Abstract
Anthraquinone dyes are extensively used in the textile, cosmetic, and food industries because of their bright colors, stability toward chemicals, and resistance against fading. Yet their large-scale application has resulted in great environmental problems. Anthraquinone dyes are extremely recalcitrant, that is, resistant to natural degradation processes, and have toxic impacts on aquatic life and human health. Traditional dye removal methods like adsorption and chemical oxidation tend to be inefficient and expensive, which has necessitated novel approaches. Enzymatic bioremediation is a novel method that has gained popularity due to its sustainability and high catalytic efficiency. This research examines the potential of two enzymes from Flavobacterium sp.9AF—Putative Multifunctional Dye Peroxidase DyP2 and Glycoside Hydrolase Family 5 Protein—to perform degradation through in silico approaches. Molecular docking showed that DyP2 has greater binding affinities for Solvent Green 3 (−12.1 kcal/mol), Vat Blue 6 (−11.4 kcal/mol), and Reactive Blue 19 (−11.3 kcal/mol) than Glycoside Hydrolase Family 5 Protein. Interaction analysis showed that DyP2 has stronger hydrogen and hydrophobic bonds with pollutants, indicating its better catalytic ability for dye degradation. These results present DyP2 as an attractive biocatalyst for remediation of anthraquinone dye, which has potential for enzyme-based applications in industrial wastewater treatment and conservation.
Introduction
Anthraquinone dyes are one of the most recalcitrant dyes used in industry, attributed to the fact that their fused aromatic ring systems tend to resist natural degradation processes. 1 Their widespread use has raised immense environmental issues. When released into water bodies without treatment, anthraquinone dyes are capable of lowering sunlight penetration, inhibiting photosynthesis, and lowering the level of dissolved oxygen, thus causing damage to aquatic ecosystems. 2 They are also toxic to aquatic life and carcinogenic and mutagenic to humans and animals. 3
Enzymatic anthraquinone dye degradation is mainly through the action of oxidoreductases like dye-decolorizing peroxidases (DyPs) and laccases. These enzymes catalyze the degradation of the dye molecules by acting on their chromophore groups. DyPs, for example, break the conjugated dye bonds by oxidation reactions, causing the decolorization of the dye and further degradation into smaller aromatic compounds. 4 These intermediates are also metabolized to simpler molecules such as carbon dioxide and water in aerobic conditions. In the same way, glycoside hydrolases can break down glycosidic bonds of dye substances or their derivatives, thereby degrading them. 5 Their catalytic efficiency has made them good candidates for use in bioremediation processes. Even with progress in enzymatic bioremediation, issues persist in knowing the molecular processes behind enzyme–dye interactions as well as in improving enzyme operation for use in industry. Most of the research has been conducted on model dyes or straightforward systems, and there are knowledge gaps regarding the degradation pathways of intricate anthraquinone dyes. 6
In this study, two enzymes from Flavobacterium sp. 9AF—Putative Multifunctional Dye Peroxidase (DyP2) and Glycoside Hydrolase Family 5 Protein—were examined using computational tools to assess their potential for anthraquinone dye degradation. Ligand-binding sites were modeled by Prank-Web software, showing stable catalytic pockets in both enzymes. 7 Molecular docking assays performed with CB-Dock evaluated the binding affinities of these enzymes toward 13 industrial anthraquinone contaminants. 8 DyP2 showed higher binding affinities for dyes such as Solvent Green 3 (−12.1 kcal/mol), Reactive Blue 19 (−11.3 kcal/mol), and Vat Blue 6 (−11.4 kcal/mol). Glycoside Hydrolase Family 5 Protein also demonstrated good binding against some dyes but was comparatively weaker in action than DyP2. The higher binding affinity of DyP2 could be ascribed significantly to its hydrophobic interactions, specifically with the aromatic residues such as Tyr, Phe, and Trp. These residues offer van der Waals interactions as well as π–π stacking with the aromatic rings in Solvent Green 3. However, hydrogen bonding also contributes to the protein–ligand complex stability. Toxicity profiling on the CSM-toxin platform re-asserted that the two enzymes were non-toxic and environment friendly for use. 9 Binding behavior was also analyzed through Discovery Studio to understand enzyme–substrate mechanisms at the molecular level. 10 DyP2 primarily established hydrogen and hydrophobic interactions with impurities such as Solvent Green 3, and whereas Glycoside Hydrolase Family 5 Protein exhibited comparable interaction trends with Vat Blue 6. 11
The present study seeks to investigate the biocatalytic potential of Flavobacterium sp. enzymes for the degradation of recalcitrant anthraquinone dyes, gaining insights into enzyme–dye interactions at the molecular level. Incorporating machine learning models with in-silico-based tools represents the ability to predicted new dye or pollutant degradation pathways. The combination of known degradation mechanisms and ML models can introduce potential methods rather than just molecular docking. In addition to these, ML models based on molecular characteristics such as binding energies, hydrophobic interactions etc. can aid in catalytic site identification, enzyme–dye interactions and mutational effect anticipation. Future research ought to aim toward experimental verification of computational results through in vitro enzymatic assays, enzyme activity optimization under industrial settings (e.g. fluctuating pH and temperature), immobilized enzyme system construction for large-scale wastewater treatment, and investigation into synergistic relationships between two or more enzymes or microbial consortia for increased dye degradation. 12 The introduction of immobilized enzyme systems for large-scale treatments brings several challenges such as pH shifts, thermal instability and diffusional limitations. By tackling these factors, enzymatic bioremediation may become a possible solution for alleviating industrial dye contamination, providing a sustainable remedy for conventional remediation techniques. 13 Flavobacterium sp. 9AF was selected based on prior genomic annotations indicating the presence of oxidoreductase and hydrolase enzymes potentially involved in xenobiotic degradation pathways. This study contributes novel insight by focusing on Flavobacterium sp. 9AF DyP2, an underexplored bacterial source, for its potential in anthraquinone dye degradation, while most previous studies emphasize fungal DyPs or model bacterial systems. Moreover, we systematically compared DyP2 with Glycoside Hydrolase Family 5 Protein using a multi-parameter in silico pipeline including toxicity profiling, ligand pocket prediction, and comparative docking with structurally diverse industrial dyes integrating data absent in prior studies. This study offers novelty by exploring Flavobacterium sp. 9AF, an underexplored bacterial source, for anthraquinone dye degradation using a multi-parameter in silico approach. The comparative evaluation of DyP2 and GH5 enzymes and integration of structural and toxicity data sets a foundation for subsequent experimental work.
Methodology
Enzyme Selection and Retrieval of Protein
Two specific enzymes were obtained from Flavobacterium sp. 9AF in order to examine the potential of bacterial enzymes degrading anthraquinone dyes. In this study, AlphaFold is employed to determine the three-dimensional structure of the target enzymes, facilitating structural and functional analysis. 14
DyP2 was annotated based on UniProt (ID A0A653TX29) and NCBI (accession no. VXB84822.1) entries, which classify the protein within the DyP-type peroxidase family based on conserved sequence motifs and domain structures. These include the distal His–Arg catalytic pair and GXXDG motif. AlphaFold structural predictions further confirmed a canonical DyP fold, validating its functional annotation as a putative dye-degrading peroxidase.
Additionally, BLASTp alignment was performed against the NCBI non-redundant protein database to verify homology with known dye-degrading enzymes. The alignment confirmed the close similarity of DyP2 to bacterial DyP-type peroxidases, justifying its selection for in silico studies.
Physiochemical Properties of Protein
The ExpaSy-ProtParam tool (https://web.expasy.org/protparam) was employed to analyze the sequence’s physicochemical properties. In this study, the tool was used to evaluate key parameters, including molecular weight, theoretical pI, instability index, and the Grand Average of Hydropathicity (GRAVY). By inserting the sequences in FASTA format, these properties were computed. 15
Protein Toxicity Assessment
CSM-toxin (https://biosig.lab.uq.edu.au/csm_toxin/) was used to assess the toxicity associated with the Dye Peroxidase DyP2 and Glycoside Hydrolase Family 5 Protein. CSM-toxin, an in silico protein toxicity identifier, requires the protein’s primary sequence in FASTA format. This server utilizes structural bioinformatics and machine learning methods to classify an enzyme as either toxic or non-toxic by comparing it with the known toxic proteins. The enhanced classification accuracy feature of CSM-toxin makes it a valuable toxicity screening website. CSM-toxin is a peer-reviewed platform using machine learning and structural bioinformatics for toxicity classification. It is validated against curated datasets and has been used in environmental and pharmaceutical contexts.
Retrieval of Pollutants
Pollutants containing an anthraquinone structure were downloaded from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) in structural data files (SDF) format for in silico analyses. PubChem is one of the large open-access databases and provides detailed knowledge on chemical molecules including structures, molecular weight, formula, and their biological activities. Vast and accurate molecular data can be collected from this repository for various research studies. PubChem structures may lack optimized geometries. To mitigate this, all compounds were energy-minimized before docking using Discovery Studio. The dyes were selected based on high usage frequency in the textile and food industries and PubChem annotations of persistence or bioaccumulation risk. The selected dyes cover a broad range of molecular weights, aromatic substitutions, and functional groups, reflecting real-world structural diversity. All ligands were energy-minimized using Discovery Studio with MMFF94 force field to ensure realistic conformations. Docking scores were generated using the same CB-Dock scoring protocol and cavity predictions, ensuring consistency. Top poses were further analyzed using Discovery Studio for validation of interaction geometry and plausibility.
Ligand Binding Site Prediction
Prank-Web (https://prankweb.cz/) was performed to determine the ligand binding pockets of the peroxidase and glycoside hydrolase enzymes. The PDB structures of each enzyme were uploaded on the server that permitted an in-depth in silico assessment of catalytic sites. Prank-Web is a new version of the P2Rank tool that employs techniques like machine learning to enhance its efficacy of identification procedure. It can accept 3D structures, PDB IDs, or UNIPROT-IDs and computes evolutionary conservation scores to predict the functionally conserved regions across homologous proteins. 16 Prank-Web uses evolutionary conservation and machine learning to predict pockets and has demonstrated high reliability across bacterial systems in recent benchmarks.
Comparative Molecular Docking of Proteins and Ligands
The molecular docking study used the CB-Dock tool to investigate interactions of anthraquinone dyes with two Flavobacterium sp. 9AF enzymes: Putative Multifunctional Dye Peroxidase DyP2 and Glycoside Hydrolase Family 5 Protein. The PDB structure of DyP2 was initially loaded into the CB-Dock tool, and then sequential docking protocols were performed with each anthraquinone dye ligand. The same protocol was used for the Glycoside Hydrolase Family 5 Protein, with each ligand treated separately. Binding affinities and docking scores were calculated for all protein–ligand complexes. The three highest binding affinities and docking scores were used for each group of enzymes to allow a comparative study of interaction patterns in the two enzymatic systems. 3 CB-Dock was selected for its blind docking capability, integrated cavity prediction, and automated scoring, which streamline multi-ligand comparison across unknown binding sites. CB-Dock’s default exhaustiveness and clustering settings were used to ensure reproducibility. The top poses were consistently re-evaluated in Discovery Studio for validation. Key residues were identified by visual inspection of hydrogen bonds, π–π stacking, and hydrophobic contacts in 2D/3D diagrams generated in Discovery Studio’s Ligand Interaction module.
Comparative Molecular Interaction Analysis
Interaction analysis was done using Discovery Studio to visualize the residues, bond distances their types and categories of the best of three docked complexes of both enzymes of Flavobacterium sp. 9AF. It is one of the most extensively used open-source software for high-quality 3D. 4
Results
Enzyme Selection and Retrieval of Protein
Alphafold was utilized for the retrieval of the three-dimensional structure of Glycoside Hydrolase Family 5 Protein (UniProt ID: A0A653W2V4) and Putative Multifunctional Dye Peroxidase DyP2 (UniProt ID: A0A653TX29. Putative Multifunctional Dye Peroxidase DyP2 consists of 429 amino acids whereas Glycoside Hydrolase Family 5 Protein consists of 315 amino acids (Figures 1 and 2).
The 3D structure of Putative Multifunctional Dye Peroxidase DyP2 retrieved from AlphaFold.
The 3D structure of Glycoside Hydrolase Family 5 Protein retrieved from AlphaFold.
The three-dimensional structures were predicted using AlphaFold with high model confidence. AlphaFold-predicted structures typically achieve over 90% residues in favored regions for proteins with known folds, such as DyPs and GH5 enzymes.
Physiochemical Properties of Proteins
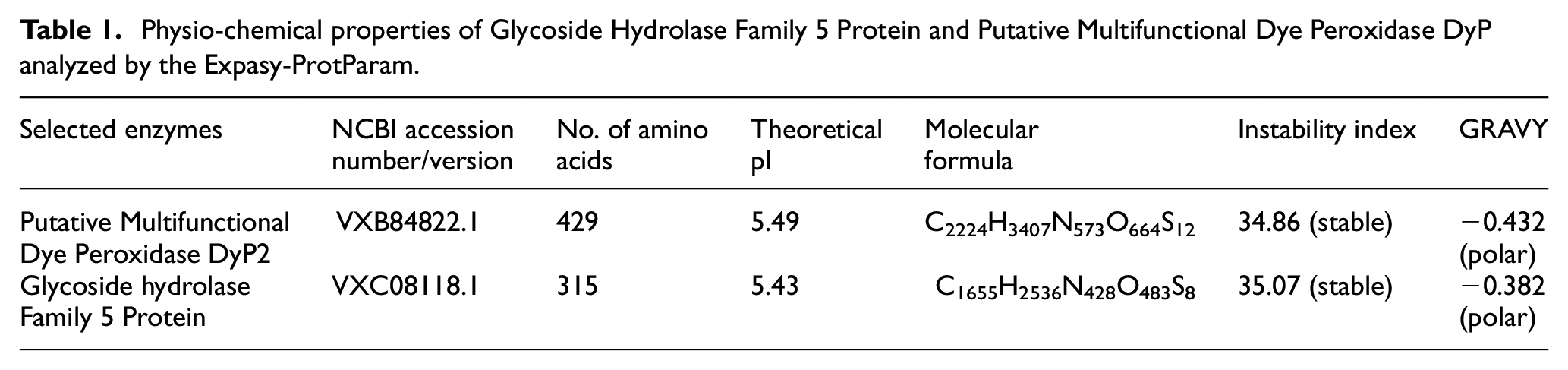
The physiochemical properties of the enzymes were analyzed and the result shows that enzymes are stable. The instability index of Putative Multifunctional Dye Peroxidase DyP2 is 34.86 and Glycoside Hydrolase Family 5 Protein is 35.07, both are less than 40, indicating its higher stability. The score of GRAVY was determined for both enzymes as −0.432 and −0.382, both of them are polar because negative GRAVY values suggest a hydrophilic nature. The lower GRAVY scores of both enzymes reveal improved high protein hydrophilicity that can facilitate with the enhanced hydrogen bonding and electrostatic interaction of DyP2 and the Glycoside Hydrolase Family 5 Protein. Table 1 provides the results of each of these outcomes.
Physio-chemical properties of Glycoside Hydrolase Family 5 Protein and Putative Multifunctional Dye Peroxidase DyP analyzed by the Expasy-ProtParam.

Protein Toxicity Assessment
The CSM-toxin aids in analyzing whether the provided protein is safe and non-lethal on application across various areas or not. Peroxidase and glycoside hydrolase were predicted to be non-toxic with zero probability. Moreover, it also provided insights into the physiochemical properties of each enzyme including amino acid composition, length, molecular weight, net charge, and hydrophobic values. The hydrophobic values of peroxidase (−0.43) and hydrolase (−0.38) revealed the hydrophilic nature of these enzymes.
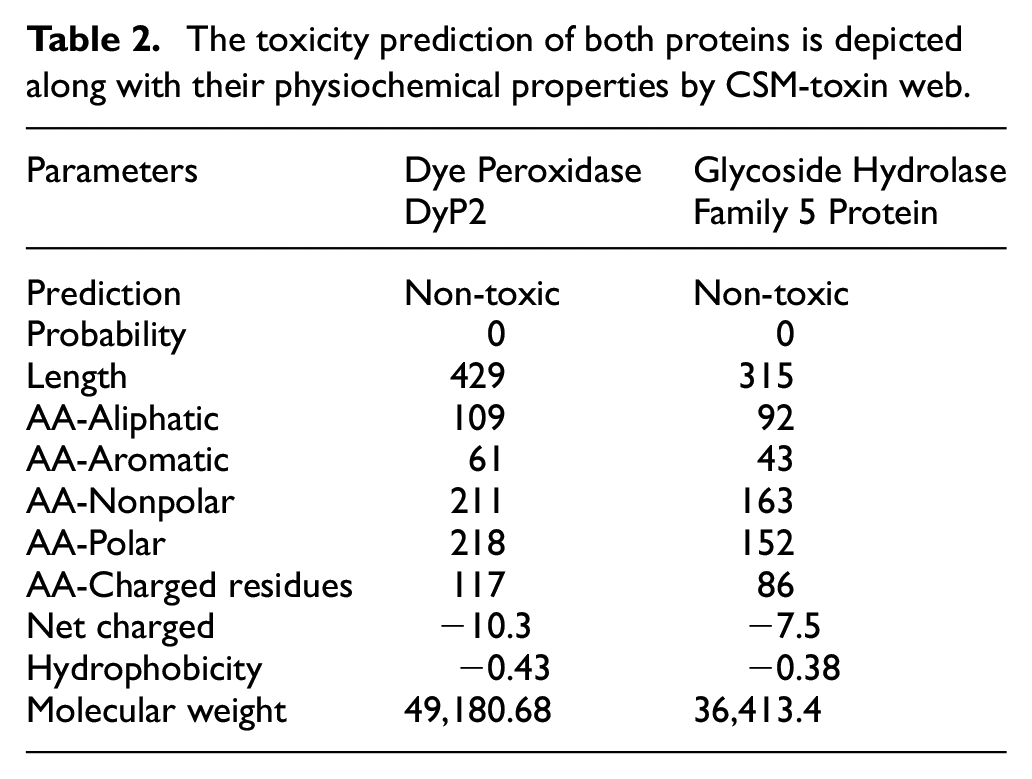
The net negative charge on the enzymes (DyP2: −10.3 and GH5: −7.5) have been reported to play an essential role in interfering with substrate selectivity process with the aid of electrostatic pre-orientation activities. Negatively charged surfaces of the enzymes electrostatically attract the positively charged substrates (i.e. dyes) offering orientation of substrates to the key catalytic sites. On the other hand, pocket depth or residue composition define the catalytic chemistry of an enzyme and a ligand, and the electrostatic nature of the enzyme surface is crucial for initial specificity and binding. Table 2 provides the information on the results obtained for both proteins.
The toxicity prediction of both proteins is depicted along with their physiochemical properties by CSM-toxin web.
Retrieval of Pollutants
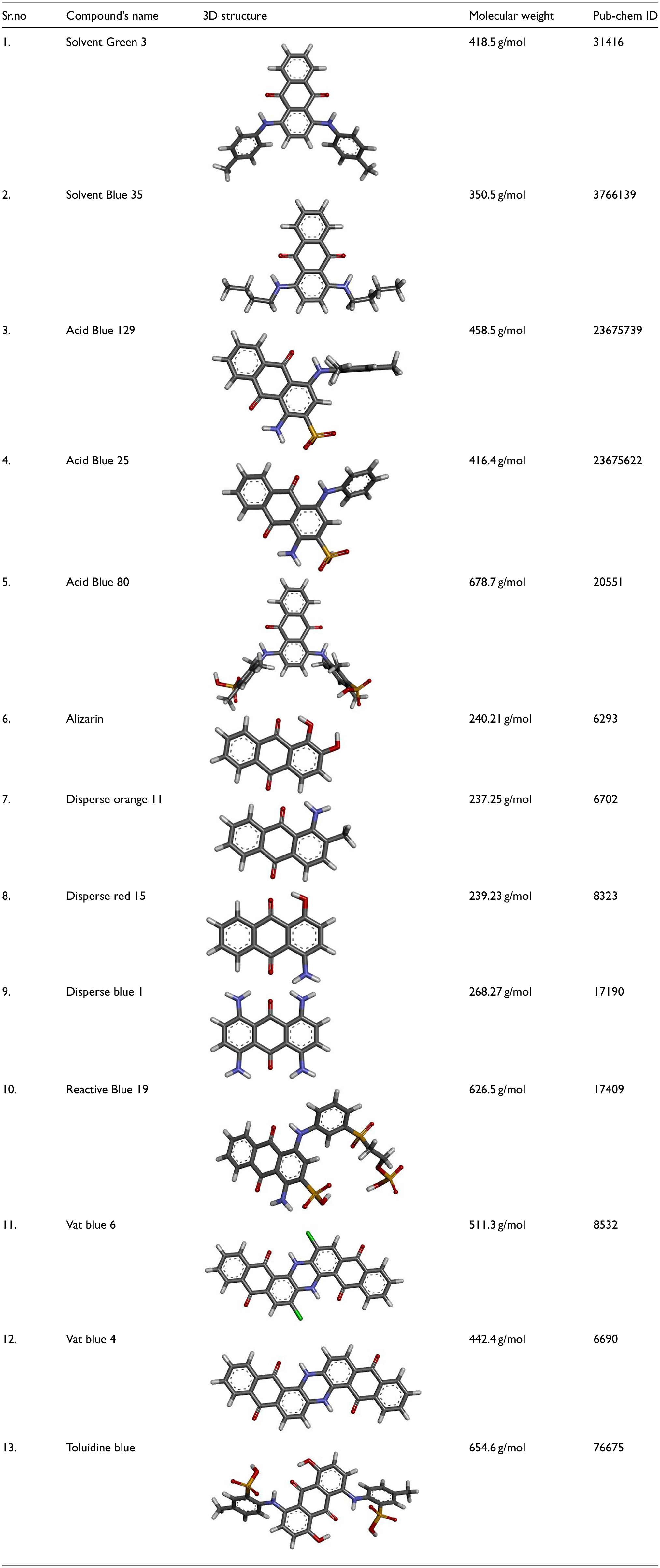
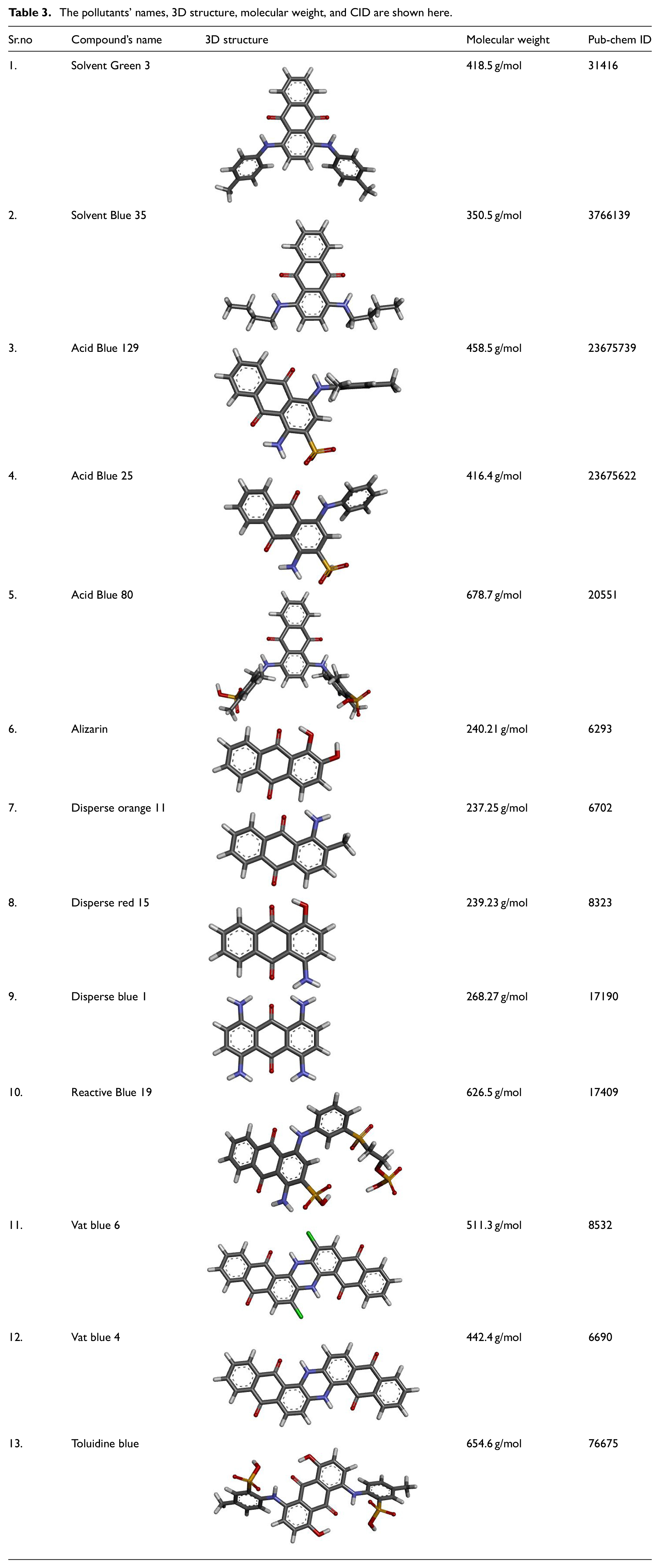
The selected 13 pollutants are widely used in the food, textile, and cosmetic industries. The SDF structures were converted to PDB format as shown in Table 3, using Discovery BIOVIA studio. These chemical molecules will be subjected to docking with peroxidase and hydrolase to determine the potential degrading ability of both enzymes. The higher molecular weight dyes such as Acid Blue 80 (678.7 g/mol) and Toluidine Blue (654.6 g/mol) have limited penetration to the catalytic sites of the enzymes, that is, DyP2 and GH5, due to the resulting steric hindrance. Such dyes possess higher surface complexity, molecular volume and conformational rigidity that leads to incorrect spatial alignment of the main reactive units of the dyes to the active sites of the enzymes.
The pollutants’ names, 3D structure, molecular weight, and CID are shown here.
Ligand Binding Site Prediction
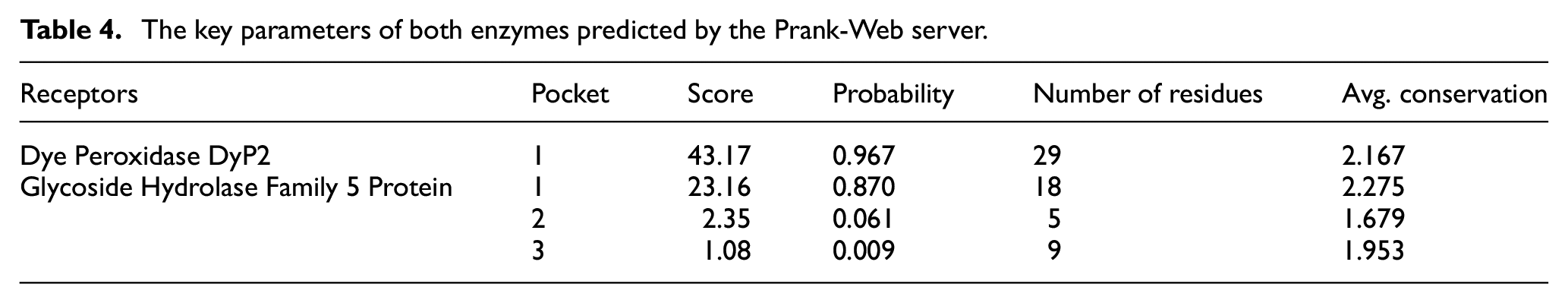
Prank-Web identified one binding pocket in peroxidase and three in glycoside hydrolase, each with their relative confidence scores. The high confidence score (43.17) and binding probability (0.97) of peroxidase’s binding pocket suggest it to be the most functionally stable site for ligand interactions. On the other hand, the primary pocket (1) is a highly functional and conserved binding region of the hydrolase enzyme with a 23.16 score and 0.87 probability as compared with pockets 2 and 3 (Figure 3). The confidence score and binding pocket probabilities of both enzymes show their robustness and substrate-binding potential, which in turn are essential for performance in the challenging environmental situations. Prank-Web reports a confidence score of 43.17 and a 0.967 probability for DyP2’s pocket, indicating strong predictive reliability. These values are consistent with known catalytic pocket thresholds (>0.9). Table 4 highlights the score, probability, residue number, and average conservation of each pocket, defining the structural properties of each enzyme.
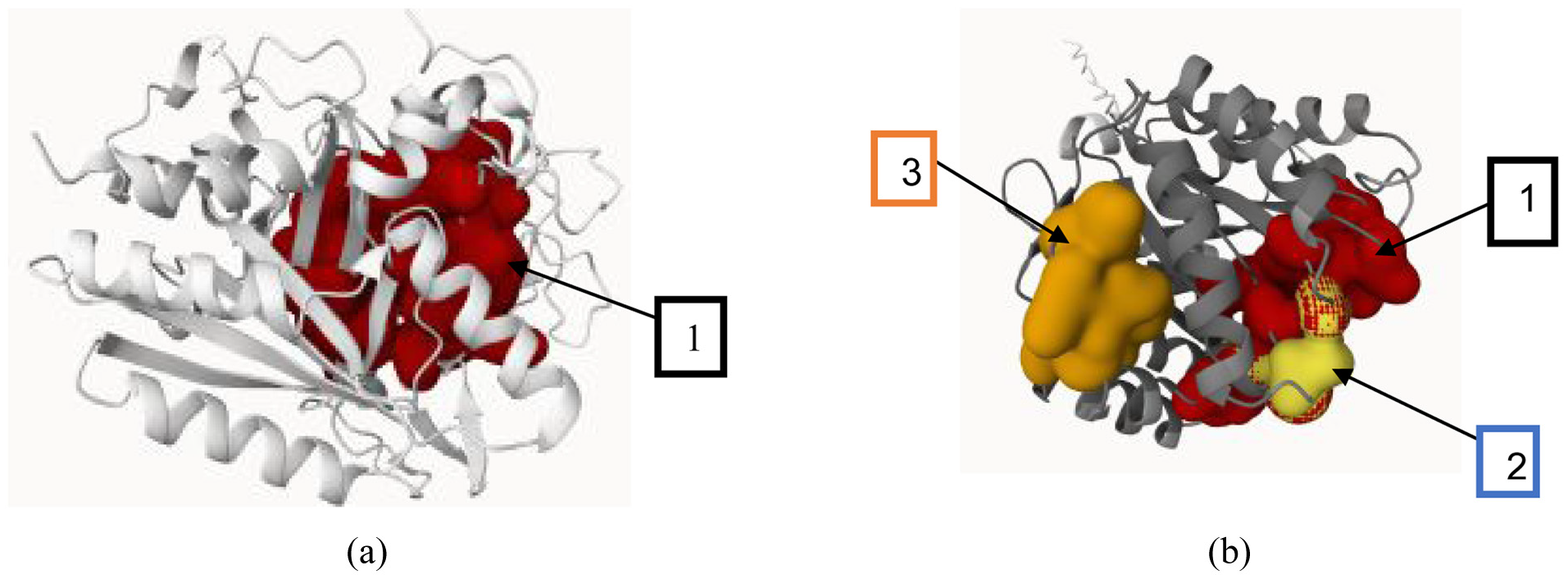
Prank-web predicts one binding site in peroxidase (a) and three distinct colored sites in glycoside peroxidase (b).
The key parameters of both enzymes predicted by the Prank-Web server.
Comparative Molecular Docking
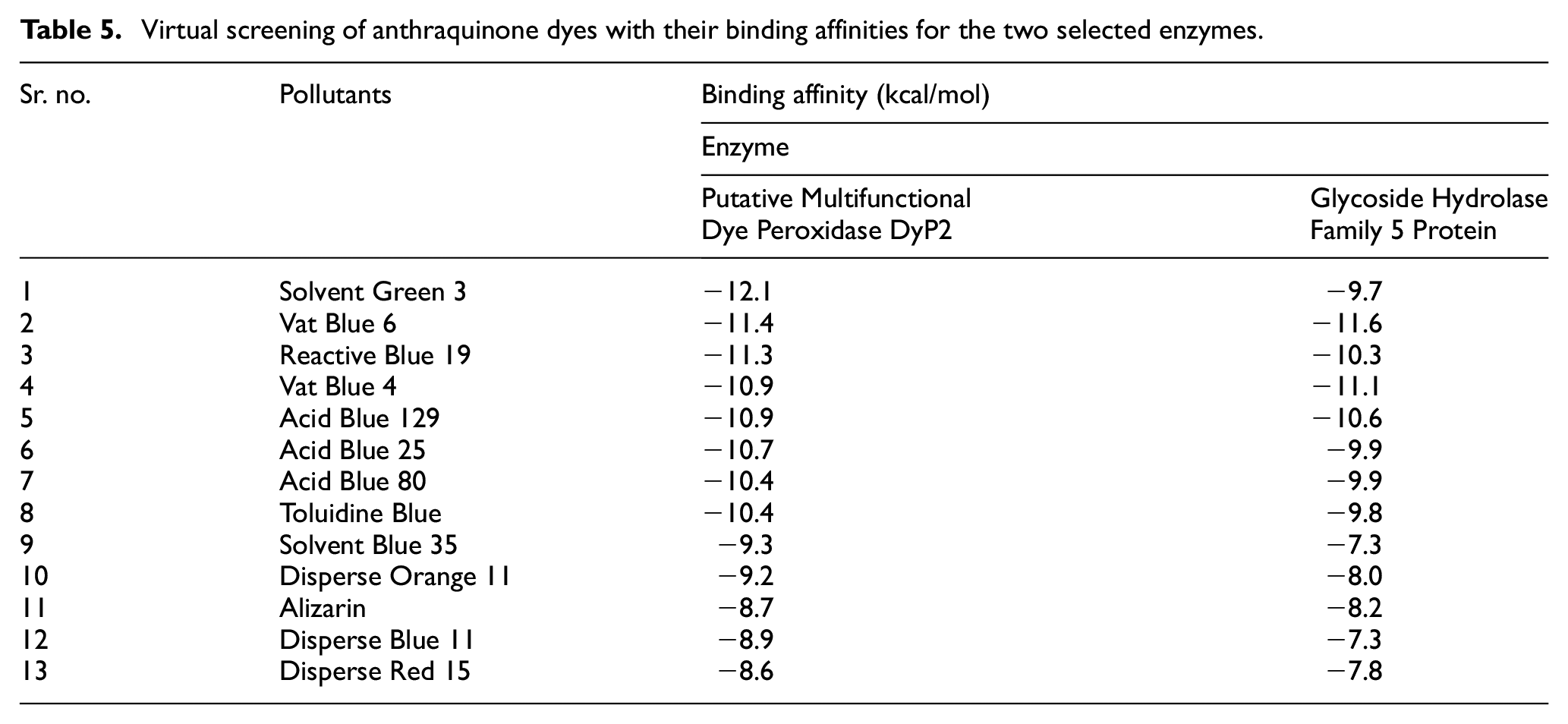
Comparative molecular docking was performed between two enzymes, Putative Multifunctional Dye Peroxidase DyP2 and Glycoside Hydrolase Family 5 Protein, and 13 pollutants through the CB-Dock tool. In the case of DyP2 docking with pollutants, the best binding affinity was found to be with Solvent Green 3 (−12.1 kcal/mol), Vat Blue 6 (−11.4 kcal/mol), and Reactive Blue 19 (−11.3 kcal/mol). The lowest energy was found to be −8.6 kcal/mol with Disperse Red 15. Regarding the Glycoside Hydrolase Family 5 Protein, the highest energies were shown in Vat Blue 6 (−11.6 kcal/mol), Vat Blue 4 (−11.1 kcal/mol) and Acid Blue 129 (−10.6 kcal/mol). The presence of abundant conventional hydrogen bonds between DyP2 and hydrolase with the first four ligands including Acid-Blue 129 indicates defined substrate orientation that increases the catalytic efficiency of each enzyme. The ligand orientation optimizes proximity to the catalytic elements which maximizes reaction kinetics and electron flow. The highest energies of both enzymes show that both enzymes of Flavobacterium sp. 9AF have the ability to degrade anthraquinone dyes. But Putative Multifunctional Dye Peroxidase DyP2 has a greater ability to degrade pollutants as compared to other Glycoside Hydrolase Family 5 Proteins. Binding affinities in the range of −10 to −13 kcal/mol are typical for strong enzyme–ligand interactions and consistent with experimental degradative DyP systems. Docked poses aligned well with the highest-confidence predicted pockets, particularly in DyP2, confirming structural congruence. No major deviations were noted. Table 5 shows the binding affinities determined by the CB-Dock between enzymes and pollutants. The observed hydrogen bonds (1.6–3.3 Å) and hydrophobic contacts (<5.5 Å) fall within stable physiological ranges, indicating likely enzyme–substrate complex stability.
Virtual screening of anthraquinone dyes with their binding affinities for the two selected enzymes.
Comparative Molecular Interaction Analysis
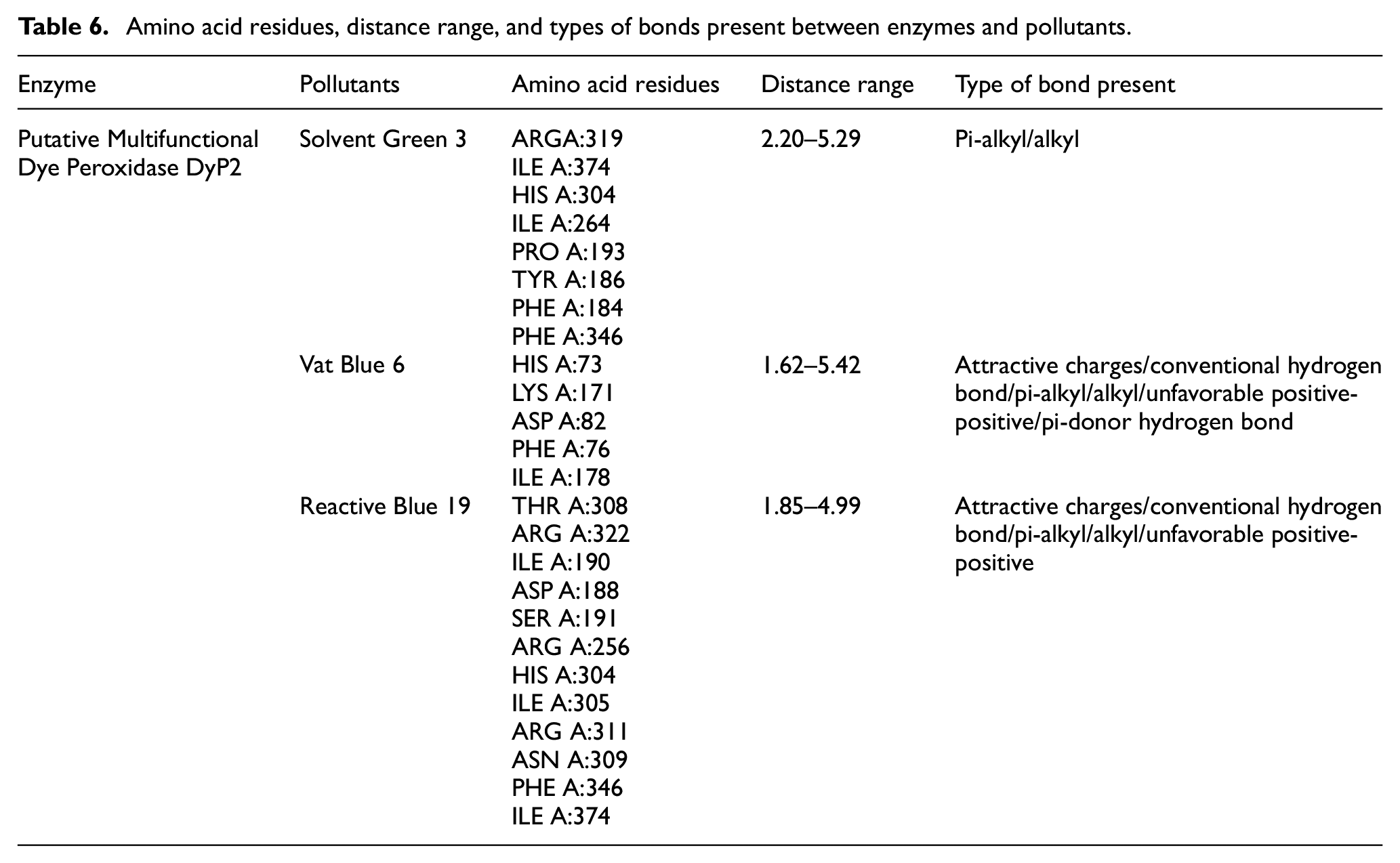
The molecular interactions between the enzymes Putative Multifunctional Dye Peroxidase DyP2 and Glycoside Hydrolase Family 5 Protein with their respective best three docked pollutant models were analyzed using Discovery Studio (Table 6). Interactions were analyzed for the top three complexes per enzyme, based on highest binding affinities. While molecular docking is a common approach, our study uniquely applies it to compare two native enzymes from Flavobacterium sp. 9AF for the same pollutant group. This strategic use of in silico tools provides comparative insights to prioritize candidates for in vitro validation. For DyP2, the interactions with Solvent Green 3, Vat Blue 6, and Reactive Blue 19 were examined. Solvent Green 3 formed two conventional hydrogen bonds (2.20–2.36 Å) and several alkyl and pi-alkyl bonds (3.88–5.29 Å). Vat Blue 6 showed conventional hydrogen bonds with HIS A:73 and ASP A:82 residues (1.62–3.12 Å) and alkyl/pi-alkyl bonds (3.96–5.24 Å). Reactive Blue 19 primarily formed conventional hydrogen bonds (1.85–3.28 Å) and alkyl/pi-alkyl bonds (3.98–4.88 Å), with an attractive charge interaction at ASP A:188 (4.70 Å). However, if targeted mutagenesis is applied on non-polar residues such as Ile 305, and Phe 346 of DyP2 could improve the hydrophobic interactions with the aromatic residues of Reactive Blue 19. Such a mutation could also enhance the stability of enzyme–ligand complexes and the degradation rate of pollutants like Reactive Blue 19. The unfavorable positive–positive clashes are majorly due to the electrostatic repulsions that reduce the enzyme’s ability to catalyze the pollutant. As electrostatic repulsions destabilizes the enzyme–substate complex structure and displaces the substrate from its accurate position. These interactions highlight the diverse bonding patterns involved in enzyme–substrate interactions, which are crucial for understanding the degradation mechanisms of these pollutants (Figure 4(a)–(c)).
Amino acid residues, distance range, and types of bonds present between enzymes and pollutants.
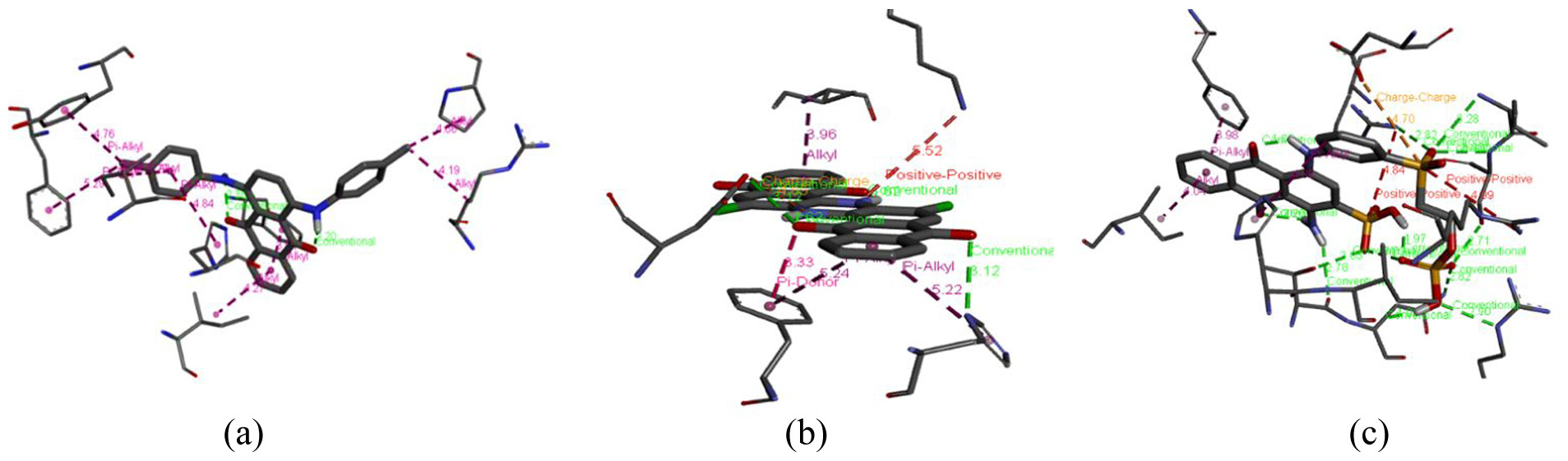
Interaction analysis of Putative Multifunctional Dye Peroxidase DyP2 with (a) Solvent Green 3, (b) Vat Blue 6 and (c) Reactive blue 19 pollutants along with their bond types and distances between them.
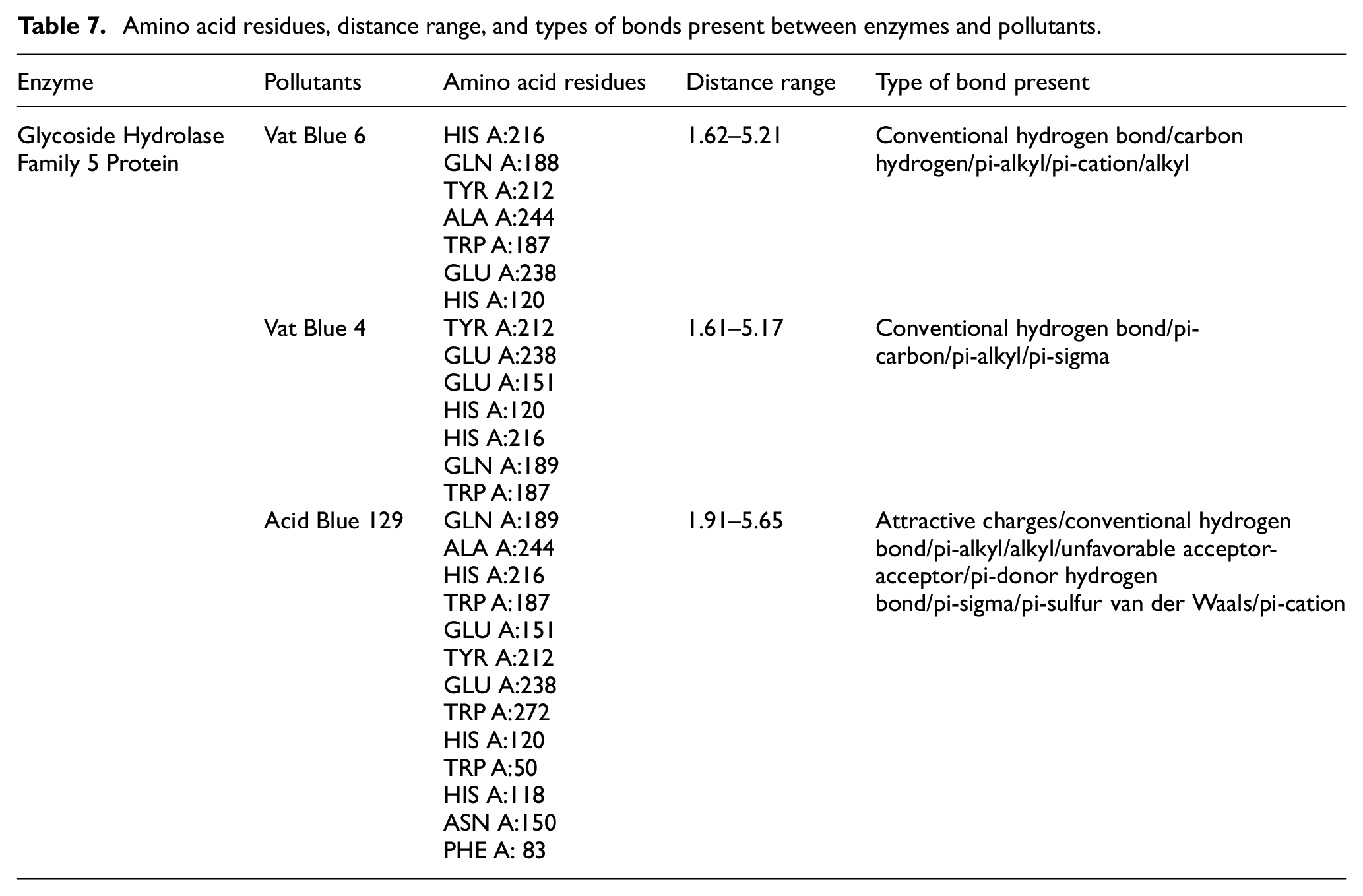
The interaction analysis of the enzyme Glycoside Hydrolase Family 5 Protein with the best three docked complexes was performed sequentially. In the case of Vat Blue 6, interaction with the enzyme shows that five conventional hydrogen bonds were formed with distances ranging from 1.62 to 3.34 Å. Only one alkyl and four Pi alkyl bonds were formed with the distances 4.07 Å, and 4.60–5.21 Å respectively. In the case of Vat Blue 4, seven conventional hydrogen bond interactions were predicted within distance ranges from 1.61 to 3.33 Å. Only one pi-alkyl and one pi-sigma bond is formed with distances of 5.17 Å and 3.66 Å respectively. The pi-cation interaction shows distance ranges from 4.00 to 4.22 Å. In Acid Blue 129, five conventional hydrogen bond interactions are shown with distance ranges from 1.62 to 3.34 Å. The pi cation interaction shows distance ranges from 4.01 to 4.34 Å (Figure 5(a)–(c)). One alkyl and four pi-alkyl bonds are formed with a distance range from 4.07 to 5.21 Å as shown in Table 7. The diversity in the bonding interactions observed among enzyme–dye complexes, for example, conventional hydrogen bonds, van der Waal forces, π–π stacking, and salt bridges, assists in rational enzyme tailoring by offering a structural draft. Researchers can predict the main units interfering in dye selection or transition state stabilization through evaluating the structural fingerprint of high binding affinity complexes. Moreover, mutagenesis and building robust protein structures can be done through these data that would enhance the catalytic efficacy or substrate binding affinity.
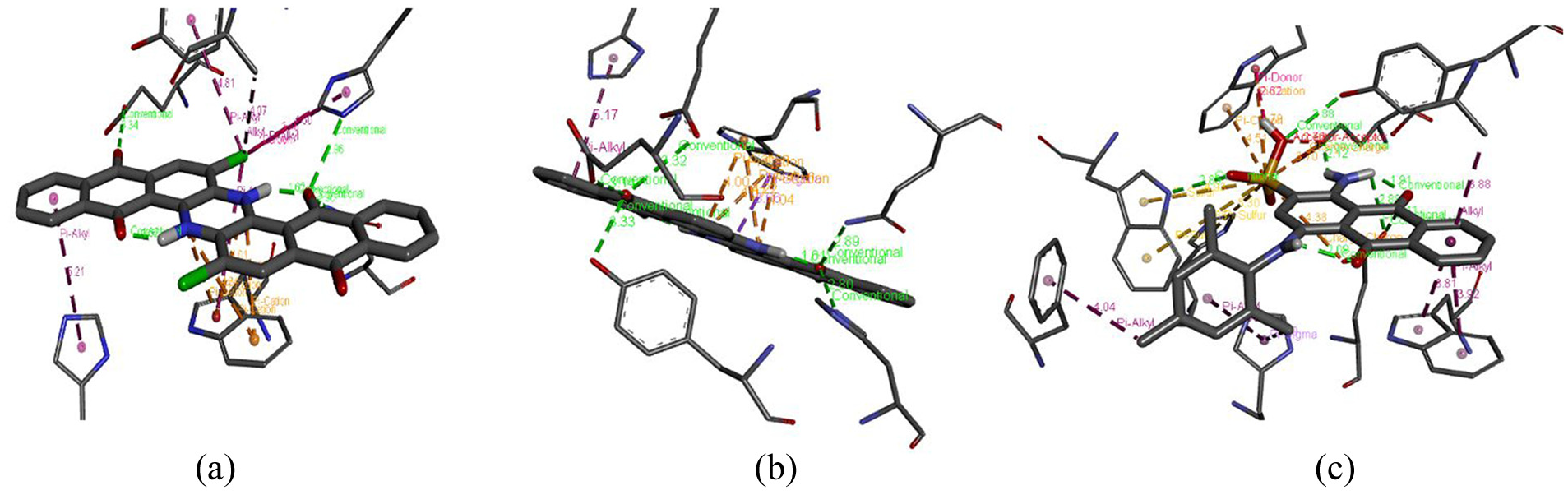
Interaction analysis of Glycoside Hydrolase Family 5 Protein with (a) Vat Blue 6, (b) Vat Blue 4, and (c) Acid Blue 129 pollutants along with their bond types and the distances between them.
Amino acid residues, distance range, and types of bonds present between enzymes and pollutants.
Discussion
The extensive use of anthraquinone dyes has led to significant environmental challenges, including toxicity to aquatic life and potential hazards to humans and animals.17,18 These dyes are highly persistent in the environment, making their degradation a critical area of research in environmental biotechnology. Enzymatic bioremediation has emerged as a promising solution for addressing this issue, with enzymes such as dye peroxidases and glycoside hydrolases demonstrating high catalytic efficiency and eco-friendliness. 19 This study evaluates the degradation potential of two enzymes from Flavobacterium sp. 9AF—Putative Multifunctional Dye Peroxidase DyP2 and Glycoside Hydrolase Family 5 Protein—using in silico approaches. To ascertain these enzymes’ safety for environmental use, toxicity was predicted via the CSM-toxin platform, which confirmed both enzymes as non-toxic. Anthraquinone dyes act as pollutants; their structures were downloaded from the PubChem database, a large database of chemical data, and ligand binding sites were predicted by Prank-Web. This research integrates toxicity profiling, ligand binding site prediction, molecular docking, and interaction analysis to provide insights into enzyme-based bioremediation methods.
The findings of this study align with previous research on enzymatic bioremediation but offer notable advancements. Key catalytic residues (e.g. His, Arg) in DyP2 align with conserved residues found in other DyP-type enzymes, supporting its functional annotation. Molecular docking revealed that DyP2 exhibited superior binding affinities for anthraquinone dyes compared to Glycoside Hydrolase Family 5 Protein. Structurally, DyP2 from Flavobacterium sp. shares key conserved residues typical of class A DyPs, such as distal His and Arg residues involved in peroxide activation. Functionally, our docking data reveal binding affinities comparable to or exceeding those reported for DyPs in
The superior performance of DyP2 is not solely due to hydrophobic interactions but also due to structural and mechanistic differences. 19 DyP2 exhibits a well-defined active site pocket with high Prank-Web confidence (score: 43.17, probability: 0.967), suggesting better substrate accommodation. Additionally, DyP2 contains conserved redox-active residues (His73, Arg311, Asp82) that enable oxidative cleavage of dye chromophores, unlike GH5, which lacks such residues and primarily targets glycosidic bonds. These structural features collectively enhance DyP2’s catalytic efficiency toward anthraquinone dyes.
Furthermore, the toxicity assessment via the CSM-toxin platform confirmed that both enzymes are non-toxic, supporting their environmental safety—a finding consistent with earlier studies that emphasized the importance of non-toxic enzymes for sustainable bioremediation. The hydrophilic nature of these enzymes (negative GRAVY values) also corroborates previous research indicating that hydrophilic proteins are more effective in interacting with water-soluble pollutants. Molecular docking assays were conducted with CB-Dock to evaluate the binding affinities of the enzymes toward 13 anthraquinone-derived industrial pollutants. Docking assays revealed that DyP2 showed the greatest binding affinities toward Solvent Green 3 (−12.1 kcal/mol), Vat Blue 6 (−11.4 kcal/mol), and Reactive Blue 19 (−11.3 kcal/mol). Similarly, Glycoside Hydrolase Family 5 Protein showed high binding affinities toward Vat Blue 6 (−11.6 kcal/mol), Vat Blue 4 (−11.1 kcal/mol), and Acid Blue 129 (−10.6 kcal/mol). The observed binding affinities (e.g. −12.1 kcal/mol for Solvent Green 3) are within the range reported in studies correlating strong docking scores with high degradation rates in DyPs. For example, fungal DyPs with similar binding energies have demonstrated >80% decolorization efficiencies in vitro. While in vitro validation was not part of this study, the docking results were obtained using validated platforms (CB-Dock, Discovery Studio) and cross-verified against known structural motifs. Binding affinities (e.g. −12.1 kcal/mol for Solvent Green 3) align with experimental data from comparable DyPs, which show similar degradation efficiency. Additionally, docking poses corresponded well with predicted Prank-Web pockets, supporting reliability.
The results demonstrate that DyP2 outperforms Glycoside Hydrolase Family 5 Protein in degrading anthraquinone dyes due to its higher binding affinities and stable catalytic pocket. Interaction analyses further revealed that DyP2 forms stronger hydrogen and hydrophobic bonds with pollutants like Solvent Green 3 and Reactive Blue 19, enhancing its degradation efficiency. Compared to Glycoside Hydrolase Family 5 Protein, which relies on conventional hydrogen bonds and fewer hydrophobic interactions, DyP2’s diverse bonding patterns contribute to its superior performance. To further understand enzyme–substrate interactions at a molecular level, Discovery Studio was used to look at binding residues, bond length, and types of bonds for the top three docked structures of each of the enzymes. DyP2 preferred hydrogen and hydrophobic bonds with contaminants like Solvent Green 3, while Glycoside Hydrolase Family 5 Protein preferred these same patterns with Vat Blue 6 as well as with other dyes. The literature reports DyP enzymes exhibiting Km values in the low micromolar range for anthraquinone dyes. These align with our docking-based substrate affinity estimates. Binding affinities offer preliminary insight into enzyme-substrate complex stability but cannot directly predict degradation rates. However, strong affinities (−10 to −13 kcal/mol) often correlate with low Km values in DyPs, suggesting efficient catalysis. These findings form the basis for selecting enzyme–dye pairs for experimental kinetic analysis under industrially relevant conditions.
This study underscores the potential of DyP2 as a powerful biocatalyst for anthraquinone dye degradation, offering an environmentally friendly solution to industrial dye pollution. The integration of advanced computational tools such as CB-Dock and Discovery Studio enabled detailed visualization of enzyme–substrate interactions, providing a deeper understanding of molecular mechanisms underlying dye degradation. These findings suggest that DyP2 could be further optimized for large-scale applications in wastewater treatment systems. This research indicates the biocatalytic potential of Flavobacterium sp. 9AF enzymes for the degradation of anthraquinone dye. Increasing the enzymatic degradation rate in the bioreactors results in constraints like substrate competition, lesser enzyme turn-over rate, product inhibition due to feedback regulation and mass diffusion limitation in immobilized systems. Real-world challenges include enzyme denaturation under fluctuating pH/temperature, inhibition by heavy metals or surfactants, and substrate diffusion limitations. These factors can reduce catalytic efficiency. Future experimental phases will include enzyme engineering (e.g. mutation of surface residues for stability), formulation buffers, and protective immobilization matrices to address these challenges. So, these challenges need empirical modeling and real-time monitoring to correspond with the in silico predictions. Ecological deployment may affect microbial community structures and nutrient dynamics. However, the non-toxic nature of these enzymes and their specificity minimize risks of non-target effects. Through the integration of toxicity profiling, ligand binding site prediction, molecular docking, and interaction analysis, this research offers insightful information on enzyme-based bioremediation methods for preventing industrial dye pollution. Site-directed mutagenesis targeting non-polar residues (e.g. Ile305, Phe346) could improve hydrophobic contacts with Disperse Red 15 and stabilize enzyme–ligand interactions. Mutants can be computationally screened prior to expression to predict improved binding affinity. This strategy could significantly enhance DyP2’s performance across a broader dye spectrum. Due to the exploratory nature of the study, in silico methods were employed as a cost-effective, high-throughput preliminary screening approach. These analyses are foundational for prioritizing enzyme-dye combinations for future in vitro validation. The use of CB-Dock, Prank-Web, and Discovery Studio enables mechanistic predictions that can later inform targeted biochemical assays and mutagenesis strategies.
Conclusion
In conclusion, this study provides significant in silico insights into the potential of Flavobacterium sp. 9AF enzymes for anthraquinone dye degradation. Through comprehensive analyses, Putative Multifunctional Dye Peroxidase DyP2 and Glycoside Hydrolase Family 5 Protein were evaluated, revealing DyP2 as a more promising candidate due to its higher binding affinities toward key pollutants like Solvent Green 3, Vat Blue 6, and Reactive Blue 19. Molecular docking studies, supported by favorable toxicity predictions and stable catalytic pocket assessments, highlighted DyP2’s superior interaction patterns, particularly its ability to form strong hydrogen and hydrophobic bonds with target dyes. Enzyme solubility and stability under different industrial wastewater conditions can be significantly enhanced through prioritizing enzyme polarity, particularly with the addition of hydrophilic based-residues. Enhanced polarity increases the protein–solvent contact that in turn decreases aggregation along with denaturation under environmental stress (e.g. PH shifts, high ionic strength, and the heavy metals presence). The findings suggest that DyP2 holds significant potential as a biocatalyst for enzyme-based bioremediation strategies, offering a sustainable and environmentally friendly approach to mitigate anthraquinone dye pollution from industrial wastewater. Moreover, tailoring enzymes like DyP2 can strategically fine tune the microenvironment surrounding the active site of DyP2 that could improve substrate interaction with reactive moieties and catalytic turnover for a number of pollutants functional groups. The 13 selected dyes represent a broad spectrum of anthraquinone derivatives differing in molecular size, substituents, and structural complexity. While sufficient for initial screening, future work will expand the dataset to include other dye classes (azo, indigoid, triphenylmethane) to explore DyP2’s broader catalytic scope, as its active site characteristics suggest possible cross-class activity. Electrostatic, hydrophobic and pKa of catalytic residues can be modified through techniques like targeted mutagenesis or in-silico remodeling of structures. DyP2 and GH5 act via complementary mechanisms—oxidative and hydrolytic—and thus their co-immobilization could improve degradation coverage. Precedents exist in laccase-peroxidase or peroxidase-hydrolase combinations enhancing dye breakdown. Immobilized enzyme consortia offer sequential action: DyP2 cleaves chromophores while GH5 hydrolyzes residual dye conjugates, potentially improving overall degradation efficiency. The Co-immobilization of DyP2 and hydrolase enzymes could yield synergistic catalytic mechanisms for better degradation and expanded dye substrates coverage. Simultaneously, DyP2 can catalyze the oxidative breakage of chromophores and then GH5 can breaks the glycosidic bonds present in dye conjugates that results into easily degradable smaller components. Prediction models may introduce bias. To minimize this, validated tools like Alpha Fold, CB-Dock, Discovery Studio, and plan to cross-check with alternative algorithms. This research supports further investigation into DyP2’s application in real-world bioremediation settings.
Footnotes
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors express their gratitude to Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2025R23), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia. The authors are also thankful to the Deanship of Scientific Research (DSR) at King Faisal University under project no. [KFU250840].
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of data and materials
All the data generated in this research work has been included in the manuscript.