Abstract
Macrodomains recognize intracellular adenosine diphosphate (ADP)-ribosylation resulting in either removal of the modification or a protein interaction event. Research into compounds that modulate macrodomain functions could make important contributions. We investigated the interactions of all seven individual macrodomains of the human poly(ADP-ribose) polymerase (PARP) family members PARP9, PARP14, and PARP15 with five mono-ADP-ribosylated (automodified) ADP-ribosyltransferase domains using an AlphaScreen assay. Several mono-ADP-ribosylation-dependent interactions were identified, and they were found to be in the micromolar affinity range using surface plasmon resonance (SPR). We then focused on the interaction between PARP14 macrodomain-2 and the mono-ADP-ribosylated PARP10 catalytic domain, and probed a ~1500-compound diverse library for inhibitors of this interaction using AlphaScreen. Initial hit compounds were verified by concentration–response experiments using AlphaScreen and SPR, and they were tested against PARP14 macrodomain-2 and -3. Two initial hit compounds and one chemical analog each were further characterized using SPR and microscale thermophoresis. In conclusion, our results reveal novel macrodomain interactions and establish protocols for identification of inhibitors of such interactions.
Introduction
ADP-ribosylation is a posttranslational modification that regulates a multitude of vital processes.1,2 Intracellular mono- and poly-ADP-ribosylation (MARylation and PARylation) are catalyzed by two different subclasses of the poly(ADP-ribose) polymerase (PARP) family, enzymes also known as ADP-ribosyltransferases with homology to diphtheria toxin (ARTDs).3–5 Protein MARylation has emerged as an important signaling event in the nucleus and cytosol.6–9 One way to mediate and modulate ADP-ribosylation signaling is by means of a universal structural module called a macrodomain (Pfam ID PF01661). 10 Lately, specific macrodomains have been reported as readers and regulators of MARylation,2,11–13 whereas others have been linked to the recognition of PARylation,14,15 breakdown of PAR chains,16,17 or turnover of ADP-ribose metabolites.18–20
Among the 11 human macrodomain-containing proteins are PARP9/ARTD9, PARP14/ARTD8, and PARP15/ARTD7, all with documented MARylation activity ( Fig. 1A ).20,21 The macrodomains of these proteins do not possess ADP-ribosyl glycohydrolase activity; 10 they bind free ADP-ribose,12,14 and PARP14 macrodomains bind several proteins that are subject to MARylation by PARP10 as well as auto-MARylated PARP10 itself. 12 The spectrum of ADP-ribosylated target proteins for these macrodomains, and also the consequences of the binding events, are yet poorly understood. It is clear, however, that these proteins are involved in a multitude of cancer-related processes and that they are potentially interesting from a drug discovery perspective.10,22,23
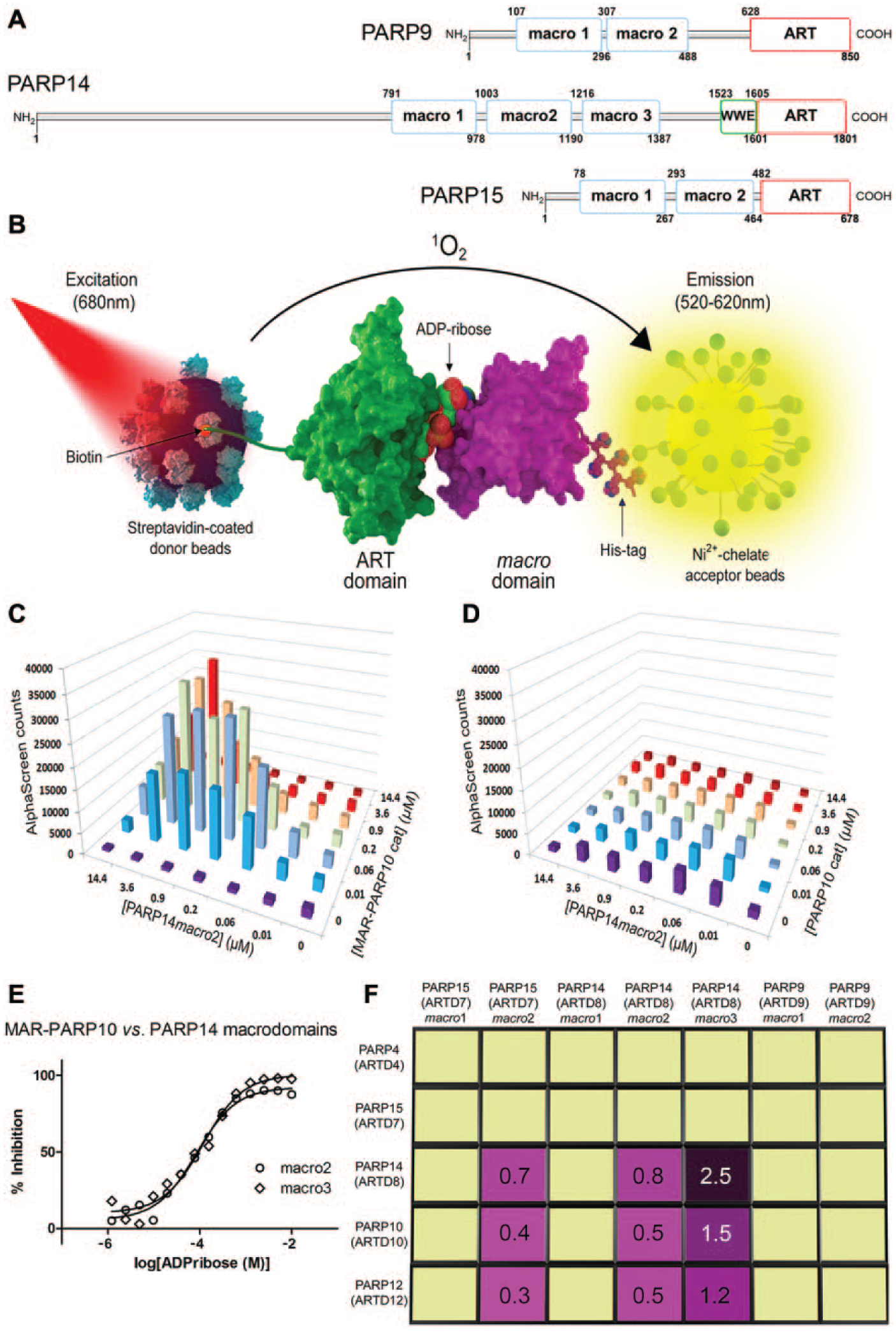
Probing the interactions between macrodomains and automodified poly(ADP-ribose) polymerase (PARP) catalytic domains. (A) Domain organization of the human macrodomain containing PARP family members. Macro, macrodomain; WWE, WWE domain; ART, adenosine diphosphate (ADP)-ribosyltransferase domain. (B) Schematic illustration of the bead-based proximity assay for macrodomain interactions with mono-ADP-ribosylated (MARylated) target proteins. PARP catalytic (ART) domains were site-specifically biotinylated in vivo to allow immobilization on streptavidin donor beads. Hexahistidine-tagged macrodomains were captured on nickel chelate acceptor beads. The interaction between macrodomain and PARP catalytic domain generated an AlphaScreen signal. (C) Cross-titration of PARP14 macrodomain-2 and auto-MARylated PARP10 ART domain to illustrate establishment of optimal assay conditions. (D) Cross-titration of PARP14 macrodomain-2 and unmodified PARP10 catalytic domain to document the absence of interaction in the absence of MARylation. (E) Competition with free ADP-ribose of the interaction between PARP14 macrodomains-2 and -3 and auto-MARylated PARP10 ART domain to further illustrate the MARylation dependency of the interaction. Data were normalized based on the signal observed in the absence of free ADP-ribose and presented as % inhibition. (F) Schematic illustration of the results of the interaction screening matrix. Positive interactions shown to be MARylation dependent (magenta) were further analyzed by SPR to determine dissociation constants (in µM; determined from the kinetics of the binding and unbinding events;
A homogeneous high-throughput assay of macrodomain–target interactions would be a valuable method for identification of novel targets, for the study of macrodomain physiological functions, and for discovery of small-molecule ligands that break these interactions. Therefore, we set out to develop an assay that could report the stable interaction between a macrodomain and its ADP-ribosylated target protein. The resulting AlphaScreen method enabled us to map the interactions between all PARP family macrodomains and a set of auto-MARylated PARP catalytic domains. Subsequently, we used this assay to screen for compounds that could block the interaction between PARP14 macrodomain-2 and MARylated PARP10. We discovered some of the first macrodomain ligands and used complementary binding methods to determine their affinities in the mid-micromolar range.
Materials and Methods
Expression and Purification of Macrodomain Protein Constructs
DNA fragments encoding the individual macrodomains of human PARP9, -14, and -15 (PARP9G102-K298, PARP9E300-M489, PARP14S794-D984, PARP14A994-S1196, PARP14F1208-G1388, PARP15S53-K251, and PARP15G265-N448) were inserted into pNIC-Bsa4 (Genbank EF198106) containing an N-terminal hexahistidine tag with a tobacco etch virus (TEV)-protease cleavage site (MHHHHHHSSGVDLGTENLYFQ*SM, where the asterisk indicates the site of cleavage). All proteins were produced in
Expression and Purification of In Vivo Biotinylated PARP Catalytic Domain Constructs and GRB14
The catalytic domain fragments PARP4E252-E581, PARP10A809-G1017, PARP12D489-S684, PARP14D1611-K1801, and PARP15N459-A656 and the SH2 domain of human growth factor receptor-bound protein-14 (GRB14) were subcloned into either pNIC-Bsa4, as above (for purposes other than AlphaScreen), or pNIC-Bio3 (GenBank JN792439.1) containing an N-terminal hexahistidine tag as above, as well as a C-terminal biotin acceptor sequence (GLNDIFEAQKIEWHE). Proteins were produced as above. For AlphaScreen, all proteins were produced and in vivo biotinylated by co-expression with BirA ligase in
AlphaScreen Protein–Protein Interaction Assay
In vivo biotinylated PARP catalytic domains were diluted to 2 mg/ml in 50 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) pH 7.5, 100 mM NaCl, 4 mM MgCl2, 0.05% Tween20, and 0.2 mM tris(2-carboxyethyl)phosphine (TCEP), and automodification was induced by incubation with 200 µM β-NAD+ at 30 °C for 30 min. Assay beads were prepared using the Histidine Detection Kit (PerkinElmer, cat. 6760619M), with streptavidin donor beads and nickel chelate acceptor beads. The MARylated, biotinylated catalytic domains were immobilized on streptavidin donor beads (20 µg/ml). Macrodomains were captured onto nickel chelate acceptor beads (20 µg/ml). The beads were mixed and incubated for 1 h in light gray AlphaPlate-384 (PerkinElmer, cat. 6005350) prior to detection in an EnSpire plate reader (PerkinElmer), according to the manufacturer’s protocol ( Fig. 1B ). Optimal assay concentrations were identified for each interacting pair of PARP macro- and catalytic domains by cross-titration experiments (each partner at 0–15 µM; e.g., Fig. 1C and 1D ). Positive interactions were confirmed by competition experiments in which ADP-ribose was used to block the interaction between macrodomains and catalytic domains (e.g., Fig. 1E ). Prism (GraphPad Software) was used to evaluate titration experiments by four-parameter regression analysis and curve fitting with no further constraints.
Characterization of Protein–Protein Interactions by Surface Plasmon Resonance
PARP10, -12, and -14 catalytic domains (without biotin tag) were diluted to 2 mg/ml in 50 mM HEPES pH 7.5, 100 mM NaCl, 4 mM MgCl2, and 0.2 mM TCEP, and automodification was induced by incubation with 200 µM β-NAD+ at 30 °C for 30 min. A Biacore 3000 instrument (GE Healthcare) was used to characterize the biospecific interactions identified by AlphaScreen assays. A pH scouting was performed to find optimal coupling conditions for each protein. MARylated PARP14 was diluted to 20 µg/ml in 100 mM HEPES pH 6.6, MARylated PARP10 was diluted to 20 µg/ml in 10 mM NaAc pH 5.0, and MARylated PARP12 was diluted to 20 µg/ml in 10 mM NaAc pH 4.5; the proteins were then covalently coupled to Sensor Chip CM5 (GE Healthcare, cat. BR100012) using the Amine coupling kit (GE Healthcare, cat. BR100050) with N-hydroxysuccinimide (NHS) and 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide hydrochloride (EDC). Macrodomains were diluted in running buffer [50 mM HEPES pH 7.4, 150 mM NaCl, 3.4 mM ethylenediaminetetraacetic acid (EDTA), and 0.005% Surfactant P20] to concentrations ranging from 0 to 30 µM. Experiments were performed with a buffer flow rate of 10 µl/min, an association phase of 3 min followed by a dissociation phase of 3 min, and regeneration with 1 M NaCl. The dissociation constants were calculated based on kinetic data using a 1:1 Langmuir binding model and the BIAeval software (GE Healthcare).
Screening Compounds and Analogs
A 1584-compound diversity set was provided by the Chemical Biology Consortium Sweden (CBCS). Primary hit compound
AlphaScreen Compound Screening Assay
Compounds (200 µM) were screened for breaking the interaction between MARylated PARP10 on donor beads (150 nM) and PARP14 macrodomain-2 on acceptor beads (3 µM). The final concentration of DMSO was 2%, and control experiments showed that the protein–protein interaction was unaffected by DMSO at concentrations up to 5%. Wells containing 2% DMSO were used as negative controls (69,325±10,725 counts among five plates), and wells lacking PARP10 as positive controls (197±146 counts). The overall Z’-factor was 0.47, and the coefficient of variation (CV) was 15.5%. A selection cutoff of 3 SD was applied. 257 compounds were evaluated twice in a counterscreen experiment containing AlphaScreen beads crosslinked using in vivo biotinylated and hexahistidine-tagged Grb14 protein, but neither PARP10 nor a macrodomain. 203 compounds either were found to quench the AlphaScreen signal in this test or could not be retrieved from the library. 53 primary hit compounds were tested in concentration–response experiments: Selected compounds were serially diluted from a 10-mM stock solution in DMSO. The highest compound concentration was 200 µM, and the final DMSO concentration was 2%. Compounds were evaluated in triplicate concentration–response experiments with both PARP14 macrodomain-2 and macrodomain-3.
Compound Characterization by Surface Plasmon Resonance
SPR (Biacore T200; GE Healthcare) was used to analyze the interactions between the screening hits and their analogs and PARP macrodomains. Different macrodomain immobilization strategies were evaluated: Direct covalent coupling to Series S Sensor Chip CM5 (GE Healthcare, cat. BR-1005-30) was compared to a hexahistidine capture strategy with an anti-His antibody (His capture kit; GE Healthcare, cat. 28-9950-56) and to oriented immobilization to Series S Sensor Chip NTA (GE Healthcare, cat. 28-9949-51). Reproducible ligand binding was observed when PARP14 macrodomain-2 was diluted to 10 µg/ml in 10 mM NaOAc pH 5.5 and when PARP14 macrodomain-3 was diluted to 10 µg/ml in 10 mM NaOAc pH 5.0 and directly coupled to the CM5 chip using the Amine Coupling Kit (GE Healthcare, cat. BR100050) with NHS and EDC. 13000RU PARP14 macrodomain-2 and 15400RU PARP14 macrodomain-3 were immobilized. HBS-EP (100 mM HEPES, 150 mM NaCl, 3 mM EDTA, 0.05% v/v surfactant P20, and pH 7.4) was used as running buffer. Compounds were serially diluted in running buffer from 20 mM stock solutions in DMSO. Concentrations ranged from 200 to 0 µM, with a final DMSO concentration of 2%. Binding experiments were performed in HBS-EP with 2% DMSO. The flow rate was 30 µl/min, injection time was 120 s, and each cycle contained a 30-s regeneration phase. All binding experiments were run twice using freshly prepared sensor chips. Dissociation constants were calculated using the BIAeval software (GE Healthcare) and a 1:1 Langmuir binding model.
Microscale Thermophoresis
Experiments were performed on a Monolith NT.115 instrument. PARP14 macrodomains-2 and -3 were labeled using the Monolith NT Protein Labeling Kit Blue–NHS (NanoTemper Technologies, cat. L0003), according to the manufacturer’s protocol. Compounds were serially diluted in 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 10 mM MgCl2, 0.05% Tween, and 5% DMSO, and mixed 1:1 with proteins. Despite extensive buffer scouting, inclusion of 5% DMSO was necessary to obtain reliable data. Measurements were carried out in Premium Coated Capillaries (NanoTemper Technologies, cat. MO-K005). Binding data were collected with 40% microscale thermophoresis (MST) power and 20% light-emitting diode (LED) power. Data analysis was performed in the MO.Affinity Analysis software (NanoTemper Technologies, version 2.1.2).
Results and Discussion
Assay Development and Interaction Mapping
The automodified (i.e., MARylated) PARP10 catalytic ADP-ribosyltransferase (ART) domain is a binding partner for PARP14 macrodomain-2. 12 In the present study, we set out to study this interaction using AlphaScreen technology ( Fig. 1B ). First, optimum protein concentrations were determined by cross-titration of the two proteins ( Fig. 1C ). The corresponding experiment using nonmodified PARP10 confirmed that the interaction with the macrodomain was strictly MARylation dependent ( Fig. 1D ). Next, we verified that addition of free ADP-ribose, which is also a ligand for macrodomains, 12 inhibited the interactions in a competitive, concentration-dependent manner ( Fig. 1E ). This confirmed once more that the macrodomain specifically recognized ADP-ribosylated PARP10 and that the interaction detected by AlphaScreen was the physiologically relevant one. The dissociation constant for the interaction was determined by SPR to be 0.5 µM.
Using the bead-based proximity assay procedure, we then tested all other PARP macrodomains (
Fig. 1A
) for their interactions with five automodified PARP catalytic domains. Pairs of auto-MARylated PARP catalytic domains and macrodomains for which a signal was detected were considered further only when cross-titration led to identification of a hook effect (as illustrated in
Fig. 1C
). Those pairs were then cross-titrated as before, but using the nonmodified PARP catalytic domains (as illustrated in
Fig. 1D
). Protein–protein interactions were considered further only when this control experiment verified that the interaction was dependent on automodification of the catalytic domain. The nine resulting pairs of auto-MARylated PARP catalytic domains and macrodomains were then subjected to competition experiments with titration series of free ADP-ribose, shown in
The results of this analysis comprise seven novel interactions (
Fig. 1F
): PARP14 macrodomains-2 and -3, in addition to their confirmed interactions with PARP10, also recognized MARylated PARP12 and -14, but did not interact with any of the other MARylated ART domains in the study. PARP15 macrodomain-2 also interacted with the MARylated catalytic domains of PARP10, -12, and -14. Quantification of the binding affinities using SPR (
Fig. 1F
) and MST (not shown) indicated Kd values in the high nanomolar to low micromolar range for all interactions. No interactions were observed between any MARylated ART domain and PARP14 macrodomain-1, PARP15 macrodomain-1, or either of the two macrodomains of PARP9. Automodified PARP4 and PARP15 ART domains were not recognized by any macrodomain in the study. We detected only weak interactions between some macrodomains and the PARP16 full-length protein expressed in
Our results show that PARP macrodomains are capable of discriminating between MARylated target proteins. Thus, these macrodomains apparently require ADP-ribose in a structure-dependent context to mediate specific protein–protein interaction. This would suggest that macrodomains also are capable of discriminating between identical targets MARylated at different sites. This concept will have to be tested, for example by using site-specific mutants of target proteins once MARylation sites have been identified.
Our results also illustrate the general utility of AlphaScreen in the identification of novel macrodomain–target interactions. The assay format is homogeneous and particularly well suited for studying low-affinity protein–protein interactions. 26 Thus, our method development is a step toward a better understanding of the specific roles of individual macrodomains in epigenetic signaling and disease etiology.
Discovery of PARP14 Macrodomain Inhibitors
Having established this assay for PARP–macrodomain interactions, we were interested in its application in screening for inhibitors of target recognition events. We acquired a screening set of diverse compounds and carried out screening as described in Materials and Methods, using the interaction between PARP14 macrodomain-2 and the MARylated PARP10 ART domain. Primary hit compounds were identified and subjected to a counterscreening experiment to remove unspecific signal quenchers, followed by concentration–response experiments. The resulting 17 compounds were evaluated using SPR. Only compounds
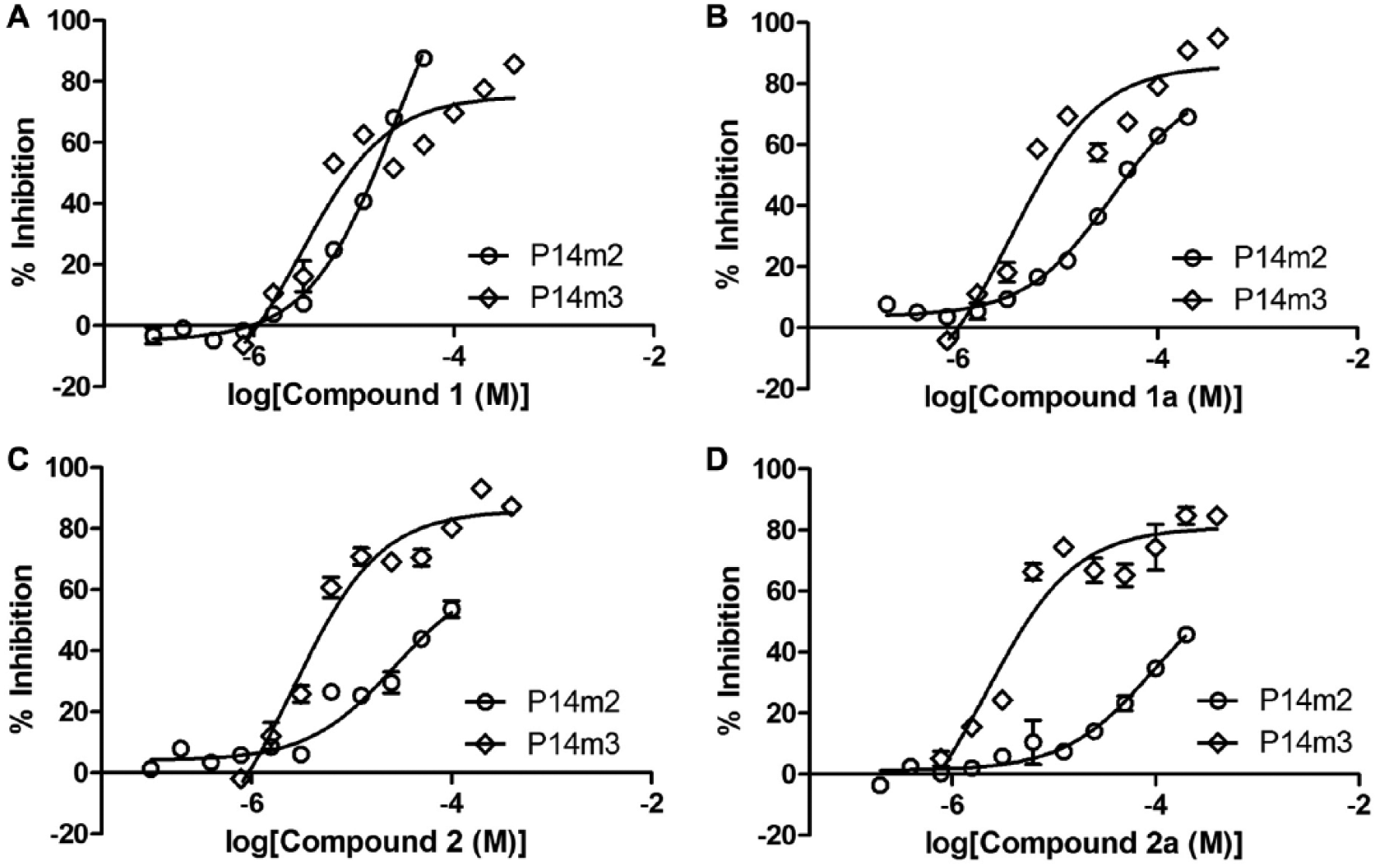
Discovery of inhibitors of PARP14 macrodomains-2 and -3. AlphaScreen concentration–response experiments for the validation of primary screening hits
Characterization of ligand binding to PARP14 macrodomains-2 and -3.
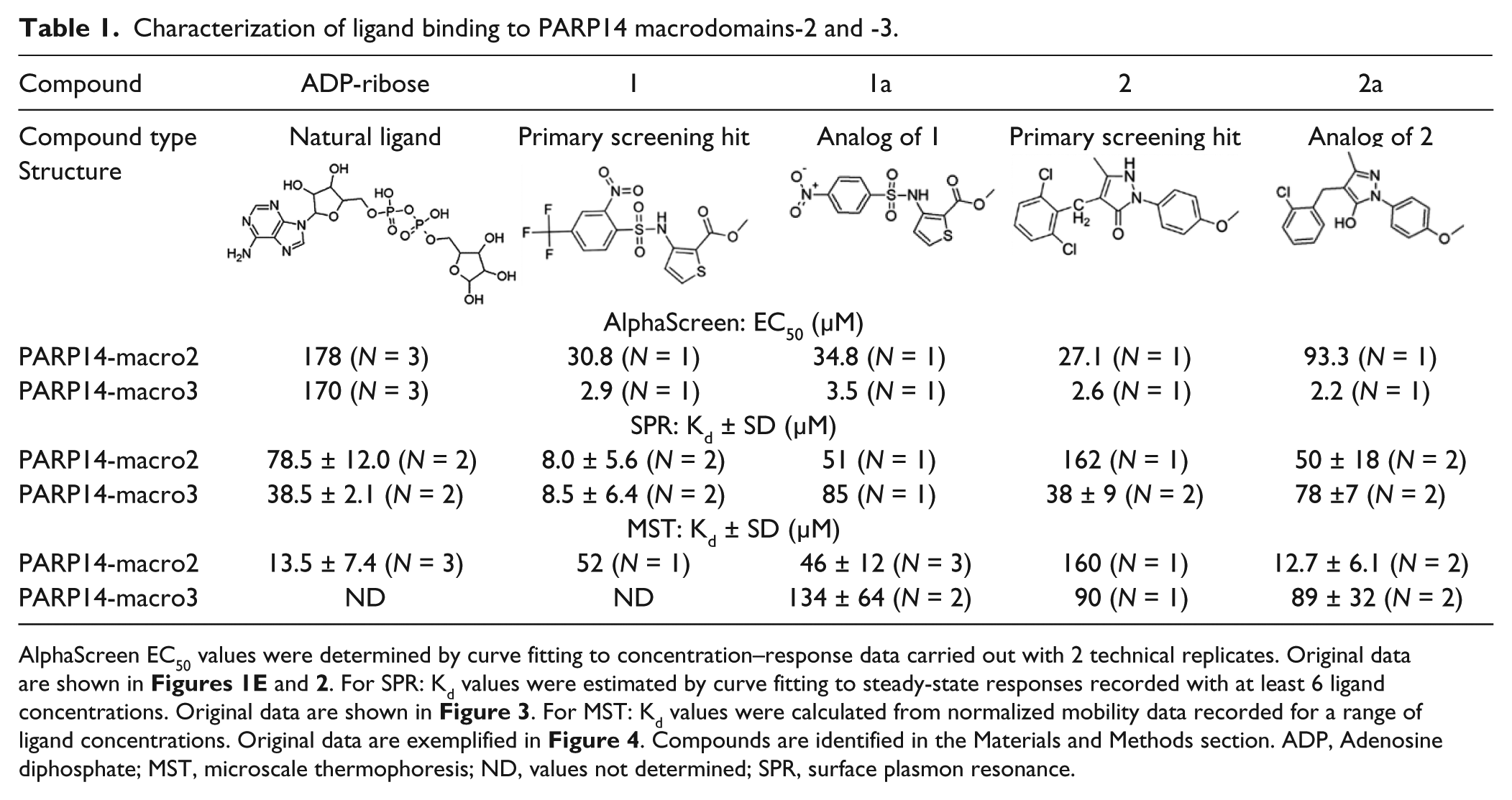
AlphaScreen EC50 values were determined by curve fitting to concentration–response data carried out with 2 technical replicates. Original data are shown in Figures 1E and 2 . For SPR: Kd values were estimated by curve fitting to steady-state responses recorded with at least 6 ligand concentrations. Original data are shown in Figure 3 . For MST: Kd values were calculated from normalized mobility data recorded for a range of ligand concentrations. Original data are exemplified in Figure 4 . Compounds are identified in the Materials and Methods section. ADP, Adenosine diphosphate; MST, microscale thermophoresis; ND, values not determined; SPR, surface plasmon resonance.
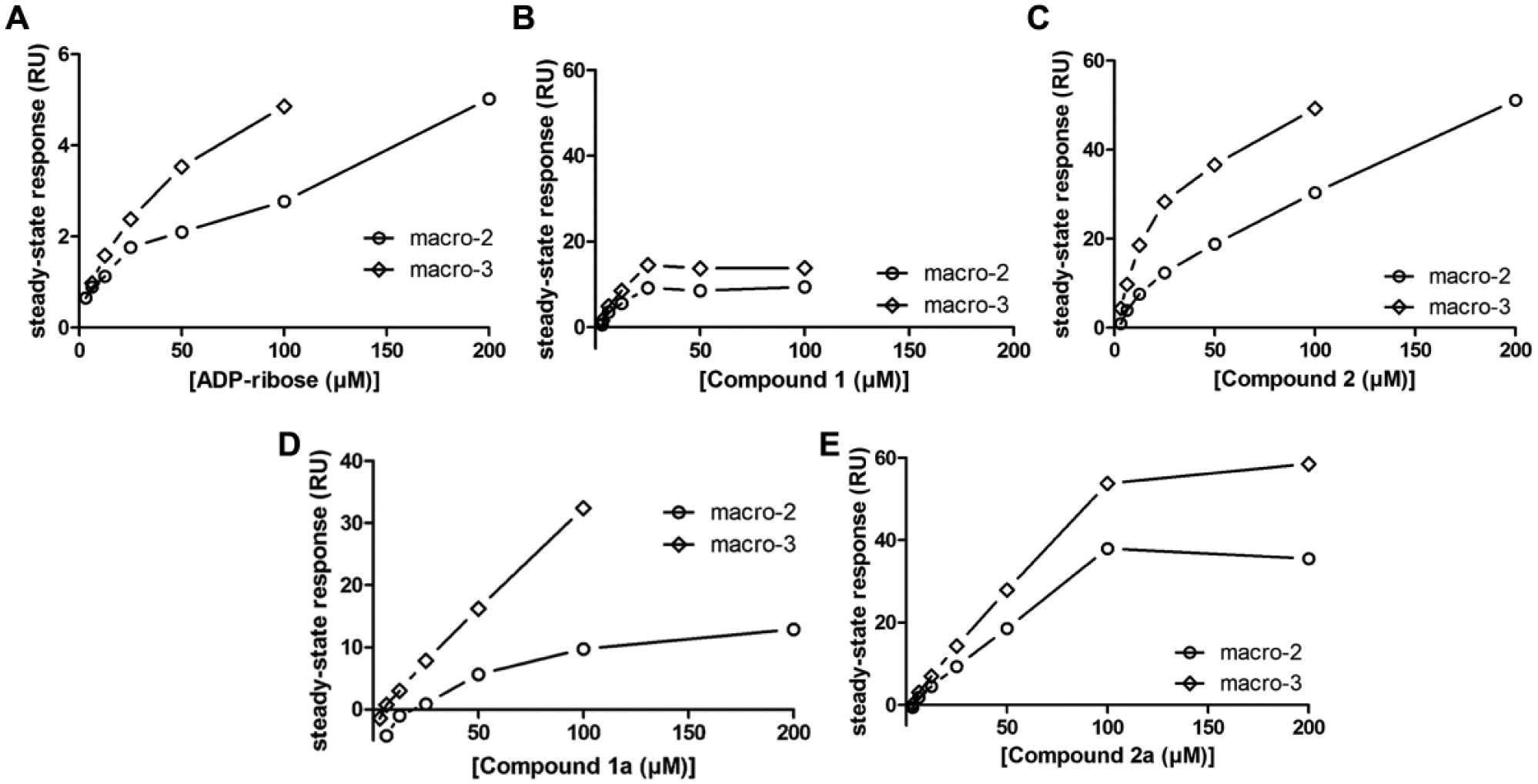
Characterization of primary screening hits and their analogs using surface plasmon resonance (SPR). Biacore steady-state response levels in dependence of ligand concentration for (A) adenosine diphosphate (ADP)-ribose and (B–E) the four macrodomain inhibitors. PARP14 macrodomains-2 and -3 were examined. Apparent dissociation constants are reported in Table 1 .
For the following analyses,
In parallel, chemical substructure searches were performed, and a small set of chemical analogs of compounds
For an independent confirmation of the binding affinities of the four novel ligands for the two PARP14 macrodomains, MST was used. In essence, the Kd values calculated from these experiments confirm the results of the SPR analysis (
Table 1
and
Fig. 4
). Together, the data highlight
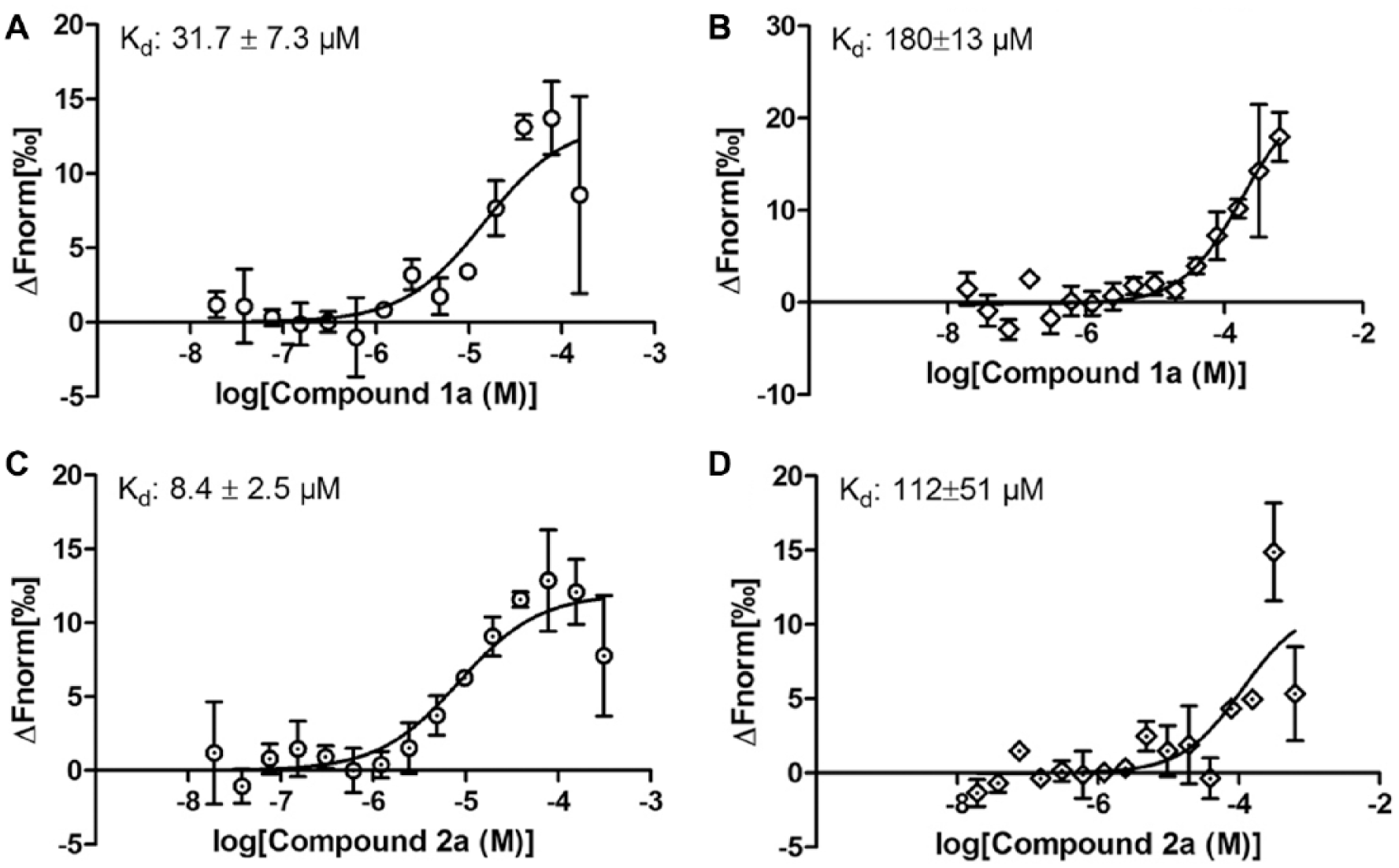
Characterization of screening hit analogs
Although the PARP macrodomains can bind free ADP-ribose with intermediate affinity (
Table 1
),
12
our results show that in binding of ADP-ribosylated target, the protein context can strengthen the interaction by at least one order of magnitude. We were interested in exploring this further by using the newly discovered macrodomain inhibitors in the context of the domain interaction matrix shown in
Figure 1F
. For this, concentration–response experiments were carried out with the four macrodomain inhibitors, probing eight different combinations of MARylated ART domains (PARP10, -12, and 14) and macrodomains (PARP14 macrodomains-2 and -3 and PARP15 macrodomain-2). The resulting AlphaScreen EC50 values (
Fig. 5
) represent a qualitative comparison of the inhibitors’ ability to disrupt the respective interactions. The results provide further evidence for a structure context-specific recognition of MARylation by the PARP macrodomains. For instance,
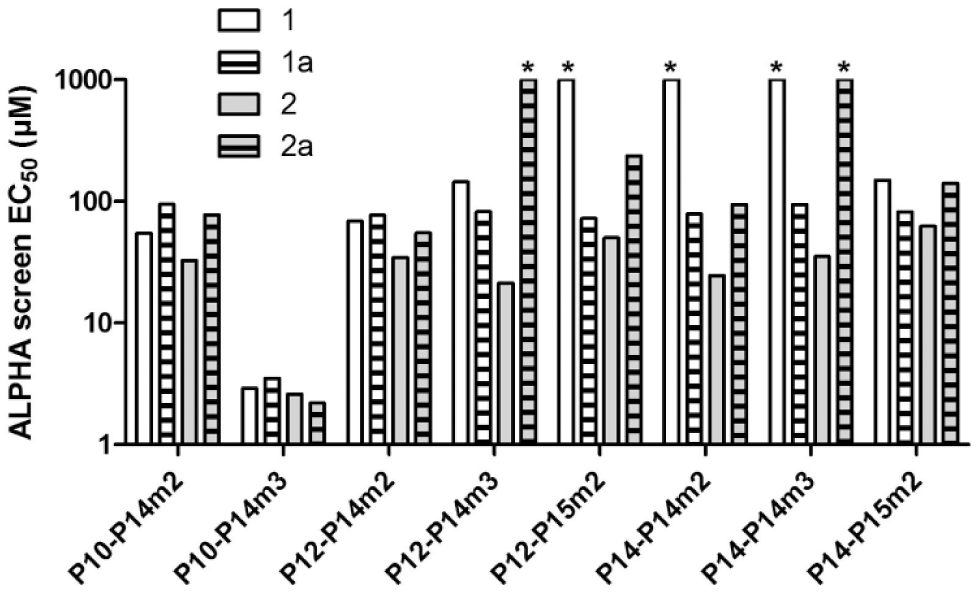
Relative potential of four macrodomain inhibitors to displace eight different macrodomain–target interactions. Concentration titrations were conducted with compounds
To summarize, we have developed protocols for efficient screening of macrodomain–target interactions and of compounds that prevent these interactions. Contrary to recently published similar methods,27,28 our protocols interrogate interactions between physiological binding partners. We have used these protocols to discover some of the first inhibitors of PARP macrodomains. Another inhibitor of PARP14 macrodomain-2 was recently identified.
27
Together, these compounds provide an entrance into development of chemical probes directed at these macrodomains, and they can provide positive controls for in vitro and
Footnotes
Acknowledgements
We thank the Protein Science Facility at Karolinska Institutet/SciLifeLab for molecular cloning; Anna-Lena Gustavsson, Åsa Slevin, and Thomas Lundbäck [Chemical Biology Consortium Sweden (CBCS)] for providing the diverse screening set of compounds; C. David Andersson (Umeå University) for suggesting chemical analogs of primary screening hit compounds; Åsa Frostell (GE Healthcare) for advice on Biacore T200 experimentation and data analysis; Luca Jovine (Karolinska Institutet) for granting access to the Monolith instrument; and Teresia Hallström (NanoTemper Technologies) for extensive advice on MST experimentation and data analysis.
Supplementary material is available online with this article.
Author Contributions
TE purified proteins, established all assays, contributed to macrodomain target mapping, performed all remaining analyses, and contributed to writing the article. PV performed molecular cloning, purified proteins, and contributed to macrodomain target mapping. AEL synthesized compound
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financed by the IngaBritt och Arne Lundbergs Research Foundation (no. 403), the Swedish Cancer Society (2012/313 and 2014/716), the Swedish Foundation for Strategic Research (RBc08-14), and the Swedish Research Council (2012-5247 and 2015-4603).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
