Abstract
The molecular structure of many proteins contains disulfide bonds between their cysteine residues. In this work we demonstrate the utilization of the disulfide bond structure of proteins for their label-free determination by surface-enhanced Raman spectroscopy (SERS). The new approach for label-free SERS detection of proteins is demonstrated for human insulin. The protein was selectively extracted from spiked plasma samples using target-specific functionalized nanomaterial. Enzyme-linked immune assay (ELISA) was used to detect insulin in the blood plasma and cross-validate the SERS method. The disulfide bonds in the molecular structure of the protein were chemically reduced and used for their chemisorption onto the gold-coated copper oxide substrate in a unified orientation at a very short distance from the hotspots. The oriented chemisorption of the protein caused significant enhancement to the signal intensity of its Raman vibration modes. This is attributed to the strong short-range electromagnetic and chemical enhancement effects that are experienced by the immobilized protein. Using this approach, label-free and reproducible SERS detection of insulin, down to 10 zM (relative standard deviation [RSD] = 5.52%), was achieved. Sixty-five percent of proteins contain disulfide bonds in their molecular structure. Therefore, the new label-free SERS detection method has strong potential for the determination of ultralow concentrations of proteins at pathology labs and in biology research.
Introduction
The rapid and sensitive screening of proteins is critical for the diagnosis of diseases and monitoring of therapeutic efficacy, as well as the discovery of new protein biomarkers in biomedical research. Analytical methods such as immunoassay, mass spectroscopy, and turbidimetric analysis are utilized for the detection of proteins. However, these methods are slow and complex and use error-prone protocols.1–4 The analysis of proteins by label-free methods offers the advantages of low cost, minimum sample preparation time, and the absence of the potential side effects of chemical labeling.5–7
Within the last decade, surface-enhanced Raman spectroscopy (SERS) has emerged as one of the most sensitive bioanalytical assay techniques. The Raman signal enhancement in SERS is attributed to the electromagnetic (EM) field that is experienced by the analyte molecules when they become trapped within the gaps between the closely spaced nanostructures of a metallic substrate, and the formation of a charge transfer complex between a chemisorbed analyte and the substrate. 8 Park et al. demonstrated that a combined charge transfer and EM enhancement factor (EF) of 106 to 108 is sufficient for the detection of a single molecule of the analyte by SERS. 9
The determination of biomolecules by SERS is carried out by either label or label-free methods. In the label-based methods, a Raman probe that is attached to a metallic SERS substrate is used for the indirect detection of the biomolecule where the Raman vibration modes of the probe are monitored rather than those of the biomolecules themselves. In contrast, the label-free methods do not use a Raman probe and directly detect the Raman vibrations of the target analyte. Label-free SERS detection of proteins requires that the biomolecule structure contain a chromophore moiety that provides a strong Raman signal due to surface-enhanced resonance Raman scattering (SERRS). 10 However, many proteins lack the presence of a chromophore moiety in their molecular structure and have a relatively low Raman scattering cross section. In addition, the weak physical adsorption of the protein leads to random orientation of the biomolecules onto the substrate, which undermines the chemical enhancement in the SERS measurement of the protein.11,12
The EM and charge transfer enhancements in SERS can be maximized by the oriented immobilization of analyte molecules at a very close distance from the nanostructured surface of the SERS substrate.8,11 Many proteins, such as insulin, contain a disulfide bond structure to maintain their stability. 13 These bonds can be chemically reduced to provide free sulfhydryl (SH) groups that form stable Au–S bonds with a gold surface. 14 Therefore, in this work, we reduced the disulfide bond structure of human insulin to provide free SH terminal groups and assemble the biomolecules, in a unified orientation, at a very short distance from the nanostructures of gold-coated copper oxide SERS substrate.
Insulin is a naturally occurring hormone that is responsible for keeping blood sugar within its normal level of ~79 pM. 15 An abnormally high insulin concentration in blood leads to diseases such as hyperinsulinemia and increases the risk of Alzheimer disease.16,17 Low insulin levels are mainly associated with developing a diabetic medical condition due to the functional damage of the pancreatic beta cells. 18 Therefore, a simple and sensitive method for the determination of insulin is valuable for the clinical diagnosis of insulin-related diseases and assessing the quality of pancreatic islets for the treatment of diabetes. 19 In addition, it can also support the ongoing research on insulin-derived amyloidosis. 13 In this work, human insulin was selectively extracted from a spiked blood plasma using extractor magnetic gold nanoparticles that are functionalized with an anti-insulin antibody. The protein was recovered from the extractor nanoparticles, reduced, and detected by SERS down to 10 zM. The SERS quantification of the protein was cross-validated against enzyme-linked immune assay (ELISA).
Materials and Methods
Chemicals and Reagents
Magnetic gold nanoparticles (particle size = 51.8 ± 6.1 nm) were purchased from Nanoimmunotech (Zaragoza, Spain). Anti-insulin antibody was purchased from Abcam (Cambridge, UK). Human insulin, phosphate-buffered saline (PBS), an insulin ELISA kit, glycine, and potassium hydroxide were purchased from Sigma Aldrich (St. Louis, MO). Neutral tris(2-carboxyethyl)phosphine (TCEP) solution was purchased from Thermo Fischer Scientific (Waltham, MA). LoBind tubes were purchased from Eppendorf AG (Hamburg, Germany). Gravity flow size-exclusion columns (Illustra NAP-5) were purchased from GE Healthcare Life Sciences (Sydney, Australia). Deionized water (ultrapure Millipore, 18.2 MΩ cm at 25 °C) was used in all preparations. Gold-coated copper oxide substrates (substrate diameter = 4 mm) were obtained from FLEW Solutions (Brisbane, QLD, Australia) and used for the SERS measurements. Blood plasma samples were provided under Queensland University of Technology’s research ethics clearance exemption 1800001209.
Preparation of Human Insulin Standard Solutions
Insulin standard stock solution (10.5 mg/mL) was used to prepare 20 zM, 20 fM, 100 pM, 160 pM, 300 pM, 450 pM, and 600 pM insulin working solutions by serial dilution using 1× PBS buffer (pH 7.4).
Preparation of Extractor Magnetic Gold Nanoparticles
Magnetic gold nanoparticles were used for the development of the extractor nanoparticles due to their magnetic property, which allows for their collection from solutions using an external magnet. Target-specific antibody fragments were prepared by reducing the disulfide bonds in the hinge region of the insulin antibody using TCEP. 3 Briefly, 90 µL of 0.4 mM neutral TCEP solution was mixed with 90 µL of the antibody (0.2 mg/mL) in LoBind Eppendorf tubes and incubated for 30 min at 37 °C for the formation of the antibody fragments. Antibody fragments (180 µL) were mixed with 200 µL of magnetic gold nanoparticles and left to stand for 1 h to form Au–S bonds between the fragments and the nanoparticles. To avoid nonspecific binding from interfering molecules, the remaining bare sites on the nanoparticle surface were blocked by adding 38 µL of 1-butanethiol (1 × 10−6 M in 1× PBS, pH 7.4) and allowing the mixture to stand for 1 h to the complete attachment of the alkane thiol onto the gold surface of the nanoparticles via Au–S bonds. The functionalized and backfilled nanoparticles were then collected from solution using an external magnet, washed three times with 100 µL of 1× PBS buffer (pH 7.4), redispersed into 100 µL of 1× PBS buffer (pH 7.4), and stored at 4 °C for future use.
Selective Extraction of Insulin from Blood Plasma
Human insulin (250 pM) was spiked into a blood plasma sample and the antibody-functionalized nanoparticles were used to extract the protein from the spiked sample. Spiked blood plasma (100 µL) was mixed with 100 µL of the antibody-functionalized nanoparticles and left to stand for 15 min at room temperature with occasional shacking to bind the protein. The nanoparticles were then collected from the plasma matrix using an external magnet and washed five times with 100 µL of 1× PBS buffer (pH 7.4) to remove any weakly adsorbed interfering molecules. To recover the bound protein, the nanoparticles were reconstituted into 100 µL of 0.1 M glycine HCL buffer (pH 2.5) and left to stand for 5 min at room temperature. The nanoparticles were collected using an external magnet, and the buffer solution containing the released protein was loaded onto a size-exclusion column to remove the glycine buffer. The purified protein was eluted from the column using 500 µL of 1× PBS.
ELISA Measurements
To confirm the identity of the extracted protein, an aliquot of the eluted sample was screened using an insulin-specific ELISA kit and a linear calibration curve was constructed in the concentration range of 0–4 ng/mL, by recording the average optical density (OD) at 450 nm against the concentration of the protein standard. The regression equation was found to be
Rapid Reduction of Human Insulin
Different concentrations of standard human insulin (in PBS, pH = 7.4) were mixed with equal volumes of 20 mM TCEP solution (in PBS, pH = 7.4) into LoBind Eppendorf tubes in the presence of gold-coated copper oxide substrate. The mixtures were left for 15 min to allow for the reduction of the disulfide bonds within human insulin and the binding of the biomolecules to the metallic substrate through Au–S bond formation. The solution was then removed and the substrate washed with 1× PBS buffer to remove the excess TCEP. Finally, the protein-bearing substrate was dried under N2 stream and scanned with a handheld Raman spectrometer.
Fabrication of Gold-Coated Copper Oxide Substrates
Gold-coated copper oxide sensors, which had been reported earlier by Balčytis et al., 20 were used as SERS substrates. Briefly, an aqueous solution that consisted of 1.25 M NaOH and 66 mM Na2S2O8 was used for the wet chemical etching of polished Cu foil, for 5 min at 65 °C, and the development of nanostructured cupric oxide layers. Magnetron sputtering was then used for the plasma vapor deposition (PVD) of atomic gold onto the developed copper oxide surface layers. 20
SERS Measurements
The SERS measurements were carried out using a handheld Raman spectrometer (spectral resolution = 12 cm−1, spot size = 2.5 mm; Ocean Optics, Largo, FL) in the raster orbital scanning (ROS) mode over the wavelength range 700–1700 cm−1. 3 A 785 nm laser source (laser power = 5 mW) was used for excitation. Ten accumulations with a total acquisition time of 1 s were used for the SERS measurements. The software algorithm of the instrument (OceanView Spectroscopy 1.5.07) automatically averages and corrects the background noise and fluorescence within the acquired spectra.3,21
Results and Discussion
Synthesis of Extractor Nanoparticles and Extraction of Human Insulin
Insulin-specific extractor nanoparticles were synthesized by functionalizing gold-coated magnetic nanoparticles with anti-insulin antibody fragments. The antibody fragments were synthesized by simple chemical reduction of the disulfide bond structure within the antibody hinge region. The reduction process generates free terminal SH groups within the molecular structure of the antibody fragments. The SH terminals were used to attach the antibody fragments onto the gold surface of magnetic nanoparticles in a unified orientation through the formation of stable Au–S bonds. The remaining empty gaps on the surface of the nanoparticles were blocked with inert alkane thiol molecules to prevent the nonspecific adsorption of interfering molecules from the sample matrix.
The extractor nanoparticles were used to selectively bind insulin from blood plasma (
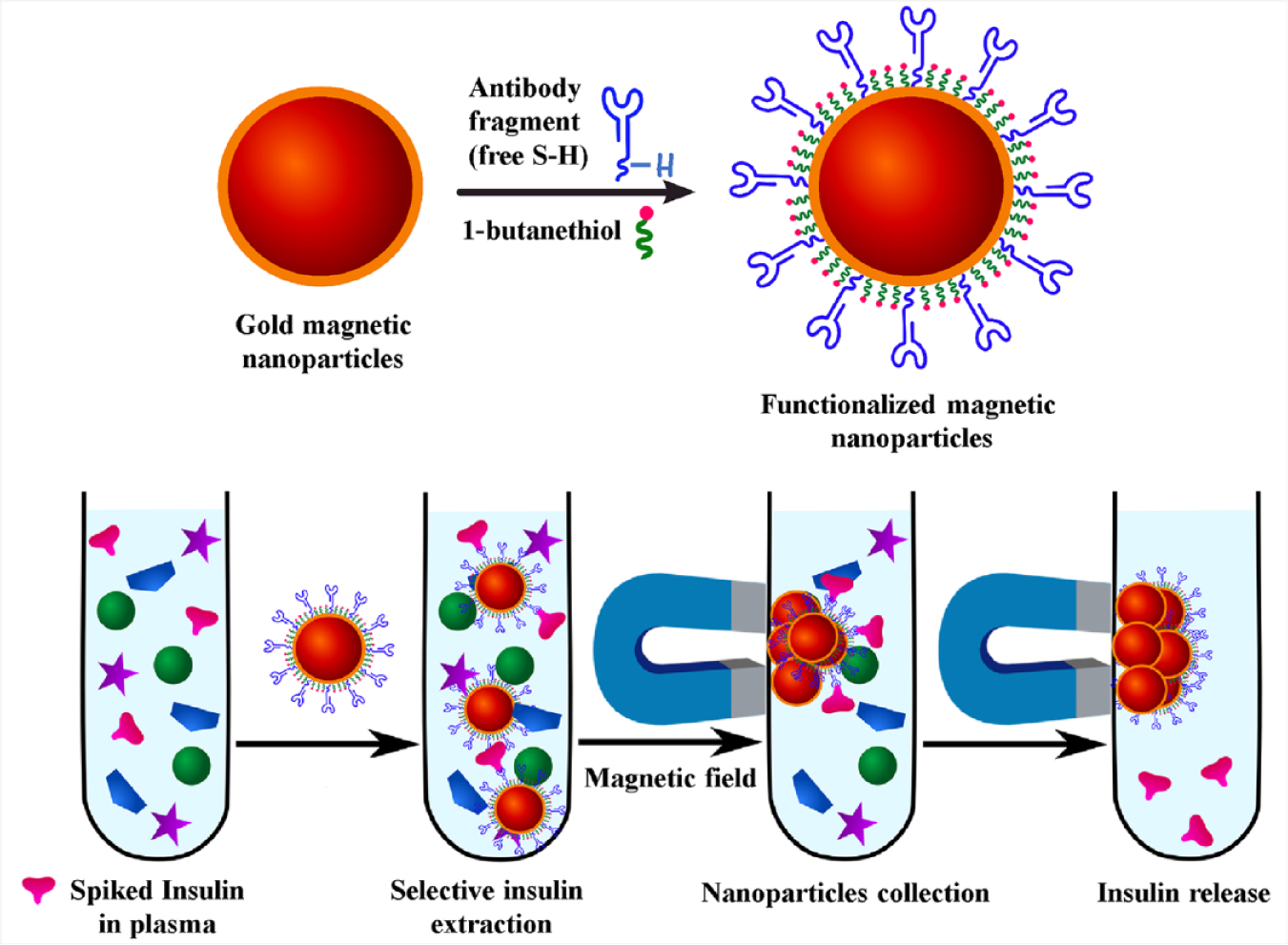
Schematic representation of the selective extraction of human insulin from spiked blood plasma by antibody-functionalized magnetic nanoparticles.
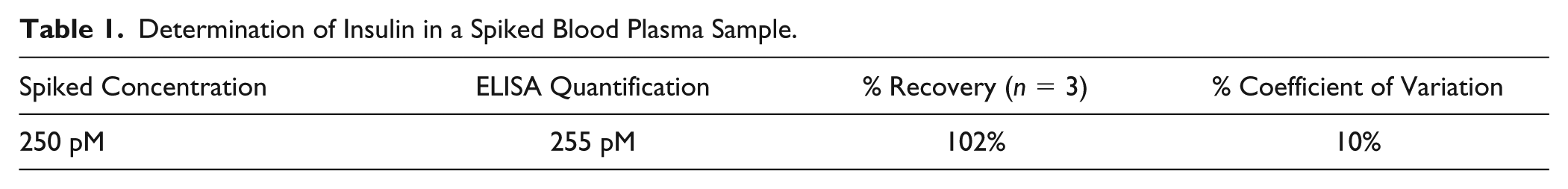
Determination of Insulin in a Spiked Blood Plasma Sample.

Fabrication of Gold-Coated Copper Oxide Substrates
The scanning electron microscopy (SEM) image and energy-dispersive x-ray (EDX) measurement of the gold-coated copper oxide SERS substrate are depicted in
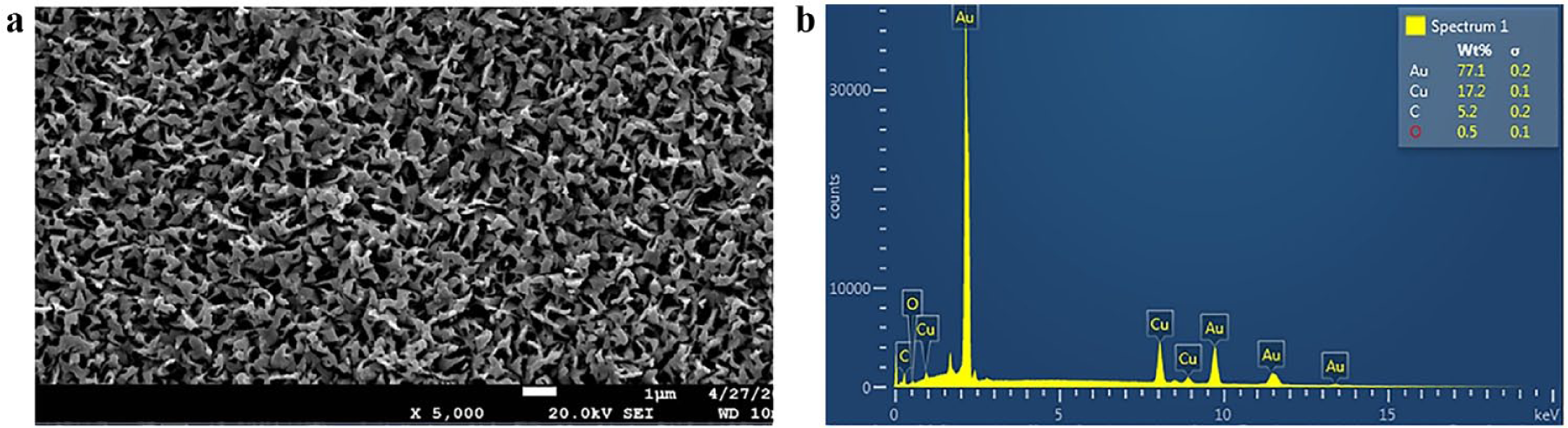
(
SERS Detection of Unreduced and Reduced Human Insulin
Unreduced human insulin (80 pM) was loaded onto the gold-coated copper oxide substrate and screened by SERS. The Raman spectrum of the physically adsorbed protein is depicted in
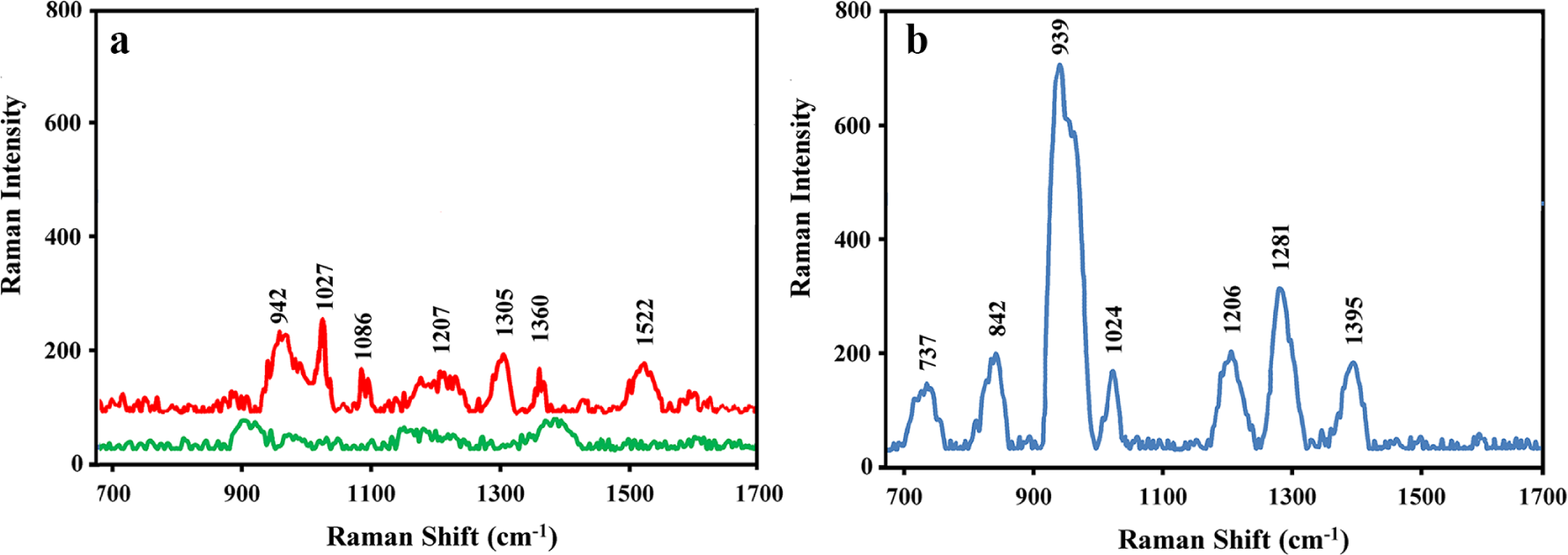
SERS spectra of (
The Raman spectrum of reduced insulin is depicted in
Effect of Reduction Time on the Raman Signal Intensity
To investigate the effect of the reduction time on the Raman signal intensity of the insulin, 80 pM insulin was reduced with TCEP for 15, 30, 60, and 120 min, respectively. As indicated by
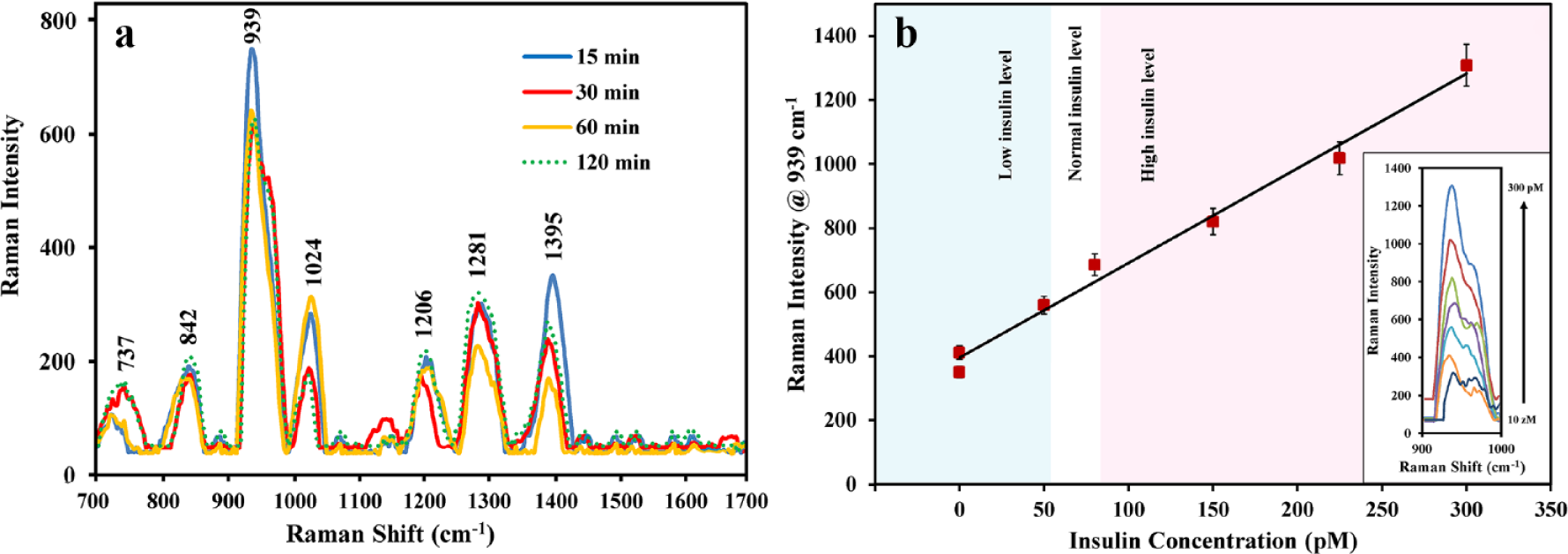
(
Label-Free SERS Quantification of Insulin
For the label-free SERS quantification of the protein, the intensity of the Raman band at 939 cm−1 was monitored at different concentrations of the reduced insulin (
The reproducibility of the SERS measurements of the protein was investigated by comparing the Raman spectra of nine different concentrations of the protein (on nine independent substrates). As indicated by
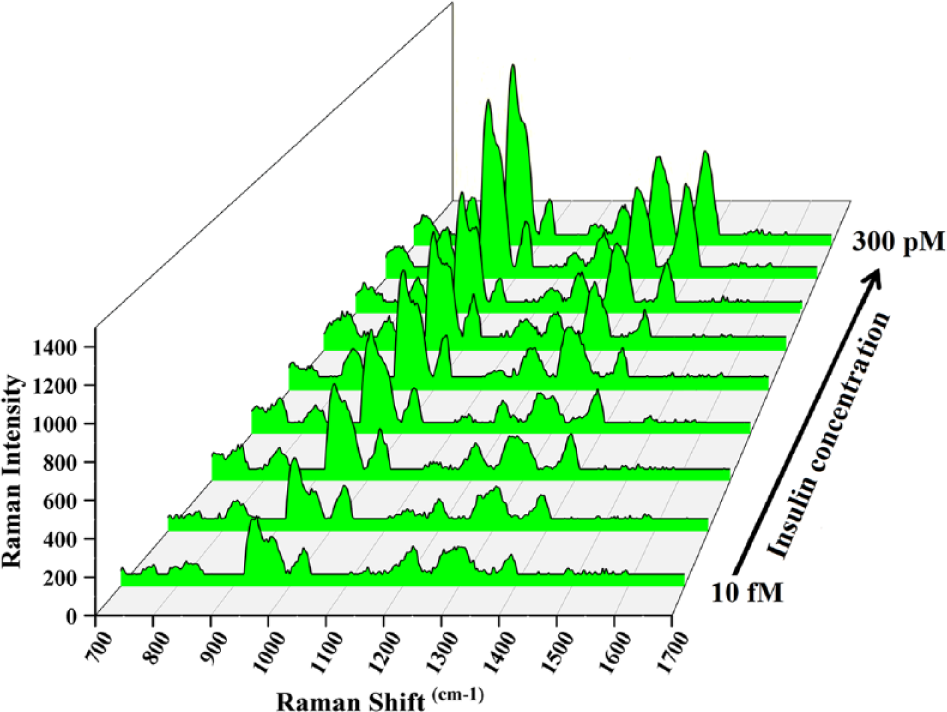
SERS spectra of reduced human insulin at different concentrations. The spectra were collected on nine independent gold-coated copper oxide substrates.
Recently, Cho et al. reported a method for the Raman detection of insulin where a gold-coated ZnO substrate was synthesized and utilized for the SERS screening of the protein. 25 The substrate was prepared using a hydrothermal method to grow ZnO nanowires on a Si wafer. The ZnO nanowires were then coated with gold nanostructures. The process was repeated five times to form dense Au–nanoparticle stacks in pillar forms and increase the uniformity and SERS activity of the substrate. Therefore, the substrate fabrication required several hours and multiple chemical processes, and the SERS measurement relied mainly on the EM enhancement provided by the substrate geometry. In the present work, we utilized an inexpensive and easy method to manufacture gold-coated copper oxide substrate and, instead of relying on the geometry of the substrate, used the protein’s own disulfide bond structure to achieve highly sensitive SERS measurements of insulin down to 10 zM (approximately ninefold lower than that demonstrated by Cho et al.). The chemisorption and oriented immobilization of the protein onto the gold-coated copper oxide substrate maximized the chemical and EM enhancement effects in the SERS measurement.
We have demonstrated functionalized gold nanomaterials for the selective extraction of proteins from blood plasma.30,31 Therefore, by combining selective extractor nanomaterials with the new method for biomolecules, selective and label-free SERS screening of many proteins and disease biomarkers can be achieved for molecular diagnosis applications and biology research.
In conclusion, we demonstrated a proof of concept on a simple, reproducible, and ultrasensitive label-free SERS detection of insulin by using the protein’s own disulfide bond structure. The reduction of the disulfide bond structure of the protein allowed for its rapid chemisorption in a unified orientation onto a gold-coated copper oxide substrate via Au–S bond formation. This causes the biomolecules to form a charge transfer complex with the SERS substrate at a very short distance from the hotspots on the plasmonic surface of the substrate. Therefore, the Raman vibration modes of the immobilized protein experience maximum EM and chemical enhancement effects, and a dramatic increase in the protein Raman signal intensity is observed. Using this approach, reproducible SERS detection of insulin, down to 10 zM, was achieved. Since 65% of proteins have a disulfide bond molecular structure, the demonstrated method can be utilized for their label-free SERS determination at pathology labs, at points of care, and in biology research, especially when combined with a selective extraction method for the target protein.
Supplemental Material
DS_DISC875102 – Supplemental material for Toward Label-Free SERS Detection of Proteins through Their Disulfide Bond Structure
Supplemental material, DS_DISC875102 for Toward Label-Free SERS Detection of Proteins through Their Disulfide Bond Structure by Waleed A. Hassanain and Emad L. Izake in SLAS Discovery
Footnotes
Acknowledgements
The authors thank FLEW Solutions Pty Ltd Australia for donating gold-coated copper oxide substrates for this work.
Supplemental material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
