Abstract
Doxorubicin (DOX) is a leading anthracycline drug with exceptional efficacy; however, little is known about the molecular mechanisms of its side effects, which include heart muscle damage, noncancerous cell death, and drug resistance. A total of 17,950 human proteins expressed in HEK293 cells were screened and yielded 14 hits. Competitive and binding experiments further verified the binding of DOX to UMP/CMP kinase 1 (CMPK1), and microscale thermophoresis showed that DOX binds to CMPK1 with a Kd of 1216 nM. In addition, we observed that the binding of DOX to CMPK1 activated the phosphorylation of CMP, dCMP, and UMP. A significant activation was observed at the concentration of 30 µM DOX and reached plateau at the concentration of DOX 30 µM, 150 µM, and 100 µM, respectively. DOX would add up stimulation of CMPK1 by DTT and overcome inhibition of CMPK1 by NaF, EDTA. In summary, we showed that DOX might bind to the nonactive site of CMPK1 and regulate its activity with magnesium.
Introduction
Doxorubicin (DOX; also called Adriamycin) is an anthracycline compound isolated from Streptomyces peucetius. Since its clinical application in 1970s, DOX has become the first-line drug for breast cancer. Now it is widely used for the chemotherapy of acute myeloid leukemia, endometrial tissue carcinoma, esophageal cancer, and non-Hodgkin’s lymphoma. 1 DOX has shown great efficacy in killing cancerous cell and is regarded as one of the most potent chemotherapeutic drugs among those approved by the Food and Drug Administration. Despite its wide clinical use, DOX causes potential side effects (heart muscle damage, 2 noncancerous cell death, 3 and drug resistance 4 ). However, little is known about the molecular mechanisms of its side effects. Because the mechanism of DOX involves its intercalation between DNA base pairs, inhibition of topoisomerase II, 5 DNA adduct formation, and production of reactive oxygen species, it is possible that it may act on multiple molecular targets to produce a range of cytotoxic effects.1,6 In addition, DOX is a zwitterion. It enters the cells by diffusion or transporters (organic anion transporters [OAT]s; organic cation transporters [OCTs]) 7 and is eliminated from the cells by ATP-binding cassette–related transporters. 8 Many kinases (AKRs, ubiquitous cytoplasm carbonyl reductase, etc.)9,10 were found to participate in the metabolic pathway of DOX.
CMPK1 is crucial for cellular nucleic acid synthesis by supplying precursors, which is part of pyrimidine nucleotide interconversion pathways, and phosphorylating CMP, UMP, and dCMP to their respective diphosphate forms by catalyzing γ-phosphoryl, usually adenosine 5′-triphosphate (ATP), with the presence of magnesium in both de novo and salvage pathways. 11 UMP/CMP kinase 1 (CMPK1) plays an important role in the activation of deoxynucleoside analogs, many of which are important antiviral and anticancer agents. For example, gemcitabine (2′, 2′-difluordexorycytidine) is clinically used to treat pancreatic cancer and several other cancers. Analogs such as L-ODDC (β-L-dioxaolanecytidine), 1-β-D-arabinofuranosylcytosine, and ddC (β-D-2′, 3′-dideoxycidine) are all active anticancer or antiviral agents. CMPK1 is reported as a prognostic marker for non–small-cell lung cancer 12 and for pancreatic cancer patients when treated with a gemcitabine-based chemotherapy. 13 CMPK1 has also been identified as a prognostic marker for lymph node–negative and adjuvant systemic chemotherapy–naive triple-negative breast cancer patients. The nuclear localization of CMPK1 may be related to dysregulation of the extracellular matrix and cell cycle molecules. 14 Here, we report for the first time the interaction between DOX and CMPK1 in vitro.
To further understand the molecular mechanisms of DOX within the cell, we used a high-density protein microarray to screen a total of 17,950 unique mammalian-expressed proteins with biotinylated-DOX (Bio-DOX). The use of streptavidin/biotin (Ka = 1015) interaction supplied fast streptavidin-Cy3/5 interaction, allowing the capture of biotinylated drug-protein complex. We further proved CMPK1 as a novel DOX target and exhibited a Kd of 1216 nM by microscale thermophoresis (MST). Kinase activity assay showed that 30 µM of DOX significantly stimulated the CMPK1 activity. Additional DOX can superimpose the activation of CMPK1 by DTT and reverse the inhibition of CMPK1 by EDTA, NaF, and gemcitabine.
Materials and Methods
Reagents
High-density protein array (the development of the high-density protein array was described by Ma et al. 15 ), 4C5 monoclonal anti-DDK antibody, CMPK1 antibody, and cDNA ORF clones of hits were gifts from Origene (Rockville, MD); DOX was purchased from Meilun (Tianjin, China); Bio-DOX synthesis related agents were from Sigma (St. Louis, MO); streptavidin and steptavidin-Cy3/5 were purchased from Jackson Laboratory (Bar Harbor, ME); anti-mouse IgG Fc capture surface sensor was purchased from FortéBio (Menlo Park, CA); cytidine phosphate high-performance liquid chromatography (HPLC) marker, CMP, dCMP, UMP, and ATP were purchased from Sigma. CMPK1 protein was purchased from Sino Biological (Beijing, China).
Synthesis of Bio-DOX
The biotinylated DOX with a linker chain of 6-carbon was synthesized as described earlier16,17 with some optimization ( Fig. 1A ). The reductive alkylation was substituted by direct acylation catalyzed by dicyclohexylcarbodiimide and 4-dimtehylamino-pyridine.
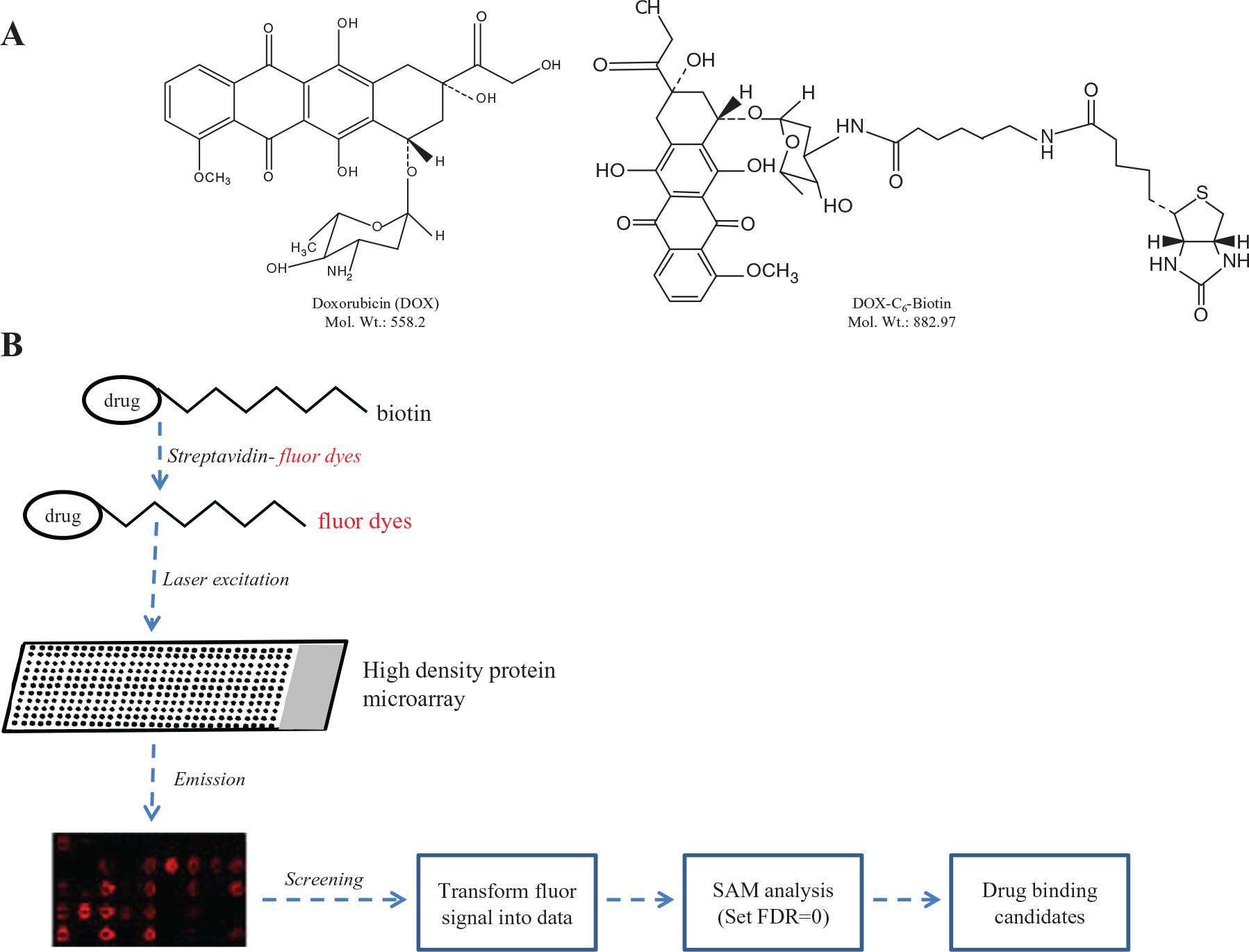
Overall strategy for the identification of binding protein hits using protein microarrays. (
Methods Performed in Protein Arrays
The array contains a total of 38,400 spots with 17,950 lysates in duplicates (35,900 spots) and 2500 control spots. The whole slide is divided into 48 subarrays (4 × 12). The detailed protocol is described in Ma et al. 15 Every cDNA clone used for overexpression lysate production contains a DDK epitope tag. The confirmation and pretreatment of the protein array were performed as reported. Bio-DOX (25 µM) and streptavidin-Cy3/5 (2 µg/mL) were incubated at 4 °C for 1 h to form Bio-DOX-streptavidin-Cy3/5, and then Bio-DOX-streptavidin-Cy3/5 was incubated with the array at 4 °C overnight. The next day, the solution was discarded and the array was washed in 20 mM phosphate-buffered saline (PBS) gently for 1 min. The fluorescence signals of Cy3/5 were detected by GenePix 4100A (Molecular Devices, Sunnyvale, CA). The pixel intensities of each spots were determined using GenePixPro 6.0 software. The mean value of replicates was used for subsequent significant analysis for microarray (SAM) assay. The false discovery rate was set to 0. The mean signal-to-noise ratio was calculated in the same block, and the Student t test of positive spots must achieve p < 0.5 ( Fig. 1B ).
Competitive Method
DOX, Bio-DOX, and streptavidin-Cy5 were mixed with the ratio [DOX]: [Bio-DOX]: [streptavidin-Cy5] = 200:1:1, and the mixtures (DOX group) were incubated together with the microarrays as described in the method performed in protein arrays; the microarray incubated with the ratio [Bio-DOX]: [streptavidin-Cy5] = 1:1 was set as control. The arrays were scanned for Cy5 (Red). The signal of each dot was normalized according to orientation marker control in the array. Each subarray of the array contains an autofluorescent BSA Cy5 and Cy3 mixture spot and a mixed IgG spot (mixture of human, mouse, and rabbit IgGs). The ratio changes of control group/DOX group were greater than 1.5-fold confirmed as positive hits.
Production of DOX Hits for Binding Experiment
Plasmid DNAs of 14 hits and one positive control (AKR1C3) were obtained from OriGene Technologies Inc. and transformed into competent DH5α Escherichia coli cells, followed by selection on Luria-Bertani medium with 50 µg/mL kanamycin at 37 °C for 14 to 16 h. After extraction (Generay Biotech, Shanghai, China), the concentration and purity of DNA were determined using NanoDrop (Thermo Scientific, Waltham, MA). The DNAs were then stored at −30 °C for further use. HEK293 cells were cultured in DMEM supplemented with 10% fetal bovine serum and 1% antibiotics (penicillin and streptomycin) in 15-cm dishes. When the confluence reached 70% to 80%, cells were transfected with a mixture of liposome 2000 (Invitrogen, Carlsbad, CA) and each individual plasmid DNA of 14 hits and AKR1C3. After 48 h in culture, cells were scraped and washed with PBS at 4 °C. After the last wash, cells were sonicated (80 w, 10 pulses of 1 s each, with 1 s interval between pulses) in 500 µL PBS and centrifuged at 14,000 rpm, 4 °C for 2 h. The supernatant was collected, its protein concentration was determined using NanoDrop, and stored at −80 °C for later use.
Binding Experiment
The interaction between DOX and 14 hits was tested by the FortéBio Octet RED96 (FortéBio). 18 Because every cDNA clone used for overexpression lysate production contained a DDK epitope tag, the 4C5 monoclonal anti-DDK antibody was used to capture the recombinant proteins. The biosensors (anti-mouse IgG Fc capture surface sensor) were wetted in dialysis buffer for 15 min prior to use. A total of 30 µg/mL DDK antibody was immobilized to the biosensors and then loaded protein lysate (all expressed protein contains a C-terminal DDK tag) to the biosensors. The process of association and dissociation of the proteins with DOX was monitored in parallel. The biosensor without loading-expressed protein was used as a negative control, and the biosensor loading AKR1C3 was used as a positive control. The experiment was carried out at 30 °C. The data were analyzed by the software in the FortéBio Octet RED96.
Microscale Thermophoresis
Serial dilutions of DOX or bovine serum albumin were incubated with 20 nM of a NT647-labeled CMPK1 in MST buffer (1% DMSO) at ambient temperature for 30 min. Microscale thermophoresis measurements were performed using a Monolith NT.115 instrument (NanoTemper Technologies GmbH, Munchen, Germany). The measurements were performed using standard capillaries at 40% LED excitation and 20% MST power, with a laser-on time of 30 s and a laser-off time of 5 s. NanoTemper Analysis 1.2.20 software was used to fit the data and to determine the Kd.19,20
CMPK1 Activity Assay
The enzyme assay was performed in 300 µL of 50 mM Tris-acetate, pH 7.5, 2 mM MgCl2. ATP (2 mM) was used as phosphate donor. 11 (d) CMP, (d) UMP, and enzymes were added sequentially. Influence factors of the reaction such as gemcitabine (1 mM), EDTA (1 mM), NaF (5 mM), and DTT (2 mM) were tested. Dexrazoxane (600 µm) was also tested because it is a treatment for DOX-induced cardiotoxicity. The reaction mixture was incubated at 37 °C for 2 min–120 min and stopped by the addition of 100 µL of 45% trichloroacetic acid and chilled on ice. The activity was determined by analysis of the product by HPLC (Agilent 1260) with a ZORBAX C18 column (4.6 × 250 mm, 5 µm; Agilent, Santa Clara, CA). Methanol with 0.6% phosphoric acid (6:94, pH adjusted to 6.6 by triethylamine) was used as mobile phase. The nucleotide phosphates were detected by ultraviolet light absorbance at 271 nm and quantified by comparison to standards.
Results
Identifying DOX Hits from High-Density Protein Microarrays
We first applied biotin alone to exclude the endogenous reaction, and only two proteins positively responded to biotin alone: pyruvate carboxylase and propionyl-CoA carboxylase alpha subunit, both reported to contain biotin-binding regions. Thus, we inferred that the method of conjugating C6-biotin to drug for the detection of interacting proteins was effective. After SAM assay further excluded the biotin-binding proteins and nonspecific binding proteins, there were 14 hits remaining in the microarrays ( Table 1 ).
Corresponding Positive Protein Hits with Related ACCN Numbers and GO.

CMPK1 and SLC22A6 Were Confirmed as DOX-Binding Targets
Only MGAT1, CMPK1, and SLC22A6 positively responded by competitive method. DOX itself showed weak green florescence; we could see the red florescence signals of the three dots were attenuated, accompanied by the weak green florescence appearance ( Fig. 2A ). The 14 plasmids with corresponding cDNAs and AKR1C3 were transfected into the HEK293 cells individually. The lysate was collected for binding analysis with AKR1C3 used as the positive control. Lysate of HEK293 cells transfected with empty pCMV6 plasmid vector was defined as the negative control. The response of SLC22A6, TRMT112, DYNLL1, and CMPK1 was much stronger than AKR1C3 ( Fig. 2B ).
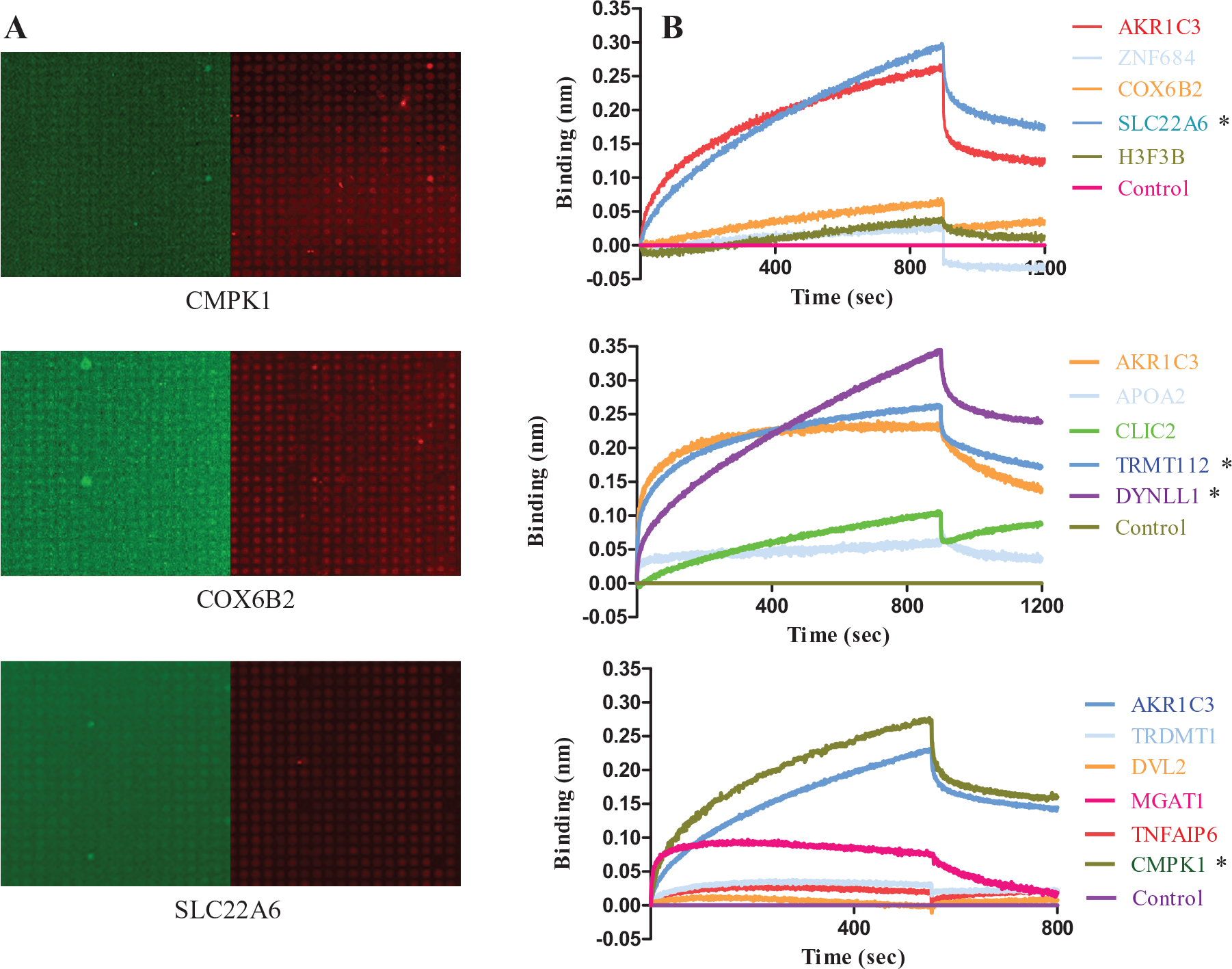
SLC22A6 and CMPK1 are the most possible doxorubicin (DOX)–binding targets. (
DOX Can Bind CMPK1 Directly In Vitro
CMPK1 and SLC22A6 had shown positive responses to DOX in both methods mentioned above. The relationship between the SLC22 family and DOX has been reported. 7 Here, we chose to study the relationship between CMPK1 and DOX. The binding strength of DOX to CMPK1 was further analyzed by MST. DOX exhibited binding when titrated to fluorescently labeled CMPK1 with a Kd of 1.2 ± 0.5 µM ( Fig. 3 ).
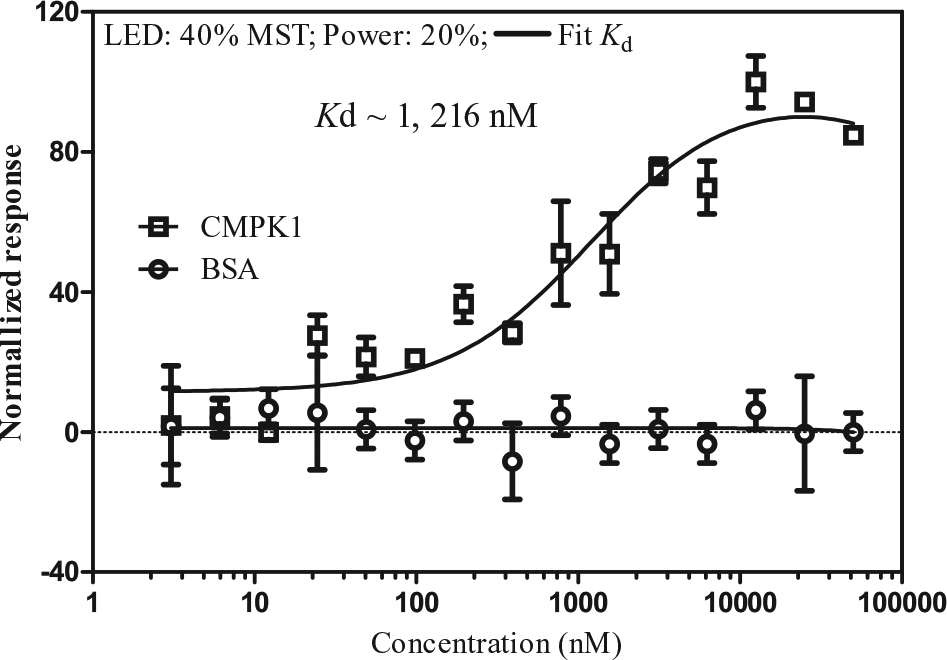
Interaction of doxorubicin (DOX) with CMPK1. Microscale thermophoresis binding curves for DOX titrated against NT-647–labeled CMPK1 (20 nM). Bovine serum albumin (BSA) was used as a negative control. Error bars show standard deviations from a triplicate experiment (duplicate experiment). Kd, dissociation constant.
Activity of CMPK1 Was Regulated by DOX
The activity of CMPK1 was up-regulated by DOX, which used CMP, dCMP, and UMP as substrate while ATP serves as phosphate donor. The EC50 activation by DOX was 66.6 µM, 29.7 µM, and 10.1 µM when using CMP ( Fig. 4A ), dCMP ( Fig. 4B ), and UMP ( Fig. 4C ) as substrate, respectively. The pH of the system before and after the reaction was not changed. By fixing the concentration of DOX at 60 µM while increasing that of CMPK1, the effect of up-regulation by DOX peaked at 2 nM and diminished when CMPK1 concentration reached 10 nM ( Fig. 4D ).
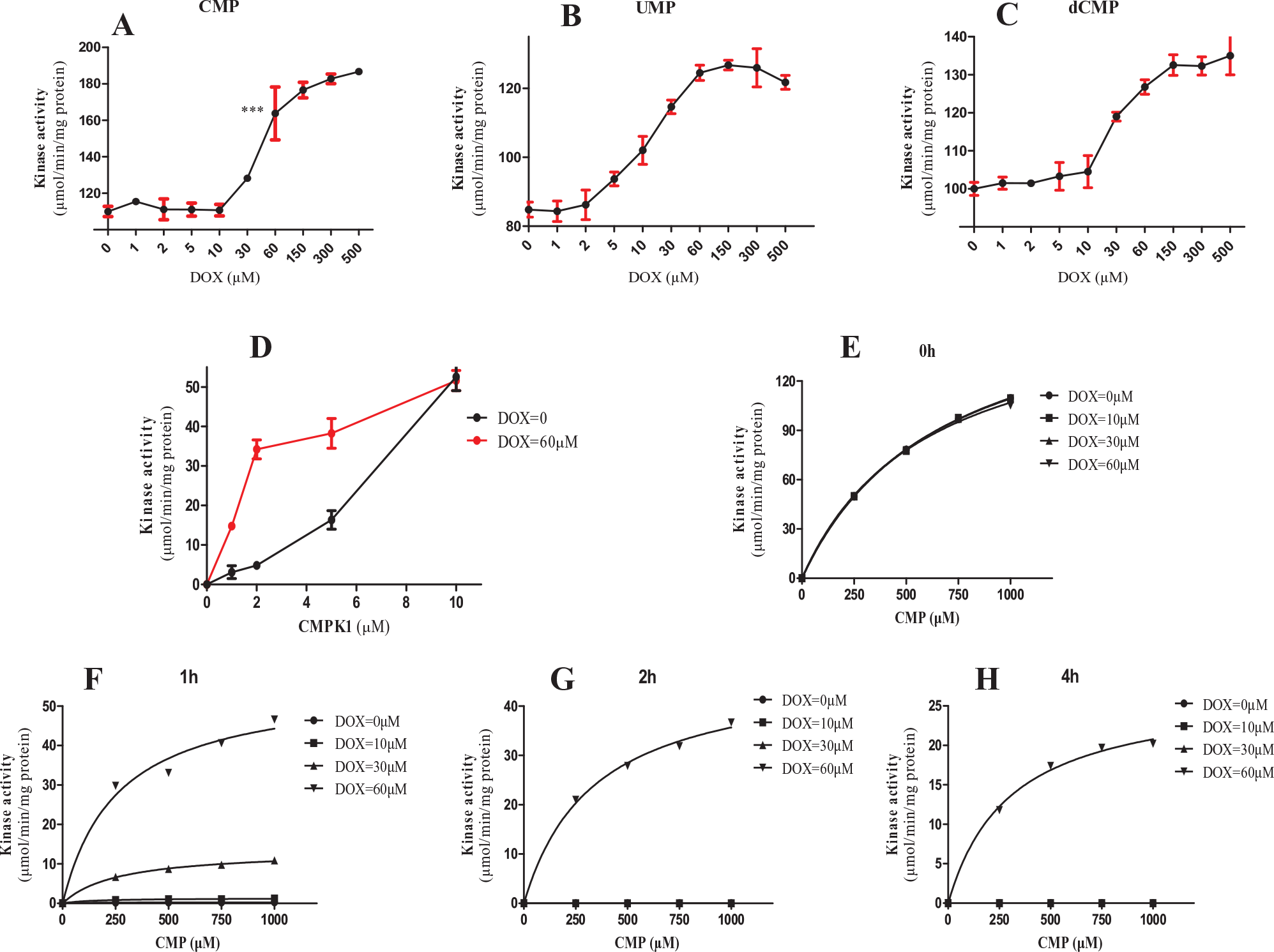
CMPK1 was recognized as the binding target of doxorubicin (DOX) and the activity regulated by DOX. (
We further examined the effect of DOX concentration on CMPK1 activity in a time-dependent manner. CMPK1 (10 nM) was incubated with various concentrations of DOX (0 µM, 10 µM, 30 µM, 60 µM) at 4 °C for 0, 1, 2, and 4 h. The enzyme reaction was initiated by the addition of CMP and ATP and stopped by trichloroacetic acid. The amount of products was analyzed by HPLC as described in the Materials and Methods section. The CMPK1 activity continued decreasing until it could not be detected after 1 h at 4 °C. In contrast, the activity of CMPK1 remained in a dose-dependent manner with addition of DOX ( Fig. 4E–H ). In summary, the enzyme activity remained at 0.72%, 7.5%, and 32.7% in the presence of DOX at 10 µM, 30 µM, and 60 µM, respectively, after 1 h incubation and at 32.7%, 28.0%, and 15.9% after 1 h, 2 h, and 4 h incubation ( Suppl. Table S1 ).
Regulation of DOX to CMPK1 in the Presence of Many Representative Factors
Gemcitabine is an analog of CMP and competitively inhibits the enzyme reaction. EDTA inhibits the CMPK1 activity by removing metal ions (Ca2+, Mg2+). Dexrazoxane was reported to affect the formation of free radicals mediated by ferric ion, which is the main cause of cardiac toxicity of DOX. 2
Gemcitabine, EDTA, and dexrazoxane were added to each experimental group, respectively. CMP and ATP were used to initiate the reaction. The activity was calculated from the amount of change of CMP. As expected, the enzyme activity was inhibited by gemcitabine, and the inhibition could be reversed by addition of DOX. EDTA showed inhibition on CMPK1 activity, and this effect could be overcome partially by addition of DOX. Dexrazoxane showed no effect on enzyme activity when compared with the control group ( Fig. 5A–D ; Suppl. Table S2 ). We also calculated the Vmax of each experiment group. Gemcitabine and EDTA decreased the Vmax of the reaction as reported, 21 and it could be reversed by addition of DOX. The Vmax of the gemcitabine group was even equal to the control group with/without DOX ( Fig. 5E ). The Vmax of the EDTA group was still below the control group regulated by DOX. DTT can also activate the reaction by reducing the CMPK1 to low-molecular form, and NaF was reported to inhibit the reaction. 21 We could see the enzyme activity loss was reversed by DOX in a dose-dependent manner, and DOX can still activate the reaction, which has been accelerated by DTT ( Fig. 5F ). In a word, the activation and/or stabilization of the enzyme by DOX was not affected by these factors, and magnesium was necessary for the reaction, which cannot be replaced by DOX.
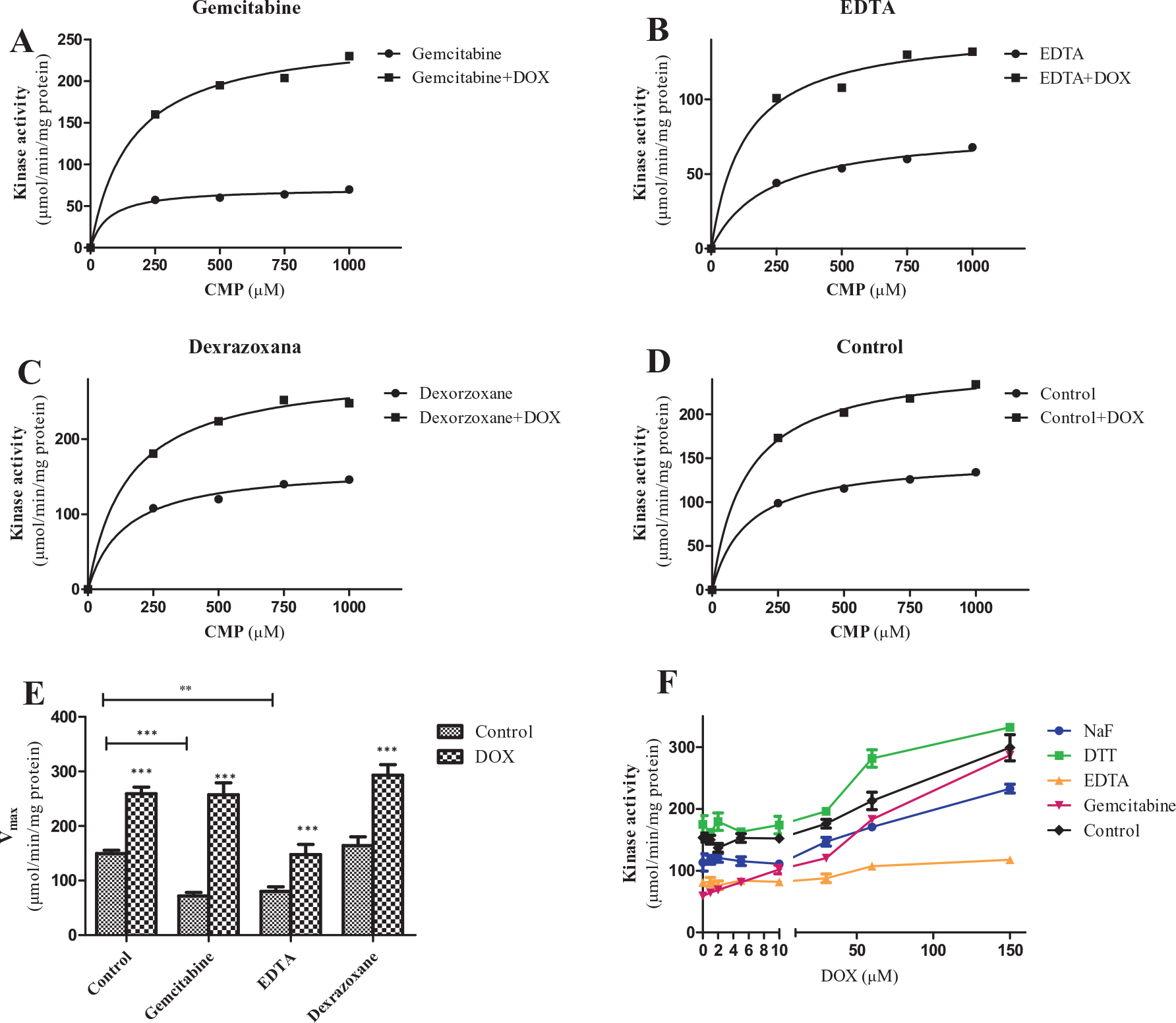
The regulation of doxorubicin (DOX) to CMPK1 in the presence of many representative factors. (
Discussion
In this study, we identified CMPK1 and 13 other proteins that positively bind to DOX in high-density microarray assays and confirmed that DOX could definitely bind to CMPK1 in vitro. We also tested whether DOX could affect CMPK1 enzyme activity, and the results showed that it could stimulate the activity in a dose-dependent manner.
We tried both streptavidin-Cy3 and streptavidin-Cy5 for the screening procedure. Cy5-conjugated streptavidin was more detectable for this experiment, so we chose Cy5-conjugated streptavidin for our further work. The wash step was first done as in Ma et al, 15 but we failed to discover any interactions. We used the moderate wash method as mentioned above to help discover low-affinity interactions.
Cardiotoxicity is a major side effect that arises during chemotherapy with DOX, ranging from arrhythmia to heart failure. 22 The activation of CMPK1 in the presence of a high concentration of DOX might cause depletion of ATP as CMPK1 uses it as a donor to form CDP and UDP, which might cause mitochondrion dysfunction in the heart, as described. 23 It had been reported that gemcitabine as a CMPK1 substrate was more effective in killing cancer cells, simply by increasing the expression level of CMPK1. A similar effect had also been observed when elevated CMPK1 expression helped DOX efficacy (unpublished data). Currently, there are many ongoing trials combining gemcitabine and DOX for treating cancers, which has shown promising activity compared with other combination therapies. 24 Our findings support the notion of combinational therapies.
Extracellular binding was often transient and exhibited lower affinity than that of endogenous compounds (hormones) in a screening assay of protein array.25,26 We could not find significant responses in the array when surface active reagent (Tween 20, 0.5%) was first used in the wash step; thus, we switched to milder phosphate buffer (20 mM) for washing. This modification enabled us to identify a total of 14 proteins as potential DOX-binding targets. We studied only CMPK1 in detail for proof of concept; the potential functions of other identified DOX-binding proteins need to be investigated. First, SLC22A6 and APOA2 have been reported as transporters,7,27 suggesting they may be involved in drug resistance. SLC22A6 is a member of the SLC22 family, which has been reported to uptake DOX into cells and heighten the sensitivity to DOX. In contrast, APOA2 has been reported to carry liposoluble drugs out of the cell, and we found there had been much less DOX in APOA2-overexpressed cells, suggesting APOA2 might anticipate DOX elimination (unpublished data). ZNF684 belongs to the ZNF family, which has been reported to be involved in drug resistance 28 and H3F3B to modulate BRCA1 to regulate drug sensitivity. 29 Second, the target proteins might cause cardiotoxicity. CMPK1 has been analyzed for the possibility of causing such an effect. CLIC2 was reported to inhibit Ca2+ release from cardiac lipid bilayers and sarcoplasmic reticulum vesicles. 30 Dvl2 is essential for normal cardiac morphogenesis, somite segmentation, and neural tube closure. 31 COX6B2 has been reported to have cytochrome-c oxidase activity, which is part of the electron transfer chain. 32 The binding of DOX to COX6B2 may cause the dysfunction of energy production and ultimately cause heart failure. Third, the target proteins might participate in cell migration and metastasis. MGAT1 is an enzyme for the formation of the N-glycan branch that enhances the cell surface residency of growth factor receptors. 33 Tumor invasion and metastasis may be affected by the interaction of MGAT1 with DOX. TNFAIP6 is a secret protein that contains a hyaluronan domain and is known to be involved in extracellular matrix stability and cell migration. 34 DYNLL1 is a light chain of cytoplasmic dyneins and is also found to regulate autophagy and apoptosis through its interaction with BIM, which is a switch protein between autophagy and apoptosis. The anoikis resistance of MCF7/DOX can cause cell migration and metastasis. 35 Lastly, the functions of TRMT112 and TRDMT1 are related to protein methylation (TRMT112) 36 or tRNA methylation (TRDMT1). 37 The association of DOX with these proteins may regulate the on and off expression of genes, and many diseases such as cancer, aging, and Alzheimer’s disease are caused by improper methylation. COASY is a kinase that uses ATP as a phosphate donor in the catalyzed reaction. 38
In summary, we set up a screening platform using high-density protein microarray and recognized CMPK1 as one of the DOX-binding targets; the enzyme activity was stimulated by the binding of DOX. In later work, we aim to research the function of those DOX-binding target proteins and screen the binding targets of other drugs based on this platform.
Footnotes
Acknowledgements
We gratefully acknowledge Yihong Deng from Quantum Design China (Shanghai) for technical support of MST experiment.
Supplementary material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of International (Regional) Cooperation and the exchange program (grant 81361168001), science and technology project of clinical medicine of Jiangsu province (BL 2014019), and Chinese National High Tech Program for Biotechnology grant 863 (2014AA020909).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
