Abstract
Inhibition of NK and effector T-cell functions and activation of regulatory cell populations are the main immunosuppressive effects of indoleamine-2,3-dioxygenase1 (IDO1). By converting tryptophan (Trp) into kynurenine (Kyn), IDO1 is involved in the immune response homeostasis, and its dysregulated expression is described in immune-related pathologies, as tumors that hijack it to evade immune destruction. Thereby, IDO1 inhibitors are being developed to stimulate antitumor immune responses. Existing and standard quantitation methods of IDO1 substrate and metabolite(s) are based on the total level of Trp and its metabolites determined by liquid chromatography tandem mass spectrometry analysis in human plasma, cerebrospinal fluid, and brain. Here, we describe the detection, localization, and absolute quantitation of Trp and Kyn by quantitative mass spectrometry imaging (qMSI) in transfected murine tumor models expressing various levels of IDO1. Myeloid, glycolysis metabolic signatures, and correlation between IDO1 expression and Trp to Kyn conversion are also shown. High-definition IDO1 and GCN2 immunostainings overlaid with Kyn molecular images underline the tumor metabolism and heterogeneity. The development of immunotherapies such as IDO1 inhibitors requires a deep understanding of the immune system, the interplay of cancer cells, and biomarker characterization. Our data underline that qMSI allows the study of the spatial distribution and quantitation of endogenous immune metabolites for biology and pharmacology studies.
Introduction
The immune system consists of a large number of different types of cells and proteins that distinguish between normal and abnormal cellular components and between “self” and “nonself.” Its main role is to protect the body against illnesses and infections. 1 Immune cells play a key role in controlling tumor growth, while cancer cells try to develop strategies to evade the immune surveillance. 2 To boost the immune system to strengthen its response and to effectively fight the tumor, immunotherapy has been studied for decades and was first used for patients with advanced cancer. Since then, it has been translated to the adjuvant setting of patients with high-risk operable disease and for postoperative recurrence. Different immune strategies have been developed to improve or restore immune system functions, including antibodies that either stimulate immune cells or block immune inhibitory pathways, cancer vaccines, and adoptive transfer of ex vivo activated T or natural killer cells. 3 Immunotherapy has now been clinically validated as an effective approach for cancer therapy. Clinical trials with antibodies blocking cytotoxic T lymphocyte–associated antigen 4 and programmed cell death protein–1 or programmed cell death ligand–1 have led the way to a second generation of immune checkpoint inhibitors. 4
Despite all these promising existing and currently developed therapeutic modalities, and because of the immune system’s complexity and the diversity of strategies used by tumors to escape host immunity, challenges to successful immunotherapy remain, with the need for further understanding of the pathways and biomarkers involved in the tumor immunology. 5 Many findings have already indicated that indoleamine-2,3-dioxygenase 1 (IDO1) may be a reliable and promising prognostic indicator for the treatment of human cancers.6–9 IDO1 is a natural endogenous mechanism of immune suppression acting through modulation of the Trp degradation pathway. IDO1 catabolizes Trp into Kyn, which could be further metabolized into Kyn pathway metabolites. 10 The fact that IDO1 has a role in enabling tumors to escape the host immune system, it is defined as an attractive new target for cancer immunotherapy. Therefore, IDO1 inhibitors are currently being developed in immuno-oncology. 11
Trp and Kyn are key biomarkers of IDO1 functions, and the balance between these 2 metabolites drives immune cell homeostasis. 12 The existing and standard quantitation methods are based on the total level of Trp and its metabolites determined by liquid chromatography tandem mass spectrometry (LC-MS/MS) analysis in plasma, cerebrospinal fluid, and brain. However, this methodology does not take into account important parameters such as spatial distribution and tumor heterogeneity.
In this study, we describe the setup, development, and application of a new quantitative method based on mass spectrometry imaging (MSI) to detect, localize, and quantify Trp and Kyn in biological tissues. Transfected murine tumor models expressing different levels of IDO1 were used as proof-of-concept models for this study. Basically, MSI is able to simultaneously record the distribution of hundreds of biomolecules directly from tissue, without labeling and without prior knowledge. 13 MSI based on matrix-assisted laser desorption/ionization (MALDI) applied with different tissue preparation procedures can be used to analyze proteins, peptides, glycans, lipids, metabolites, and drugs.14–18 Often, the large number of molecules detected within a single experiment often includes isobaric species that have the same nominal mass but different exact mass. It has been demonstrated how high-mass-resolution instruments, such as Fourier transform ion cyclotron resonance (FTICR) and Orbitrap mass spectrometers, are able to resolve many of the isobaric species, whereas the more commonly used time-of-flight systems amalgamate the signals.
Here, we report a quantitative ultra-high-mass-resolution study, performed using a 7T MALDI-FTICR that allowed us to highlight key metabolites of the Trp pathway that are responsible for the immunosuppressive tumor microenvironment.
Materials and Methods
Chemicals and Reagents
All chemicals including 1,5-diaminonaphtalene (1,5-DAN), tryptophan, kynurenine, tryptophan-d5 and kynurenine-d4, and formic acid were purchased from Sigma-Aldrich (St. Louis, MO). Methanol, acetonitrile, and Optima LC-MS water were purchased from Fisher Scientific (Waltham, MA). Indium tin oxide (ITO)–coated glass slides were purchased from Bruker Daltonics (Bremen, Germany).
Primary rabbit anti-mouse IgG IDO (106134) and GCN2 (137543), secondary goat anti-rabbit IgG H&L (HRP) secondary antibody (205718), and the detection system through horseradish peroxidase followed by steady DAB/plus (brown chromogen) were purchased from Abcam (Cambridge, UK).
Sample Collection and Tissue Preparation
IDO1-positive P815 murine mastocytoma clones were generated as described in Uyttenhove et al. 19 Two P815-IDO1 clones were selected according to the level of IDO1 expression (one IDO1-low and one IDO1-high P815 clone). 19 P815-IDO1 cells were subcutaneously grafted in DBA/2 6- to 8-week-old mice (Charles River Laboratories, France). After 10 to 15 days of tumor growth, mice were sacrificed, and tumors were sampled and snap frozen in liquid nitrogen for 15 s. The samples were kept at −80 °C until use. Ten micrometer thick tissue sections were obtained using a cryostat microtome (CM-3050S, Leica, Germany) with a microtome chamber and a specimen holder chilled at −23 °C. They were thaw mounted onto ITO-coated slides for downstream MALDI imaging and serial sections on SuperFrost slides for histological and immunohistochemistry (IHC) analysis. Biological triplicates in duplicate (distant serial sections) were performed for analytical reproducibility. For LC-MS/MS analysis, five tissue sections of 10 µm thickness were harvested to perform calibration curves and quantitation of both Trp and Kyn.
All animal experiments were compliant with the 2010/63/UE European Directive on Laboratory Animal Welfare and were approved by an Ethical Committee.
LC-MS/MS Analysis
Five tissue sections of 10 µm thickness liver were collected to perform the calibration curve. Then, two calibration curves were used in a range of 0 to 500 nM that contain 5 nM of internal standard each (Trp-d5 and Kyn-d4). Afterward, between 0.5 and 2 mg of serial tumor sections was collected in an acidified aqueous extraction solution (H2O 0.1% formic acid = mobile phase A) that contains 5 nM on internal standards. An overnight extraction at 4 °C, centrifugation at 3000 × g, for 4 °C/15 min allowed us to recover 500 µL of supernatant from both calibration curves of Trp/Kyn and all the IDO1-low and high P815 sections that were used for the LC-MS/MS analysis. A total of 5 µL was injected into the LC-MS/MS system. No additional filtration step was necessary. The LC-MS/MS system consisted of an ultra-high-performance liquid chromatography–focused+ LC system composed of RS column compartment, RS pump, and RS autosampler (Dionex Ultimate 3000, Thermo Fisher Scientific) that contains a Thermo Scientific column (Hypersil Gold C18 50 × 21, particle size = 1.9 µ) coupled to a TSQ Quantiva Thermo Scientific (Thermo Fisher Scientific). Data acquisition and processing were carried out using TSQ Quantiva software version 2.0 1292 and Xcalibur 3.0 software.
Quantitative MALDI Analyses
Quantitative MALDI analyses were performed using our internal newly developed software Multimaging 1.2 software (ImaBiotech SAS, France). This multimodal imaging platform combines quantitative mass spectrometry imaging (qMSI) and microscopy platform with statistical analysis for the understanding of the Omics information at cellular levels. Regarding the quantitative part, internal standards optimal concentrations were first determined using deuterated compounds (Trp-d5 and Kyn-d4). Using isotopic modified compounds should allow getting the same suppression effect if present within the tissue regarding the endogenous molecules of interest. These compounds were sprayed over tissues of interest (IDO1-low and -high P815 tumors) to calculate the ratio between internal molecule and deuterated molecule intensities that should not exceed 10 (5 µM of Trp-d5 and 10 µM of Kyn-d4 were chosen for following experiments). Afterward, calibration curves were measured following two steps (large and restricted ranges). Following these, low limit of quantitation, upper limit of quantitation, and limit of linearity parameters were defined for each calibration curve that has been performed using at least 10 spiked concentrations on 10 µm thickness liver tissue sections. The concentration range was 125 to 0.5 pmol for both Trp and Kyn, which has been validated as a linear regression model. On the same big ITO slide, calibration curves and tissue sections of interest (IDO1-high and -low P815 in triplicates and duplicates) were deposited and matrix containing internal standards sprayed over. Then, using Multimaging software, normalized Trp and Kyn intensities over calibration curves and tissues of interest were performed. Finally, using the calculated equations, different quantities in µg/mm2, then converted to µg/g of tissue, were obtained in selected regions of interest (ROIs; triplicates of IDO1-high and -low P815 tumors analyzed in duplicates).
MALDI-FTICR Imaging
For MALDI MSI of metabolites, a uniform layer of filtrated 1,5-diaminonaphtalene (1,5-DAN) prepared at 10 mg/mL with 50/50 ACN/H2O matrix was deposited onto the tumor tissue sections using an HTX-TM sprayer device (HTX Technologies, LLC, Carrboro, NC) with a nozzle heated to 60 °C and high of 40 mm from the sample slide. Ten passes at 0.1 mL/min flow rate and 1200 mm/min velocity with 3 mm track spacing and a pressure of 10 psi and 2 L/min of gas flow rate. Then, MALDI MSI analysis was performed using 7T MALDI-FTICR (SolariX XR, Bruker Daltonics, Bremen, Germany) with a SmartBeam II laser. MSI data were recorded in negative ion mode (m/z range 100–1000) and at 120 µm and then 30 µm of spatial resolutions with an online calibration.
Data acquisition was performed using the Flex software suite (FtmsControl 2.1.0 and FlexImaging 4.1x64) from Bruker Daltonics and data processing and visualization using Multimaging 1.2 software (ImaBiotech SAS, France). This multimodal imaging platform combines qMSI and microscopy platform with statistical analysis for understanding the Omics information at cellular levels.
MSI data were acquired from each tissue section as well as matrix control areas adjacent to the tissue sections to check for analyte dispersion/delocalization during sample preparation.
IHC Analyses, Digital Scan Image, and High-Definition Overlay
Sections were stained for IDO1 and GCN2 using rabbit polyclonal antibodies purchased from Abcam (Cambridge, UK) and adapted to fresh frozen tissue sections. Sections were first exposed to 0.5% Triton-X for 15 min at room temperature and washed with phosphate-buffered saline prior the addition of the primary anti-IDO1 and anti-GCN2 antibody (1:50 and 1:100 dilutions, respectively) and processed with goat anti-rabbit IgG H&L (HRP) secondary antibody (1:2000). The detection system was through horseradish peroxidase followed by steady DAB/plus (brown chromogen).
After MSI data acquisition, any residual matrix was removed with a 70% ethanol wash, and the tissue samples were then stained with hematoxylin and eosin (H&E) solution.
High-resolution histological images of H&E stain or IHC were then recorded using a digital slide scanner (3D Histech Pannoramic) then loaded in Multimaging technology to perform the high definition overlays with convoluted molecular images.
Results
MALDI MSI of Tryptophan Pathway: An Insight in the Tumor Metabolism
As expected, we detected decreased Trp in IDO1-high compared with IDO1-low P815 tumors. This low Trp level in the IDO1-high tumors was consistent with the detected high Kyn expression ( Fig. 1A ). Average intensities of Trp and Kyn were extracted from all ROIs and ratios (Kyn/Trp) calculated and raised in both models. Mouse models with high IDO1 expression showed higher Kyn/Trp ratios ( Fig. 1B ). Figure 2 summarizes the Trp pathway metabolites that we detected using 7T-MALDI-FTICR MSI in IDO1-P815 tumor tissue sections (biological triplicates) and in which the ultra-high-resolution and mass accuracy enabled an elemental formula to be assigned and isotope patterns to be matched (MS spectra in Suppl. Fig. 1 SF1 and full MS list in Suppl. Table ST1 ). As shown before, low Trp level in the IDO1-high tumors was consistent with high Kyn expression, which in turn was associated with an overexpression of anthranilic acid, 3-hydroxy-kynurenine, quinolinic acid, nicotinamide, and 5-hydroxyindole-3-acetic acid. H&E stain showed the intratumor heterogeneity that is consistent with differential metabolic activity.
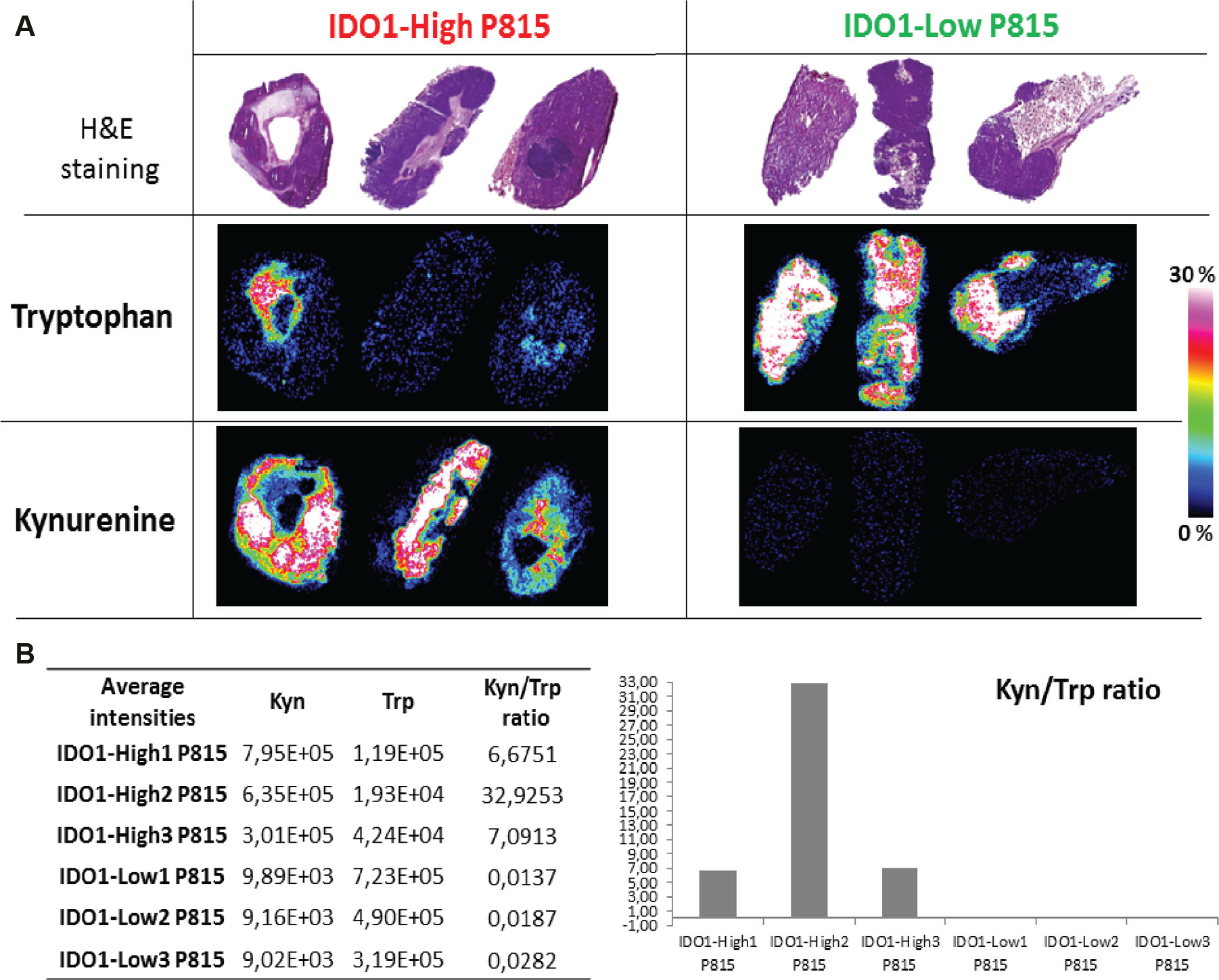
Matrix-assisted laser desorption/ionization (MALDI) mass spectrometry imaging (MSI) of tryptophan pathway: an insight in the tumor metabolism. IDO1-high and -low P815 tissue sections of 10 µm thickness in triplicates were analyzed by MALDI MSI after 1,5-DAN matrix deposition. Serial tissue sections were stained with hematoxylin and eosin. Tryptophan and kynurenine histological localization is shown in triplicate in both models (
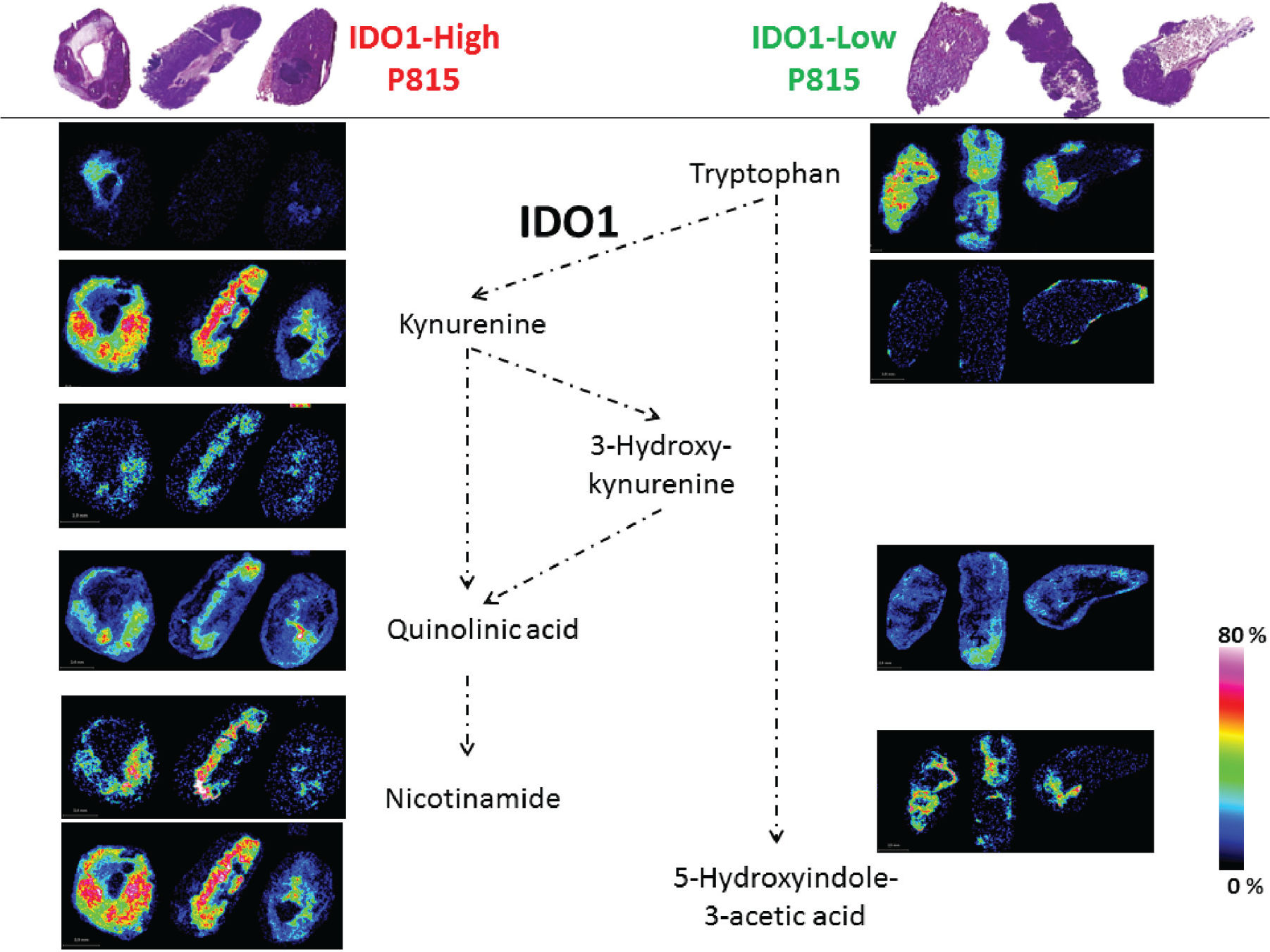
Matrix-assisted laser desorption/ionization (MALDI) mass spectrometry imaging (MSI) of tryptophan pathway: an insight in the tumor metabolism. Implicated metabolites in Trp catabolism pathway were detected using 7T-MALDI Fourier transform ion cyclotron resonance MSI in biological triplicates of IDO1-high and -low P815. Scale colors and bars are shown for each section.
Quantitative MALDI MSI of Tryptophan and Kynurenine
For more biological and analytical accuracy, an absolute quantity evaluation of both Trp and Kyn was performed in both P815-IDO1 models. An assay method was developed and applied to both tumor models using biological triplicates (each in duplicate). First, internal standards concentration was set following a first test on which we obtained an isotopic labeled compound (ILC) ratio >0.1 (endogenous molecule intensity/internal standard intensity >0.1; Suppl. Table ST2 ). Figure 3 shows the spots of molecules of interest Trp ( Fig. 3A ) and Kyn ( Fig. 3B ; spiked on liver tissue) that were analyzed after normalization. Then, Multimaging software calculated the calibration curve equations and correlation coefficient corresponding to both metabolites. The two calibration curves were found to be linear (R2 > 0.993) in the range (125–0.5 pmol) after IS normalization, followed by the calculation of limits of detection and quantitation. Finally, both calibration curves were used to perform the absolute quantitation on ROIs (IDO1-low and IDO1-high tumors). For each model, biological triplicates (triplicates 1, 2, 3) were used in duplicate (duplicates 1 and 2). Following the found equation, for each normalized average intensity, a corresponding value of quantity was found for all the ROIs. Thus, the results demonstrated a significant average level increase of Trp in IDO1-low (1.27–4.35 µg/g) compared with IDO1-high on which signal is detectable but not quantifiable (below limit of quantitation; Fig. 3C ). Kyn expression is increased in IDO1-high (25.51–4.1 µg/g) compared with IDO1-low, on which there was no signal detection (below limit of detection; Fig. 3D ).
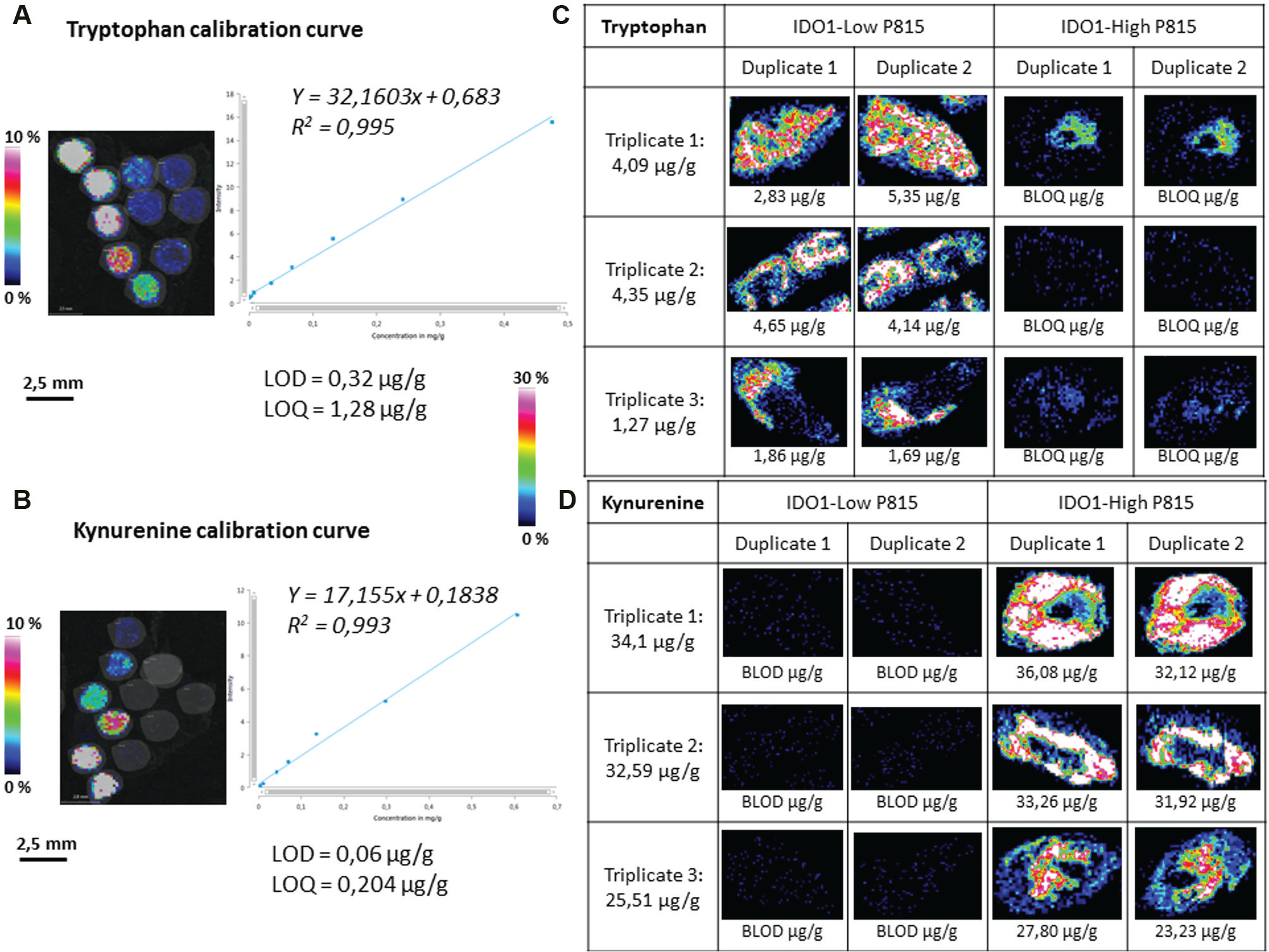
Quantitative matrix-assisted laser desorption/ionization (MALDI) mass spectrometry imaging (MSI) of tryptophan and kynurenine. Calibration curves were performed using spiked analyte on 10 µm liver tissue sections. Internal standards mixed to the matrix were sprayed over all the regions of interest. Multimaging software was then used to normalize and calculate the calibration curve equations and correlation coefficients specific to both metabolites. Linear (R2 > 0.993) calibration curves in the range (125–0.5 pmol) after IS normalization were followed by limit of detection (LOD) and quantitation (LOQ) calculation (
Confirmation of absolute endogenous Trp and Kyn tissue levels was also performed using standard LC-MS/MS analysis after metabolites extraction from serial tumor tissue sections ( Suppl. Fig. SF2 ).
The intersample variability and intratumor heterogeneity were also noticed between the triplicates and even the duplicates (distant serial sections).
High-Resolution Imaging and Definition: An Access to the Cell Level
Using Multimaging software, high-definition stained images (H&E or IHC) were overlaid to the high-resolution images performed at 30 µm resolution. Figure 4A shows a high-definition overlay of an ROI with the convoluted molecular localization of Kyn. The molecular convoluted images of Kyn and Trp are shown in Figure 4B to be complementary (Kyn in red and Trp in green colors). We performed IHC staining of IDO1 and GCN2 markers. As expected, Kyn was shown to co-localize with IDO1 expression and Trp deprivation marker GCN2. Indeed, we showed a high-resolution overlay image with co-localization of low Trp level, high Kyn level, and GCN2 expressions ( Fig. 4C ). In these same specific tumor areas, low Trp and high Kyn fits with an overexpression of IDO1 ( Fig. 4D ). In parallel, a high Trp expression co-localizes in other tumor regions with an absence of GCN2 and IDO1 staining.
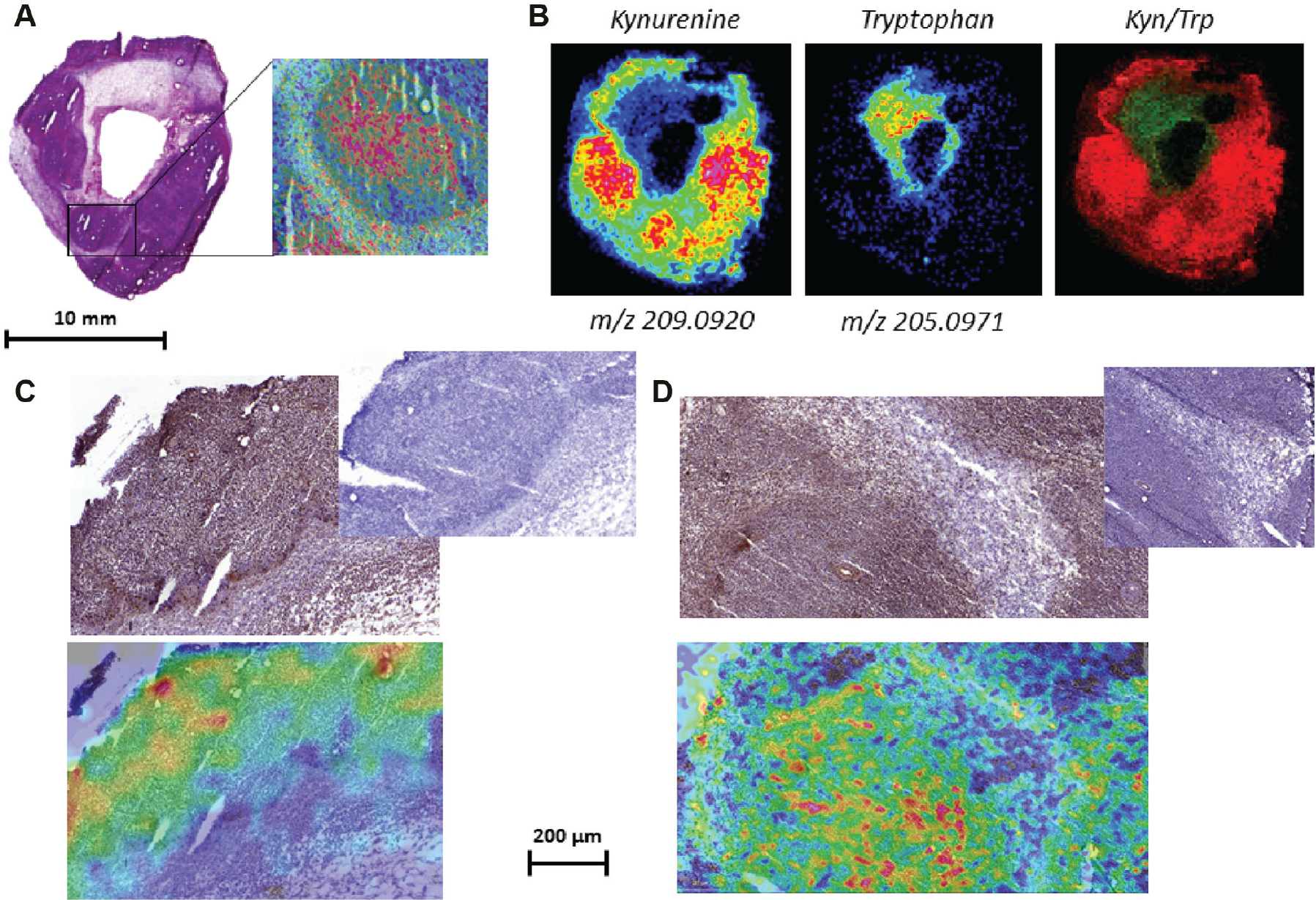
High-resolution imaging and definition: an access to the cell level. High-definition scan of a hematoxylin and eosin–stained IDO1-high P815 tissue section was overlaid with a Kyn molecular image (
Characterization of the Tumor Microenvironment Metabolism by MALDI MSI
Proliferating cancer cells alter the metabolic composition of the extracellular tumor microenvironment. Figure 5 highlights the histological region (cellular) typing following metabolite characterization such as lactate, glucose, glutamine as glycolysis signature; arginine, ornithine, proline as myeloid cells markers; and inosine and adenosine cloud. Use of extracellular metabolites such as glucose and glutamine by cancer cells during glycolysis results in the accumulation of extracellular lactate. This figure shows the correlation between tumor cell phenotype and metabolism. For instance, IDO1-positive tumor cells consume Trp and glucose and are characterized by low Trp and low glucose patterns by MALDI MSI. IDO1-negative tumor cells reflect the weak glycolysis activity that perfectly correlates with Trp overexpression. By overexpressing tryptophan-degrading enzymes such as IDO1, cancer cells modify the tumor microenvironment. Trp depletion and Kyn increase will affect the endogenous metabolism by inhibiting T-cell effector function and metabolism while promoting the regulatory T-cell phenotype and M2-macrophages and generating tolerogenic dendritic cells.
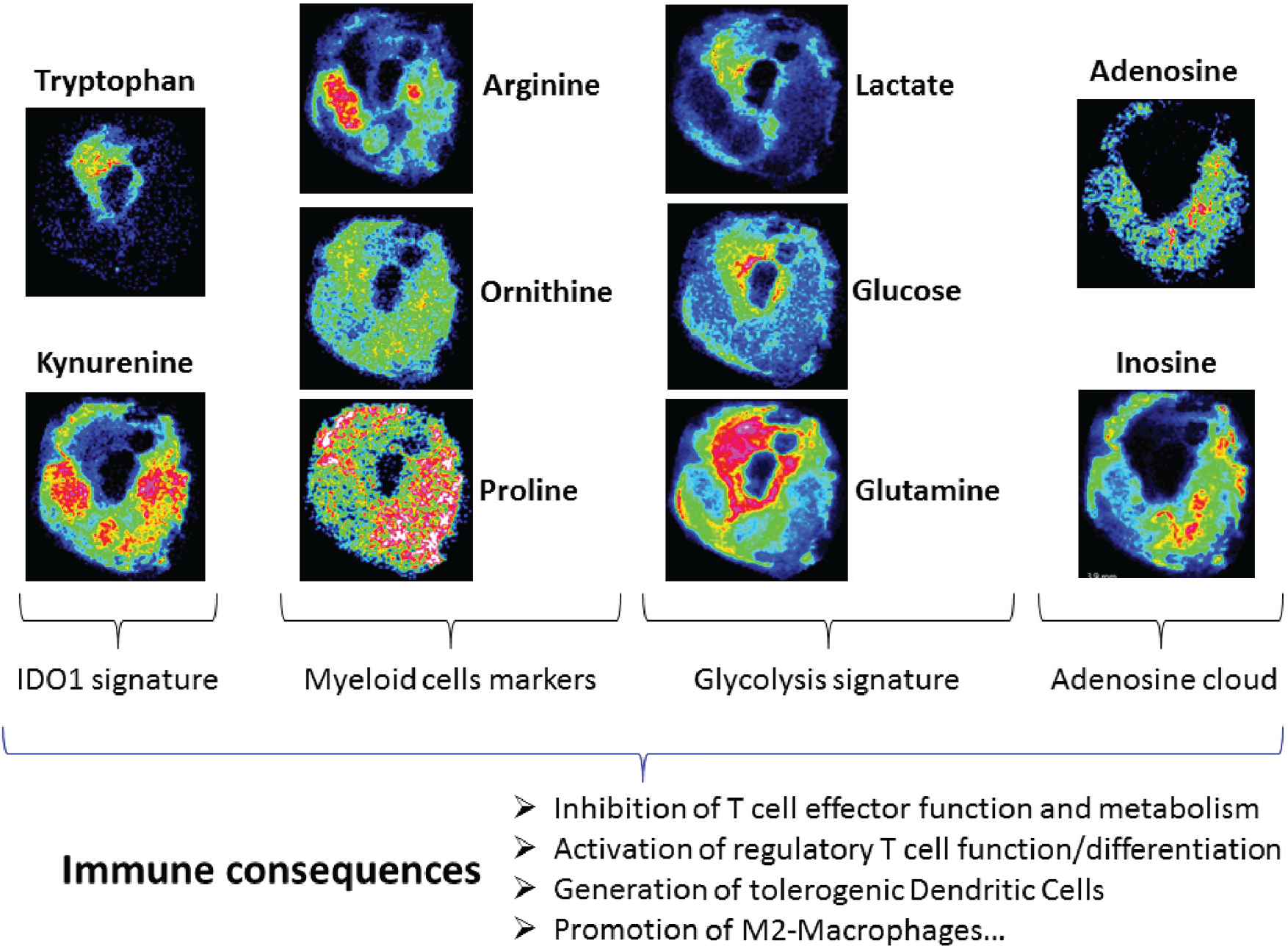
Tumor metabolic interactions with the microenvironment highlighted by matrix-assisted laser desorption/ionization mass spectrometry imaging. IDO1-mediated Trp degradation results in inhibition of T-cell effector proliferation, apoptosis, de novo hyperactivation of T-reg, generation of tolerogenic dendritic cells, and promotion of M2-macrophages. Cancer cells alter the chemical composition of the extracellular medium, which exerts pleiotropic effects on the phenotypes of normal cells that reside in the vicinity of the tumor as well as the extracellular matrix. Reciprocally, the microenvironment affects the metabolism and signaling responses of cancer cells themselves.
Some other metabolites involved in other metabolic pathways were also detected using 7T-MALDI FTICR MSI of the same tissue section ( Suppl. Fig. SF3 ) and in which the ultra-high-resolution and mass accuracy enabled elemental formula to be assigned and isotope patterns to be matched.
Discussion
The P815 cell line is an experimental tumor model that has led to pivotal advances in tumor immunology, culminating in the preclinical development of human cancer vaccines for melanoma. 20 In this study, we investigated IDO1 metabolic role by characterizing IDO1-low and IDO1-high in transfected P815 mouse tumor models. Qualitative comparison between IDO1-low and -high tumor tissue sections confirmed the differential metabolic profiles of the two models. Tissue Kyn/Trp ratios using average intensities were raised confirming the high and low IDO1 activities of both models, respectively.
The histological localization of IDO1-dependent metabolites from the Trp degradation pathway (kynurenine, anthranilic acid, 3-hydroxy-kynurenine, quinolinic acid, nicotinamide, and 5-hydroxyindole-3-acetic acid) was well highlighted by our MSI results. As expected, substrate (Trp) and products (Kyn and its derivatives) of IDO1 enzymatic activity were distinctively expressed depending on the expression level of IDO1. MALDI MSI showed and confirmed the role of IDO1 as a pivotal molecular switch controlling the functional status of Trp and Kyn metabolites pathway.
Based on the fact that all variations in tumor cellular metabolism are a hallmark of cancer, 21 the impact of IDO1 function on its metabolites expression was of high interest and importance. For this purpose, absolute quantitative MALDI MSI was performed on biological triplicates and in duplicate to support qualitative results with higher analytical confidence about the real endogenous levels of Trp and Kyn.
Despite the lack of histological localization, the high sensitivity of other analytical methods such as LC-MS/MS has already been used in patients with various cancers, including acute myeloid leukemia, in which the serum kynurenine/tryptophan ratio (Kyn/Trp) was raised. This analysis allowed for the detection of a higher IDO1 activity than in healthy people, suggesting that patients with higher Kyn/Trp ratios showed lower survival. 22 Serum concentrations of Kyn have also been reported to predict overall survival in diffuse large cell lymphoma 23 and to correlate with treatment response in adult T-cell leukemia/lymphoma. 24 Moreover, measuring kynurenine, tryptophan, and downstream metabolites in patients’ sera has been assessed in a cohort of myelodysplastic syndrome patients. 25 More recently, the first-in-human phase 1 study investigating the maximum tolerated dose, safety, pharmacokinetics, pharmacodynamics, and antitumor activity of epacadostat (INCB024360), a potent and selective inhibitor of IDO1, followed kynurenine/tryptophan ratio by LC-MS/MS analysis. 26 Thus, Trp catabolism can be easily monitored in patient sera and may have predictive value. However, biomarker histological localization is still of high importance given the correlation between the immune infiltrate and the metabolite signature (beyond the Kyn/Trp quantitation). Regarding this high tumor heterogeneity, many notable applications in cancer research are effectively based on the determination of marker levels. This could characterize a patient population with a better likelihood to respond to some treatments, such as the expression of human epidermal growth factor receptor 2 (HER2) for trastuzumab (Herceptin) therapy. HER2 has been considered since 2006 as a biomarker in patients diagnosed with invasive breast cancer. 27 In this specific example, the two tests for HER2 status, IHC and fluorescence in situ hybridization, rely on a highly subjective manual interpretation step. By employing MALDI MSI, Walch and colleagues 28 constructed a model for the classification of breast tumor HER2 status with a sensitivity of 83% and specificity of 92%. For this purpose, MALDI MSI is considered as a very promising ex vivo tool to investigate, using molecular lenses, histological biomarkers of diagnosis, prognosis, and response to treatments. 29
Furthermore, MALDI MSI is particularly suited to biomedical research because the high-resolution MSI-analyzed tissue section can be aligned with high-definition histological images (such as IHC, ISH, and IF images). Even if using conventional immunohistochemical staining (IHC) allows for obtaining high-resolution distribution images of targeted proteins, its significant limitation is still the need for labeling, which means that the target molecules must be known prior to the experiment. Alternatively, MSI has evolved as a powerful tool for the analysis of a wide range of molecules (label-free), mainly using MALDI-MS but also using other ionization methods such as desorption electrospray ionization (DESI) mass spectrometry and secondary ion mass spectrometry (SIMS). Thus, molecular signatures from specific histopathological entities may be extracted from the often-heterogeneous tissues encountered in biomedical research, especially in oncology. Although the underlying concept is simple, factors such as choice of ionization method, sample preparation, instrumentation, and data analysis must be taken into account for successful and randomized applications of MSI. In addition to that, a strong correlation was observed between the Kyn and Trp concentrations determined from the LC-MS/MS and the intensities calculated in the MSI method, suggesting the future potential of MSI for quantitative analysis in multiple organs simultaneously and of its applicability in clinical settings. To do that, this method intends to respond to the main challenges encountered in quantitation through MALDI imaging dedicated to whole-body or single heterogeneous organ samples (brain, eye, liver), as the high dependence of the detected signal on the matrix deposition, the MALDI ionization yield of specific target molecules, internal reference standard, and lastly, the ion suppression effect on the tissue. Many strategies have been, and continue to be, tested to address limitations in qMSI. Suitable software has been developed to interpret complex imaging data produced by quantitation experiments. In the coming years, more applications in preclinical and clinical research will be developed for qMSI to accelerate discovery and development of new drugs and validation of biomarkers. 30
At this stage, our study showed the spatial correlation between IDO1 expression and the Trp/Kyn levels and localization. Implication of the integrated stress response kinase GCN2 was also shown. This kinase is activated by amino acid deprivation and is recognized as an important effector of the IDO1 pathway. 31 Given the contradictory recent data about GCN2 involvement 32 in another tumor model, our results warrant further studies of the activation status of GCN2. Indeed, Sonner et al. 32 investigated whether GCN2 attenuates tumor rejection in experimental B16 melanoma using a T-cell–specific GCN2 knockout mice model system. Despite high tryptophan turnover by using immune checkpoint inhibitors, maintenance of intratumoral tryptophan levels was demonstrated using MALDI time-of-flight (ToF) MS imaging of B16 melanomas. Their use of MALDI-ToF to assess ex vivo tryptophan levels informs about the analyte signal that is high relative to other locations. Moreover, a comparison between similar samples (e.g., series of plasma samples sections) should be constant because of the constant background for a given sample run repeatedly. Contrariwise, comparing different samples (e.g., series of different tumors), where the studies may have been performed at different times, with different instrument conditions/settings or even on different instruments by different operators requires incorporating internal standards. In our study, internal standards of Trp (Trp-d5) and Kyn (Kyn-d4) were included and, in addition, we have used a high mass resolution MALDI instrument. Using 7T-FTICR MALDI, many isobaric species are easily resolved, whereas the more commonly used ToF systems amalgamate the signals. As an illustration, Supplemental Figure SF1 shows the high resolved average spectrum extracted from an IDO1-high P815 tumor section with zooms on specific Trp catabolism pathway metabolites detected with 7T-MALDI FTICR MSI in negative ionization mode with 1,5-DAN as a matrix.
Moreover, Trp catabolism by IDO1 is known to alter inflammation and favor T-cell tolerance in cancer, but the underlying molecular mechanisms remain poorly understood. 31 MALDI MSI allowed the identification of multiple metabolites from various metabolic pathways. Our study showed tumor metabolic interactions with the microenvironment resulting in cell-typing analysis that could be summarized in three principle states: proliferation of tumor cells, suppression of the effector functions and metabolism of tumor-associated T-cells, and promotion of a tolerogenic microenvironment.33,34 Metabolites promoting the emergence of an immunosuppressive microenvironment were identified and localized on tumor tissue. The high utilization of extracellular glucose and glutamine by proliferating cancer cells results in the accumulation of extracellular lactate, which is shown to affect a number of cell types within the tumor microenvironment. Increased lactate levels promote the emergence of an immune-permissive microenvironment by attenuating dendritic and T-cell activation and monocyte migration.35–37 In addition, lactate stimulates the polarization of resident macrophages to a so-called tolerogenic M2 state, which is consistent with an overexpression of some specific metabolites as arginine, ornithine, and proline. 38
Overall, the use of IDO1-overexpressing tumor models provided proof-of-concept of the usefulness of MALDI-MSI technology to study not only the Trp/Kyn pathway but also multiple metabolic pathways in the tumor microenvironment.
Finally, small-molecule inhibitors of IDO1 are currently under preclinical and clinical development. 39 We could position our quantitative MALDI MSI methodology at different levels during drug discovery, development, and marketing:
to monitor proximal pharmacodynamics biomarkers of drug efficacy, by measuring the early metabolic response (Trp and Kyn endogenous levels);
to study the histological PK/PD relationship, by running a pharmacokinetic study of the drug and showing the correlation between drug presence and target modulation in tumor microenvironment;
to further characterize the mechanism of action of the drug at the immune and metabolic levels, by following the qualitative or quantitative expressions of other metabolites that are involved later in the T-cell effector and T-cell regulatory functions and macrophages polarization (such as arginine, ornithine, proline, glutamine, etc.). The high-resolution imaging and the ability to produce a high-definition overlay between metabolites and some biomarkers of angiogenesis or expression factors will be of great added value for the identification of intratumor heterogeneity depending on the immune microenvironment areas;
to explore the putative toxicity of the drug; and
to mediate the relationship between inflammation, immune state and tumor diagnosis by highlighting indications for tryptophan degradation, measured by Kyn/Trp linked to other metabolites.
To summarize, our developed methodology could be applied to other pharmacodynamics immuno-oncology studies in preclinical and clinical samples, toward enhancing the efficiency of immunotherapies.
Footnotes
Acknowledgements
The authors thank Reece Marillier for his great assistance in providing biological materials, and Hélène Cazier for her precious preliminary results regarding mass spectrometry methods optimizations.
Supplementary material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors thank the grant obtained for Tumor Microenvironment Characterization and Applications in Immuno-oncology (A1408004N) sponsored by la MEL (Metropole Europeenne de Lille), la Region Nord Pas de Calais et BPI France. The authors also thank the grant obtained from the Wallonia Region.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
