Abstract
Analysis of interactions between molecules is of fundamental importance in life science research. In this study, we applied weak affinity chromatography, based on high-performance liquid chromatography and mass spectrometry, as a powerful tool for direct analysis of the components of a chemical reaction mixture for their binding to a target protein. As a demonstration of the potential of this method, we analyzed the binding of the compounds of the reaction mixture to the chaperone heat shock protein 90 (Hsp90). It was possible to analyze quantitatively the binding of the components of the mixture to the target independently from each other without any preceding process such as purification. This feature has wide implications in biological sciences as crude mixtures, either natural or synthetic, can be analyzed directly for their possible binding to a target. This method could lead to savings in costs and labor through shortening chemical research project development time.
Introduction
The interactome is probably the most complex situation to analyze. In a biological context, the intrinsic interplay of molecules in a living cell is tremendous and encompasses a panorama from very transient interactions to strong binders. 1 Despite vast progress in biological sciences, our understanding of the number of all the components and how they interact in a biological entity is certainly beyond our current technology, and most likely, we have so far discovered only a small fraction of it. The challenge for life science researchers is to work with these complex biological mixtures in the study of biological systems such as cell machinery. Concomitantly, there has been a rapid development of parallel organic synthetic procedures in all areas of chemical science that has resulted in the production of large complex compound libraries that are very expensive to screen as single compounds. The complexity is even higher when natural product broths and mixtures are considered as viable analytes.
As a result, there is a great need to study and process crude and purified mixtures of compounds emanating from biological and chemical sources. A huge amount of effort and resources has therefore been spent on developing tools to purify and isolate molecules of interest, which ultimately has become a significant bottleneck within biological and chemical research. An analytical tool to study directly how the components of a mixture interact with a target, such as a protein, without processing in terms of purifying individual components, would result in sizable savings in terms of cost and labor during the initial screening phase. And, most important, this tool should be able to analyze all types of interactions occurring between the target and the component, including the more transient ones. Of special importance is the interaction between protein to protein and small molecule to protein, as these interactions in many cases are subtle and demonstrate transient (weaker) binding (lower than µM affinities).
Affinity chromatography is a powerful technique for purifying a biomolecule from a crude mixture of substances. 2 By combining it into a high-performance liquid chromatography (HPLC) format and further developing the technology for analytical purposes, it was introduced as an efficient tool to study the specific interactions between a protein target and a binding analyte.3–5 Furthermore, developments showed that this technology could be applied to the whole panorama of biological interactions, including the more transient ones. In the latter case, this is termed weak affinity chromatography (WAC). 6 High-performance analytical affinity chromatography has now been applied successfully in a number of areas in bioscience.7–9
The combination of affinity chromatography (HPLC based) and mass spectrometry (MS) may offer an efficient approach (affinity-LC/MS) to the binding characterization of all interactors, including transient or weak binders, in a mixture to a target. Identification of analytes from their mass spectra is a viable approach in affinity chromatography, enabling multicomponent mixtures to be efficiently studied. 10 As presented in this study, we illustrate our method of affinity-LC/MS with the analysis of an organic crude reaction mixture of inhibitors to a chaperone protein, Hsp90. Hsp90 is an important target in cancer research. We analyzed the affinity (KD) of the components of a crude reaction mixture of analogues of the clinical stage Hsp90 inhibitor BIIB021 to the target Hsp90.11–13 For validation, we compared the affinities of the compounds from the crude mixture with a mixture of the pure compounds.
Materials and Methods
Chemicals and HSP90 Bioassay
Hsp90 protein (11.1 mg/mL; storage buffer: 20 mM 2-[4-(2-hydroxyethyl)-1-piperazin-1-yl] ethanesulfonic acid [HEPES], pH 7.5, 300 mM NaCl, 10% glycerol, 2 mM tris[2-carboxyethyl]phosphine [TCEP]) was received from Protein Production Platform at Nanyang Technological University, Singapore (www.proteins.sg). The Hsp90 bioassay was carried out by Reaction Biology Corp (www.reactionbiology.com) using FITC-labeled geldanamycin (FITC-Gd). Commercially available analytical reagent-grade solvents and anhydrous solvents packed in resealable bottles were used as received from local vendors without further purification, distillation, or drying. Compound
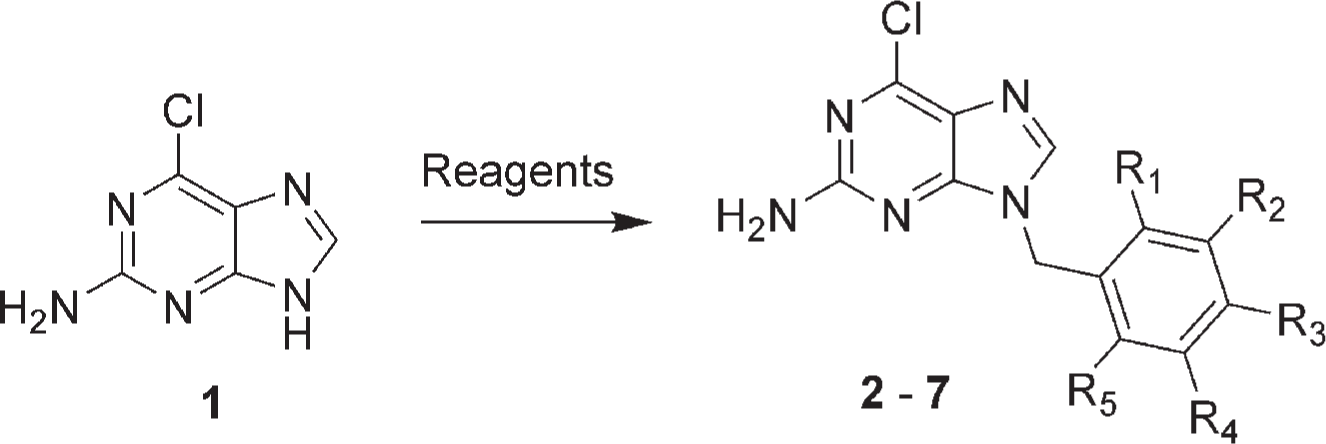
Reaction scheme for synthesis of benzyl substituted purine-based Hsp90 inhibitors. Ar-Br represents the benzyl bromide reagents. R1-5 represents the substituents of the benzyl group ( Table 1 ). Reagents and conditions: Ar-Br, K2CO3, MeCN.
Synthetic Procedures for Individual Compounds
2-Amino-6-chloro-9-substituted purine analogues (
Comparison of Affinity Values (KD) from the Crude Reaction Mixture with the Corresponding Mixture of Pure Compounds.
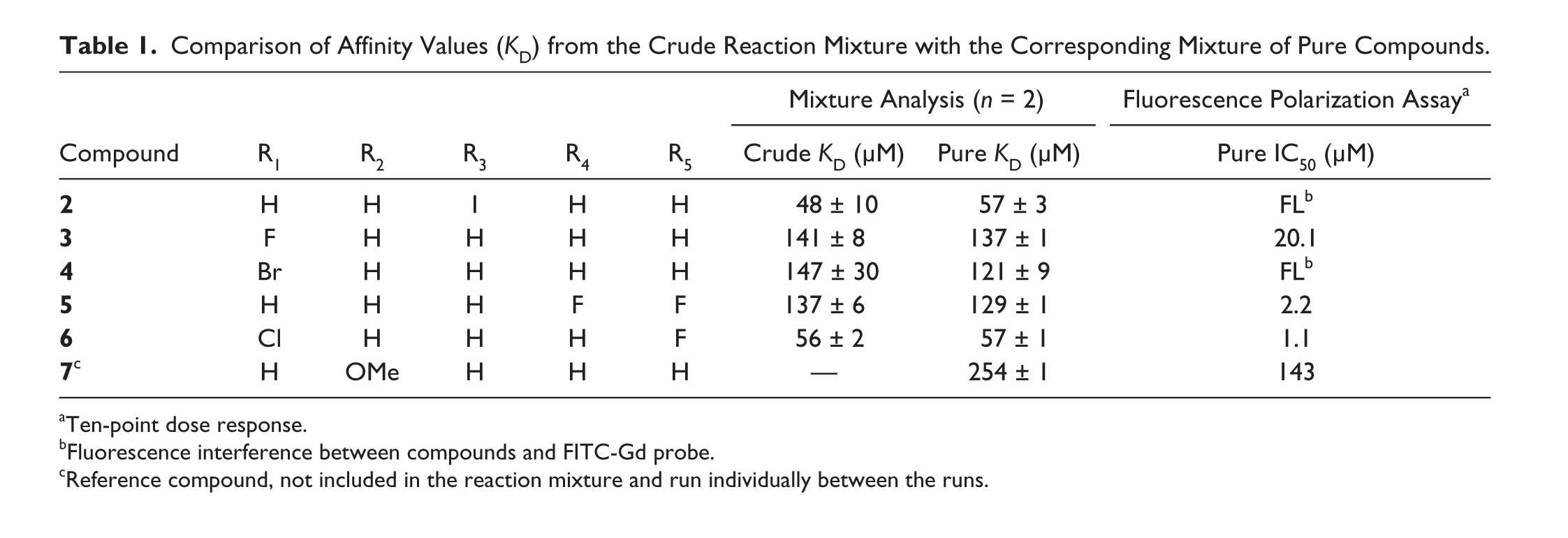

Ten-point dose response.
Fluorescence interference between compounds and FITC-Gd probe.
Reference compound, not included in the reaction mixture and run individually between the runs.
Purification
Samples were purified by flash column chromatography on silica gel 60 (0.040–0.063 mm) purchased from SiliCycle Inc. (Quebec, Canada) or Merck Millipore. The organic layer was concentrated and purified using hexane/EtOAc (2:1) to obtain the final 2-amino-6-chloro-9-substituted purine analogues. The purity of the compounds was assessed using an Agilent 1200 series HPLC system with UV detection at 254 nm on a Zorbax SB-C18 (pore size = 80 Å, particle size = 5 µm) column with dimension 250 × 4.6 mm using a gradient elution starting from a 5% solution of MeCN with 0.1% trifluoroacetic acid (TFA; and 95% solution of 0.1% TFA in water) to a 100% solution of MeCN with 0.1% TFA at 1 mL/min over 20 min. Yields refer to chromatographically and spectroscopically homogeneous materials, unless otherwise stated.
Characterization of Pure Compounds
Mass spectra were obtained on an Agilent 6130B Quadrupole LC/MS in atmospheric pressure ionization electrospray (API-ES) mode with an Agilent 1260 Infinity HPLC system using a Thermo Scientific Hypersil column with the dimension of 150 × 2.1 mm and particle size of 5 µm.
1H-NMR and 13C-NMR spectra were obtained using a Bruker AMX400 (400 MHz; Billerica, MA) nuclear magnetic resonance (NMR) spectrometer at ambient atmosphere. Chemical shifts are reported in parts per million, and residual undeuterated solvent peaks were used as internal reference: proton (7.26 ppm for CDCl3, 2.50 ppm for DMSO-d6), carbon (77.0 ppm for CDCl3, 39.52 ppm for DMSO-d6). 1H-NMR coupling constants (J) are reported in Hertz (Hz), and multiplicities are presented as follows: s (singlet), d (doublet), t (triplet), m (multiplet), and br (broad). Details of the NMR and MS characterization of the pure compounds are presented in supplementary data.
Synthesis of Crude Reaction Mixture
To a suspension of compound
Production of Affinity Columns
Spherical porous diol silica particles (Kromasil, EKA Chemicals, Bohus, Sweden) with an average diameter of 5 µm, 300 Å pore size, and a surface area of 100 m2/g were used as HPLC support for the batch immobilization of Hsp90 protein. 10 The original storage buffer for Hsp90 interferes with the immobilization; hence, it was exchanged with immobilization buffer, 0.1 M sodium phosphate buffer pH 7.0 (PB). An Amicon Ultra-0.5 centrifugal filter with a molecular weight cutoff of 3k (Merck Millipore) was used for buffer exchange. Ultrafiltration by centrifugation at 14,000 × g (10 min) was repeated three times using immobilization buffer. The concentrate was collected by inversion of the filter device into a concentrate collection tube followed by centrifugation for 2 min (1,000 × g). The final concentration of the collected concentrate was measured using UV light at 280 nm (Nanodrop 1000; Thermo Scientific, Waltham, MA). For immobilization, diol silica was oxidized into aldehyde using periodic acid (100 µL; 100 mg/mL) by rotation at ambient temperature (21–24 °C) for 2 h. The aldehyde silica slurry was washed with (300 µL × four times) PB to remove any remaining unreacted periodic acid. The aldehyde silica was then mixed with the solution containing Hsp90 protein (84.30 µL; 9.96 mg/mL) to covalently bind the primary amines of the protein with the aldehyde group of diol silica. This resulted in formation of a Schiff’s base as an intermediate. As a last step, reductive amination was carried out with 100 mg/mL NaBH3CN, which was added as a reducing agent to the slurry to a final concentration of 10 mg/mL. Immobilization was conducted at room temperature (21–24 °C) by rotation of the protein-silica slurry for 68 h after addition of NaBH3CN to the slurry.
After immobilization, the reaction was stopped by centrifugation at 2,000 × g for 2 min followed by removal of the supernatant. The protein-silica slurry was carefully washed using PB (300 µL × five times). The eluate from the washings was collected, and the yield of bound Hsp90 was estimated using UV absorption at 280 nm of both the eluate and the initial protein solution. The silica with immobilized protein was packed into a stainless-steel column (50 × 0.5 mm) using an air-driven liquid pump (Haskel MS-110, Burbank, CA) at 340 bar packing pressure. Immobilization buffer was used as mobile phase for packing. A reference column with the same dimensions was packed with silica produced using the above immobilization method except for the use of protein. Columns were stored in PB at 4 °C when not in use.
Affinity Analysis on Single Quad LC/MS
Crude reaction mixture and the mixture containing pure compounds were analyzed in duplicates on the Hsp90 column and the reference column. The column compartment was maintained at 22 °C. Each mixture was analyzed for 30 min on the reference column and for 180 min on the Hsp90 column. Compounds were analyzed with an injection volume of 0.4 µL in isocratic mode at a flow rate of 15 µL/min. For the present study, AmAc (20 mM) pH 6.8 was used as mobile phase. Agilent 1260 infinity LC system (Agilent Technologies, Germany) combined with a diode array detector (DAD) and an Agilent 6130B single quadrupole MS detector was used for analysis. A wavelength of 254 nm (with the reference wavelength of 360 ± 4 nm) was used for UV detection. Compounds were ionized by API-ES with a nebulizer pressure of 25 psig in positive mode. A flow rate of N2 was 8 L/min at a temperature of 300 °C. The capillary and fragmentor voltage were set at 3000 V and 110 V, respectively, in positive mode. Chromatograms of individual compounds in mixtures were acquired by selected ion monitoring (SIM) mode, where m/z values were set corresponding to the mass ([M+H]+) of compounds present in the mixtures. The peak apex of extracted ion chromatograms (EIC) was used to measure the retention times. The Agilent OpenLab version A.02.01 (1.3.3) chromatography data system was used for data acquisition and analysis.
Compounds were analyzed with the final concentration of each component at approximately 50 µM or more (with 0.5% DMSO) and in the pure mixture as 50 µM (1.25% DMSO), respectively. Reference sample
Data Acquisition
The dissociation constant, KD (mM), of the inhibitors was based on their binding with Hsp90 and reference column and was estimated according to equation 1.
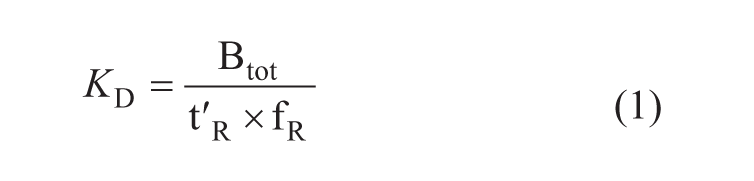
where Btot is the number of binding sites on the Hsp90 affinity column (nmol), fR is the flow rate (µL/min), and
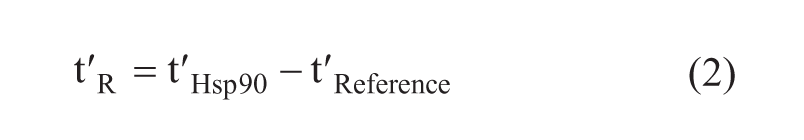

where
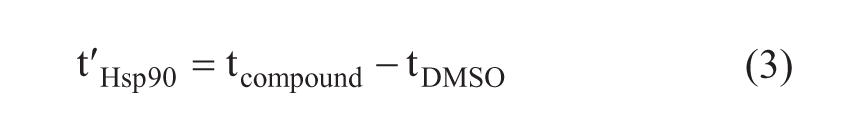
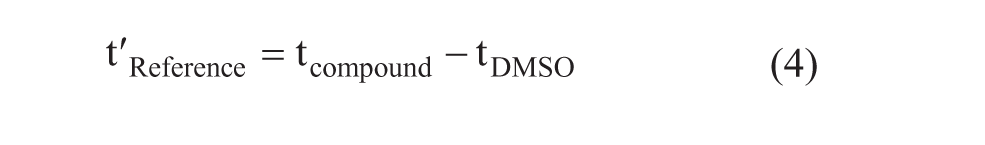
Equation 1 is valid under linear isotherm conditions when the concentration of the compound (Ca) is low compared with the KD value of the analyte (Ca << KD). Btot in the column was estimated to be 5 nmol, which is 50% of the immobilized protein content. This is a reasonable approximation based on other affinity-LC/MS studies conducted with various target proteins. 14
Results and Discussion
We selected 2-amino-6-chloropurine BII021 analogues because a one-pot reaction protocol combining 2-amino-6-chloropurine (
Five target compounds were synthesized in a single mixture and also prepared as individual compounds ( Table 1 ; Fig. 1 ). No purification was performed for the crude mixture synthesis. The resulting solid was determined to have the five desired products present in a ratio of 12:24:19:29:7. It is important to note that products do not form uniformly because of different rates of reaction of different benzyl bromides. This is typical of the organic synthesis of mixtures and presents significant problems for analysis. However, using the affinity-LC/MS method, even highly disproportionate mixtures can be analyzed with ease. Following the isolation of the crude product mixture, a mixture of pure compounds was prepared for comparison. In this latter mixture, the concentration of the individual compounds was approximately matched with the corresponding compounds in the reaction mixture.
Affinity-LC/MS was primarily performed on a single quad LC/MS system. Some additional analyses were also performed on a quadrupole time-of-flight LC/MS system in which the objective was to confirm the affinity values obtained with the single quad LC/MS (data not shown). In this study, a single quad MS was sufficient as the number of components was limited and they have unique masses. With more complex synthetic mixtures, a higher-resolution MS would be preferred. The apparent dissociation constant, KD (µM), of the compounds was estimated, as described in the Data Acquisition section.
Affinity-LC/MS was able to detect and measure the individual affinities of the compounds in the crude reaction mixture in the range of KD from 48 to 147 µM. When compared with the mixture of pure individually synthesized compounds, the affinities corresponded well (correlation coefficient [r] = 0.98; p = 0.0029 [significant]) with a mean coefficient of variation (CV) of 7.5% ( Table 1 ). The small discrepancies between compounds in mixtures and individuals could be attributed to, apart from experimental variation, variation in retention due to nonlinear binding behavior.
Individually purified compounds
Affinity-LC/MS allows for the monitoring of m/z of most compounds, at a wide range of differing concentrations of each analyte, which enables analysis of all the components of a molecular mixture and their binding to an immobilized target. The typical affinity range that can be directly measured with affinity-LC/MS is in the low mM to 0.1 µM range, depending on the amount of active target. Tighter binders in the nM range elute very late and may avoid detection because of extensive dilution of sample. If they are present during analysis, they can be seen by forced elution (step or gradient) by means of changing the conditions of elution such as of pH, temperature, or ionic strength. One serious consideration is that the immobilized target can withstand any such changes in elution conditions.
Furthermore, the affinity-LC/MS platform possesses the potential to monitor the reactants of an organic synthesis mixture including any contaminants or reagents and to monitor side reactions by the sole use of their molecular weights and fragmentation patterns. In this study, we were able to follow closely the reactant
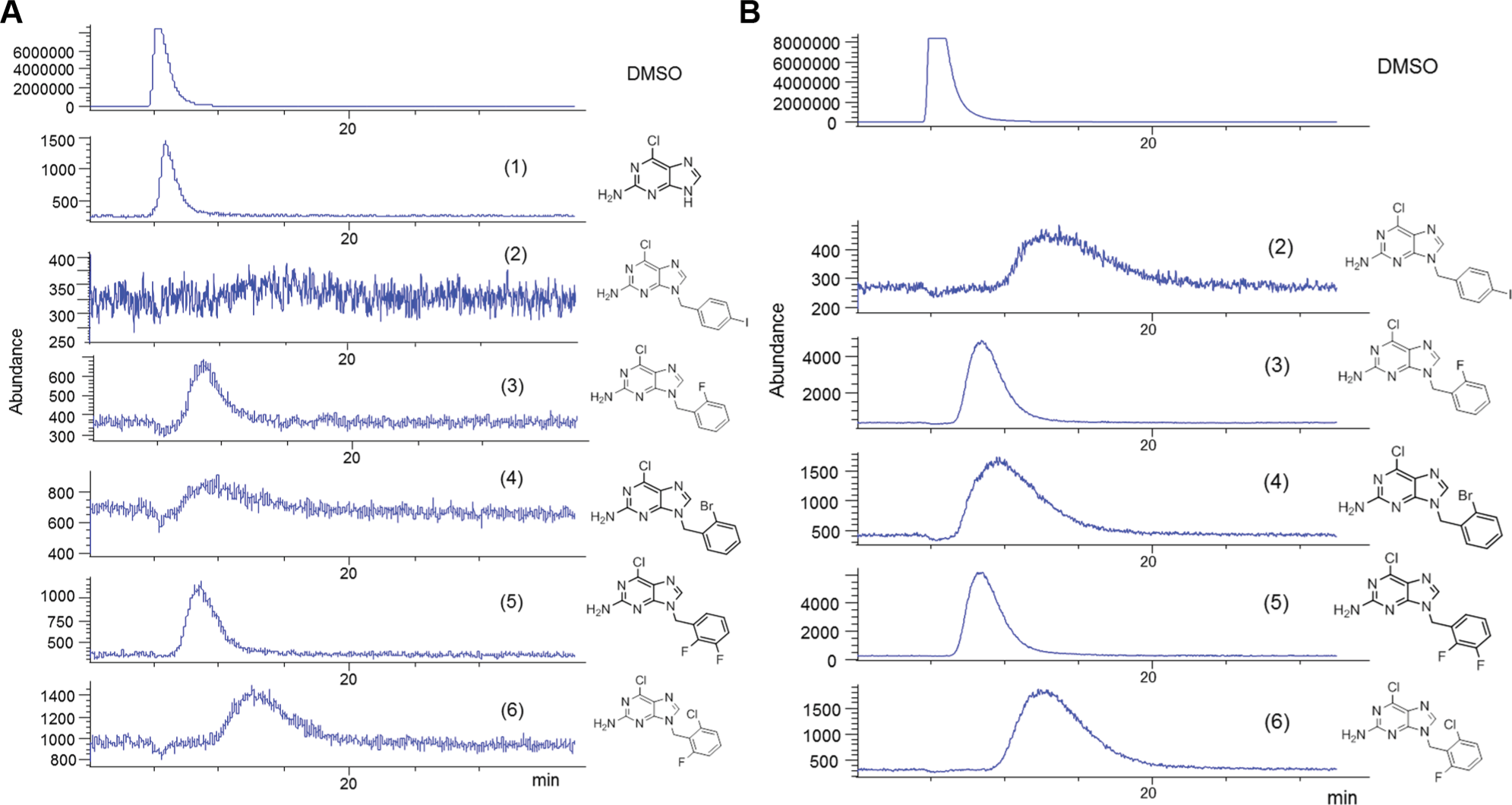
(
Compound
These studies demonstrate that LC/MS can identify many components of a mixture, although they are present at different concentrations and have different ionization potentials for mass detection. More complex mixtures can also be analyzed with this method if there is a high surplus of target that will allow the binders to bind in a linear fashion and virtually unaffected by competition. In a previous study with affinity-LC/MS, Duong-Thi et al. 10 evaluated successfully the binding of fragment mixtures (mixtures of pure compounds) with up to 65 compounds simultaneously to the same target.
Given the challenges and costs of screening in the chemical and biological sciences, we believe that affinity-LC/MS offers a cost-effective, sensitive, and broadly applicable screening method available to most laboratories with access to an entry-level LC/MS machine. Major cost savings will be evident in the early stages of affinity-LC/MS screening of mixtures as purification is no longer required. This approach is more straightforward than other methods for characterization of binding, such as the use of bioassays.
In summary, we have in this study demonstrated the potential of affinity-LC/MS technology to identify and quantify the weak interactions between the components of an organic synthesis mixture and protein target without any need for purification. Of special importance is that the methodology allows a scan of a broad spectrum of binding interactions ranging from mM to sub-µM including strong binding (nM range), and it is not sensitive to variations in the concentrations of the mixture components. Apart from the application of affinity-LC/MS in organic synthesis and combinatorial libraries, it will also be of interest to natural sources of mixtures such as biological extracts. We expect affinity-LC/MS technology will be a valuable complement to procedures such as yeast-2-hybrid systems and affinity purification-MS in proteomics as it will also include the complete analysis of transient interactions among proteins. It remains to be seen how applicable affinity-LC/MS will be for highly complex mixtures and to fully understand all their interactions to a target. Future studies on this subject will determine the potential of this technology. However, this initial work, as presented here, has clearly demonstrated the potential of affinity-LC/MS at least for simple but still crude synthetic organic mixtures.
Footnotes
Acknowledgements
The authors wish to thank the Singapore Universities, Nanyang Technological University and National University of Singapore, for giving us the opportunity and the support to conduct this research.
Supplementary material is available online with this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by NMRC CBRG grant R-148-000-189-511 and AcRF TIER1: RG 31/15.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
