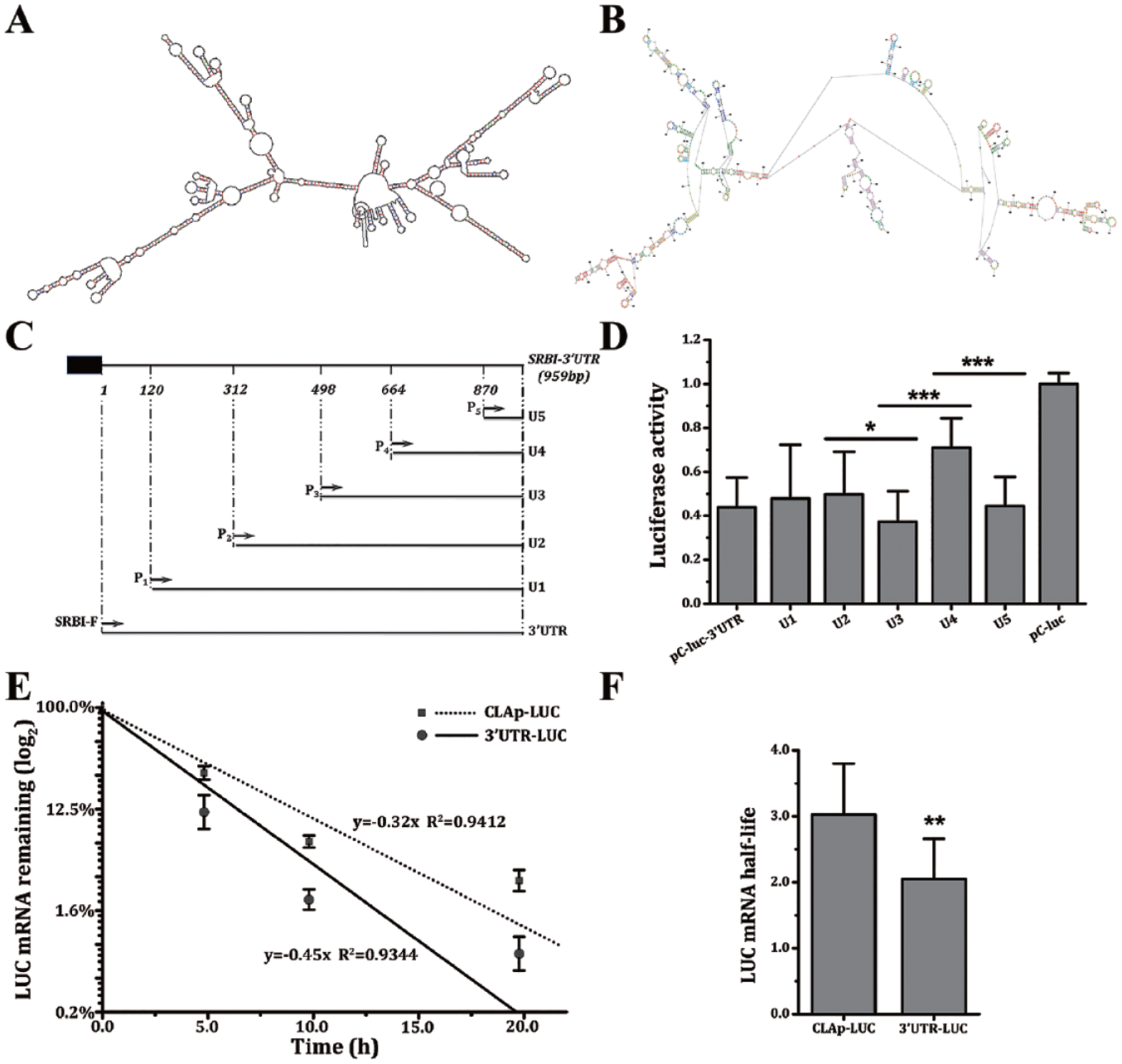
Analysis of scavenger receptor class B member 1 (SR-BI) 3′ untranslated region (3′UTR) structure and the effect of its 3′UTR on the mRNA half-life of LUC. The secondary structure of SR-BI mRNA 3′UTR predicted by RNAdraw (
The moderating effects of 3′UTR, U1, U2, and U5 on the activity of the luciferase reporter are similar. Plasmid construction has been described in our previous study. 34 Comparing U2 to U3, after the fragment of 312 bp ~ 498 bp was deleted, the activity of luciferase was greatly reduced. Comparing U3 to U4, after the sequence between 498 bp and 664 bp was deleted, the luciferase activity significantly increased. Comparing U4 to U5, after the 664 bp ~ 870 bp fragment was deleted, the expression activity of the LUC significantly decreased ( Fig. 1D ). This result indicated there are different cis-elements at 3′UTR influencing the stability of SR-BI mRNA.
CLAp-LUC and 3′UTR-LUC HepG2 cells were treated with Act D to block mRNA transcription, and then the rate of LUC mRNA degradation was investigated. After 5 h of treatment with Act D, the levels of mRNA were significantly changed. The LUC mRNA half-life of the 3′UTR-LUC cell line was 2.05 h, and the LUC mRNA half-life of the CLAp-LUC cell line was 3.02 h, which shows that the inserted SR-BI 3′UTR segment was able to significantly lower the stability of LUC mRNA ( Fig. 1E, F ).
High-Throughput Screening
Establishment and evaluation of the screening model
CLAp-LUC and 3′UTR-LUC HepG2 cells are used for the establishment of HTS assays for discovery of small molecules enhancing SR-BI mRNA stability. The activity of luciferase is regulated by the promoter and 3′UTR of SR-BI in the HTS assays using the cell line 3′UTR-LUC. After deducting the effect of the compound on the SR-BI promoter region in CLAp-LUC cells, the increase in luciferase activity in the 3′UTR-LUC HepG2 cells indicated the enhancement of human SR-BI mRNA stability upregulated by positive compounds through targeting SR-BI 3′UTR.
Cells were seeded in 96-well plates at a cell density of 5 × 105 cells/mL and 5 × 104 per well, and the concentration of compounds used in the primary 5-in-1 screen was 2 µg/mL.
Currently, there is no small molecular compound that has been verified to enhance SR-BI mRNA stability through targeting SR-BI 3′UTR. Thus, we initially used miR-185 inhibitor,33,34 an antisense oligonucleotide to miR-185, as the positive control, and evaluated the parameters of the HTS assay system. The results of the evaluation showed that the screening model used in this study is reliable and sensitive and can be applied in large-scale screening to find new molecules capable of enhancing the stability of human SR-BI mRNA ( Suppl. Table S2 ).
Application and reevaluation of the model
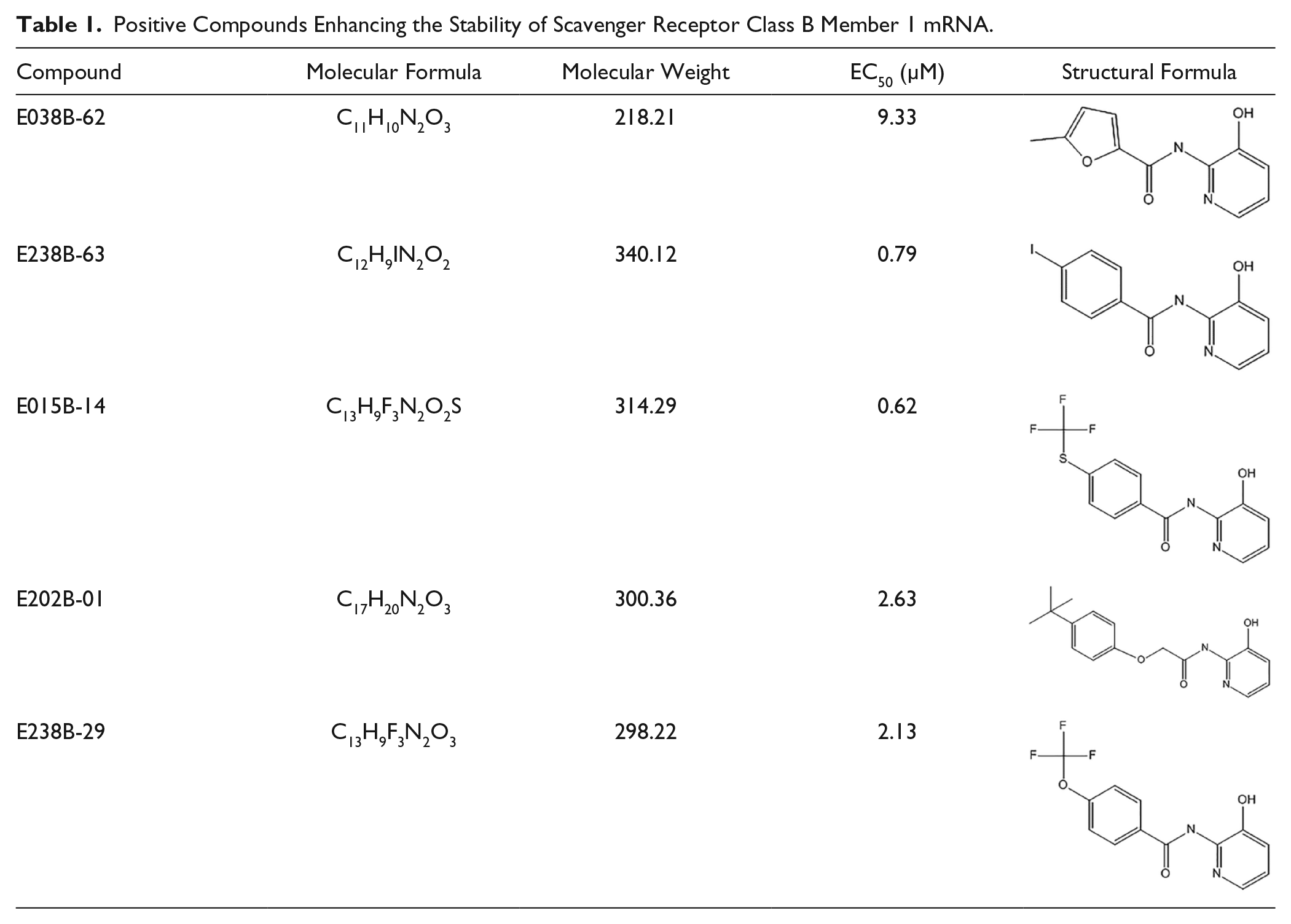
Applying the HTS system, large-scale screening was carried out on 25 755 compounds. Through primary screening, secondary screening, and further verification, we obtained five active compounds, with a positive rate of 0.19% ( Table 1 ). These 5 positive compounds were diluted 10-fold, then their activity was verified and their EC50 calculated. E238B-63 is the one of the most active compounds, and its dose-response relationship curves are shown in Figure 2A .
Positive Compounds Enhancing the Stability of Scavenger Receptor Class B Member 1 mRNA.
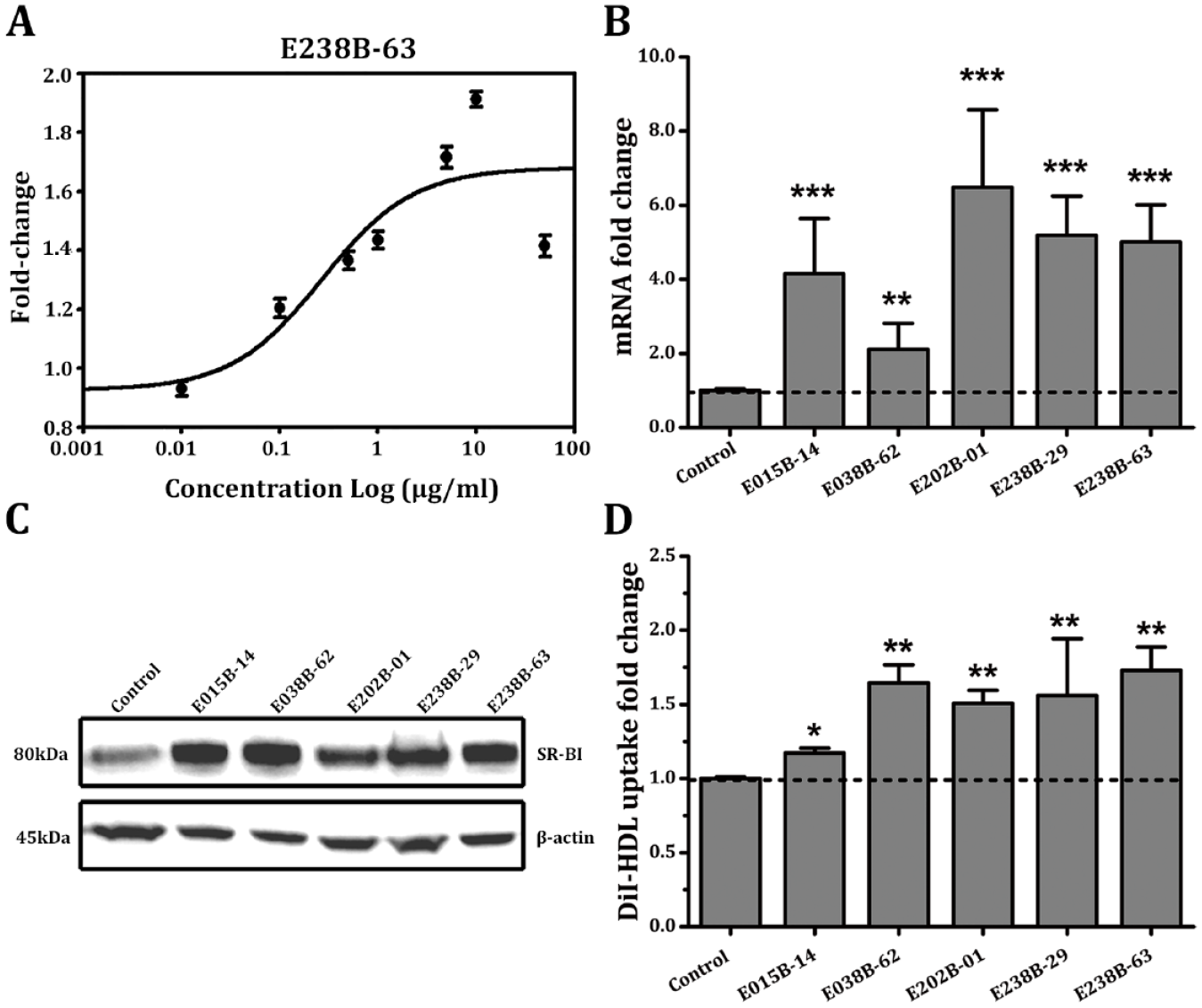
Effects of positive compounds on scavenger receptor class B member 1 (SR-BI) expression and high-density lipoprotein (HDL) uptake in hepatocytes. The dose-response relationship curve of E238B-63 (
The positive compound E238B-63 was used to reevaluate the parameters of the HTS assay system, resulting in an improved Z′ factor, and the relative parameters were conformed to the requirements of HTS assays ( Suppl. Table S3 ).
Effect of the positive compounds on hepatocyte SR-BI expression and HDL uptake
After the treatment of HepG2 cells with five positive compounds at a final concentration of 1 µg/mL for 24 h, respectively, the mRNA and protein expression levels of SR-BI and the uptake of DiI-HDL in the hepatocytes were tested. The results showed that the five positive compounds can significantly upregulate SR-BI mRNA and protein expression level by more than twofold and significantly increase the uptake of DiI-HDL in HepG2 cells ( Fig. 2B–D ). From this comprehensive investigation of the effect of the five positive compounds on SR-BI expression and HDL uptake in HepG2 cells, E238B-63 was chosen for further study.
Dose-response relationship of E238B-63
To determine whether E238B-63 affects mRNA and protein expression and the uptake of DiI-HDL in hepatocytes in a dose-dependent manner, concentrations of 0.1 µg/mL, 1 µg/mL, and 10 µg/mL of the compound were selected to treat HepG2 cells for 24 h, and then the mRNA and protein expression level and DiI-HDL uptake activity were investigated ( Fig. 3A–C ).
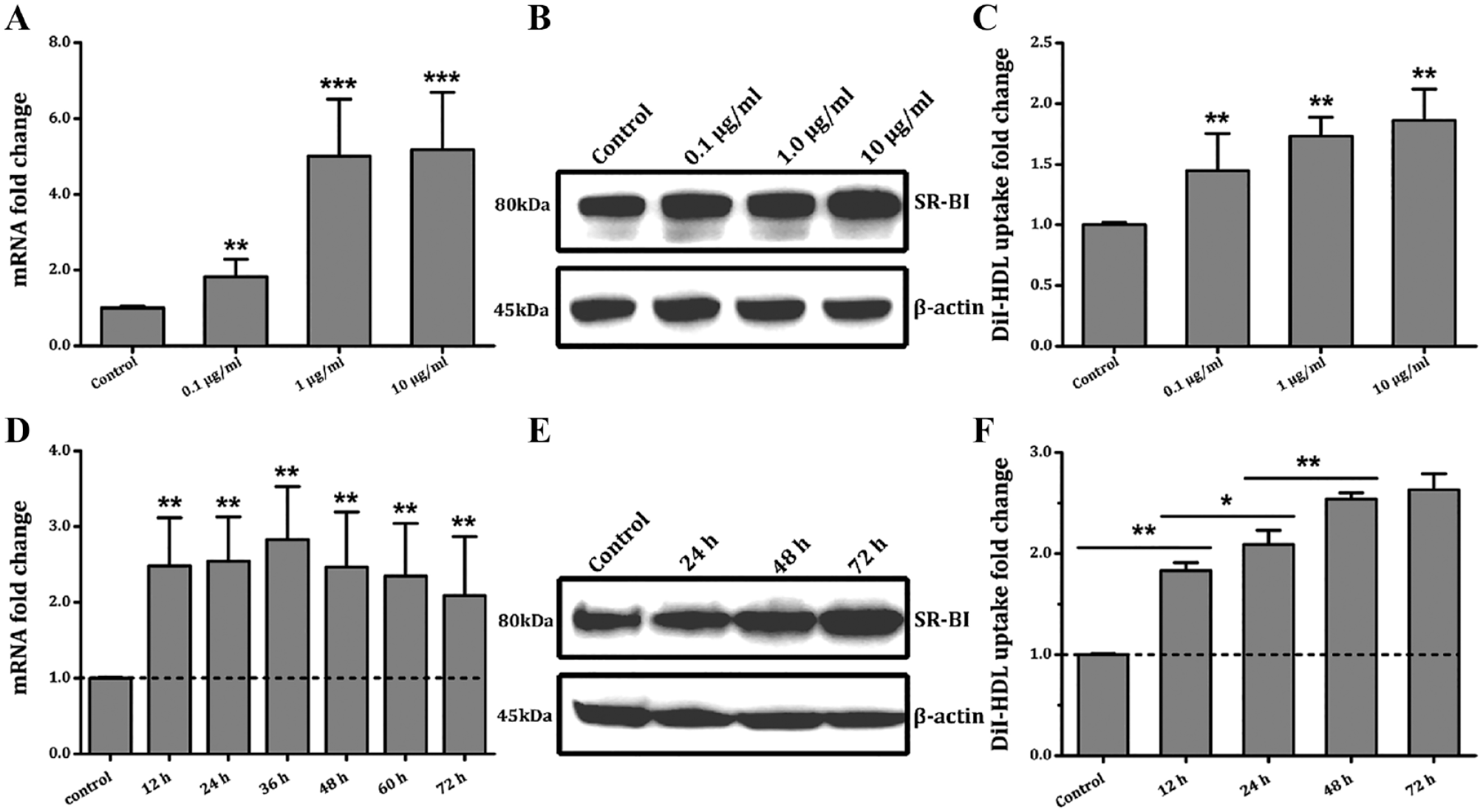
The compound E238B-63 affects the expression of scavenger receptor class B member 1(SR-BI) and high-density lipoprotein (HDL) uptake in hepatocytes in a dose-dependent and time-dependent manner. The effects of E238B-63 on SR-BI mRNA (
At E238B-63 concentrations of 0.1 µg/mL, 1 µg/mL, and 10 µg/mL, SR-BI mRNA levels reached 1.82, 5.01, and 5.17 times control levels, respectively ( Fig. 3A ). The Western blotting results showed that the protein expression of SR-BI in HepG2 cells also had a certain positive correlation with the concentration of E238B-63 ( Fig. 3B ). DiI-HDL uptake was also increased in EB238B-63–treated HepG2 cells in a concentration-dependent manner, as indicated by 1.45-, 1.73-, and 1.87-fold increases in Dil-HDL uptake at increasing concentrations of this compound ( Fig. 3C ).
Time-effect relationship of E238B-63
The effects of E238B-63 on the SR-BI expression level and DiI-HDL uptake activity in HepG2 cells at different treatment times were investigated ( Fig. 3D–F ). mRNA levels were significantly increased after exposure at 1 µg/mL for 12 h, which persisted for at least 72 h ( Fig. 3D ). The protein expression level was highest with exposure to compound E238B-63 at 1 µg/mL for 72 h ( Fig. 3E ). Consistently, after exposure to E238B-63 at 1 µg/mL for 48 h, the HDL uptake by HepG2 cells reached almost the highest level ( Fig. 3F ).
Effect of E238B-63 on mRNA Half-Life
Effect on LUC mRNA half-life
In the above experiment, we demonstrated that SR-BI 3′UTR could significantly reduce the stability of its upstream coding gene LUC mRNA ( Fig. 1E, F ). To investigate whether E238B-63 regulates the stability of the corresponding transcript through the SR-BI 3′UTR fragment, we measured the changes in half-life of LUC mRNA of CLAp-LUC cells and 3′UTR-LUC cells after exposure to E238B-63.
After the E238B-63 treatment of CLAp-LUC and 3′UTR-LUC HepG2 cells at a concentration of 5 µg/mL and blockage of mRNA transcription by Act D, mRNA levels at different times were detected by qRT-PCR. The results showed that, for the CLAp-LUC cells without SR-BI 3′UTR downstream of the reporter gene ( Fig. 4A, B ), the mRNA half-life of the LUC after exposure to E238B-63 (2.18 h) was not significantly different from that of the control (3.02 h). For the 3′UTR-LUC HepG2 cells with SR-BI 3′UTR downstream of the LUC, the mRNA half-life of the control was 2.05 h; after exposure to E238B-63, the half-life of LUC mRNA was 8.99 h, which was significantly increased ( Fig. 4C, D ). This suggests that E238B-63 can regulate the corresponding transcript stability through the SR-BI 3′UTR fragment.
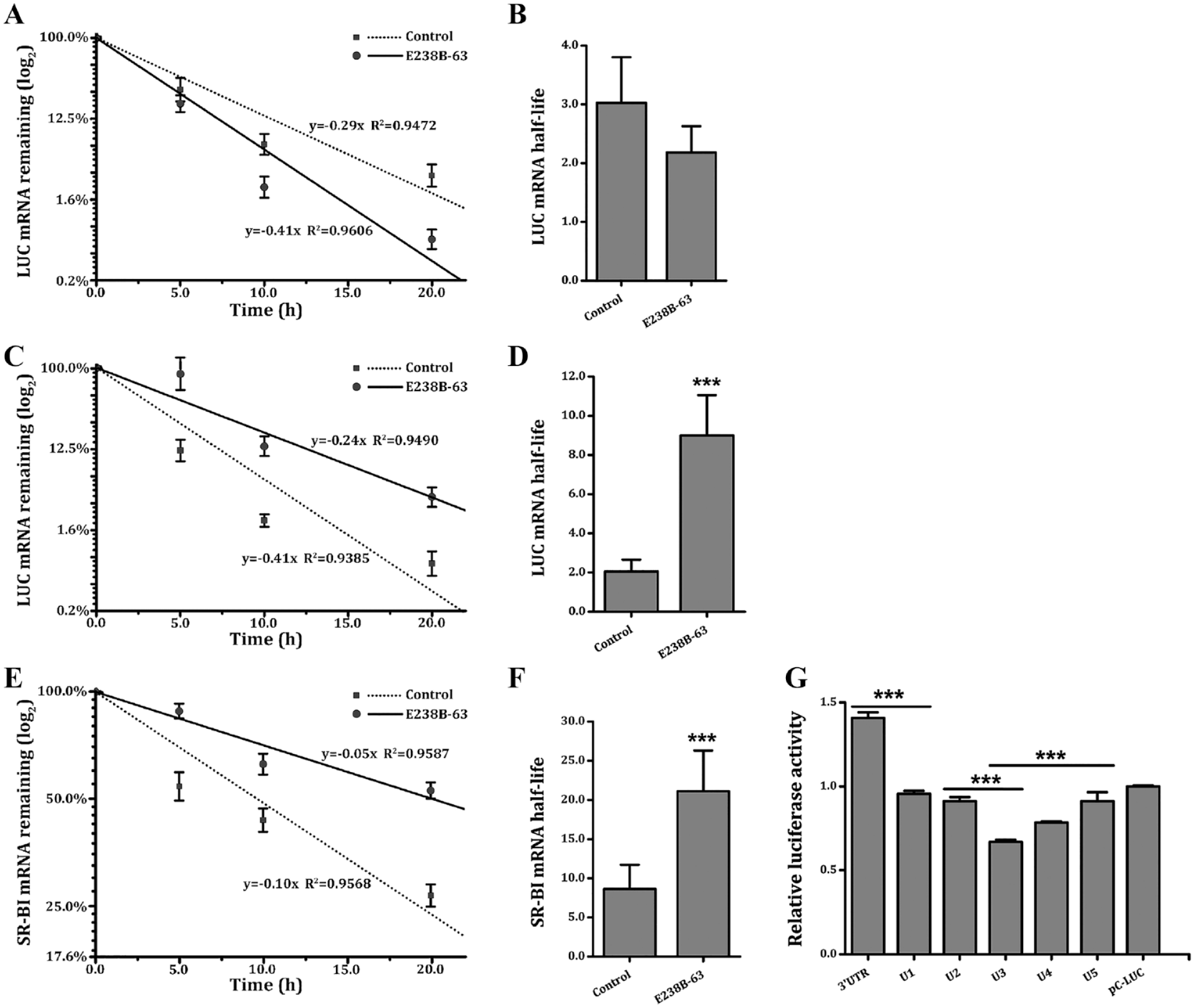
Effect of E238B-63 on mRNA half-life. The effect of compound E238B-63 on the mRNA half-life of LUC in CLAp-LUC HepG2 cells (
Effect on SR-BI mRNA half-life
The changes of SR-BI mRNA half-life in HepG2 cells were further investigated after E238B-63 treatment to determine the effects of the compound on SR-BI mRNA stability in hepatic cells.
After HepG2 cells were exposed to 5 µg/mL E238B-63 treated with Act D, mRNA decay across different time points was detected by qRT-PCR. In control HepG2 cells, the half-life of SR-BI was 8.64 h, whereas after exposure to compound E238B-63, its half-life was 21.10 h, which was significantly increased. This indicated that E238B-63 could significantly prolong the mRNA half-life of SR-BI and enhance the stability of SR-BI mRNA ( Fig. 4E, F ).
We also investigated the effect of compound E238B-63 on different fragments of SR-BI 3′UTR. Overall, the results raise two possibilities: first, that the difference between full-length SR-BI 3′UTR and U1 was responsible for the upregulation effect; alternately, the full-length 3′UTR enhanced mRNA stability because the upregulations of U1~U5 were lower than that of pC-LUC. The results showed that the compound produced a significant effect on 3′UTR-U1 and U2-U3, indicating that E238B-63 may affect the stability of 3′UTR through the sequences between the sites 1-120 and 312-498 ( Fig. 4G ).
Analysis of the structure-activity relationship
It is surprising that all of the five positive compounds obtained by HTS contain the structure N-(3-hydroxy-2-pyridyl)-carboxamide; thus, we inferred that this structure may be the key pharmacophore for prolonging the half-life and enhancing the stability of SR-BI mRNA. According to this hypothesis, we searched the compound library from the National Center for Microbial Drug Screening using the structure fragment N-(3-hydroxy-2-pyridyl)-carboxamide ( Fig. 5 ), to find compounds containing this fragment. Thirty-one compounds (including the five positive compounds) with the common structure unit N-(3-hydroxy-2-pyridyl)-carboxamide were found ( Suppl. Table S4 ). Two concentrations, 5 µg/mL and 10 µg/mL, were chosen to detect the effects of the 31 compounds on the luciferase activity using the HTS assay established in this study.
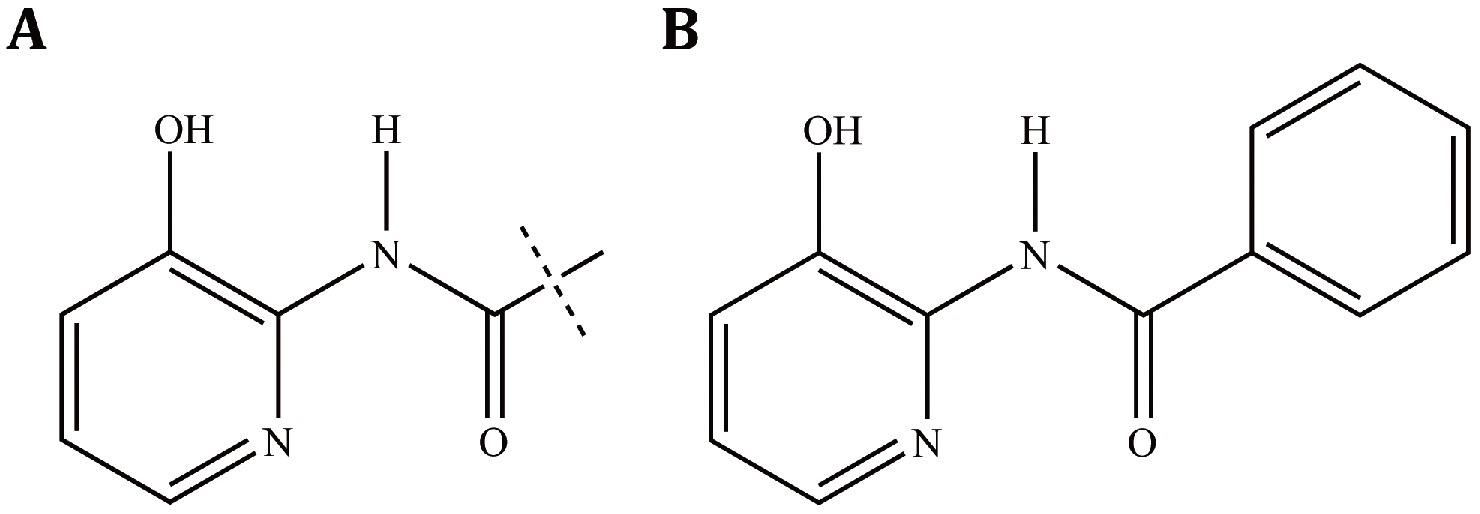
The predicted active structure of positive compounds. (
Most of the 31 compounds had an enhancing effect on SR-BI mRNA stability in the HTS assay at both 5 µg/mL and 10 µg/mL, and the upregulation ratios of the compounds at 10 µg/mL were significantly higher than at 5 µg/mL. At 5 µg/mL, 58.1% (18/31) of the compounds showed >1.3-fold induction of SR-BI stability, 29.0% (9/31) showed >1.5-fold induction, and 6.5% (2/31) showed >twofold induction. At 10 µg/mL, 96.8% (30/31) compounds showed >1.3-fold induction of SR-BI stability, 80.7% (25/31) showed >1.5-fold induction, and 16.1% (5/31) showed >twofold induction. These results support our hypothesis that N-(3-hydroxy-2-pyridyl)-carboxamide is an important structure unit for the compounds to play the role of enhancing SR-BI mRNA stability.
On the other hand, the side groups conjugated to the formyl group have important influences on the activity of the compounds. When the formyl group is directly conjugated to a five- or six-membered ring structure, the activity of the compound is generally better ( Suppl. Table S4 ). Phenyl ring (E238B-63, E238B-29, E042B-39, E015B-14, and E042B-27), cyclohexane ring (E014B-21 and E158B-13), pyrazine ring (E194B-14), furan ring (E038B-62 and E105B-09), thiophene ring (E170B-44), and triazole ring (E224B-16) are important active pharmacophores ( Suppl. Table S4 ). The characteristics and position of the substituent on the five- or seven-membered ring might also influence the activities of the compounds, and E238B-63, which shows the highest activity, has a para-substituted iodine atom on the phenyl ring. In addition, compounds E170B-49 and E192B-44 showed high activity ( Fig. 5 ; Suppl. Table S4 ).
Discussion
Regulatory mechanisms in eukaryotic mRNA stability are complex and diverse.17,18,39 The 3′UTR of eukaryotic mRNA is an important functional element, and the key factor that affects the mRNA life is mainly located in this region.17,20,40,41 For some genes, the mRNA 3′UTR sequence often contains the necessary conserved RNA sequence and special structure for regulating RNA stability.17,20,21,42–44 On one hand, the 3′UTR in eukaryotic mRNA can control the in vivo stability and degradation rate of mRNA, adjust the mRNA subcellular localization and translation level, and decide the fate of special mRNA. Furthermore, mRNA 3′UTR can determine the type of cells expressed by mRNA and control mRNA utilization efficiency to help identify particular codons. In addition, 3′UTR mutations may also affect the expression of one or more genes, leading to diseases.17,20,21,45 Moreover, 3′UTR is also an important site of intracellular interactions of other factors, playing an important role in cell phenotype control and growth and differentiation.17,20,21,28
It is of great theoretical and practical significance to enhance mRNA stability and increase the expression of hepatocyte SR-BI to promote the selective uptake of HDL-C and RCT in liver cells. This should reduce adverse inflammation and maintain the redox balance and ultimately reduce atherosclerotic vascular diseases. The SR-BI is regulated by a variety of levels of transcription and posttranscription. The expression of genes such as SR-BI that can rapidly respond to external signals during the cell metabolism process, which may depend on mRNA stability, is generally controlled by the 3′UTR. Our results showed that when the 3′UTR was intact, the mRNA half-life of LUC was significantly reduced, indicating that the SR-BI 3′UTR may act as a decisive factor affecting the mRNA instability.
Currently, there is no known small molecular compound regulating the mRNA stability of SR-BI by targeting SR-BI 3′UTR. Therefore, the first difficulty in the establishment of the HTS assay is selecting the positive modifier to evaluate the HTS assay system of SR-BI mRNA stability. The 3′UTR of human SR-BI mRNA contains two miR-185 seed regions. 34 miR-185 can significantly reduce the mRNA and protein expression of SR-BI and reduce the uptake of DiI-HDL in hepatocytes. An miR-185 inhibitor increased the mRNA and protein expression of SR-BI and the uptake of DiI-HDL in hepatocytes.33,34 Therefore, this was chosen as the positive control to evaluate the HTS system. The Z′ factor is a simple and widely known characteristic parameter for evaluating the quality of HTS assays, and it shows the dynamic change scope of the signal in the test and data variations related to signal measurement. It is applicable to evaluating and verifying the screening test quality, as it assesses signal change scope, offset of specimen, and reference substance. 35 The evaluation results showed that parameters such as signal-to-background ratio, signal-to-noise ratio, coefficient of variation, and Z′ factor are within a reasonable range. We then found five positive compounds by performing this HTS assay. In addition to significant upregulation of luciferase activity on the screening model, these five compounds can significantly upregulate the mRNA and protein expression level of SR-BI and increase the uptake of DiI-HDL in hepatocytes. After obtaining the active compounds, we selected E238B-63 as a positive compound to further evaluate several screening parameters, and the results showed that the screening system of human SR-BI mRNA stability established by targeting 3′UTR fully met the requirements of the HTS assay and hence can be used in large-scale screening experiments.
The compound E238B-63 significantly increased SR-BI mRNA and protein expression in a dose-dependent and time-dependent manner. E238B-63 could also significantly increase HDL uptake in HepG2 cells in a dose-dependent and time-dependent manner. The uptake of HDL in hepatocytes was affected by a variety of complicated factors. E238B-63 incorporation did not induce obvious cytotoxicity. In addition to the increase in SR-BI expression, high-concentration compounds may cause a series of other effects and affect the uptake of HDL in liver cells; the exact causes need further experimental study. In addition, studies on the correlation between the duration of E238B-63 exposure and SR-BI mRNA and protein expressions showed that the effects of the compound on SR-BI expression exhibited a time-dependent manner.
Whether eukaryotes can translate mature mRNA into protein to meet growth and development needs is associated with mRNA stability. The half-life of mRNA is the gold standard for evaluating mRNA stability. The half-life of mRNA can be influenced by many factors. mRNA half-life varies widely across different cell types, ranging from a few minutes to several hours, and therefore, the half-life of an mRNA is not a deterministic value. The widespread short-lived mRNA is sensitive to ribonuclease and can be quickly degraded. The long-lived mRNA possesses features that can free it from attack of ribonuclease and direct degradation. In highly differentiated cells, some mRNA is extremely stable and some can live for several days. 40 The results showed that E238B-63 can significantly prolong the mRNA half-life of the reporter gene and enhance the stability of SR-BI mRNA by interacting with the SR-BI 3′UTR. Furthermore, compound E238B-63 may affect the stability of 3′UTR through the sequences at the sites 1 to 120.
Analysis of the structure-activity relationship of the 31 compounds with a common fragment showed that the structure N-(3-hydroxy-2-pyridyl) carboxamide was possibly a key structural unit of active compounds. The groups connected with the formyl group also had important influences on the activity of the active compounds. The compounds whose formyl groups were directly conjugated to phenyl had high activity. The type of substituent structure and position of the substituent on the phenyl ring also seemed to influence the activity of the compound, and the parasubstituted compounds often exhibited strong upregulation activity.
In conclusion, we established and applied the new anti-atherosclerosis drug-screening system with the target of human SR-BI mRNA stability. Through HTS and further validation, we identified five positive compounds with definite effect. Compound E238B-63 could significantly prolong the half-life of SR-BI mRNA, enhance the stability of SR-BI mRNA, and affect the expression and functions of SR-BI in hepatocytes in a dose-dependent and time-dependent manner. Thus, the reported HTS assay targeting the SR-BI 3′UTR represents a promising method for the discovery of a novel SR-BI upregulator by enhancing its mRNA stability.
Supplemental Material
Supplementary_information_191009 – Supplemental material for Identification of Novel Compounds Enhancing SR-BI mRNA Stability through High-Throughput Screening
Supplemental material, Supplementary_information_191009 for Identification of Novel Compounds Enhancing SR-BI mRNA Stability through High-Throughput Screening by Xiao-Jian Jia, Yu Du, Hua-Jun Jiang, Yong-Zhen Li, Yan-Ni Xu, Shu-Yi Si, Li Wang and Bin Hong in SLAS Discovery
Footnotes
Acknowledgements
We appreciate Nicola Chen’s help in revisions and discussions.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of China (81473214, 81402929, and 81621064), CAMS Innovation Fund for Medical Sciences (2016-I2M-2-002, 2016-I2M-1-011, and 2017-I2M-1-008), the Drug Innovation Major Project of China (2018ZX09711001-003-006, 2018ZX09711001-007, and 2018ZX09735001-002), Beijing Natural Science Foundation (7162129), the Science and Technology Planning Project of Shenzhen Municipality (JCYJ20170306155028734, JCYJ20180306170919604, JCYJ20180507182420114), and the Natural Science Foundation of Guangdong Province (2018A030313189).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.