Abstract
The Scripps Research Molecular Screening Center (SRMSC) was founded in 2004 and comprises more than $22 million of specialized automation. As part of the Translational Research Institute (TRI), it comprises early drug discovery labs and medicinal chemistry. Together with Scripps Research at the La Jolla, California, campus, this represents one of the most competitive academic industrial screening centers worldwide. The SRMSC uses automated platforms, one a screening cell and the other a cherry-picking platform. Matched technologies are available throughout Scripps to allow scientists to develop assays and prepare them for automated screening. The library comprises more than 1 million drug-like compounds, including a proprietary collection of >665,000 molecules. Internal chemistry has included ~40,000 unique compounds that are not found elsewhere. These collections are screened against a myriad of disease targets, including cell-based and biochemical assays that are provided by Scripps faculty or from global investigators. Scripps has proven competence in all detection formats, including high-content analysis, fluorescence, bioluminescence resonance energy transfer (BRET), time-resolved fluorescence resonance energy transfer (TR-FRET), fluorescence polarization (FP), luminescence, absorbance, AlphaScreen, and Ca++ signaling. These technologies are applied to NIH-derived collaborations as well as biotech and pharma initiatives. The SRMSC and TRI are recognized for discovering multiple leads, including Ozanimod.
Introduction
Scripps Florida was founded in 2004 as a division of Scripps Research, headquartered in La Jolla, California, and is located in Jupiter, Florida (Palm Beach County). Using the latest cutting-edge technologies, researchers at Scripps Florida focus on basic biomedical research and drug discovery. More than 600 employees, with 56 full-time faculty members, work at the 350,000 ft2 campus, which comprises three state-of-the-art research buildings. Much of the work at Scripps Florida is dedicated to basic biomedical research, a vital segment of medical research that seeks to decipher the most fundamental processes of life. Additionally, researchers at Scripps Florida are developing advanced technologies and applying these tools to the discovery of new therapeutic agents for a variety of devastating human diseases. Departments represented at Scripps are Immunology and Microbiology, Integrative Structural & Computational Biology, Neuroscience, Molecular Medicine, and Chemistry. Scripps also has a graduate program offering doctoral (PhD) training in chemical and biological sciences on the La Jolla and Florida campuses.
The Scripps Florida campus is uniquely positioned to provide a multidisciplinary approach with ready access to advanced instrumentation for next-generation sequencing, proteomics, x-ray crystallography, structural biology, and high-throughput drug screening, offering unique opportunities to discover new therapeutic targets and identify drug leads. Scripps Florida builds on its current strength in medicinal chemistry for biomedical research and focuses on the development of enabling chemical technologies to turn biological insights into small-molecule probes, therapeutic leads, and ultimately drugs to solve global health challenges.
Support for drug development efforts is also provided through the Drug Metabolism and Pharmacokinetics (DMPK) Core at Scripps Florida and is staffed by experienced professionals with backgrounds in the pharmaceutical and biotech industries. The Scripps Florida Animal Resources Center (ARC) is dedicated to providing a comprehensive state-of-the-art program of animal care, comparative medicine, and associated animal-related resources in support of biomedical research performed by Scripps Florida investigators. The Scripps Florida animal facility is an Association for Assessment and Accreditation of Laboratory Animal Care (AAALAC)-accredited 25,000 ft2 vivarium, which includes animal holdings, procedures, behavioral rooms, and the Behavioral Core.
Scripps Research Molecular Screening Center
Currently under the directorship of Dr. Louis Scampavia and Timothy Spicer, the Scripps Research Molecular Screening Center (SRMSC) has been in nonstop operation since 2005 as a major academic high-throughput screening (HTS) facility and formerly as one of the four comprehensive HTS providers during the NIH Molecular Libraries Programs Center Network era.
Annually, we conduct 20–30 large-scale HTS campaigns (+665,000 compounds) that require numerous smaller-scaled screening efforts to support these large-scale HTS campaigns. This includes pre-HTS campaign support with assay development, pilot screening, and post-HTS campaign support that includes confirmation screening, counterscreening, and dose titration screening, also providing medicinal structure–activity relationship (SAR) support with dose testing for potency and selectivity.
Since establishing operations in 2005, the SRMSC has completed more than 300 primary campaigns against a large diversity of targets to generate more than 80 million data points for both industrial and academic collaborations. Numerous scientific papers (>110 from the SRMSC alone) and presentations demonstrate our ability to produce excellent research-driven data. Our screening results have led to the development of more than 72 bioactive molecular probes as novel drug targets for biomedical studies (https://hts.florida.scripps.edu/). Two probes have now been approved as investigational new drugs (INDs) by the FDA as new chemical entities (NCEs) for clinical trials in the treatment of multiple sclerosis (MS) and cancer as licensed by Celgene (Summit, NJ) and Cleave Biosciences (Burlingame, CA). 1
During the Molecular Library Probe Production Centers Network (MLPCN) era, the SRMSC was one of four comprehensive screening centers that supported screening, informatics, and medicinal chemistry. During the MLPCN, the SRMSC performed 235 HTS campaigns from 2005 through 2013. Comparisons to other MLPCN comprehensive centers are limited to data available from PubChem. These data include the total number of assays performed (multiple assays per HTS campaign), total number of substances tested, and active number of substances tested. The SRMSC ranked first for total number of assays performed and second in both tested and active substances, behind the NIH Chemical Genomics Center. Additional cross-center comparisons of more sophisticated descriptors, such as assay reporters and assay types, would be interesting to measure; however, this is currently impossible to accomplish due to a lack of standardized assay descriptors, unique assays performed, and terms in PubChem. Coupled to our HTS efforts is Scripps Florida’s expertise in medicinal chemistry and pharmacology; Scripps Florida was responsible for 14 of the 18 licensed probes ( Fig. 1 ). Moreover, Scripps Florida was the only comprehensive HTS screening center to forward probes into INDs for clinical trials.
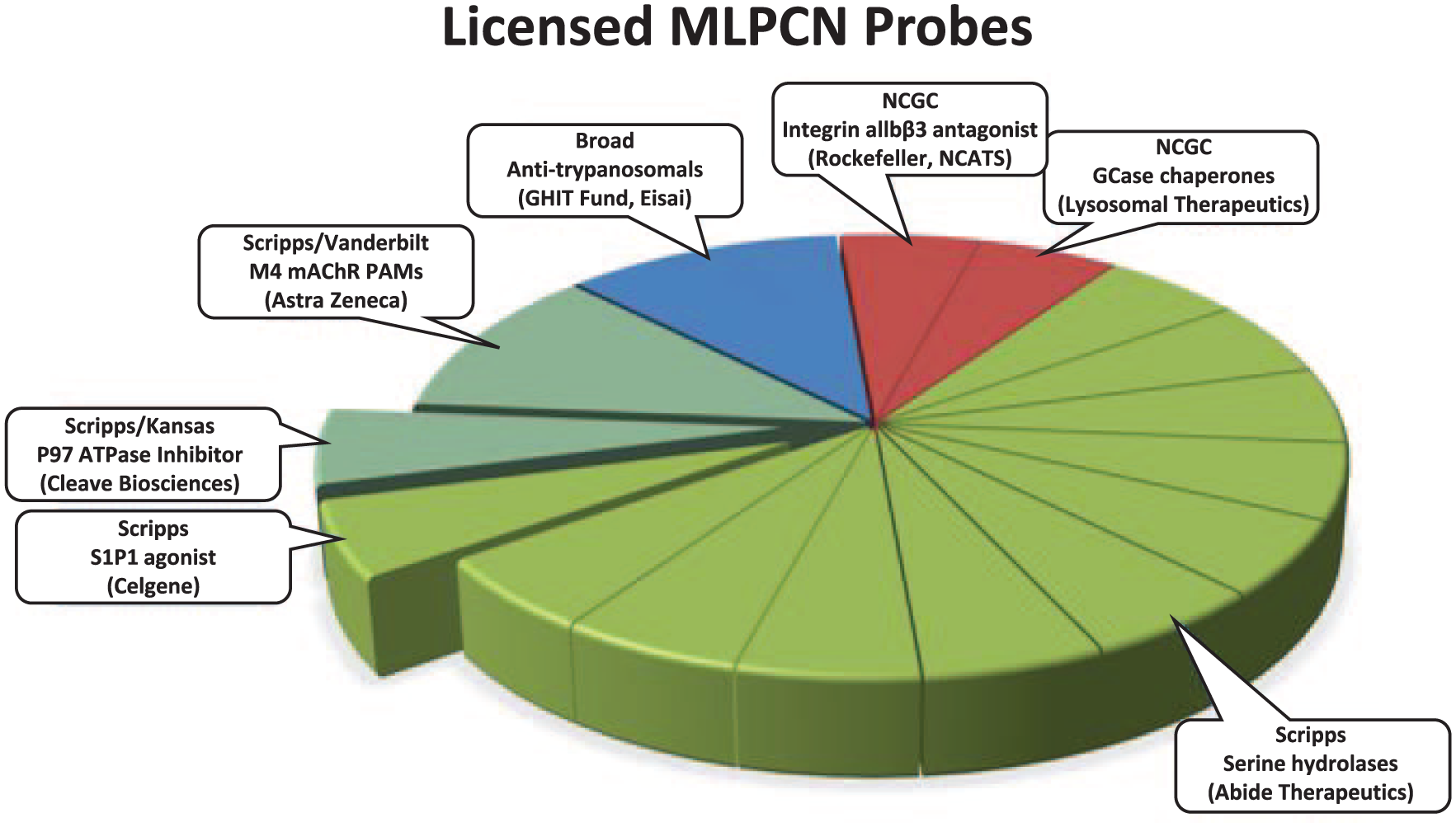
Fourteen of the 18 probes licensed from the MLPCN initiative were discovered at Scripps Florida.
The SRMSC also supports research via fee-for-service collaborations with industrial partners. These collaborations allow for flexibility in screening the SRMSC 665,000-compound library, partner-provided libraries, or both. For partner-provided libraries, the SRMSC also offers compound management services for compound registration, reformatting, quality control (QC), and cherry-picking. Typical turnaround times in fee-for-service collaborations entailing a full 665,000-compound library HTS campaign, subsequent confirmation screens and counterscreens, concentration response determination, liquid chromatography–mass spectrometry (LC-MS), and final reports deliverables are in the 4- to 6-month range. This is based on the availability of reagents, timing of assay delivery, and assay development requirements, all dictated to a degree by the assay provider. Many of the collaborations established between the SRMSC and industrial partners have also resulted in publications, including research performed for Proteostasis Therapeutics (Boston, MA), 2 Eli Lilly (Indianapolis, IN), 3 and Eutropics Pharmaceuticals (Cambridge, MA). 4
High-Throughput Screening at the SRMSC
By combining robotics, biology, chemistry, and informatics, the SRMSC is able to perform HTS, a fully automated and fully functional robotic process that tests hundreds of thousands of drug-like compounds for biological activity using biochemical interaction or cell-based assays. Currently, the SRMSC employs a 1536-well plate format to screen its Scripps Drug Discovery Library (SDDL) of more than 665,000 small-molecule entities in a rapid and economical fashion. To this end, the SRMSC has two industrial robotic platforms from GNF Systems (San Diego, CA) dedicated to running ultra-HTS (uHTS) campaigns and providing hit-picking compound management support ( Fig. 2 ). In addition to the deployment of commercial uHTS platforms, the SRMSC has also invested considerable engineering effort into further automating all aspects of the uHTS environment. This includes the development of in-house software to fully integrate the GNF HTS and hit-picking platforms to the SRMSC Laboratory Information Management System (LIMS) and custom QC platforms used to ensure data fidelity.
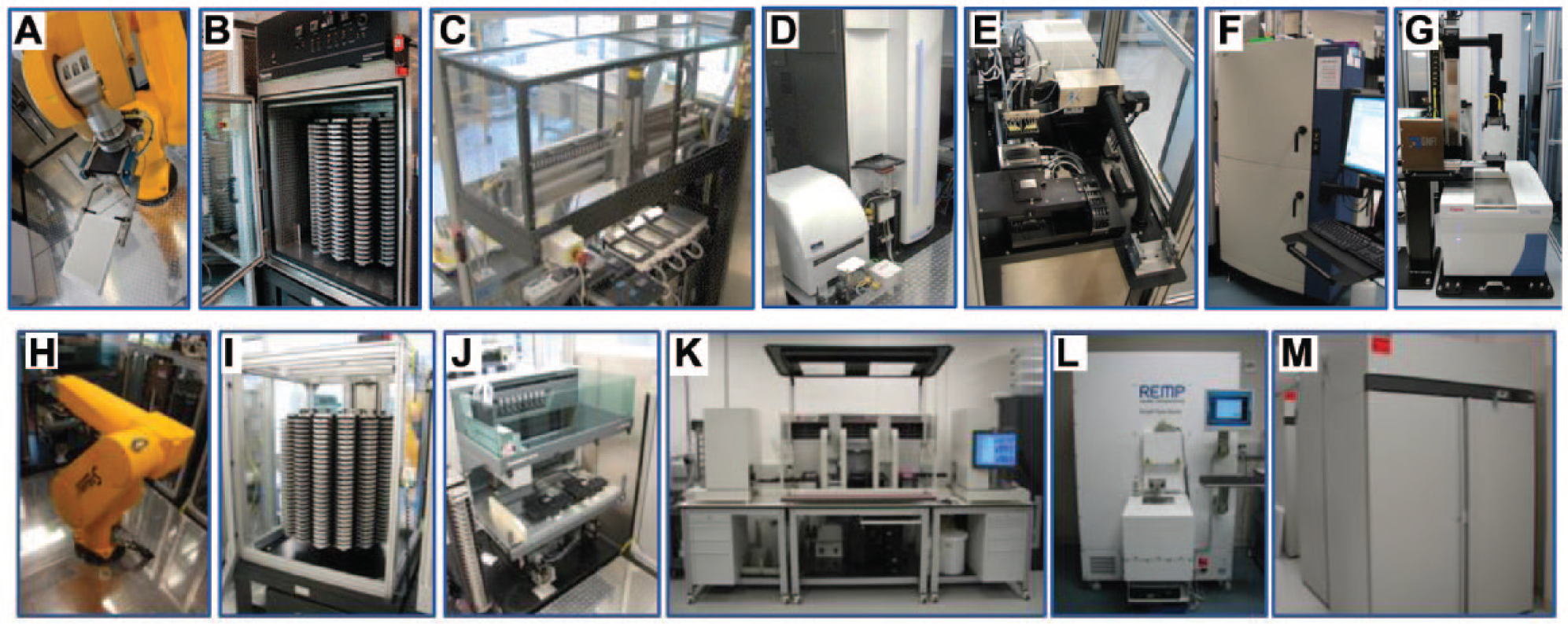

Lead identification HTS and compound management automation: (
Nearly all HTS-ready assay formats are supported, including protein-to-protein interaction (PPI) assays, biochemical-based assays, or cell-based assays. HTS campaigns are conducted as either target- or phenotypic-based readouts using all fluorescence modalities, absorbance, bio- or chemoluminescence, or high-content analysis.
HTS efforts typically begin with assay development and optimization. Successes in assay development/optimization often advance toward a small-scale pilot screen to validate HTS readiness and estimate hit rate. Pending the target type, these pilot screens may use small collections of around ~1000 compounds, such as LOPAC (1280 compounds; Sigma Life Science’s Library of Pharmacologically Active Compounds, St. Louis, MO), TOCRIS SCREEN (1070 compounds, Bristol, UK), or Prestwick (1200 compounds), or larger libraries, such as the Scripps FDA drug-approved library (~3300 compounds) or the Maybridge HitFinder (14,400 compounds, Loughborough, UK). Upon completion of a pilot screen, data are shared and discussed with the collaborating investigator to evaluate performance and make adjustments to the primary/secondary screening strategies to improve hit discovery and selection. Once an agreement is reached, the full-deck uHTS campaign is scheduled and quality assurance (QA)/QC testing of the robotic instrumentation is initiated to ensure the best results prior to any HTS campaign execution.
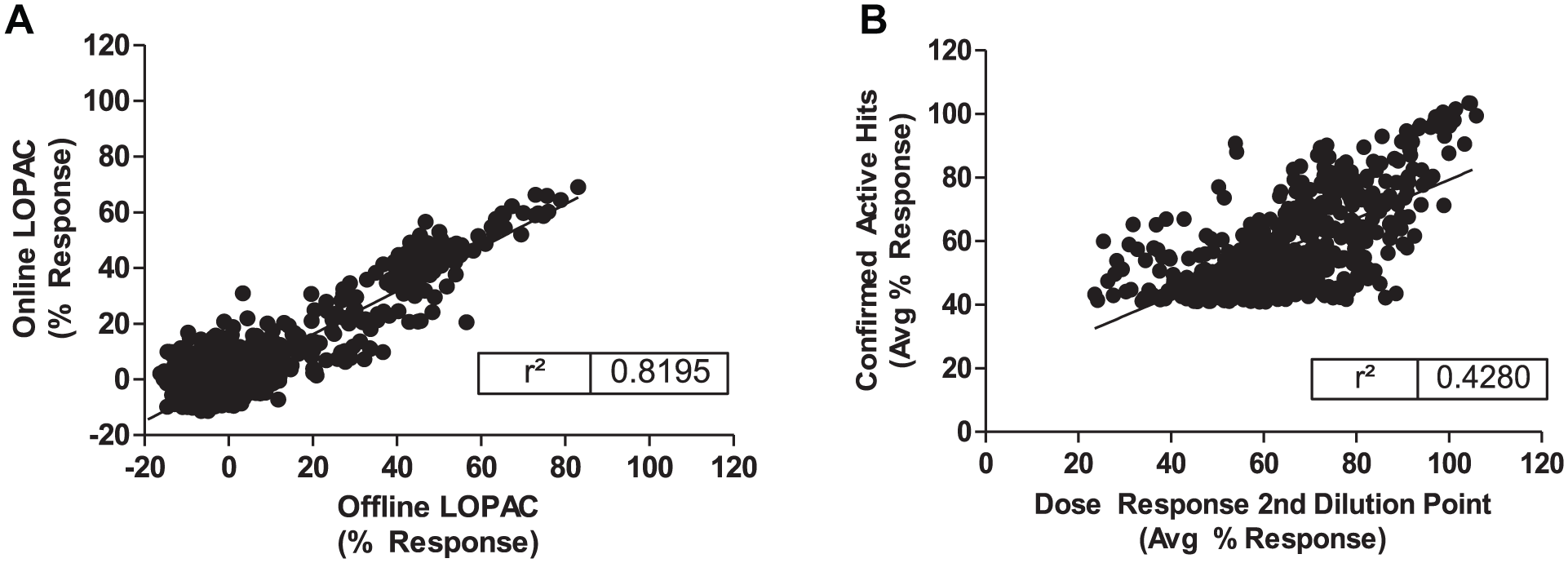
Once the assay implementation has been completed, a pilot screen is performed to validate the assay. The robotic assay validation is performed to confirm that the assay is performing well-based assay statistics and a good reproducibility of the activity of the compounds between offline and online assays is been observed (
Fig. 3
). Then the primary screen is performed in a singular (1×) testing mode against the SDDL collection of ~665,000 compounds, using good statistical separation for hit selection as determined by an acceptable Z′ factor of the controls. Typically, ~1% of the SDDL will be cherry-picked and advance to confirmation testing in triplicate (3×). In addition, secondary screenings will be used to rule out undesirable hits or false positives. Each HTS project will have a unique formulation of secondary screenings, which typically include:
(
Hits triaged from confirmation and secondary assays are forwarded to 10-point dose titration profiling to determine the IC50, hillslope, dose profile, and therapeutic window compared with a secondary assay. Confirmation, secondary, and dose titration assays are performed in triplicate, with averaged values reported along with standard deviation. Hits that demonstrate an acceptable potency and selectivity are then analyzed using in silico procedures to ascertain structural cluster profiles, pan-assay interference compounds (PAINS), rule of five compliance, and historical promiscuity score.5,6 A comparison of the activity of the compounds at the point of the secondary assay versus dose response data at the same concentration shows the selectivity of the most active compounds ( Fig. 3 ). Data are compiled and typically reviewed with contract investigators and medicinal chemists to select tractable hit scaffolds for SAR development and hit-to-lead optimization.
Compound Management at the SRMSC
The SRMSC Compound Management facility comprises software and hardware automation to enable rapid processing of samples in a QC-focused environment ( Fig. 2 ). Compound management automation includes Beckman Coulter Biomek FX and NXp platforms (Brea, CA) along with a Tecan Freedom EVO (Männedorf, Switzerland), which is integrated into a GNF hit-picking platform. All compound management automation is fully integrated with a corporate LIMS platform to eliminate the need for the manual curation of files.
In addition to support for automated sample handling in formats up to 1536-well microtiter plates, the SRMSC Compound Management facility utilizes a custom in-house-developed automated sample QC system known as the Plate Auditor. 7 Every compound plate that passes through the SRMSC Compound Management facility is processed through this instrument, which uses machine vision to automatically identify artifacts in compound plates, including empty wells, air bubbles, partially filled wells, precipitate, and variations in color between samples. Plate Auditor results are automatically cross-correlated against the SRMSC LIMS to determine if any errors exist in a plate and if sample remediation is necessary prior to releasing plates for downstream processes. If errors are detected, Compound Management team members are notified in real time. Consistent usage of this technology, when combined with in-house LC-MS capabilities, ensures that sample fidelity is tracked over time and that potential errors in compound management are captured and corrected in real time, prior to being released for downstream processes in HTS.
When not in use, all samples are stored in secure freezers. When sample access is required, labware containing samples of interest can be placed on the GNF hit-picking platform or in the REMP Small-Size Store (Chelmsford, MA), both of which provide fully automated retrieval of sample materials.
All sample records are maintained in a secure Oracle database with access restricted to necessary personnel. Full sample and plate genealogies are tracked from the entry of the sample into the lab through use in HTS campaigns, follow-up work, and delivery to external collaborators. This tracking includes full automated imaging of compound plates via the Plate Auditor, which provides a visual record of plate and sample status at each checkpoint throughout the life of a sample.
Informatics at the SRMSC
The SRMSC institutional informatics infrastructure is built around a commercial LIMS platform originally developed by Symyx/MDL (Santa Clara, CA). This infrastructure consists of three core components: Chembio for compound registration, Plate Manager for plate registration, and Assay Explorer for assay registration. These three core components are built on an Oracle database (Redwood City, CA) and are tightly integrated with one another, resulting in straightforward exchange of data without any need for intermediate manual curation or manipulation of data by end users. Compounds registered in Chembio are automatically associated with plates in Plate Manager when new plates are registered. Similarly, assays created in Assay Explorer automatically aggregate plate metadata with compound metadata and combine both with assay results to provide a single point of access for reviewing and performing QC of assay results.
Access to the institutional LIMS database and all associated software products not only is tightly restricted to relevant users, but also has specific roles assigned to individual users to ensure that access is project and function specific. Databases are also further segregated by funding source and isolated from each other by distinct login parameters, requiring specific user access to be granted for each instance.
In addition to the software platforms being fully integrated with one another, a Symyx/MDL software application programming interface (API) is available that allows the SRMSC to further extend the functionality of the existing software by developing in-house solutions that directly interact with the core Symyx/MDL platform ( Fig. 4 ). Over the past 13 years, the SRMSC has developed software to provide web-based interfaces for each of the three core components and to further extend and automate many routine compound management and HTS tasks. The web-based interfaces developed by the SRMSC have been linked to one another to enable approved users to seamlessly browse datasets across various stages of the drug discovery effort without having to load many different interfaces to access the information of interest.
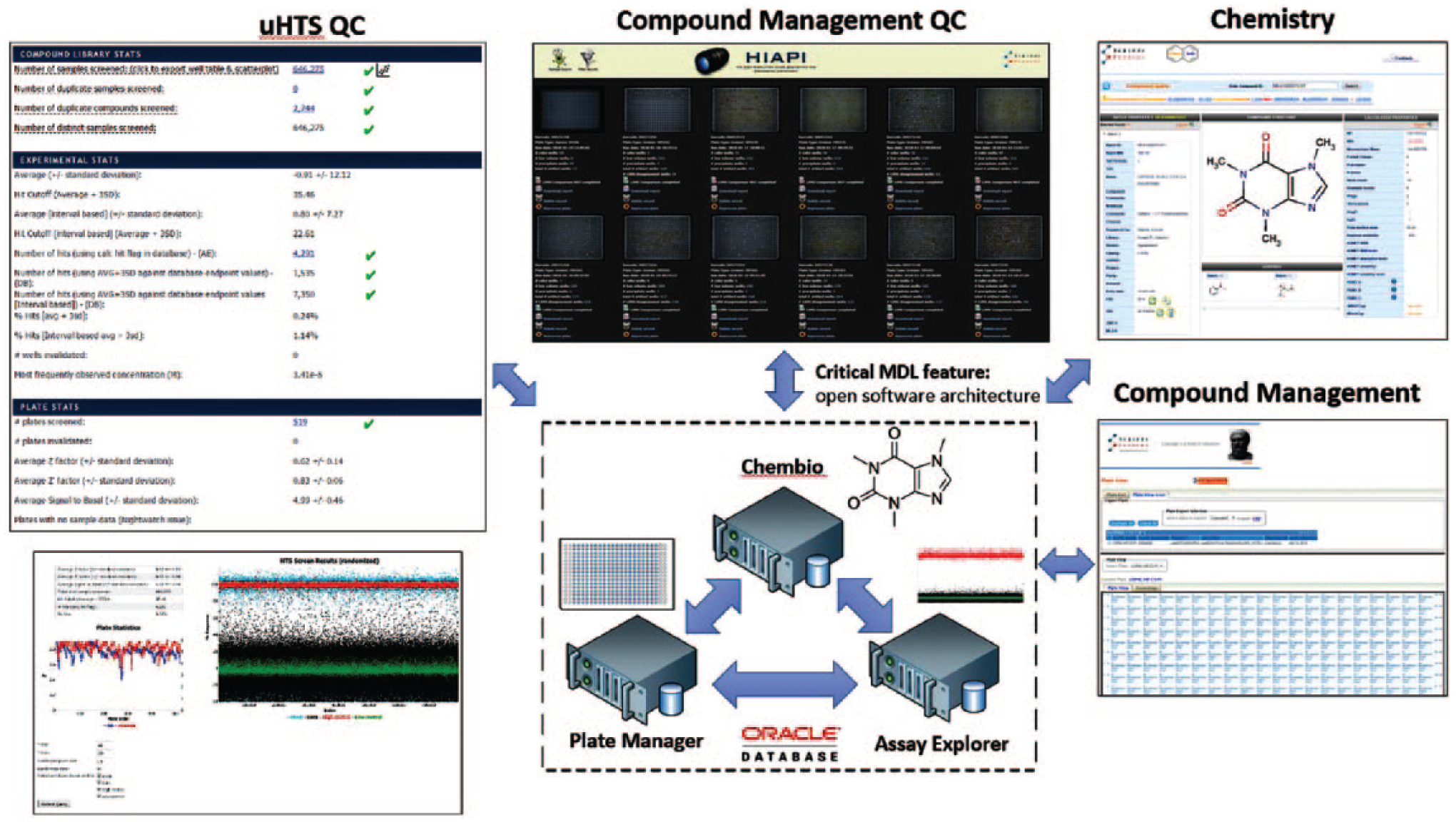
In-house software has been developed to extend the core Symyx/MDL LIMS platform functionality and provide user access via web interfaces.
Extended Software Automation
Dose Response Reporter aggregates dose–response curve results with compound metadata in an Excel format that is easily sharable with collaborators without requiring proprietary software to review. This report can be run by assay scientists via a web interface that allows for user selection of data to be brought into the report. Curves from multiple dose response assays can be plotted alongside data from primary screen, confirmation screen, and counterscreen assays to provide a report of overall compound activity over the course of a project. The SRMSC also maintains a list of approved drugs and associated metadata that is added to all reports as appropriate. The historical activity of compounds in the form of a promiscuity index is also included, providing insight into compound selectivity across assay type and target. All report generation is fully automated via the core Symyx/MDL databases and requires no manual manipulation of data by the user ( Fig. 5 ).
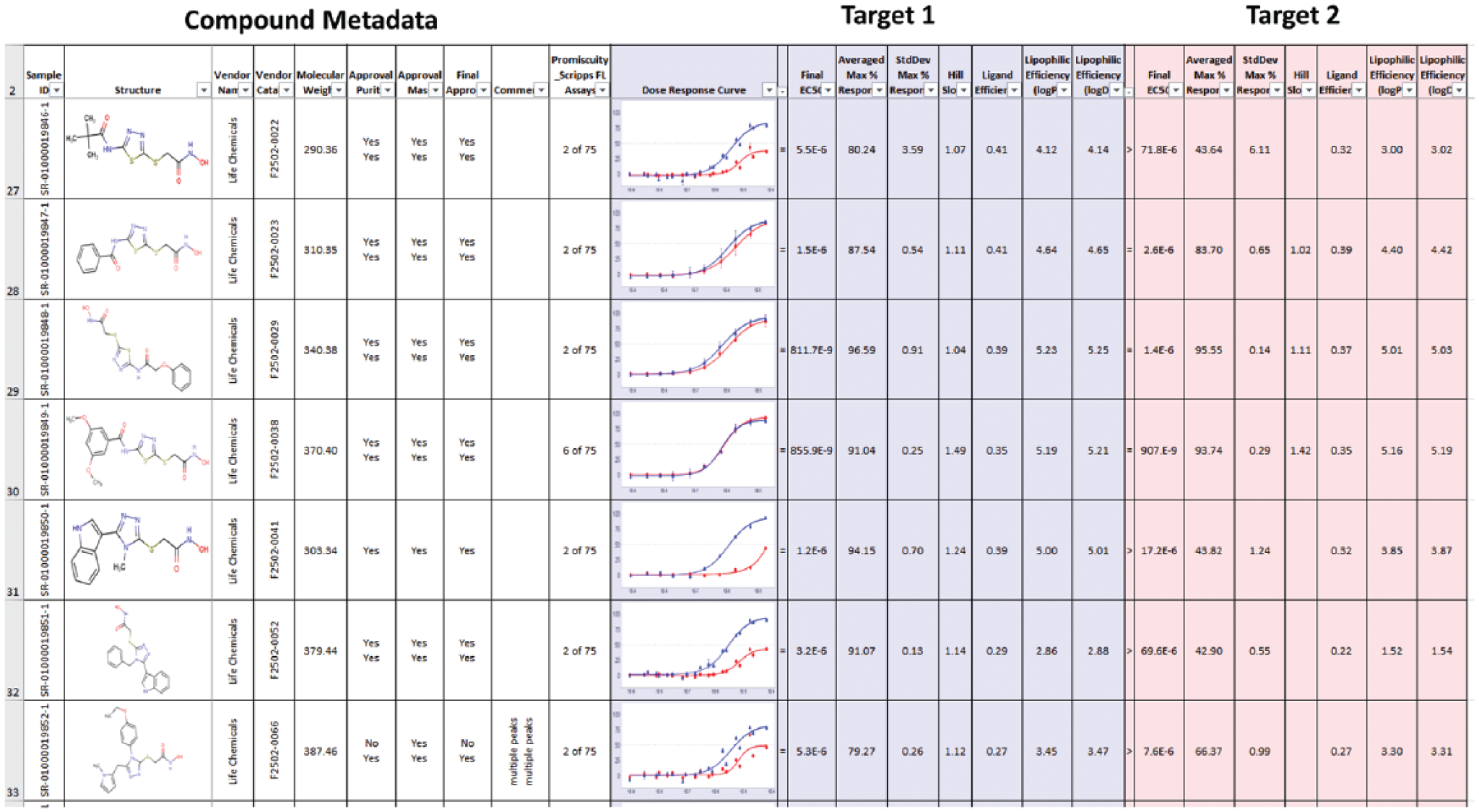
Example of a standard automated report generated without human intervention at the end of the HTS campaign, including compound metadata, dose–response curves with associated data, and LC-MS results.
Plato is a web-based interface to Plate Manager that allows users to register new plates, query for existing plates, and review plate data in a variety of formats. In addition to interfacing with Plate Manager, Plato is integrated with the GNF screening and hit-picking platforms, allowing robot inventories to be monitored in real time. Assay daughter plates from screening campaigns are automatically associated with parent compound plates, including transfer volumes, assay diluent volume, and full genealogical plate information. Additionally, Plato has been developed to support data uptake from various compound management automation, including Beckman Biomek FX and NXp units. Biomek liquid handling platforms are capable of reformatting millions of compounds with full barcode tracking and data export for registration in Plato. The SRMSC routinely processes large compound collections for HTS efforts with all data tracking and management performed in an automated fashion.
ChemInfo is a web-based interface to Chembio that allows users to review compound metadata. Assay results are also available for review, providing a full history of compound activity across all associated drug discovery activities, including HTS campaigns and DMPK studies. The ChemInfo interface also allows users to quickly browse related PubChem compound and substance summaries along with PubChem bioactivity analysis. The SRMSC has also utilized ChemInfo to provide compound metadata beyond what is available in Chembio, by including calculated properties such as PAINS descriptors, AlogP, polar surface area, H acceptors/donors, rotatable bonds, and solubility.
The HTS Dashboard automates a number of routine HTS QC and reporting tasks. From the HTS Dashboard interface, users can quickly validate all relevant HTS parameters prior to releasing data. The HTS Dashboard also automatically generates standardized visualizations for ease of reporting and QC. In addition to QC and reporting, the HTS Dashboard provides users with tools to explore compound promiscuity across multiple assays, to query for LC-MS results, and to perform compound enrichment searches when specific compound properties are desired for validation or small-scale screens.
SRMSC informatics capabilities are further extended past the Symyx/MDL platform and in-house tools with other commercial drug discovery informatics tools, including Spotfire (Somerville, MA), Pipeline Pilot (San Diego, CA), GraphPad Prism (San Diego, CA), and ChemAxon (Budapest, Hungary).
Innovation in HTS Biology and Laboratory Automation at the SRMSC
Efforts to advance screening technologies at the SRMSC include the exploration of three-dimensional (3D) cell cultures in 1536-well format, the introduction of sophisticated informatics support for high-content screens, neuron-based probe discovery platforms capable of HTS, and the development of custom software and hardware automation.8–12
Traditional high-throughput drug screening in oncology routinely has been performed on two-dimensional (2D) cell culture monolayers, which inadequately recapitulate the physiology context of cancer. 3D cell culture models have recently gained popularity thanks to their ability to better mimic the complexity of in vivo systems. 3D cell models have been described as more physiologically relevant for a variety of applications, especially with regard to cancer research and viability/cytotoxicity assessment in in vivo systems. Therefore, physiologically relevant 3D cell cultures have been recognized as a potential bridge between traditional in vitro 2D cultures and in vivo animal studies.13,14 It is thus critical to advance the development of scalable and affordable methods of producing 3D spheroids and/or organoids suitable for high-throughput cancer drug discovery. Creating 3D culture systems utilizing patient-derived tumors for rapid testing in high-density format would constitute an important achievement toward precision medicine and regenerative therapies. 3D cellular models have been developed in our screening center that are cost-effective and HTS amenable by combining the use of a cell-repellent surface with a bioprinting technology incorporating magnetic force ( Fig. 6 ). We have validated this homogenous process by evaluating the effects of well-characterized anticancer agents against four patient-derived pancreatic cancer KRAS mutant-associated primary cells, including cancer-associated fibroblasts. 8 In testing this scenario, we identified multiple compounds in the 3D assay format that had significantly different results from the traditional 2D monolayer assay format. In a direct comparison between the 2D and 3D assays, most of the tested drugs were less active in 3D, but a few drugs showed preferential cytotoxicity against 3D models over 2D culture, which proved to be both cell and drug dependent. One such example was disulfiram (DSF), which was more active in the pancreatic cancer 3D organoids than the 2D format models. As a follow-up to this research, we tested DSF and its metabolite diethyldithiocarbamate (DDC) and copper (Cu)-gluconate in PANC-1 cells in both 2D and 3D formats. While the 2D results demonstrate sensitivity to DSF or DDC, the 3D format remains completely insensitive unless Cu is added to the test at 5 µM. Thus, the presence of Cu significantly sensitizes either DSF or DDC in this 3D model, enhancing their cytotoxic effects, 15 possibly indicating a synergistic effect between the two ( Fig. 7 ). Notably, we found that DSF is currently undergoing clinical studies in metastatic pancreatic cancer, 16 prostate cancer, 17 and glioblastoma,18,19 which seems to confirm that we are using a more phenotypically relevant strategy. The future of our 3D methods could ultimately translate into the development of precision medication initiatives for oncology research.
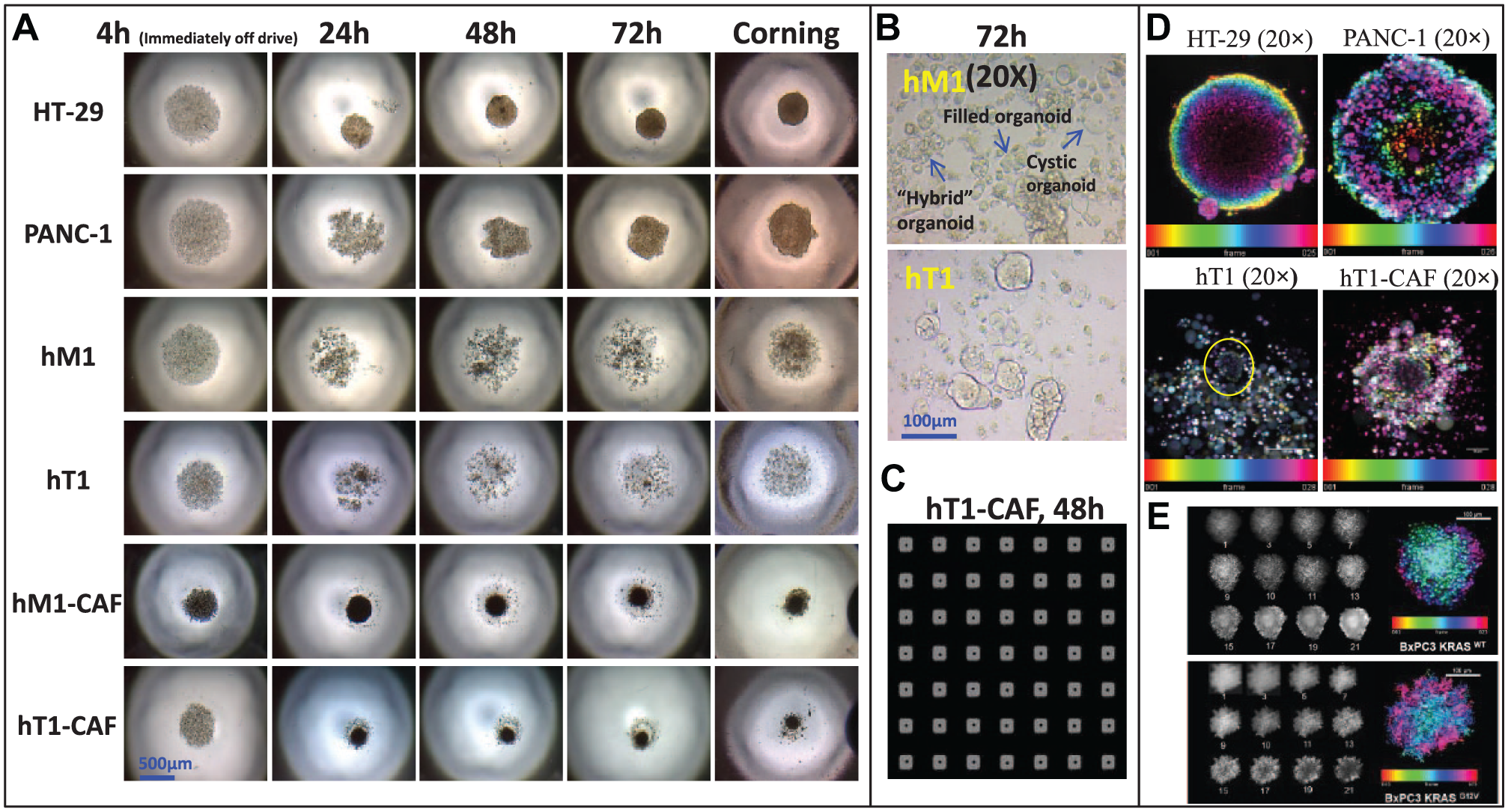
A panel of pancreatic cancer-derived cells was evaluated for their ability to form 3D structures using n3D bioprinting technology. (
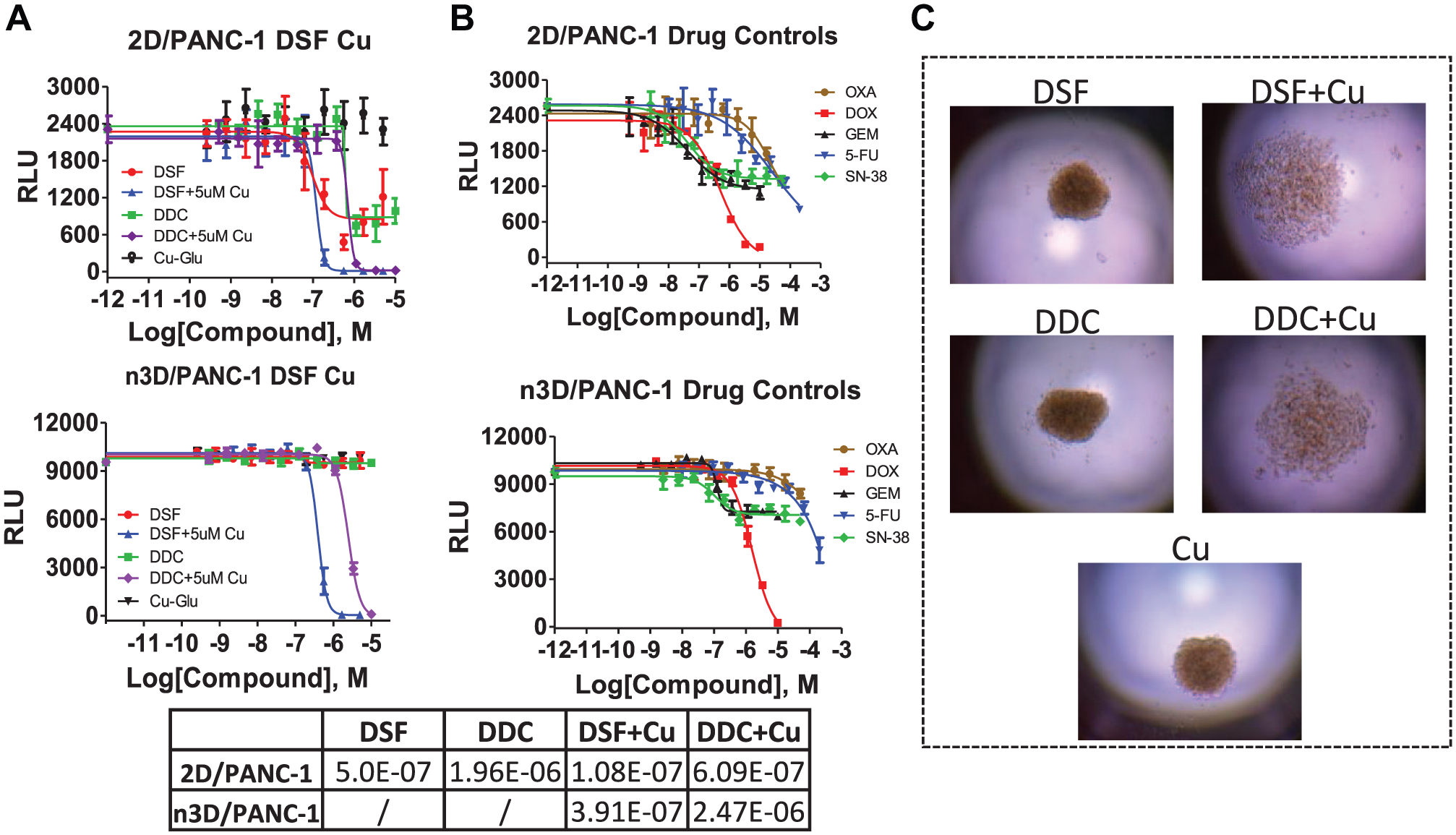
(
3D cell culture developments also included the execution of a novel 3D HTS campaign to find inducers of a mutant KRAS 20 oncogenic gene. The 3D spheroid approach facilitated the identification of a selective inhibitor of cells, proscillaridin A. The SRMSC has collaborations with industrial partners such as n3D Biosciences and Greiner Bio-One for the development of an HTS-compatible magnetic cell culture platform 8 and Corning Inc. for the development of cell-repellent surface microcavity microtiter plates for spheroid culture. 21
Additionally, high-content screening (HCS) has gained significant attention as another development in complex screening technologies. HTS provides a single readout of activity for individual assays, whereas HCS can provide data and measurements for multiple features of individual cells in assays all at once. More recently, research with HCS has even been enhanced with the introduction of 3D culture cell lines as opposed to working with the traditional 2D monolayer cell cultures. HCS has the potential to both identify lead compounds and predict compound toxicity that has specific prominence in drug discovery. 10 Predicting toxicity and efficiency is extremely powerful and part of what makes HCS so complex. In collaboration with Michael Conn from the University of Texas (Austin, TX), we performed a drug screening in a cell-based high-content assay in the 1536-well plate format studying pharmacoperones associated with oxalosis kidney disease, which is a rare metabolic disorder.
Another collaboration using cell-based HCS was performed with Scripps Professor Gavin Rumbaugh. There is an unmet need to develop cell-based assays that involve the use of induced neuron (iN)-based probe discovery platform assays using primary human cells to study neuropsychiatric disorders. 11 These methodological improvements simplify the creation of highly scalable neuron-based phenotypic assays designed to improve drug discovery in central nervous system (CNS) disorders.
Thus, a new direction is being investigated by exploiting the use of HCS and primary human cells to create induced human neurons for use in this research ( Fig. 8 ). The lack of robust phenotypic HCS assays using iNs is still a challenge due to the multistage and long differentiation protocols. We have been developing a simple method for scaling the production of human induced pluripotent stem cell (iPSC)-derived iNs. As an example, iPSCs were used as human-derived neurons in functional screening assays to measure mitochondrial activity and formation. 22 Using both HTS and HCS, these iNs could be screened for toxicity and against multiple compounds for cell activity in a miniaturized, cost- and time-efficient throughput method. As an example of such an application, we have successfully used iNs in 384-well plate format to screen the LOPAC pilot library as demonstrated in Figure 9 . The end goal is to apply the iPSC-derived lines from patients with genetic diseases to evaluate neurotoxicity in early drug discovery.
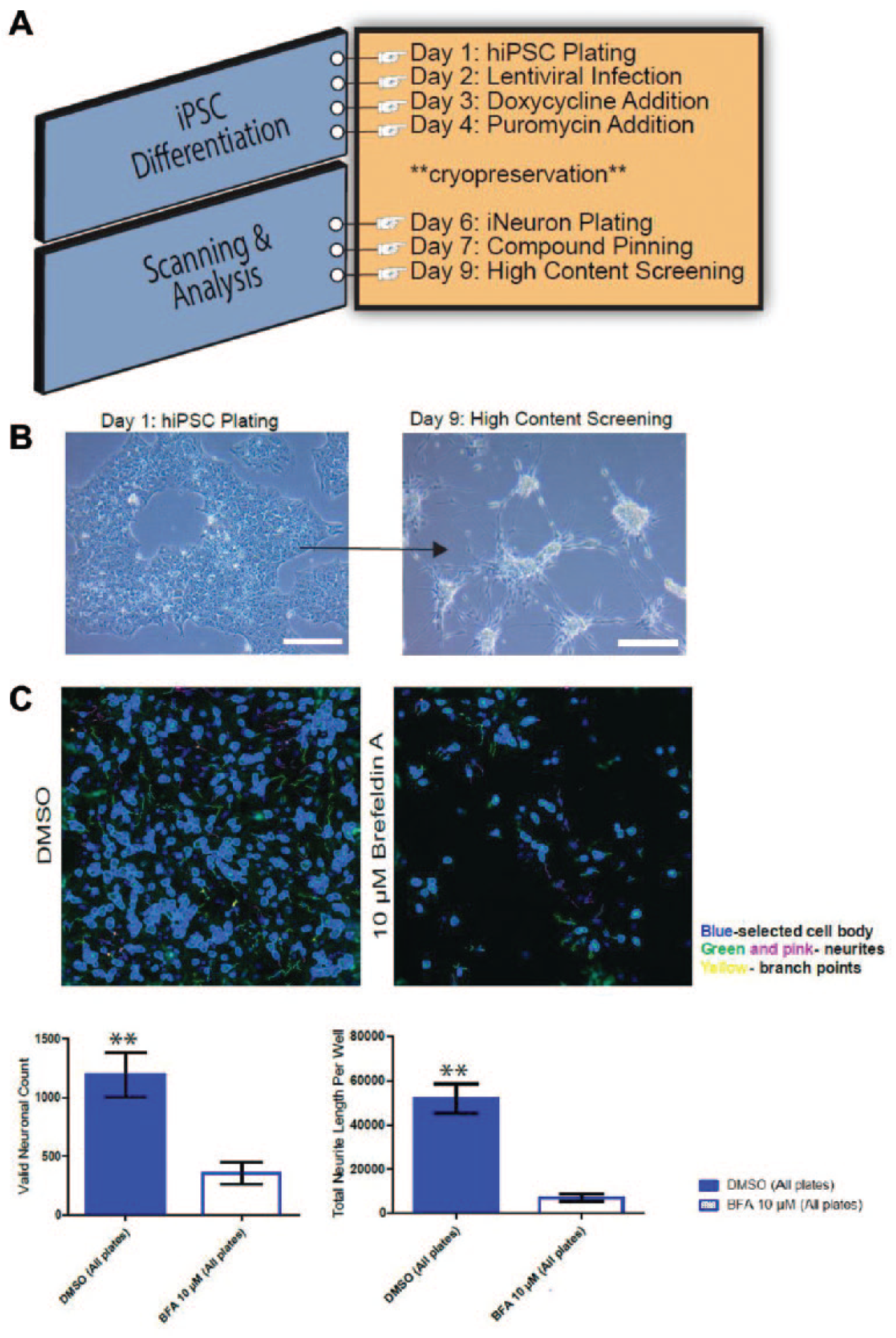
(
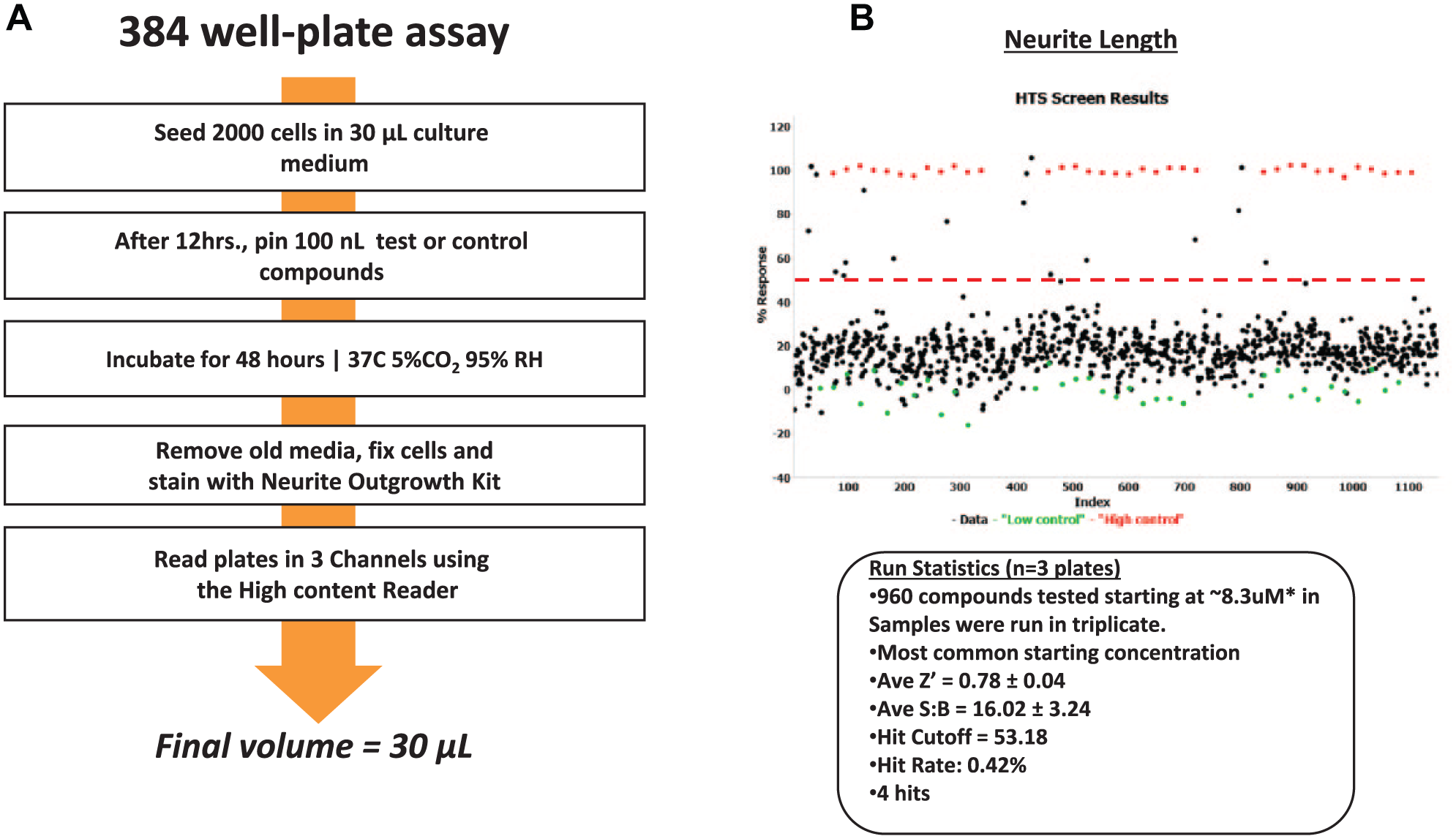
(
Finally, the SRMSC also features a modern engineering lab space for repair and maintenance of existing laboratory instrumentation and development of novel automation. The engineering lab space houses a Tormach 770 CNC mill (Waunakee, WI), an Ultimaker 2+ 3D printer (Geldermalsen, Netherlands), and an electronics workbench. These fabrication capabilities allow SRMSC engineers to provide rapid support for novel HTS workflows and to develop custom tools that improve overall lab efficiency. In addition to the previously described Plate Auditor, this also includes the development of a real-time HTS liquid dispenser QC monitoring system, 12 the creation of custom robotic incubator shelving for 3D spheroid HTS workflows, electronic pipetting LED light guides, and a wide variety of one-off automation modifications and repairs.
Footnotes
Acknowledgements
The authors would like to acknowledge Dr. Peter Hodder, Mark Southern, Dr. Franck Madoux, Peter Chase, Dr. Shurong Hou, Dr. Dmitriy Minond, and Dr. Sanjay Saldanha for their contributions and help in the initiation of the screening center and for their efforts during the MLPCN.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The SRMSC and TRI were supported by the Molecular Library Probe Production Centers Network (MLPCN) grant proposal number #5U54 MH084512 and now currently in part by the National Cancer Institute of the National Institutes of Health under award number R33CA206949. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
