Abstract
Natural extracts are complex mixtures that may be rich in useful bioactive compounds and therefore are attractive sources for new leads in drug discovery. This review describes drug discovery from natural products and in explaining this process puts the focus on ion-channel drug discovery. In particular, the identification of bioactives from natural products targeting nicotinic acetylcholine receptors (nAChRs) and serotonin type 3 receptors (5-HT3Rs) is discussed. The review is divided into three parts: “Targets,” “Sources,” and “Approaches.” The “Targets” part will discuss the importance of ion-channel drug targets in general, and the α7-nAChR and 5-HT3Rs in particular. The “Sources” part will discuss the relevance for drug discovery of finding bioactive compounds from various natural sources such as venoms and plant extracts. The “Approaches” part will give an overview of classical and new analytical approaches that are used for the identification of new bioactive compounds with the focus on targeting ion channels. In addition, a selected overview is given of traditional venom-based drug discovery approaches and of diverse hyphenated analytical systems used for screening complex bioactive mixtures including venoms.
Keywords
Introduction
Natural extracts are complex mixtures that may be rich in useful bioactive compounds with medicinal use. Natural extracts are therefore attractive sources for new leads in drug discovery. The work presented here mainly aims to review recent (technological) advances in the screening and identification of novel bioactive compounds targeting brain receptors. The focus is on the alpha-7 nicotinic acetylcholine receptor (α7-naChR) and the serotonin type 3 receptor (5-HT3R). In recent years many advances have been made in the development of analytical techniques facilitating improved identification of bioactive compounds from these natural extracts. This work is divided into three main parts: “Targets,” “Sources,” and “Approaches.” The “Targets” part discusses neurotransmitters, neurotransmitter receptors, and their function. Attention is given to the α7-naChR and 5-HT3R, their general relevance, and their relevance as a target in central nervous system (CNS) diseases specifically. In “Sources,” the importance of the environment in finding new drug leads is discussed. The use of various natural extracts, through history and their ongoing importance for current and future drug discovery purposes, is reviewed. A division is made between plant and animal extracts. The “Approaches” part gives an overview of the invaluable set of classical and novel techniques for the screening, detection, identification, and characterization of drug leads from natural extracts targeting ion channels. Included here is an overview of the venom-based drug discovery approach and the diverse hyphenated analytical systems used for complex-mixture screening.
Targets
Ligand-Gated Ion Channels in the Nervous System
The brain consists of hundreds of billions of neurons (nerve cells), which communicate with each other predominantly via chemical signaling in specialized parts of the neurons, so-called synapses. In synapses the presynaptic neurons can release endogenous compounds, neurotransmitters, which can bind to specific neurotransmitter receptors that are present on the postsynaptic neurons, or alternatively presynaptic terminals, resulting in modulation of synaptic activity. 1 The neurotransmitters can be categorized in small-molecule-type transmitters that are enzymatically synthesized (e.g., acetylcholine, GABA, or serotonin) and peptide-type neurotransmitters encoded by the genome (e.g., the opioid peptides). 2 The neurotransmitter receptors are categorized in two major groups: G-protein-coupled receptors (GPCRs) and ligand-gated ion channels (LGICs). Ion channels can be gated (i.e., opened or closed) by voltage (voltage-gated ion channels) or by a specific ligand binding to the channel (LGICs). The main role of ion channels is to generate ion concentration gradients that change the membrane potential, which may result in the propagation of electrical signals in the neurons. 3 The LGICs can be divided in subcategories based on their structure: the Cys-loop-type LGICs consist of five subunits, the ionotropic glutamate receptors (iGluRs) are formed from four subunits, and the ionotopic ATP receptors (P2X-Rs), which consist of three subunits. The categorization of LGICs is shown in Table 1 .
Classification of the LGIC Superfamily.
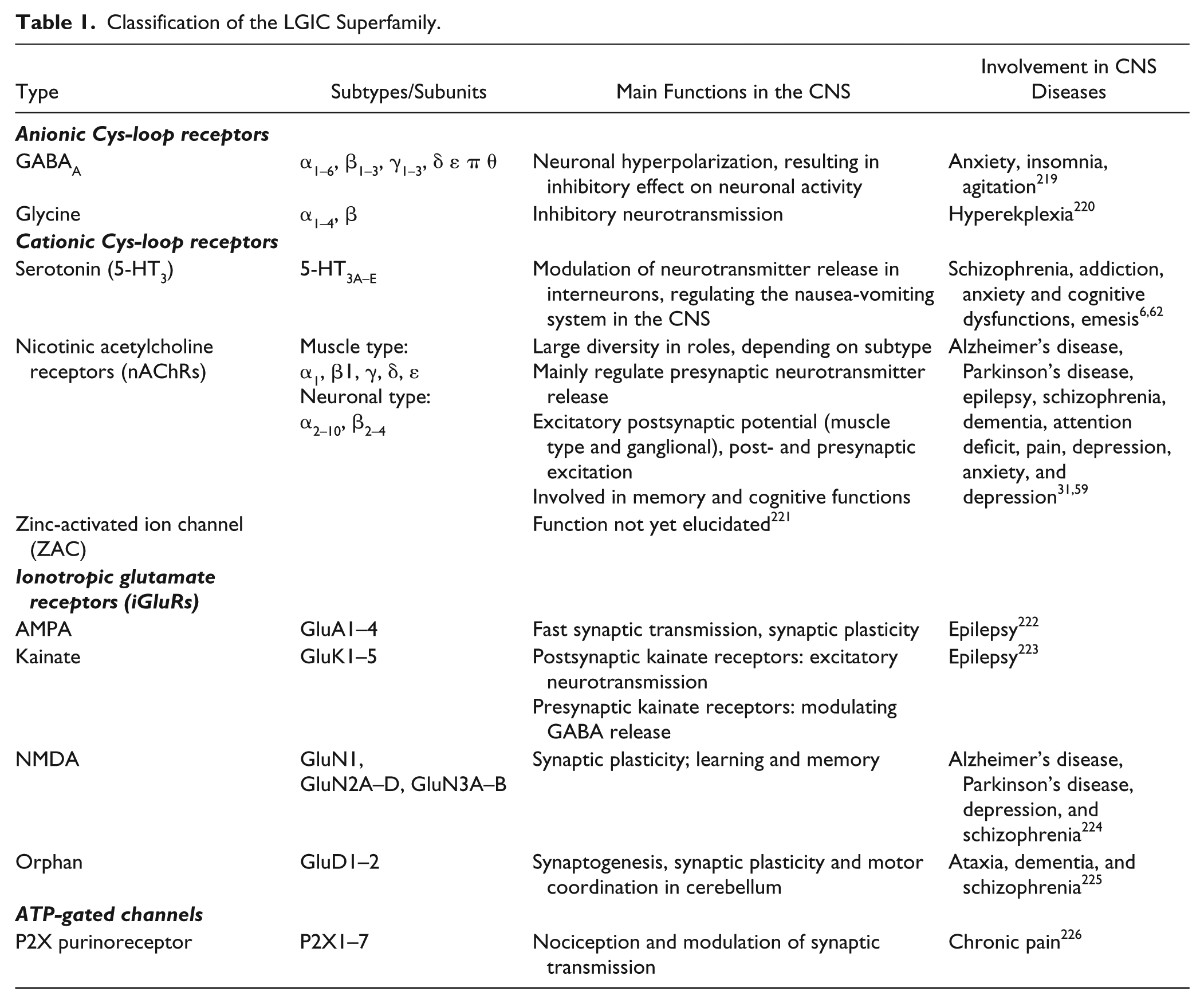
Many CNS-related diseases are caused by the altered regulation, function, or expression of neurotransmitter receptors. This review focuses on two of these receptors that are involved in several CNS diseases and therefore comprise important therapeutic targets: the α7-nAChR and the 5-HT3R (
(
nAChRs and the α7-nAChR
The nAChRs belong to the Cys-loop receptor superfamily of the LGICs. The Cys-loop receptor family is named after a 13-amino-acid loop present in these receptors formed by a disulfide bridge. The members of this receptor family are the nAChRs, the GABAA receptors, the 5-HT3Rs, and the glycine receptors (GlyRs).9–12 The nAChRs can be divided into two groups: the muscle-type nAChRs and the neuronal-type nAChRs.13,14 The muscle-type nAChRs are found in neuromuscular junctions of the peripheral nervous system (PNS), whereas the neuronal types are found in the CNS, but are also expressed in non-neuronal tissues and organs, for example, in macrophages, lung, or skin. The nAChR subunits are classified as α subunits when the C loop of the receptor contains two adjacent cysteine residues, whereas in the β subunits these cysteine residues are absent. Up to now there are nine neuronal α subunits (α2–10) and three β subunits (β2–4) identified.
15
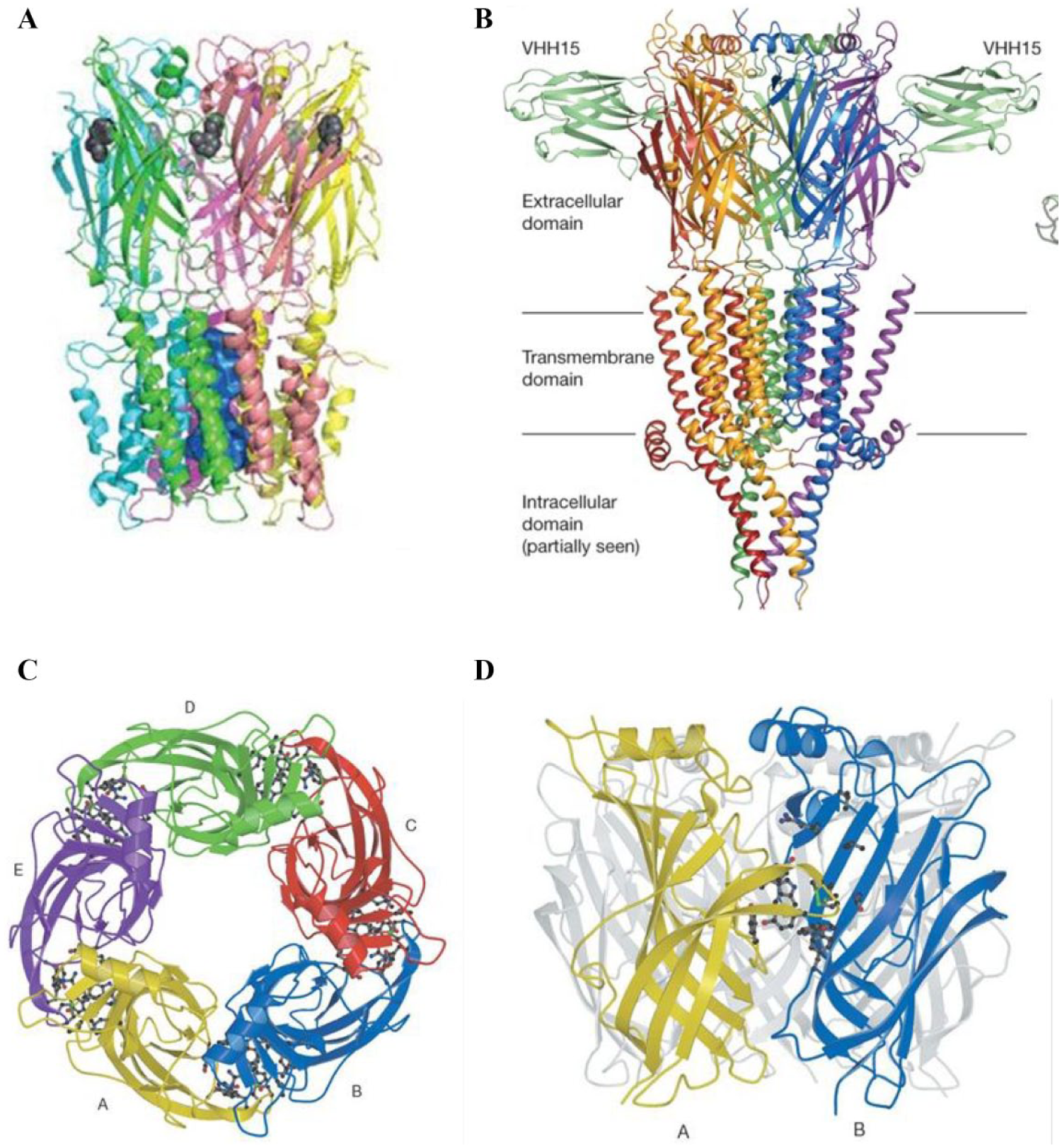
Whereas some of the α subunits can form so-called homomeric receptors consisting of five homologous α subunits (the α7- and the α9-nAChR), the other neuronal α subunits form heteromers consisting of a combination of α and β subunits (e.g., α4β2 and α3β4). Crystal structure studies initially using the acetylcholine binding protein (AChBP) provided detailed information regarding the structure of nAChRs specifically and LGICs in general
16
(
This review focuses on the homopentameric α7-nAChR, which has been implicated in CNS diseases. However, other subtypes of nAChR also have high clinical relevance. For example, the α4β2-nAChR is the predominant nAChR subtype in the brain and it is known to be involved in addiction to tobacco/smoking. For treatment of tobacco addiction, varenicline (Champix) is an approved drug targeting the α4β2-nAChR. Besides tobacco addiction, α4β2-nAChRs are also involved in cognitive disorders and in pain, and there are several compounds targeting α4β2-nAChR in clinical trials for the treatment of these.
In the brain the α7-nAChR is localized mainly in various brain regions involved in cognitive function, learning, and memory. α7-nAChRs were found in the cerebral cortex, hypothalamus, ventral tegmental area, substantia nigra, hippocampus, pineal gland, amygdala, medial habenula, olfactory bulb, and cerebellum.20–24 The α7-nAChR is also expressed in nonneuronal tissues, such as in macrophages, lymphocytes, skin, and kidney.25–30
Typical characteristics of the α7-nAChR are its high desensitization rate, calcium permeability, and the relatively low affinity of acetylcholine and nicotine toward the receptor.31,32 The most common functions awarded to the α7-nAChR are modulation of the other neurotransmitter systems, for example, modulation of synaptic plasticity in the brain (glutamate, dopamine, serotonin, GABA, and norephineprine), and the activation of messenger pathways (e.g., gene expression or neuronal survival) on postsynaptic neurons by changes in the intracellular Ca2+ concentration.33–35 The abnormal functioning or loss of nAChRs has been associated with many CNS diseases, such as to Alzheimer’s disease, 36 Parkinson’s disease, 37 epilepsy, 38 schizophrenia, 39 attention deficit hyperactivity disorder (ADHD), 40 pain, 41 anxiety, 42 and depression. 42
In schizophrenia patients, the expression level of the α7-nAChR is reduced in many brain regions compared with healthy subjects.43,44 There have been several efforts for developing drugs targeting the α7-nAChR. The partial α7-nAChR agonist Encenicline (EVP-6124), which showed promise in both Alzheimer’s disease and schizophrenia, successfully came through phase II clinical studies;45,46 however, due to severe adverse gastrointestinal effects in some patients during phase III trials, this compound has been put on hold for further studies.47,48 ATA-101, formally known as TC-5619, is a full α7-nAChR agonist that showed promise as a drug candidate in clinical phase I trials for improving the cognitive functions in schizophrenic patients, 49 and also showed promise in the treatment of ADHD and potentially Alzheimer’s disease. However, in the phase II trial for schizophrenia 50 the drug showed lack of efficacy and further development was halted for treatment of schizophrenia. Since then the manufacturer has also stated the drug candidate to be unsuccessful for treatment of ADHD; however, the compound was recently proposed as a drug candidate for treatment of acute and chronic cough. 51
The role of the α7-nAChR in Alzheimer’s disease and in its treatment has been extensively studied, as discussed in recent reviews in the literature.36,52 In Alzheimer’s disease, the loss of α7-nAChRs in the hippocampus correlates with decreased cognitive functioning. 53 Also, recent studies show that amyloid beta (Aβ) binds with high affinity to the α7-nAChR, and by this disrupts its normal functioning.54,55 However, there are still different opinions in the literature regarding the type of interaction and whether Aβ acts as an agonist or an antagonist on the α7-nAChR, and some publications state that there is no interaction for Aβ and the α7-nAChR. Nevertheless, there are α7-nAChR agonists, which are in phase I and phase II studies, that are potential new drug candidates for the treatment of the cognitive symptoms of Alzheimer’s disease.
Next to the possible involvement in CNS-related diseases, the α7-nAChR might be involved in non-CNS-related processes. Recently, the immunomodulatory and anti-inflammatory effects of the α7-nAChR were under investigation.25,56 Another recently discovered potential of non-CNS pharmaceutical targeting of the α7-nAChR is in cancer treatment. The α7-nAChR was found to modulate the chemosensitivity of gastric cancer cells, and recent studies showed that silencing α7-nAChR levels increased the sensitivity of gastric cancer cells to chemotherapy drugs.57,58 In a recent review, the role of nAChRs in disease is discussed in more detail, and an overview of the different nAChR-targeting drugs that are in clinical studies is provided. 59 As a conclusion, the α7-nAChR is potentially an important therapeutic target against several medical conditions. Studies toward further understanding the biological functioning of this receptor are ongoing; 60 however, there are still many remaining questions that need to be answered regarding the precise functions and roles of this receptor.
5-HT3R
The 5-HT3R ( Fig. 1B ) is the only serotonin receptor that belongs to the LGIC family, whereas all others are GPCRs. 61 The 5-HT3R, like the α7-nAChR, is assembled from five subunits: either a homomeric receptor composed of five 5-HT3A subunits or a heteromeric receptor composed of 5-HT3A and 5-HT3B subunits.62–64 The expression of the subunits in the CNS or PNS has not yet been completely elucidated. Also, the genes of the 5-HT3C, 5-HT3D, and 5-HT3E subunits have been described, but their function and localization has not been characterized yet. 65 The 5-HT3R has high similarity to the α7-nAChR. The two receptors share high sequence (30%) and structural homology, and therefore many ligands binding to the α7-nAChR are also interacting with the 5-HT3R. 66 A few examples of ligands that show orthosteric binding to both the 5-HT3R and α7-nAChR are varenicline, epibatidine, and tropisetron.67–69 5-HT3Rs are expressed in both the CNS and PNS, and they are expressed in different brain locations, like in the cortex, amygdala, sustantia nigra, and brain areas involved in the vomiting reflex, such as the nucleus tractus solitarius and the area postrema.
The 5-HT3R has been associated with CNS diseases, such as schizophrenia, addiction, anxiety, and cognitive dysfunctions; however, there are currently no 5-HT3R-targeting drugs used clinically for the treatment of these diseases. In the clinic, the two main applications for 5-HT3R antagonists are the prevention of chemotherapy- and radiotherapy-induced nausea and vomiting, and the treatment of irritable bowel syndrome (IBS). 5-HT3Rs are expressed in both the PNS and CNS at the chemoreceptor trigger zone of the area postrema. 5-HT3R antagonists, such as ondansetron and granisetron, are used clinically to block these receptors; however, it is not certain yet if their action is mediated peripherally, centrally, or both. The 5-HT3R antagonist alosetron is widely used in the clinic for the treatment of diarrhea-predominant IBS, targeting the 5-HT3Rs of the enteric nervous system in the gastrointestinal track.
Since the α7-nAChR and the 5-HT3R are relevant drug targets, there is a high need for identifying new ligands that modulate these receptors, which can be developed further as new drug leads.
Sources
Natural Extracts as Sources of New Drug Leads
Nature has provided a large number of medicinal products for treating diseases over the course of history. The first medications known in many cultures and civilizations were made from plant extracts. The first examples of the use of natural extracts for medication are the Ebers Papyrus (Egyptian civilization, BCE 1500, a record of more than 700 natural extracts with medicinal properties), the Chinese Materia Medica (China, BCE 1100, 52 prescriptions),
70
and the Ayurvedic (India, BCE 1000, more than 300 medicinal natural extracts described) systems.
71
Besides the Middle and Far Eastern cultures, the ancient Greeks and Romans also used natural products as medicines. Hippocrates, who is considered the father of modern medicine, built the foundation of the Western medication system and applied phytotherapy, or healing with herbs, in his treatments. In the last century, many important medicines have been derived from natural sources.
72
Well-known examples of medicines discovered from natural sources are aspirin from the
Active Compounds Identified from Microbes, Plants, and Mushrooms
Today’s pharmacology would not be the same without the discovery of nicotine from the
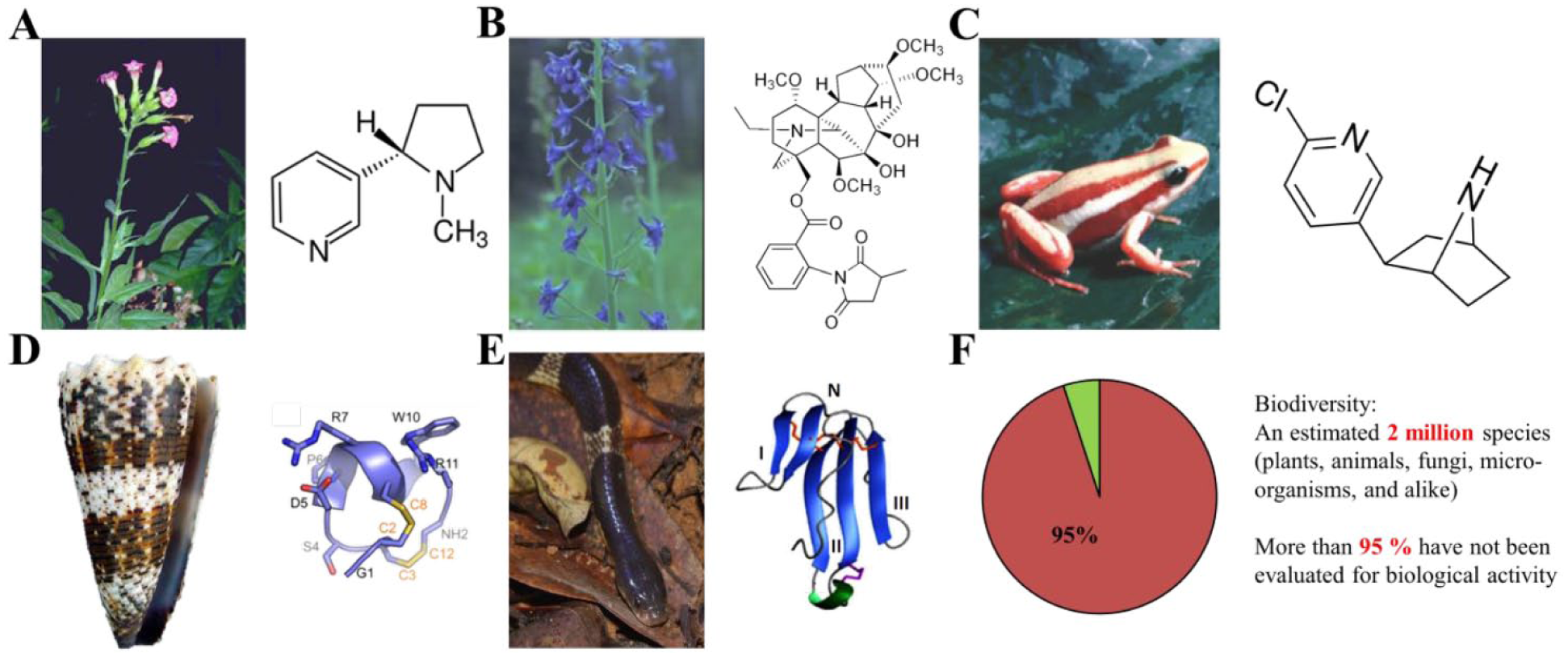
Examples of compounds targeting the α7-nAChR from natural sources. (
Active Compounds Discovered from Animal Venoms
Animal venoms are complex mixtures of biologically active compounds, which can cause various physiological effects, such as neurotoxicity and effects on the blood coagulation system, and are tissue necrotizing, when injected into a prey during the process of envenomation. The composition of venoms can range from small organic molecules to peptides and proteins, such as enzymes. Venoms can also contain metal ions, lipids, carbohydrates, and nucleosides. Diverse animal species such as the invertebrates (e.g., sea anemones, corals, mollusks, and arthropods) and vertebrates (e.g., reptiles, fish, and even mammals) produce venoms. These venoms contain a large variety of bioactive compounds, of which some are of interest as pharmacological tools and/or as new lead molecules in drug discovery.78–81
The diversity of toxins in venoms often targets a large variety of different types of ion channels, among which are the nAChRs. Venoms from marine animals in this regard are a rich source for discovering new bioactive compounds. For example, the cone snails, which are predatory sea snails, produce very potent neurotoxic venoms, each of which is a cocktail of tenths of different bioactive compounds.82–84 For instance, the α-conotoxin IMI (
Fig. 2D
) isolated from
The venoms of various types of arthropods, such as centipedes, spiders, and scorpions, contain a large variety of toxins acting mainly on different voltage-gated ion channels, but also on LGICs. Argiotoxin is an AMPA receptor blocker from the venom of the
Frogs are not typically known for injecting venoms into their prey, and therefore they are considered poisonous (the toxin acts when it is eaten, inhaled, or digested). The high-affinity nAChR agonist epibatidine was discovered from the poisonous dendrobatid frog
Snake venoms are one of the richest sources of toxins targeting the α7-nAChR. Based on their structure and function, the snake venom proteins can be divided into the following protein families: three-finger toxins, phospholipase A2s, proteinase inhibitors, serine proteinases, lectines, and metalloproteinases.
92
Most of the snake venom toxins targeting the α7-nAChR belong to the family of the three-finger toxins. α-Bungarotoxin was isolated from
Many novel bioactive compounds discovered in venoms do not pass clinical trials and never make it to a new medicine. 96 This may be due to the fact that compounds identified from natural sources often have high affinity toward target receptors and often are antagonists/inhibitors, a combination that is in most cases not an advantage. Moreover, venom-based peptides and proteins often cannot pass the blood–brain barrier (BBB) of mammals, so that they cannot reach their target in the CNS. 79 In addition, peptide and protein drugs commonly have to be administered parenterally, encompassing an increased risk on adverse immunogenic responses. A recent review on peptide therapeutics from venoms states that successfulness looks more optimistic as structure elucidation’s successes increase due to advances in proteomic and transcriptomic approaches. 96 Among others, these advances increase the potential of finding new bioactive peptides from venoms as potential therapeutics. Even if newly discovered bioactive peptides and proteins from venoms may not reach the market as medicines, they still can provide new relevant information for a better understanding of receptor expression and binding site information, and of receptor functions (physiology and structure). In addition, this information can find its way into diagnostics97,98 and allow for more efficient ways of the recombinant expression of snake venom toxins, such as recombinant viper three-finger toxins studied as potent inhibitors of nAChRs. 99 Furthermore, other applications for these compounds are found, such as in cosmetics100,101 and pesticides.102–104
Approaches
Approaches to Find New Compounds Targeting Ion Channels
Ion channels are important targets for the development of new drug lead molecules. Several “classical” approaches are used in the drug discovery process for ion-channel targets. More advanced and faster screening techniques have emerged over the last years.105–109 Classical approaches in the initial screening process and lead optimization phase include binding assays and assays measuring ion fluxes across the cell membrane (e.g., monitoring the membrane potential). More recently developed screening technologies involve automation and increase in the speed of screening. The newest assay formats can be considered high throughput, such as the fluorescence imaging plate reader (FLIPR) and calcium-flux assay. 110 Besides these high-throughput screening (HTS) assay formats, nowadays medium/high throughput can be achieved with automated patch-clamp technologies and with the newest HTS instrumentation for automated electrophysiology ion-flux assays.111–113 The following section discusses in detail the different assay formats and instrumental setups used in venom and ion-channel drug discovery, with a focus on the α7-nAChR and the 5-HT3R.
Binding Assays
Binding assays are used to detect binding of a ligand to a receptor, enzyme, macromolecule, or antibody. In order to measure the interaction between a ligand and a receptor, in most of the assay formats labeling of the ligand or the receptor is required. Depending on the type of labeling, the receptor–ligand binding assays can be classified into radioactive and nonradioactive labeling formats. 114 Assays measuring the displacement of a radioactively labeled ligand are widely used screening techniques due to their sensitivity, general use, straightforward synthesis of radioactive ligands, and easy automation in HTS. The radioactively labeled ligand assays can be performed in heterogeneous and homogeneous assay formats. In heterogeneous assays, the bound ligand needs to be separated from the nonbound ligand by filtration, dialysis, or centrifugation. Due to the labor-intensiveness of this separation step, homogeneous scintillation proximity assays (SPAs) have also been developed. In SPAs, the receptor is immobilized on a scintillation bead or scintillation well, which emits light via energy transfer when a beta-radiating radioactive ligand molecule is bound to the immobilized receptor. 115 This mechanism is based on close proximity. The beta radiation travels a short distance in water and decay is thus fast. Close to the scintillation material on a bead or coated well, the radiation is transferred to the scintillation material, which converts it into light. More recently, a study by Wittmann and Strasser (2017) investigated kinetic radioligand competition binding assays as a technique. 116 One-state models, to date, are not applicable when radioligands display biphasic binding kinetics to a receptor. In his study, Wittmann and Strasser (2017) developed a molecular model for kinetic competitive radioligand binding assays to detect two different binding orientations. In this study they targeted the GPCRs. Possible limitations of this model were studied by comparison with different traditional models targeting only one binding orientation. However, the developed model, successful as an alternative experimental method for the detection of two different binding orientations of a ligand, had some limitations as the detection of the different binding orientations was only achieved under distinct conditions. 116 Another study, by Guo et al. (2018), developed a mathematical model for determination of the binding kinetics of unlabeled ligands. A so-called two-state model for the binding kinetics determination was developed for the assessment of competitive radioligands showing biphasic binding characteristics to a receptor. 117 Radioligand binding assays for screening ligands for binding to the α7-nAChR and the 5-HT3R were extensively used in academia and industry and allowed the identification of many nAChR ligands.118–122 The disadvantages of radioligand binding assays are the high costs (especially the SPAs) and aspects related to safety, health, and disposal of radioactive waste. The filtration assays are also lower in throughput and more labor-intensive. Therefore, many efforts have been made to develop nonradioactive methods.
Nonradioactive binding assays use various types of assay readout strategies, in most cases based on fluorescence, chemo-/bioluminescence, or colorimetric detection. More specifically, fluorescence/bioluminescence resonance energy transfer (FRET/BRET),123–125 fluorescence polarization (FP),126,127 total internal reflection fluorescence (TIRF),128,129 and fluorescence enhancement assay principles can be named. 130 Fluorescence detection is often nondestructive and, due to its noninvasive character, enables live cell studies as shown by Hovius (2013). In this study, the compound GR-186741X was evaluated as a high-affinity fluorescent agonist for the 5-HT3R. 131
Nonradioactive binding assays can be performed in both homogeneous and heterogeneous assay formats, based on the different types of assay component interactions. There are various types of instrumentation for measuring binding assays, such as plate reader formats, flow cytometry,132,133 chromatographic separation formats (frontal chromatography, quantitative affinity chromatography),134–136 and surface plasmon resonance (SPR) analysis.137,138 Nonradioactive binding assays developed for screening α7-nAChR and 5-HT3R ligands include SPR-based AChBP ligand binding assays, 138 TIRF assays for studying the 5-HT3R,128,129 and fluorescence enhancement assays using AChBP as a soluble binding protein that mimics the extracellular domain of the α7-nAChR.130,139 Fluorescent ligands need sufficient affinity toward the protein target and need to give significantly high fluorescent signals for optimal in vitro (and possibly in vivo) pharmacology assessment. Often, high-affinity ligands are synthesized or modified with fluorophores to achieve this goal, as is done for 5-HT3 ligands, as discussed by the studies of Lochner et al. (2015) 140 and Jack et al. (2015). 141
Ligand binding assays in general have the limitation that only information on the binding affinity toward a receptor or ion channel is measured, and the mechanism of action (e.g., if a ligand is an agonist, antagonist, or a modulator) is not known. Also, ligand binding assays measure the interaction with a specific binding site of the receptor, and therefore ligands interacting with other sites of the receptor (allosteric modulators) are not detected. Therefore, functional cell-based assays, such as ion-flux assays and electrophysiology, are widely used for screening purposes.107,142
Ion-Flux and Membrane Potential Assays
Cell-based ion-flux assays measure the functioning, activation, and blocking of ion channels by monitoring the flux of ions passing through the channel. There are several approaches described in the literature for HTS ion-flux assays, such as flux assays based on radioactive isotopes using, for example, 86 Rb+ for studying potassium channels and nonselective cation channels,143,144 and nonradioactive Rb+-flux assays using atomic absorption spectroscopy. 145 Since only an endpoint can be measured, the kinetics of the ion-channel activation/inactivation cannot be assessed. More extensively used ion-flux assays are based on using voltage-sensitive FRET dyes or ion-selective fluorescence dyes. 146 From the ion-selective fluorescence dyes, the calcium-sensitive fluorescence dyes, such as the fluo-4 and FLIPR calcium dyes, are most widely used.147,148 These dyes, which can monitor the changes in intracellular Ca2+ ion concentrations caused by ion influxes, are highly sensitive and provide a high dynamic range. Furthermore, they provide a real-time readout (i.e., they measure calcium fluxes in time upon ligand stimulation). The disadvantage of these assays is that they can only be used for measuring calcium fluxes. After the introduction of these calcium-flux assays in high-throughput plate readers, such as the FLIPR, 110 calcium-flux assays became one of the most widely used screening techniques in the pharmaceutical industry and academic research for this type of assaying, as well as for the initial identification of new α7-nAChR ligands.149–153
A more traditional, but high-content information-providing technique to measure ion fluxes (ion currents) is electrophysiology. The most commonly used approaches for this type of measurement are the two-electrode voltage clamp (TEVC)
154
in
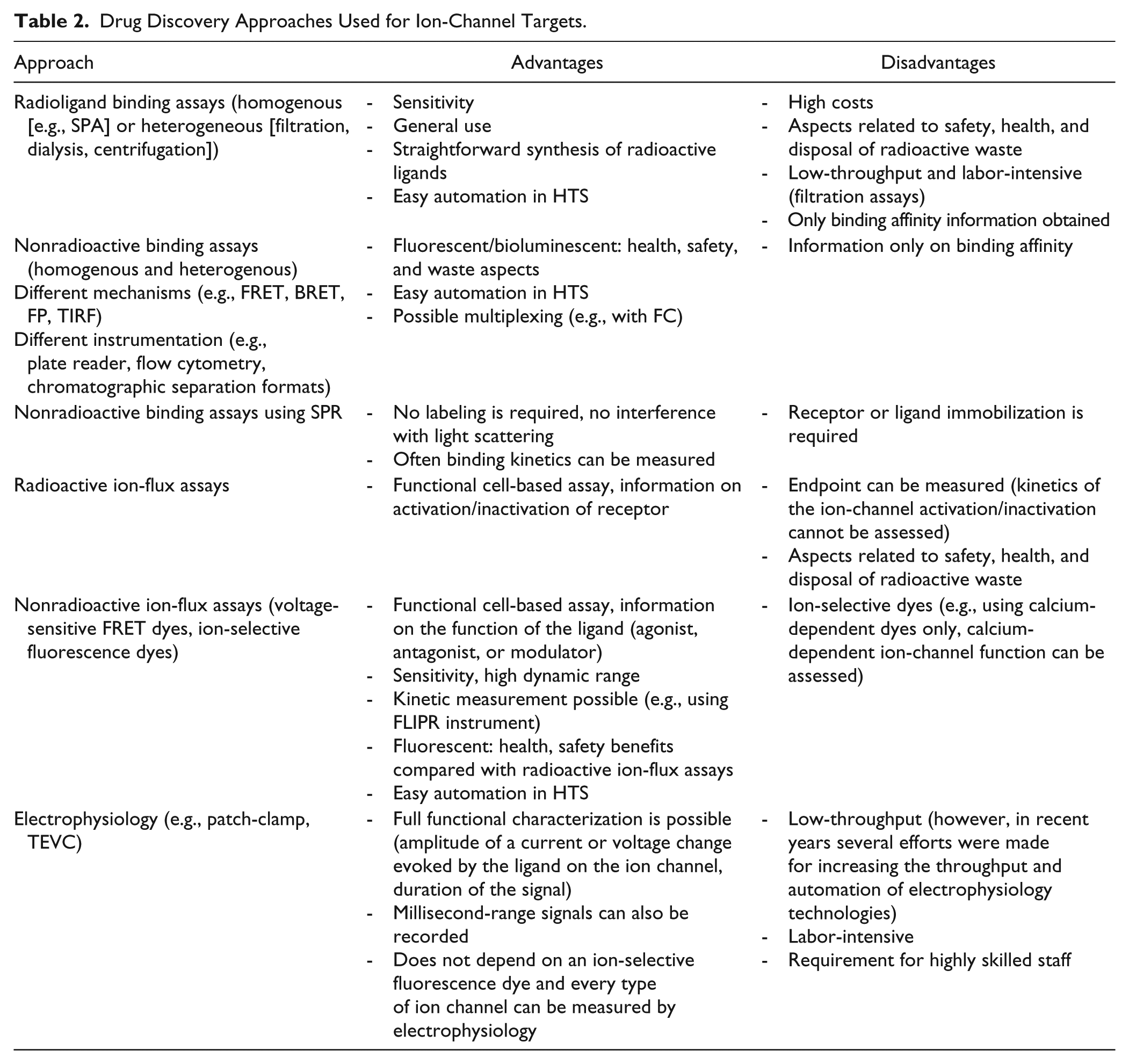
The approaches discussed in “Binding Assays” and “Ion-Flux and Membrane Potential Assays” are summarized and compared in Table 2 .
Drug Discovery Approaches Used for Ion-Channel Targets.
Classical Approaches for Screening of Mixtures: Bioassay-Guided Fractionation
A limitation of standard HTS approaches, which were discussed in “Binding Assays” and “Ion-Flux and Membrane Potential Assays,” is that they can be used only for the screening of libraries of pure compounds. When complex mixtures, such as natural extracts from plants or animal venoms, have to be screened for bioactive compounds, fractionation of the sample needs to be performed in order to allow for proper screening, detection, and identification of the bioactive compounds. This is important for the isolation of bioactive compounds in general; however, it is crucial when bioactive compounds relevant for signal transduction have to be distinguished from other compounds with other types of bioactivity. Commonly, multiple orthogonal separation steps are required in order to obtain sufficiently pure fractions to unambiguously assign the activity monitored by the applied screening assay to an individual bioactive. Also, the choice of bioassay is crucial for singling out bioactive compounds with desired activity. This traditional approach of natural extract screening is called bioassay-guided fractionation (BGF) 156 and might also be used in environmental and food analysis, where it is called effect-directed analysis (EDA).157,158 BGF screens for medicinally relevant bioactives, whereas screening for toxicants (such as endocrine disruptors) is the focus of EDA. Prior to BGF, for many sample types, such as plants and mushrooms, a pretreatment is needed in order to remove interfering matrix constituents and transfer the bioactive compounds to a solution that can be analyzed. This sample pretreatment may involve steps of prewashing, drying, or freeze-drying of the material, grinding the material to obtain a homogeneous sample, and extracting the active compounds with an adequate extraction technique (e.g., pressurized-liquid extraction, solid-phase extraction, solid-phase microextraction, supercritical fluid extraction, or a surfactant-mediated technique). 159
The isolation of bioactive compounds during the BGF process is usually performed using different separation techniques, including liquid chromatography (LC) on columns with different stationary phases, and by using thin-layer chromatography (TLC). With TLC, small quantities of complex mixtures can be separated while at the same time active compounds can be localized on the plate.158,160,161 Moreover, BGF approaches traditionally use low-resolution fractionation (fraction sizes of minutes or milliliters) after LC separation of a mixture (snake venom, plant extract, etc.), which showed bioactivity in an initial crude mixture screening. More specifically, the approach usually starts with a microfractionation step, commonly using ion-exchange chromatography (IEC) or size-exclusion chromatography (SEC) for the first separation round, after which each fraction is subjected to a bioassay of choice in order to select the bioactive fractions of interest.162,163 An orthogonal LC (often reverse-phase [RP]) analysis is then performed, including further microfractionation and bioassaying. When a bioactive compound is successfully isolated after repeated separation and bioassay steps, identification or confirmation of its chemical structure is attempted by mass spectrometry (MS), MS/MS, and/or nuclear magnetic resonance (NMR) analysis. However, even when using 2D LC, due to the low-resolution nature of the microfractionation approach, fractions collected remain complex mixtures. Consequently, this process has to be continued until eventually pure bioactive fractions (i.e., containing only one compound) are obtained for compound identification. 164 However, during these procedures bioactive compounds may get lost in the process for various reasons, such as adsorption to LC tubing or to the walls of the collection vials, precipitation, degradation/oxidation, and/or denaturation in the case of peptides/proteins. Moreover, BGF approaches are very time-consuming and labor-intensive. In addition, BGF approaches often require large quantities of initial sample. As an exception, animal venoms typically require much less sample as venom peptides and proteins often can be fully identified using LC-MS only (i.e., proteomics approaches), while for small molecular bioactives, commonly NMR is required for full structural identification. Despite these drawbacks, there are still many successful examples to be named in which bioactive compounds were discovered using BGF, and development in BGF research is ongoing.
Recently, Nothias et al. (2018) developed a workflow procedure called “bioactive molecular networking” to help in cases where BGF is not able to isolate the active compound accordingly. 165 In short, the workflow consists of the integration of MS/MS molecular networking and bioactivity scoring to assist in BGF. This would enable active compound detection directly from fractionated bioactive extracts and would make it possible to accelerate the dereplication of molecules using molecular networking prior to subsequent isolation of the compounds. Further development entails using bioactivity score prediction to help expose additional potentially bioactive molecules.
In a recent study by Lee et al. (2017), BGF was applied for the isolation of compounds from
Plant-derived bioactives, which mostly are low-molecular-weight compounds, may be a starting point for medicinal chemistry approaches to yield optimized leads with desired pharmacodynamic and pharmacokinetic profiles. Bioactives from venoms often are peptides or (small) proteins, and usually proteomics and biochemical approaches are needed for their structural and biological characterization. Structure optimization steps might include peptide synthesis of derivatives in the case of bioactive peptides and overexpression in suitable expression systems in the case of protein bioactives.
Hyphenated Approaches for Complex-Mixture Screening
Recently, BGF protocols have been redesigned in order to deal with some of the bottlenecks during the traditional BGF approach. As an example, the so-called at-line nanofractionation approaches can be mentioned. This workflow can be used for the rapid screening of medicinally relevant compounds in complex matrices, such as snake venom, and other complex mixtures. It can be applied using many different types of off-line assays, such as enzymatic and/or cell-based assays, for bioactivity assessment.167–170 In this setup, the separation column effluent is split to provide parallel MS detection and high-resolution (second-range) fraction collection. Recently, a fraction collection machine (FractioMate) was developed by the Vrije Universiteit and Spark Holland and is now commercially available.
171
A typical at-line nanofractionation setup is shown in
Figure 3
. The small second-range fractions are continuously collected onto 96-, 384-, or 1536-well plates. Usually, the plates are then vacuum-centrifuged to dryness, followed by direct on-plate pipetting of the bioassay reagents using pipetting robotics, followed by incubation and plate reader readout. In this setup, assays are developed post-LC separation toward a bioactivity of choice with the possibility of additionally adding test compounds postfractionation in order to induce targeted inhibition, activation, and/or quenching of the biochemical reaction toward obtaining more high-content information on bioactivities observed. Furthermore, by applying the high-resolution fraction collection, cleaner samples can be obtained and the parallel MS data gives the advantage of directly assigning an accurate mass to a bioactive compound. In the case of profiling for bioactive peptides and proteins from mixtures (e.g., venoms), the method also allows straightforward subsequent on-plate proteomics approaches (i.e., using the fractionated well plate) toward structure elucidation. This redesigned BGF approach further increases the possibility of successfully isolating bioactive compounds relevant in signal transduction. An example of this approach includes the development of an at-line cell-based screening methodology that combines LC followed by at-line nanofractionation and parallel MS with a functional fluorescence-based calcium-flux assay.
172
This calcium-flux assay was performed with mammalian cells stably overexpressing the α7-nAChR. This nanofractionation system was developed and optimized in assay modes for screening agonists and for allosteric modulators of the α7-nAChR. The application of this methodology was demonstrated by the screening of a hallucinogenic mushroom extract (from
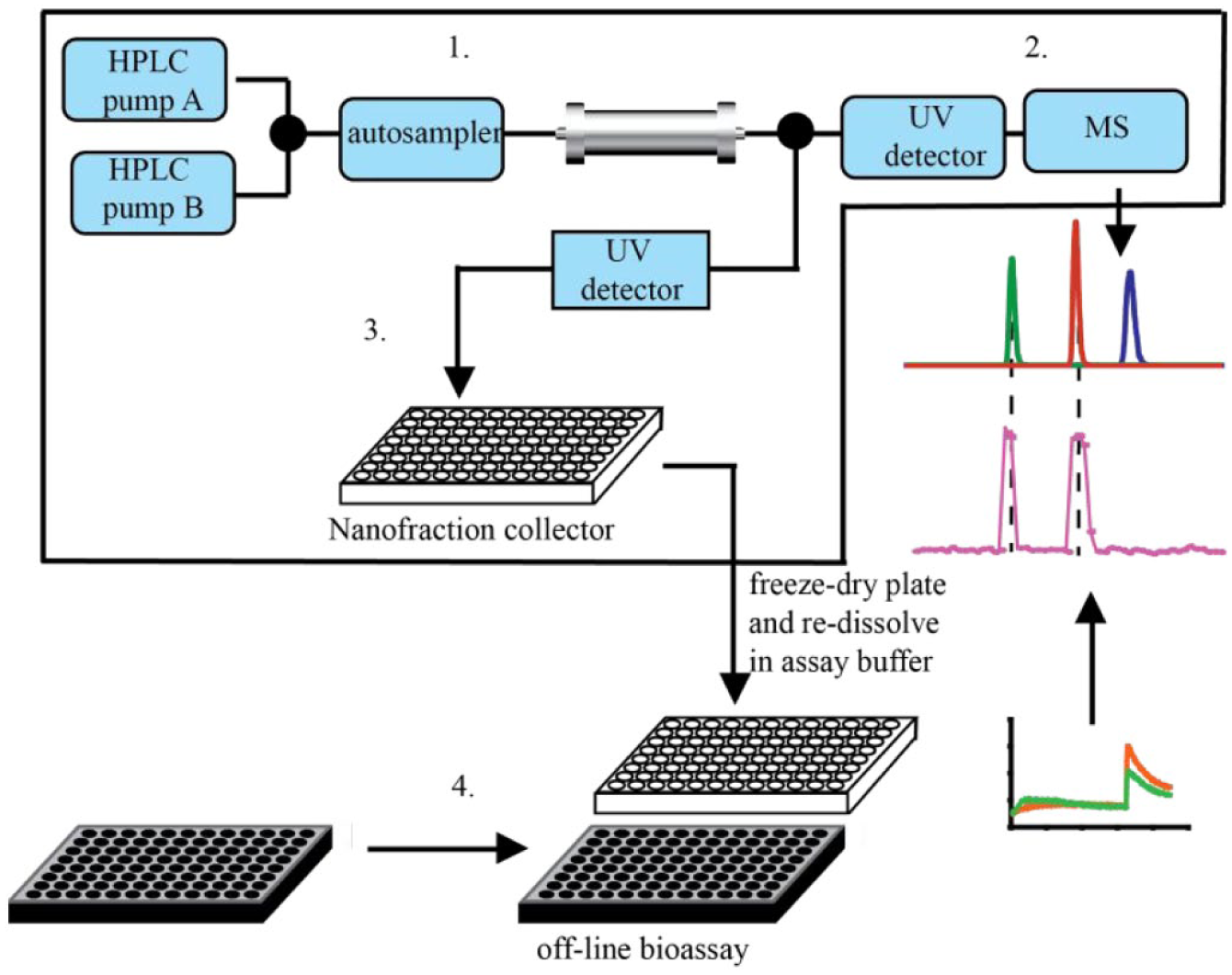
Principle of the at-line nanofractionation approach. After LC separation (1) of a complex mixture, the flow is split in two. One part of the flow is directed to a UV detector followed by MS (2), while the other part (usually the larger part, like 90%) of the flow is collected as nanofractions into well plates by a nanofraction collector (3). The well plates are usually vacuum-centrifuged after nanofractionation and followed by a bioassay that is performed off-line on the nanofractionated well plates.
Other more automated analytical techniques have been developed for the screening and identification of bioactive compounds in complex mixtures. High-resolution screening (HRS) is based on the coupling of an LC separation with a continuous-flow bioassay for direct assessment of the biological activity of eluting compounds.173,174 The assay reagent mixture is continuously mixed with the LC effluent and directed to a coil providing incubation and reaction time. HRS also commonly encompasses parallel MS detection using a postcolumn split to yield accurate mass information on eluting compounds, including the bioactive ones. A typical HRS system is depicted in
Figure 4
. Compared with the nanofractionation approach, HRS is far less time-consuming and the obtained bioassay readout has a higher time resolution. HRS is fast and allows the screening of many samples per day. Various types of biochemical assays have been successfully applied in HRS systems, although mostly enzymatic and binding assays.130,175,176 Normal bore chromatography HRS approaches require relatively large quantities of sample and biological reagents. This can be circumvented by employing miniaturized HRS systems, which consume much lower (bio)reagent amounts, and analyzing very low sample volumes. These so-called microfluidic HRS systems have been developed for profiling biologically active mixtures toward different targets including the AChBP. Using a postcolumn split, nano-LC is coupled in parallel to MS, via nano-electrospray ionization (nano-ESI), and to a microchip bioassay that utilizes a capillary bubble-cell LED-induced fluorescence detector for readout. The platform is operated in the range of nanoliter-per-minute flow rates and with nanoliter-range sample injection. The homogeneous bioassay is based on fluorescence enhancement caused by a tracer molecule bound to the AChBP. A decrease in fluorescence when the tracer is displaced by an eluting ligand binding to AChBP is used as bioassay readout. In a typical example, microfluidic HRS was developed and applied to the identification of AChBP bioactives in venoms.
177
Subsequent work demonstrated both microfluidic HRS identification and follow-up purification of the bioactive compounds identified using straightforward MS-guided isolation.
178
Subsequent bottom-up proteomics approaches were then used for chemical characterization of the bioactives. Next to snake venom profiling, other venoms and toxins were also screened using this technique.
167
In the case of screening
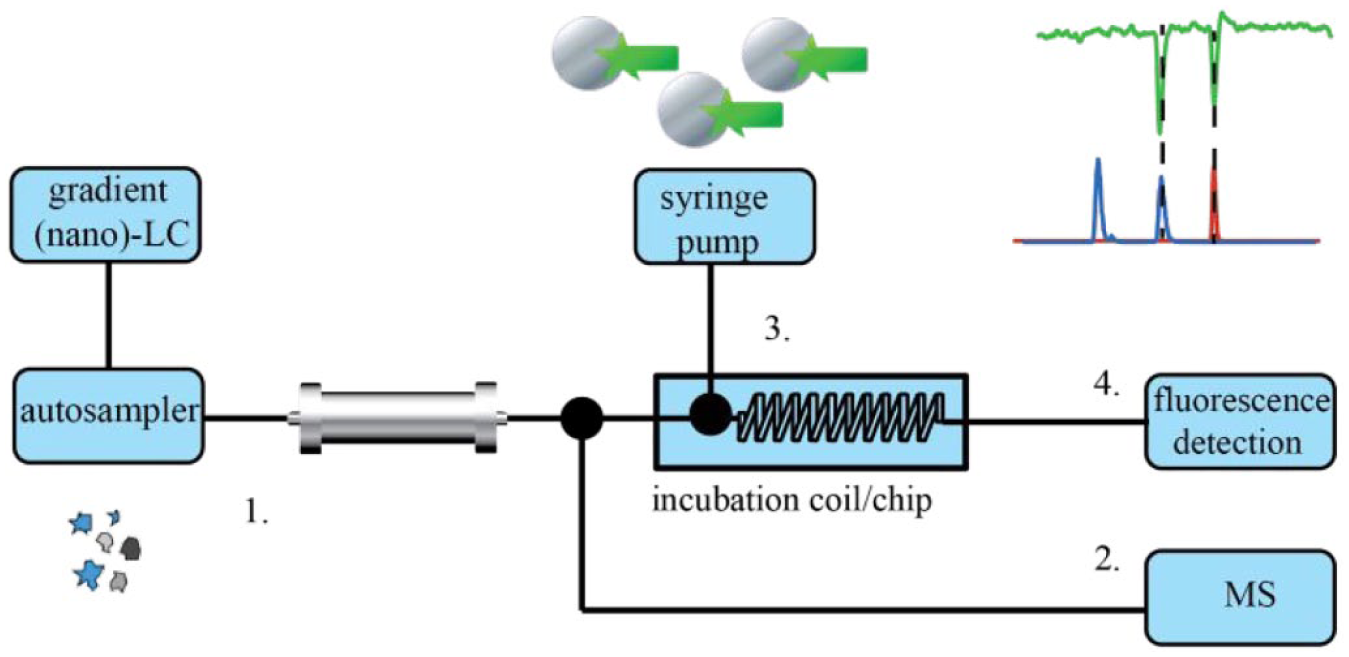
Principle of an on-line postcolumn HRS system. After LC (or nano-LC) separation (1), the eluent flow is split into two parts. One part of the flow is directed to MS (2) for the identification of the eluting compounds, and the rest of the flow is directed to a continuous-flow incubation coil (3), in which eluting compounds are incubated with bioassay reagents, followed by a fluorescence detector for bioassay readout (4) measuring the biological activity (in the case of fluorescence assays).
Next to the fluorescence enhancement assay for the AChBP, a fluorescence enhancement assay has also been developed for the serotonin binding protein (5HTBP). This was first achieved in plate reader format and then implemented in the microfluidic HRS platform. 167 The 5HTBP is an engineered binding protein that has the protein scaffold of the AChBP with the ligand recognition properties of the 5-HT3R. This fluorescence enhancement assay is a good initial screening technique for finding novel bioactives targeting the 5-HT3R. The applicability of the new technique was demonstrated by the screening of different snake venoms for compounds binding to 5HTBP.
Cell-based assays unfortunately in most cases cannot be used in HRS systems, primarily due to the technical difficulty of combining living cells with chromatographic eluents. However, for some cell lines overexpressing ion channels, this approach is feasible although technically and biologically extremely challenging. Heus et al. (2014) describe the analytical advancement to such a technique in which a Ca2+-flux assay is performed in postcolumn continuous-flow format using a flow cytometer (FC) as the readout device. 167 The screening system consisted of an LC system coupled on-line to FC and MS using PEEK tubing and so-called superloops for the infusion and mixing of the bioassay components. The bioassay applied in this system was based on a calcium-flux assay using human α7-nAChR expressing SH-SY5Y neuroblastoma cells. This LC-FC-MS screening system was developed in agonist and in mixed antagonist–agonist assay modes. The latter assay mode allows the simultaneous detection of agonists and antagonists. In proof-of-principle experiments, the application of the screening system was demonstrated by the screening of tobacco plant leaf extract in agonist mode, and snake venoms in mixed antagonist–agonist assay mode.
A disadvantage of the on-line HRS approaches is that assays have to be relatively fast (incubation times of seconds to a maximum of a few minutes) in order to obtain a measurable response. Assays with long incubation and/or assay preparation times, such as radioligand binding assays, are out of scope and need to be addressed by the more traditional BGF or recently developed nanofractionation approach.
Next to the postcolumn HRS techniques and the nanofractionation approach, which both apply bioassays after chromatographic separation, other bioactivity screening approaches of mixtures are available. These can be divided in precolumn and on-column bioactivity profiling methods. These methods in general perform either an affinity extraction of ligands from a complex mixture using a selective precolumn, or a chromatographic separation using an affinity column. 179 Using these precolumn and on-column approaches, bioactives bound to the affinity stationary phases can be separated from nonbinders before detection. Alternatively, the affinity proteins with ligands bound are separated from nonbinders predetection using ultrafiltration or size-exclusion separations. Techniques measuring protein–protein affinity and immobilized ligand–protein affinity interactions, 180 MS binding assay approaches, 181 and coupling ultrafiltration 182 or size-exclusion separations 183 to LC-MS can be named in this regard. Recently, Sichler et al. (2018) developed an MS-based binding assay for nAChRs using the ligand MB327, where a deuterated MB327 analogue was used as reporter ligand. 184 The principle of MS binding assays is similar to radio ligand binding assays; however, the readout is based on mass spectrometric detection, instead of a radiolabeled (or fluorescent) reporter ligand.
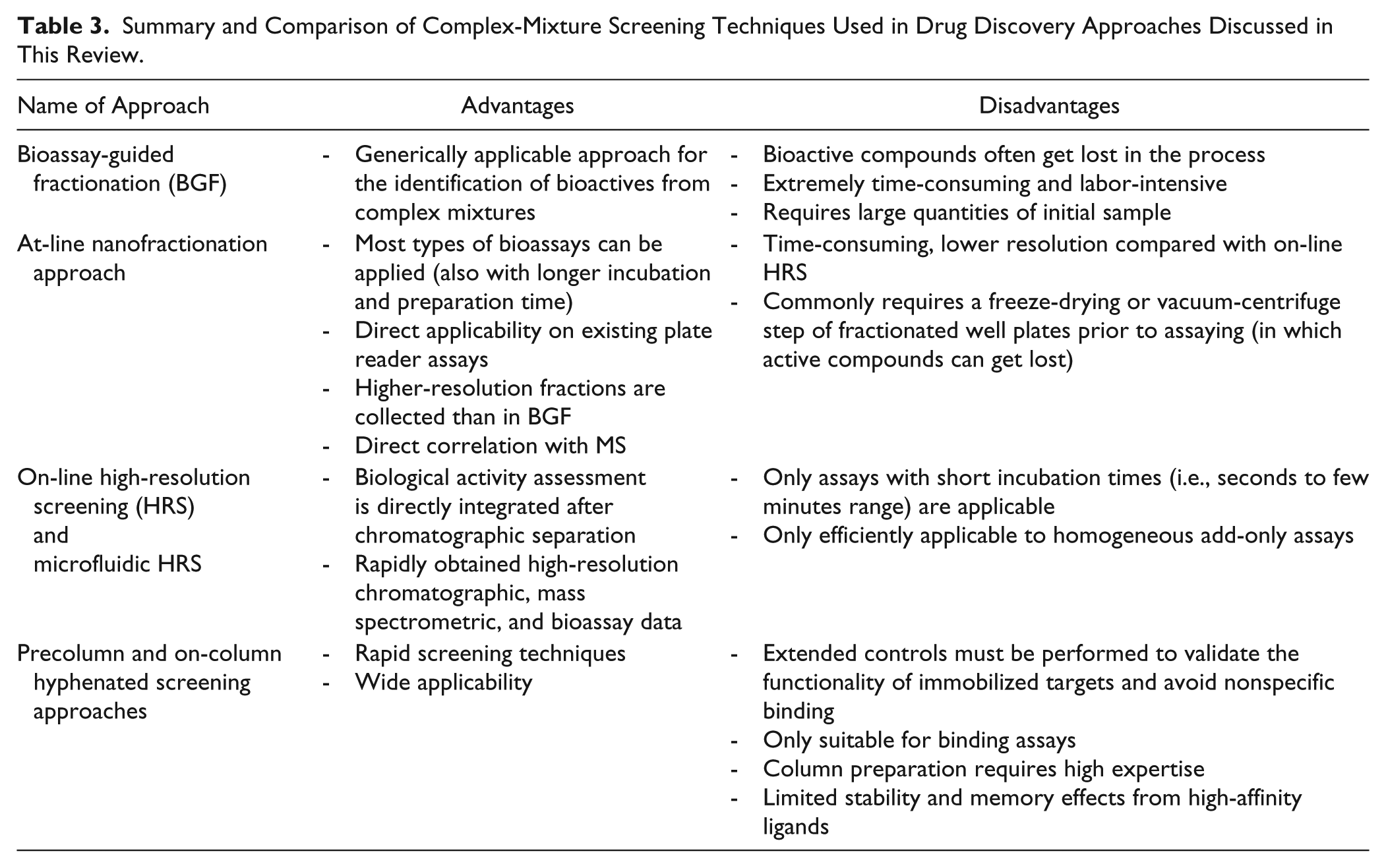
Precolumn and on-column bioactivity profiling methods are comprehensively described in reviews.173,180,185 An example of yet another precolumn bioactivity profiling methodology used in nAChR drug discovery is a magnetic bead-based affinity-selection methodology for the identification of AChBP ligands in mixtures. 135 A summary and comparison of complex-mixture screening techniques used in drug discovery approaches is given in Table 3 .
Summary and Comparison of Complex-Mixture Screening Techniques Used in Drug Discovery Approaches Discussed in This Review.
Drug Discovery Workflows toward Identification and Characterization of Venom Peptide-Derived Leads
In the previous part of this study, strategies for the screening and detection of components in natural extracts with relevant bioactivity were discussed. The focus in the following part of this review is the characterization of possible drug leads from natural extracts. The main focus is on snake venom as a matrix for novel drug leads using MS. In the previous part, MS was often mentioned as a popular, fast-growing technique that is gaining in importance in this field. Classical as well as novel approaches are also briefly discussed. Identifying and studying complete venom proteomes (i.e., all proteins and peptides in a venom and venom glands) comprehensively, by proteomics, transcriptomics, and genomics, is called venomics, an approach recently reviewed by Oldrati et al. (2016). 186 MS is a very important technique in this field.187,188 Venomics can be used to better understand venom biodiversity and therewith functioning and evolution, and also for the identification of new lead compounds in drug discovery.80,189 When applying venomics for drug discovery, as described in “Classical Approaches for Screening of Mixtures: Bioassay-Guided Fractionation,” in most cases subsequent BGF purification and characterization of toxins is of interest and often performed by using a two-step purification involving successive SEC and reverse-phase liquid chromatography (RPLC). 190 These are crucial steps for decreasing sample complexity toward successful MS measurements, by drastically decreasing ion suppression and improving the detection of low-abundance peptides and proteins. The mentioned nanofractionation with subsequent MS analysis is also a relevant technique in this field by the efficient combination of bioassaying and MS, directly after chromatographic separation. The introduction of nano-LC into the venomics workflow nowadays allows for highly reduced injection volumes, enabling the study of venoms available only in low amounts.
The most popular MS ionization approaches for life sciences research are ESI, easily combined with standard on-line high-performance liquid chromatography (HPLC) methodologies, and matrix-assisted laser desorption ionization (MALDI), where fractions are collected for subsequent off-line MALDI-MS/MS measurement. Another use here is in situ MALDI imaging-MS, as by Brunetti et al. (2018), for example, where peptides of skin tissue of a hylid tree frog were characterized, studying the difference in postsecretory peptide cleavage. This information can be valuable toward unraveling information on peptide function and mechanism of action, as some of these peptides show neuroactive and antimicrobial activities. 191 MALDI imaging-MS can also be used for neurotransmitter distribution visualization/mapping of brain tissue, specifically focused on ACh 192 and reviewed by Romero-Perez et al. (2015). 193 Several studies have suggested that using MALDI-MS/MS and ESI-MS/MS in parallel for the study of venomics gives rise to complementary mass spectral data of venom compounds and therewith a more complete view of the venom profile.186,194,195 The most widely used approach for peptide and protein identification, into which many of the relevant drug leads fall, is the bottom-up approach, using amino acid sequence determination of the peptide/protein toxins after enzymatic digestion, followed by LC-MS and MS/MS (using either crude venom samples or prefractionated venom samples) or 2D sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE).196,197 SDS-PAGE comprises separation of the venom proteins by one gel electrophoresis and subsequent staining for visualization. Alternatively, 2D gel electrophoresis of complex venom samples can be achieved in which separation occurs by isoelectric focusing (IEF) for isoelectric point separation (first dimension) and by SDS-PAGE for molecular weight separation (second dimension). After separation, individual spots can be excised and in-gel digestion can be performed, followed by LC-MS/MS analysis. Unfortunately, SDS-PAGE does not work well for low-molecular-weight peptides.187,196 After digestion of the proteins/peptides, the exact mass and fragmentation patterns determined are used for the identification of the peptides/proteins present in a sample using databases. Alternatively, N-terminal Edman sequencing can be used, but the peptides have to be analytically pure for this approach.
Identification resulting from the bottom-up approaches is nowadays reasonably automated; however, due to the absence of specific species information (e.g., genomes) in the available databases, in some cases the only possibility for identification is to assign found peptides to known protein families on the basis of similarity with existing sequence entries of other similar species for which information is available in a database. This results in incomplete sequence coverages, and generally it is difficult to differentiate between proteoforms. 198 Recently, advances were made toward top-down proteomics (TDP) approaches, so-called top-down venomics. In these approaches, intact peptides and proteins are measured by MS. When leaving the molecules intact, ideally, complete protein masses and their fragment ions can be determined. More complete information on the identification and quantification of the toxin analyzed can be achieved compared with bottom-up approaches.198–200 A successful TDP approach, in this field, includes a denaturation step and is therefore called denaturing top-down proteomics (dTDP). In this approach, denaturation is performed by either a denaturing substance (i.e., reducing agents optionally assisted by detergents and/or organic modifiers, etc.) or sometimes physical methods, toward disrupting protein interactions and quaternary conformations (e.g., by heat, pressure, etc.). 199 The dTDP approach usually consists of a prefractionation step such as IEF to reduce the complexity of the sample. The fractions are then measured with RPLC-MS/MS at low pH. To date, qualitative and quantitative analysis of intact peptides and proteins up to ~30 kDa can be achieved using this methodology. 199 Native TDP is a similar approach and is used, as by Pla et al. (2018), after reduction of the disulfide bonds. 200 However, it is not yet widely used and advances have to be made in both MS instrumentation, in order to retain potential biologically relevant noncovalent protein–protein and protein–ligand interactions, and database quality. 201
One of the challenges in these approaches is considering posttranslational modifications (PTMs), such as amidations (e.g., for scorpion and spider peptide toxins) and glycosylations (mainly venom enzymes) on the toxins present in venoms. Some toxin classes may carry PTMs that can have a great influence on their biological and pharmacological functioning, as well as on their proteoform diversity. MS analysis approaches in this regard are valuable for unraveling these PTMs toward complete characterization. 186
A relatively new technique for the analysis of binding interactions is ion mobility–mass spectrometry (IM-MS), by using ESI-IM-MS. 202 Young et al. (2015) developed an HTS approach for the screening and identification of amyloid assembly inhibitors. 202 Enabling the identification of protein–ligand interactions and providing information on the binding nature of the interacting species, this technique can be used as a tool for the identification of ligands in the drug discovery pipeline.
One of the growing research areas aiding in improved protein identification and automated MS identification is the development and advancement of comprehensive reference databases and bioinformatics algorithms/processing software. Databases are available for both bottom-up and TDP proteomics approaches. Databases exist containing, for example, genomic and transcriptomic data for the bottom-up approaches (e.g., Uniprot, National Center for Biotechnology Information [NCBI], and Indigenous Snake Species of Bangladesh [ISOB] [snake venom]). Regular updating of, for example, Arachnoserver (spider venoms) and Conoserver (cone snail venoms) will improve protein identification further.186,196
After bioactive peptide or protein screening, detection, and identification, subsequent large-quantity production can be performed. This is often done using solid-phase peptide synthesis or recombinant expression with a suitable expression system, often
The above-mentioned approach was used for peptide and protein bioactives from animal venoms. However, examples of extracting small molecular bioactive compounds can also be found from other animal sources, for example, epibatidine from the
A few case studies of the traditional approach of isolation and characterization of toxins from animal venom sources are given next. The first example deals with the isolation and characterization of an anticoagulant protein, fasxiator, from
The isolation and pharmacological characterization of a neurotoxic three-finger toxin targeting the α7-nAChR from black mamba (
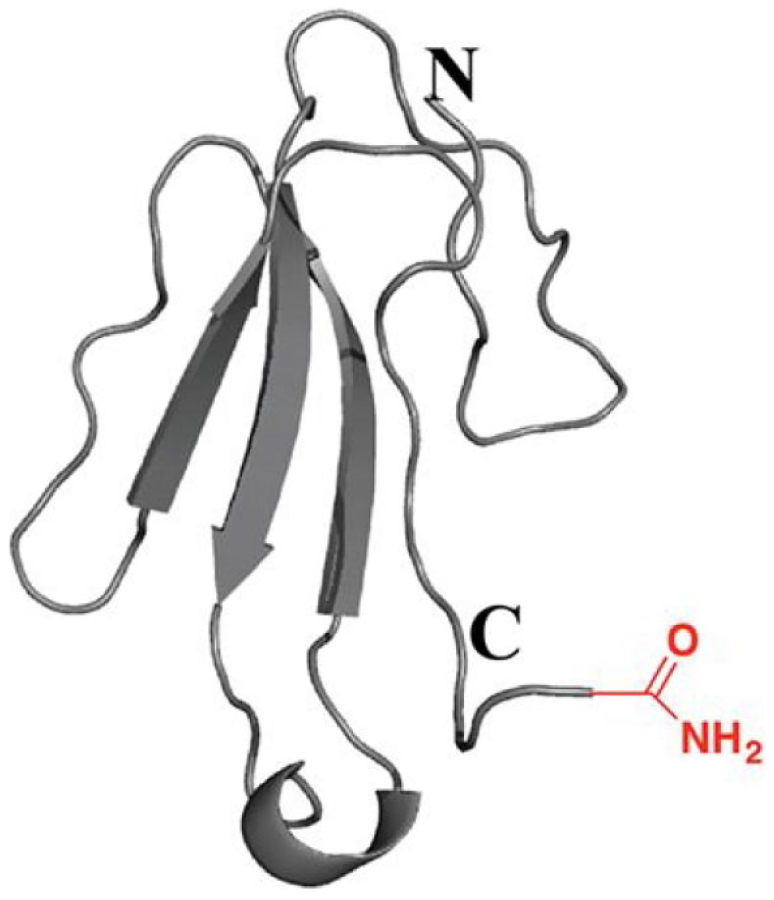
An example of a chemical structure that was isolated from natural extracts, targeting the α7-nAChR, presented in this paper as a case study example. The peptides were produced using an
Cardoso et al. isolated an inhibiting peptide of the voltage-gated sodium channel hNav1.7, which is a potential drug target for the treatment of pain disorders, from a spider venom.
210
Crude venom (1 mg) of
The ω-conotoxin MVIIA, which was the template of the approved painkiller drug ziconotide (Prialt), was discovered from the venom of the
A recent approach in the venom-based drug discovery field is the integration of traditional BGF with proteomics, transcriptomics (all mRNA present in the venom gland), and bioinformatics approaches. 213 For this approach, evidently proteome and transcriptome databases need to be available. This approach can provide efficient assignment of masses of bioactives and comprehensive elucidation of sequences of bioactive peptides and proteins.
Summary
As discussed in the “Targets” part of this review, there is a need for new compounds targeting ion channels. Besides being of interest as new drug lead molecules for CNS diseases, these compounds are also of interest as potential pharmacological and diagnostic tools. Nature is a rich source of compounds targeting ion channels such as the α7-nAChR and the 5-HT3R. There are multiple ligands of these receptors that have been identified from natural sources, as overviewed in the “Sources” part of this review. As natural extracts are complex mixtures, this makes the finding and identification of unknown bioactive components very challenging. There is thus an essential need for the development of new analytical approaches that allow for faster and more efficient screening of natural extracts for bioactives targeting ion channels. The “Approaches” part of this review aimed to discuss and review analytical techniques for the identification of bioactives from complex mixtures (such as natural extracts) acting on ion channels, with a focus on the α7-nAChR and the 5-HT3R. In this discussion, traditional and new analytical techniques were compared and evaluated for their potential and value in drug discovery from natural products in general, and for ion-channel drug discovery specifically.
Footnotes
Acknowledgements
The authors would like to thank Alexander Medvedev and Thomas Brown for providing us with Figure 2D,E, respectively.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work of Reka A. Otvos was supported by the AIMMS Bridging PhD project “Identification of Novel Bioactive Substances on Brain Receptors” (project no. 10-001-203).
