Abstract
Objective:
This cross-sectional study evaluates the association between the oral microbiome and the presence of untreated dental caries among Kenyan children and adolescents living with HIV (CALHIV).
Methods:
We collected 83 oral swab samples from CALHIV receiving medical care at Kenyatta National Hospital. We used the 16S rRNA sequencing technique on oral swab samples to profile bacterial composition. Only CALHIV with results of oral microbiome and dental examinations simultaneously were included in the further analysis. We compared the alpha and beta diversity of the oral microbiome between individuals with and without dental caries. Log-ratio LASSO regression with 2-stage estimation, 10-fold cross-validation, and adjustment for the false discovery rate was applied to select the best features that can predict caries status at the phylum level.
Results:
Of the 70 participants with both oral health data and oral microbiome data available, 66% had dental caries. The group with dental caries had higher Chao 1 alpha diversity (P = 0.046) but not beta diversity. Log-ratio LASSO regression had a higher ratio of paired features proteobacteria/SR1 (P = 0.0067), Euryarchaeota/SR1 (P = 0.0074), Acidobacteria/SR1 (P = 0.0079), and Fusobacteria/SR1 (P = 0.0125) at the phylum level and was significantly associated with a lower prevalence of untreated dental caries. A higher abundance of SR1 together with certain bacteria was associated with a higher prevalence of untreated dental caries.
Conclusions:
This is the first study to analyze the oral microbiome of CALHIV and untreated dental caries in a cohort that had received medical care since their first years of life. While this study provides information about the paired feature selection for dental caries in CALHIV on ART using cross-validation, SR1 may interact with other bacteria to increase the prevalence of untreated dental caries. Longitudinal microbiome data and samples from multiple sites in the oral cavity will be essential for understanding and confirming these findings.
Knowledge Transfer Statement:
The results of this study can help clinicians understand how SR1 interacts with other bacteria that may increase the prevalence of untreated dental caries in children and adolescents living with HIV and on long-term antiretroviral therapy. Furthermore, these findings may provide valuable insights for future research, informing longitudinal microbiome data and sampling from various sites in the oral cavity.
Introduction
The oral microbiome, defined as the collective genome of microorganisms that reside in the oral cavity (Deo and Deshmukh 2019), plays an essential role in oral and systemic health (Kleinstein et al. 2020; Xiao et al. 2020). The oral microbiome, along with salivary flow, soluble salivary components, immune cells, and mucosal epithelial cells, maintains an immune homeostatic state (Lü and Jacobson 2007; Perez Rosero et al. 2021). In a healthy state, these microorganisms consist of commensals, whose genes and phenotypes will be selected by the immune system to survive in symbiosis, through the highly regulated signals that are modulated by a network of microbial and host metabolites (Kleinstein et al. 2020). The interaction of the oral microbiome with inflammation and the immune system causing oral diseases remains unknown (Gonçalves et al. 2019; Balle et al. 2020; Coker et al. 2020; Kleinstein et al. 2020; Coker, Akhigbe, et al. 2021).
Children and adolescents living with HIV (CALHIV) on prolonged antiretroviral therapy (ART) have a high prevalence of oral mucosal inflammation and oral comorbidities such as dental caries and gingival diseases (Coker, Cairo, et al. 2021; Li et al. 2021). It is unclear why the risk of oral inflammation and oral diseases remains increased among individuals with HIV despite viral suppression (Coker, Cairo, et al. 2021). HIV infection, even when well-controlled, has been associated with the alteration of the immune response and thereby microbial composition, which could lead to low-grade inflammation and oral comorbidities (Sabella et al. 2021).
Using next-generation high-throughput sequencing techniques, a distinguishable core oral and gut microbiome has been identified among CALHIV, including those on ART (Perez Rosero et al. 2021; Beall et al. 2023; Goosen et al. 2023). ART partially reverses HIV-induced oral microbiota alterations and is associated with significantly greater healthy bacterial richness and diversity (Presti et al. 2018; Griffen et al. 2019; Fidel et al. 2021). The different effects of HIV infection and ART make it challenging to understand the interplay between oral microbial community and oral diseases, such as dental caries.
The root biological cause of dental caries is biofilm-mediated dysbiosis, during which the composition and function of the core microbiome become altered (Ribeiro and Paster 2023). This alteration fosters an acidic environment that leads to the demineralization of the hard tooth tissues. Hence, the changes in oral microbiome composition could serve as potential biomarkers for disease diagnosis and prediction (Coker, Cairo, et al. 2021; Bourgeois et al. 2022; Raksakmanut et al. 2023). As HIV infection is significantly associated with a high prevalence of dental caries (Shanmugavadivel et al. 2020; Coker, Akhigbe, et al. 2021; Akhigbe et al. 2022; Wang et al. 2023), microbial biomarkers could be a useful tool for caries management in this patient population. In addition, the oral microbiome in HIV progression is an emerging area of significant interest (Goldberg et al. 2015; Presti et al. 2018; Griffen et al. 2019; Li et al. 2021). However, due to its high dimensionality, sparsity, compositional nature, and large biological variability, microbiome data present statistical challenges for data management and modeling.
In this article, we analyzed both the counts and relative abundances of taxonomic data and compared alpha- and beta-diversity among children and adolescents with HIV infection who are on long-term ART with and without untreated dental caries. We investigated the pairwise ratios of bacteria at the phylum level to identify potential biomarkers associated with untreated dental caries in this population. This approach allows us to focus on the interactions of different taxa.
Methods
This cross-sectional study was approved by the Institutional Review Board at the University of Washington (STUDY00003298), by the Ethics and Research Committee of the Kenyatta National Hospital/University of Nairobi (KNH/ERC/R/133), and by UCLA (IRB#20-001102). The reporting in this article conforms to the STROBE guidelines.
Recruitment
Study participants were recruited from an ongoing cohort study in Nairobi, Kenya. The recruitment process and study population have been described in detail elsewhere (Seminario et al. 2022; Wang et al. 2023; Seminario et al. 2024). The study was embedded within the longitudinal cohorts of the Kenya Pediatric Studies, which recruited participants from 2 clinical trials. The first (“early ART initiation”), the Optimizing Pediatric HIV-1 Treatment (OPH) study, enrolled infants younger than 12 mo with initiation of ART at enrollment. The second (“late ART initiation”), the Pediatric Adherence Study (PAD), enrolled children aged 1.5 to 12 y with ART initiation per contemporaneous guidelines (ART initiated in CALHIV with immunosuppression). This is a convenience sample, and everyone in the cohort who agreed to participate joined the study. All parents or legal guardians provided written informed consent for participation, and all children assented to study procedures. Both consent and assent forms were provided in English and Kiswahili. All staff involved in the consent process were fluent in both languages. At the time of the oral diseases cross-sectional survey for the current analysis, children had a median age of 11.4 y and had been enrolled in OPH at a median age of 10.8 y and PAD for a median of 17.3 y.
Dental Examination
Standardized clinical assessments and questions from the World Health Organization Oral Health Surveys were used to collect demographic and clinical information. Oral examinations were conducted during their routine HIV care visits. All examinations were conducted by local community oral health officers (COHOs; aka dental therapists), who had been trained in the diagnosis of dental diseases and oral manifestations of HIV: angular cheilitis, oral hairy leukoplakia, necrotizing ulcerative gingivitis/periodontitis, necrotizing stomatitis, herpetic stomatitis/gingivitis and/or labial lesions, herpes simplex virus ulcer, herpes zoster, molluscum contagiosum, cytomegalovirus, oral warts (papillomavirus), Kaposi sarcoma lesion, aphthous ulcer, other ulcerations, dry mouth due to decreased salivary flow, unilateral or bilateral swelling of major salivary glands, and other diagnoses, such as gum disease, calculus (tartar), xerostomia (sensation of dry mouth), impacted tooth/teeth, abscess, enamel hypoplasia, dental fluorosis, and potential malignancy (Ramos-Gomez et al. 1999). All diagnoses were recorded as present or absent by these examiners, who used validated training modules provided by the University of Washington and the University of California, San Francisco (Shiboski et al., 2015). The interrater and intrarater reliability scores were 0.78 and 0.85, respectively. The Oral Health Surveys and Record Form for Oral Manifestations of HIV/AIDS (World Health Organization 2013) was used to evaluate oral diseases and HIV-associated oral mucosal lesions (Wang et al. 2023). Dental caries was evaluated based on visual examination under natural light, augmented with a headlight; gauze for cleaning and drying teeth; a probe; and dental mirrors. Each tooth was evaluated for caries, fillings/crowns, missing status due to caries/other reasons, white spots, sealants, and enamel developmental defects. Dental caries was summarized by the presence (yes/no) of caries on any tooth.
Oral Swab
The oral swab samples were collected during the dental exam by COHOs. We used an absorbent Dacron (polyester) swab to collect saliva for the assessment of oral cytokine levels. We rubbed the tip of the swab on the inside of the cheek until it was fully saturated with saliva. Then, we placed the swab into an open cryovial containing freezing media. We broke off the long swab handle so that the lid was placed on tightly. Oral swabs were stored in freezing media. The freezing media containing 1 mL fetal bovine serum and 10% DMSO media placed in Wheaton 2 mL CryoELITE sterile cryogenic vials (VWR #89125-504). Participants did not consume any food 1 h before sample collection. There were no oral hygiene restrictions before sample collection.
Oral Microbiome
Oral microbiome data were generated by 16S rRNA gene sequencing of oral swab samples using the Illumina MiSeq platform. Barcoded primer pairs (forward: 515F, reverse: 806R) targeting the V4 region of the 16S rRNA genes were used for polymerase chain reaction (PCR) amplification of bacterial DNA following the Earth Microbiome Project protocol (Gilbert et al. 2014). Amplicons were sequenced on an Illumina MiSeq desktop sequencer using 2× 150-bp v2 chemistry and in accordance with the manufacturer’s specifications (Tobin et al. 2018). We used a total of 3 plates, each including ZymoBIOMICS microbial community DNA standard as a positive control and BMCK, negative, and PCR negative control samples. All samples were randomly allocated across 3 plates for further sequencing.
Statistical Analysis
Only CALHIV who had both the results of the oral microbiome and dental examinations were included in further analysis. We used the decontam R package to distinguish the contaminants and the real bacterial community. We used the OUT-based clustering methods that can agglomerate contaminant sequences with the oral swab sample sequences. The R function Contaminate () can effectively and sensitively exclude the contaminant Amplicon Sequence Variants (ASVs). We used the default methods auto, combined DNA concentration of the samples, and prevalence of the contaminant in the control samples.
The study primarily compared the alpha and beta diversity of the oral microbiome between individuals with and without dental caries. We used the open-source computational tool FLORAL (Fei et al. 2024) to perform log-ratio LASSO regression modeling and compositional feature selection for the binary outcome. This method adapts the augmented Lagrangian algorithm for a zero-sum constraints optimization problem, and 2-stage estimation, 10-fold cross-validation, and adjustment for false discovery rate were applied to determine the association between caries status and the taxa ratio at the phylum level. For the log-ratio LASSO regression models, we ran the model 1,000 times. The P values were the average derived from these 1,000 iterations, lending robustness to our findings. A 2-stage estimation process and 10-fold cross-validation were used to select the best pairwise combination for predicting caries status.
Results
Study Population Characteristics
Our study population consisted of 70 CALHIV participants with both dental examination results and oral microbiome data available. The mean age of the participants was 13.1 y (range, 10 to <21 y). More than 65.7% had initiated ART as early as before 1 y of age. The average duration on ART was 11.6 y, with a range of 9.5 to 14.8 y. Most of the participants (68.6%) were on a nonnucleoside reverse transcriptase inhibitor–based regimen, while the remainder were on protease inhibitor–based first-line ART regimens. The number of teeth with untreated dental caries ranged from 0 to 10, with a mean number of 3.7 among those who had untreated dental caries (Table 1). Almost two-thirds of the study population (65.7%) had untreated dental caries. Age, timing of ART (early vs. late), regimen, and CD4 counts were similar between CALHIV with and without untreated dental caries. Males (65.2%) had a higher rate of untreated dental caries than females (34.8%) did (P = 0.0014) in this population, with a notably high rate of untreated dental caries (65.7%) overall.
Description of the 70 Children with Oral Swab Samples and Dental Exam Results.
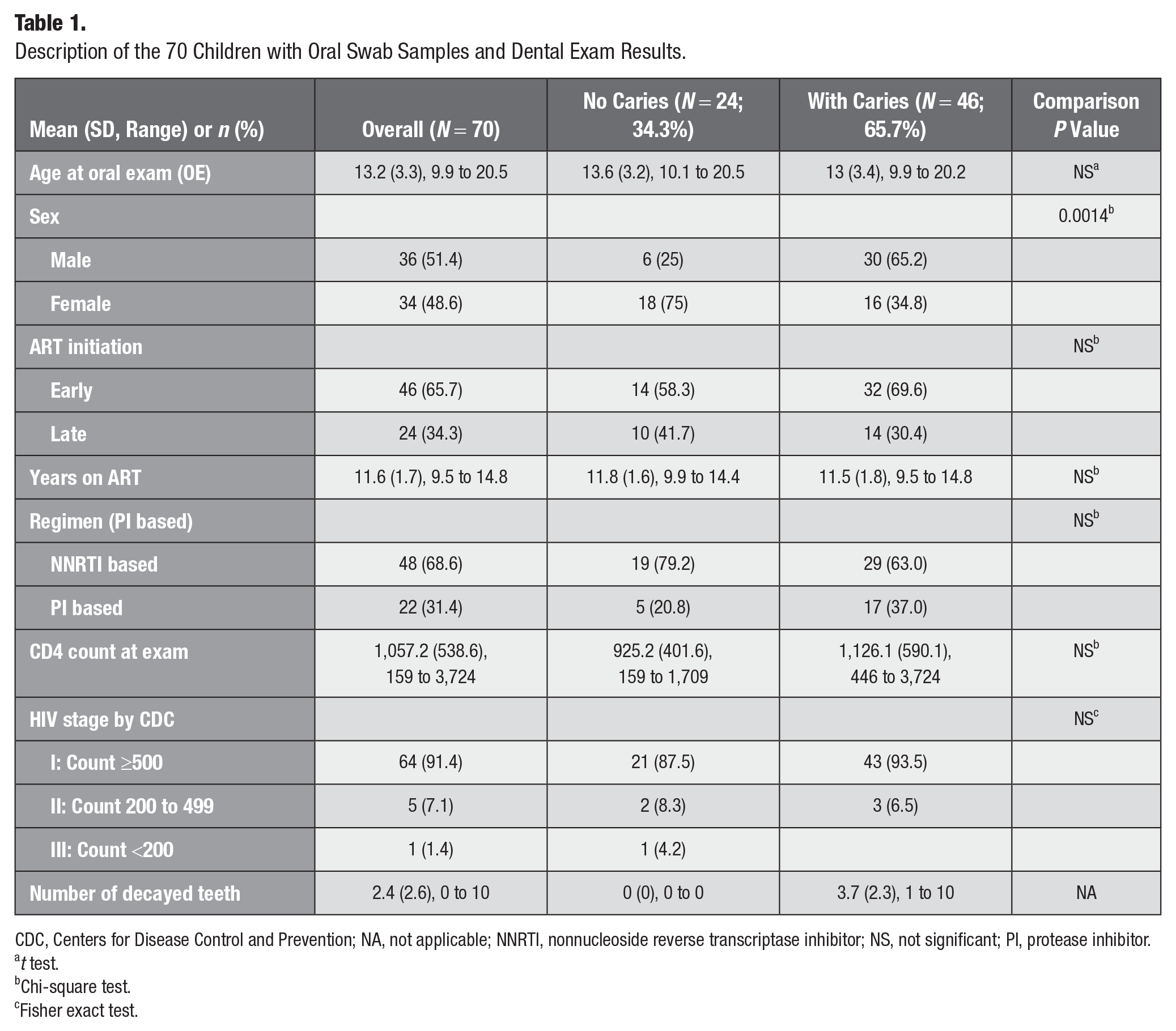
CDC, Centers for Disease Control and Prevention; NA, not applicable; NNRTI, nonnucleoside reverse transcriptase inhibitor; NS, not significant; PI, protease inhibitor.
t test.
Chi-square test.
Fisher exact test.
Microbiome Sequencing Results
We performed 16S rDNA sequencing by using the Illumina platform on a MiSeq, resulting in 9,140,516 total initial reads that ranged from 9 to 358,944 per sample with a total of 3,367 taxa and 80 samples including negative control. Supplement Figure S0 plots the community of all control samples. In the final analysis, we included a total of 70 patients with 3,057 taxa identified. The number of reads ranged from 62,386 to 358,944, with a mean of 127,754. Please refer to Supplement Figure S1 for the detailed process of removing contaminants, rarefying, and excluding control communities.
The relative abundance at the phylum level is plotted in Figure 1, categorized by groups with and without caries.
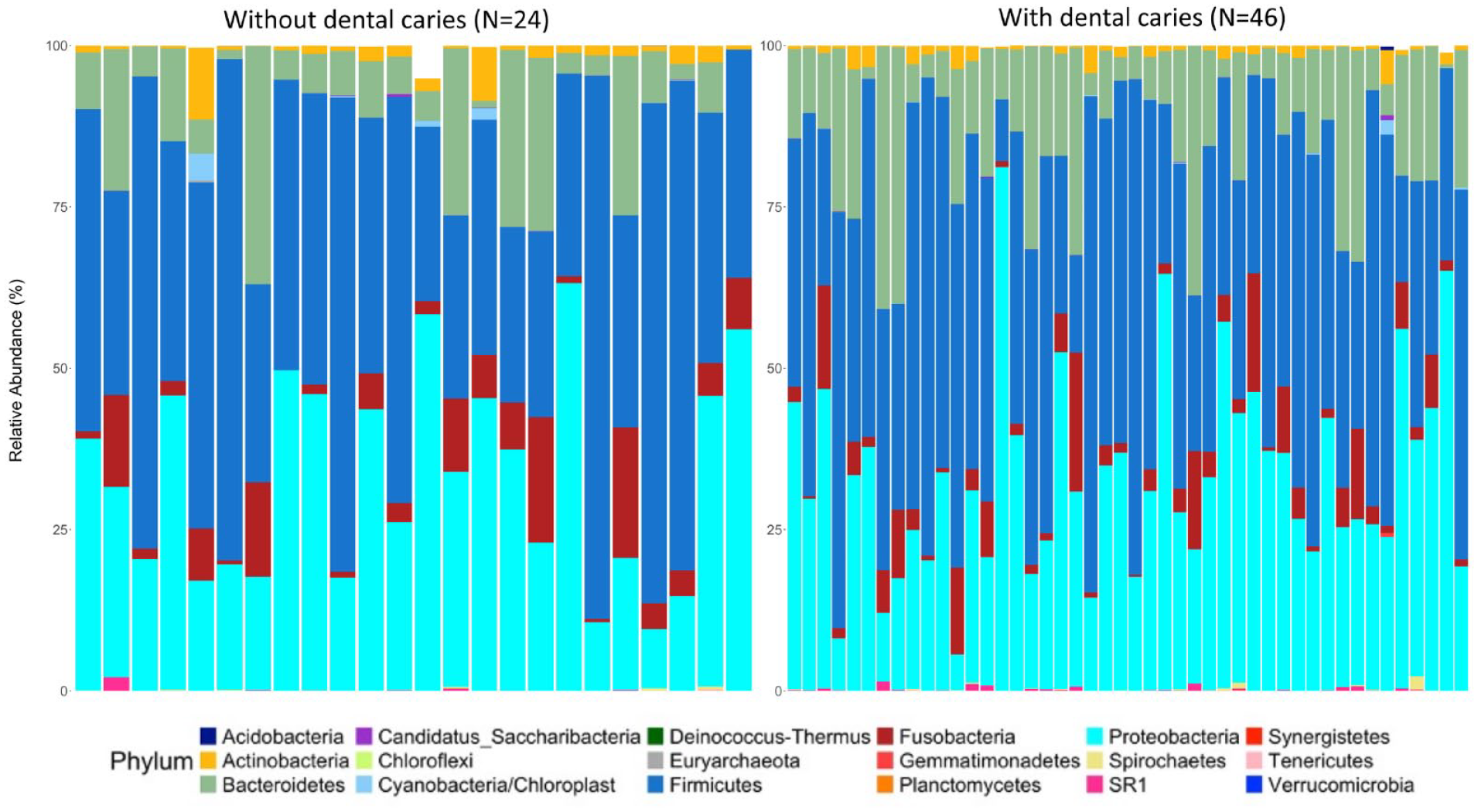
Relative abundance by caries status at phylum level.
Alpha Diversity for Richness
The richness of the bacterial community between groups with and without dental caries was compared using commonly used indices: ACE, Chao1, Observe, Pielou, Shannon, and Simpson. The Chao1 index demonstrated a higher total number of species in the caries than noncaries group (P = 0.046 by Wilcoxon test; Fig. 2), considering both observed counts and rare or unobserved counts. Both Simpson and Shannon indices suggested that the microbiota of the caries group had a very similar spread of abundances when compared with the no-caries group. The difference in alpha diversity between groups with and without caries was very small.
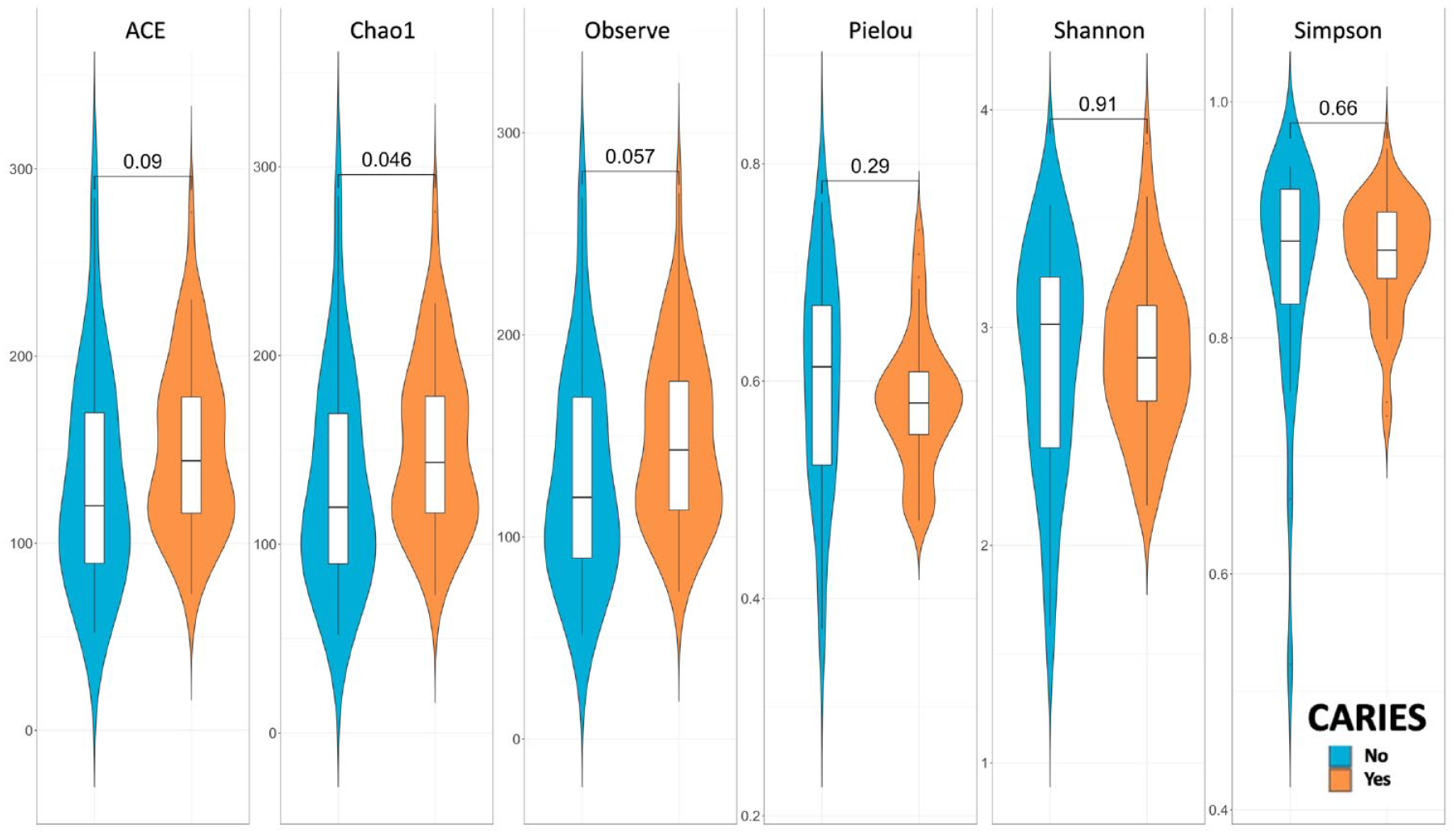
Violin plot of alpha diversity by caries and p-value by Wilcoxon test.
Beta Diversity for Similarity
We used the Bray-Curtis dissimilarity and weighted/unweighted UniFrac distance to examine the abundances of microbiome shared between groups. The Bray-Curtis distance ranged from 0 to 1, with 0 indicating that the groups share the same number of taxa at the same abundance. The microbial community between groups with and without dental caries was not compositionally different (Fig. 3). Each point represented a sample, and the distance between points represented how samples are compositionally different from each other. The points were all clustered together regardless of groups, indicating that microbial communities are similar in taxa composition. Similar findings were observed using weighted and unweighted UniFrac distance, accounting for the relative abundance of each taxon within the communities.
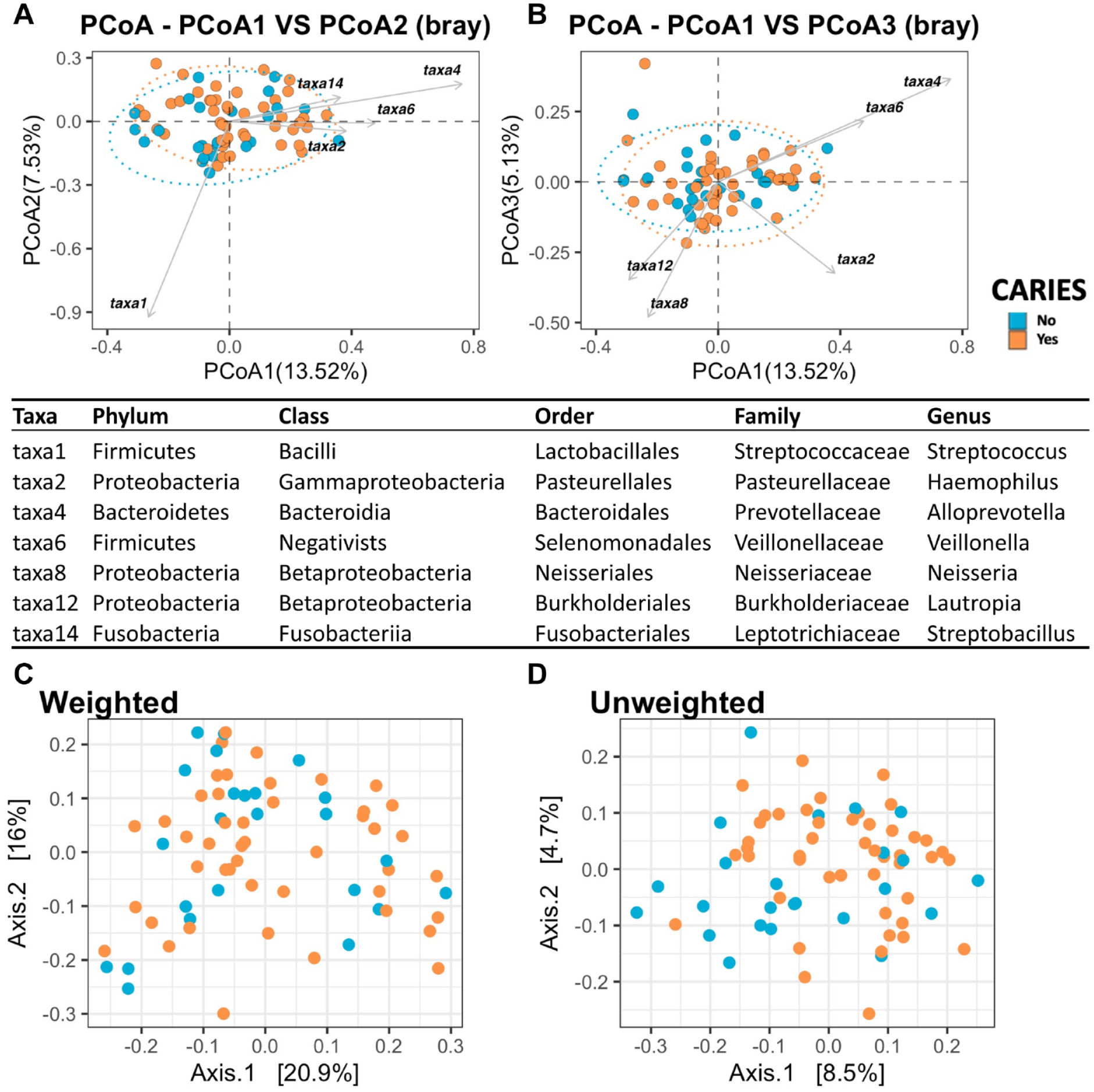
Principal coordinates analysis (PCoA) by Bray (
Log-Ratio LASSO Regression Model
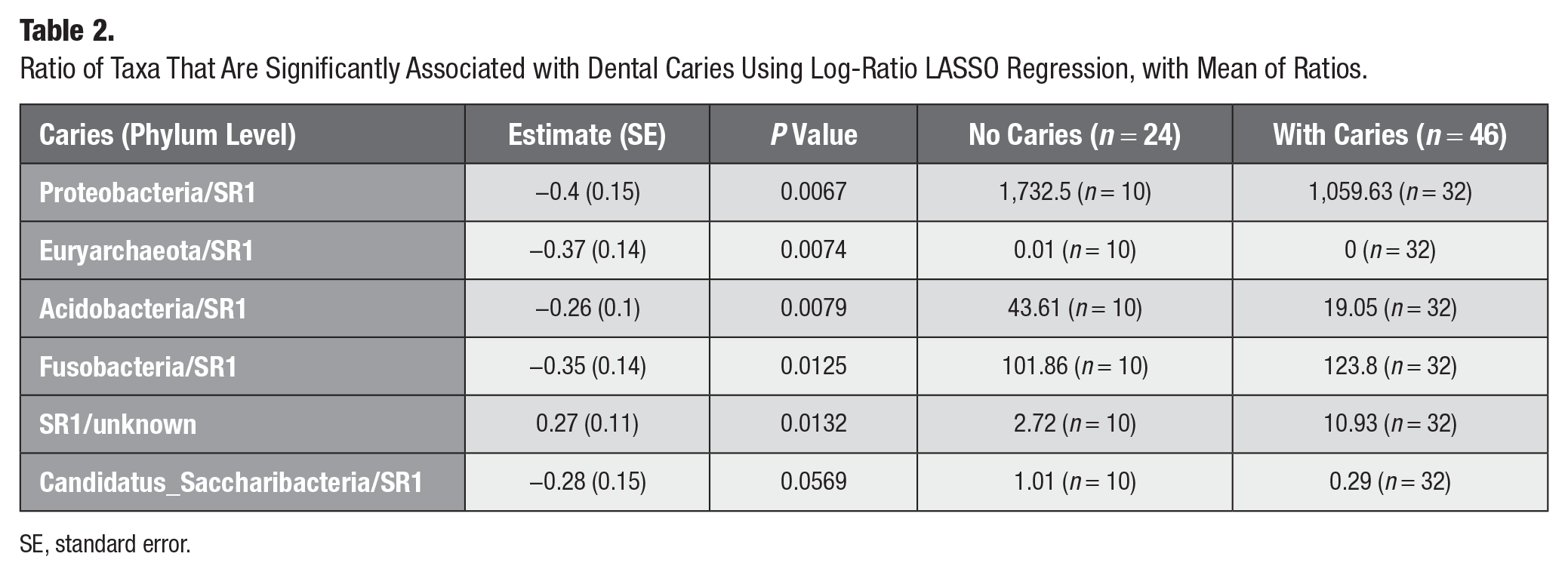
Table 2 presents the results of the log-ratio LASSO regression model at the phylum level. It reports the averages of coefficients and standard errors, average P values, and ratios of both groups. The table highlights the top ratios with an average P value from 1,000 models of less than 0.05, suggesting a significant association between higher levels of SR1 (lower value of ratios of other bacteria/SR1) and a high probability of dental caries. The top 3 ratios that are significantly associated with untreated dental caries, with mean P values among 1,000 models of less than 0.01, are Proteobacteria/SR1, Euryarchaeota/SR1, and Acidobacteria/SR1 as well as Fusobacteria/SR1 and SR1/Unknowns with P values of less than 0.05. The model presented the results that a higher ratio of paired features Proteobacteria/SR1 (P = 0.0067), Euryarchaeota/SR1 (P = 0.0074), Acidobacteria/SR1 (P = 0.0079), and Fusobacteria/SR1 (P = 0.0125) at the phylum level is significantly associated with a lower prevalence of untreated dental caries. The directions of the coefficients are consistent with the mean of the ratios between groups with and without untreated dental caries (Table 2), except for Fusobacteria/SR1, for which the mean of the ratios was higher among the groups with untreated dental caries. The interactions indicate that SR1, along with other bacteria, may together predict the prevalence of untreated dental caries.
Ratio of Taxa That Are Significantly Associated with Dental Caries Using Log-Ratio LASSO Regression, with Mean of Ratios.
SE, standard error.
Discussion
Oral biomarkers have gained increasing attention recently, especially among young children and adolescents, due to their easy management and noninvasive nature (Dewhirst et al. 2010; Lamont et al. 2018; Deo and Deshmukh 2019; Bourgeois et al. 2022). This study establishes a foundation for utilizing using swab samples, particularly focusing on the oral microbiome, to investigate a possible connection between active dental caries and bacterial profile among CALHIV. Those participants who had untreated dental caries had a slightly higher species richness than those without dental caries, as measured by the Chao1 index. Exploring these elements within the oral environment is essential in gaining vital insights for using oral biomarkers to monitor the development of dental caries and the progression of HIV. Using the log-ratio regression model, we also identified that SR1 potentially plays a role in dental caries, together with other bacteria. This model focused on examining the pairwise interactions of the oral microbiome at the phylum level that are associated with caries. To enhance the robustness of our findings, we reported the average P value from a large number of models. The higher value of the ratio of Proteobacteria, Euryarchaeota, Acidobacteria, and Fusobacteria over SR1 was significantly associated with a lower probability of dental caries status. This was consistent with findings from other studies among children without HIV infection (Kalpana et al. 2020). Further investigation will be needed to examine the paired interactions between those identified bacteria at the phylum level with unknown species that may lead to dental caries.
Consistent with other oral microbiome studies, we did not find a significant difference in other alpha diversity indices and beta diversity indices between those with and without caries (Goldberg et al. 2015; Li et al. 2020; Shanmugavadivel et al. 2020; Coker, Akhigbe, et al. 2021; Coker, Cairo, et al. 2021; Ribeiro and Paster 2023). This may be due to the small sample size. The research was carried out at the Kenyatta National Hospital in Nairobi, where this cohort of CALHIV is well-monitored and receives good care for their general health but not for oral health. The high prevalence of dental caries may reflect the continued impact of HIV infection despite ART. However, due to the lack of healthy control comparison, this hypothesis requires further investigation. The overall richness and similarity of the microbial community of CALHIV were similar to those of a healthy population, perhaps due to the long duration of ART and immune restoration (Goldberg et al. 2015; Kahharova et al. 2020). We recommend that future studies consider comparing the oral microbiome profiles of those with and without HIV infection to understand the impact of HIV infection on changes in the oral microbiome. In addition, this study included a wide range of children and adolescents with a broad age range. This factor could further bias the findings toward age differences or the initiation time on ART.
There is limited information on the interaction of the oral microbiome related to oral diseases for CALHIV on prolonged ART. Even among adults, the impact of HIV infection on the oral microbiome is controversial, where some studies support microbiome modifications and others do not (Li et al. 2021; Perez Rosero et al. 2021). This may be due to study design, detection methods, sampling location (saliva, oral wash, swab, or subgingival), or environment (Lamont et al. 2018; Deo and Deshmukh 2019; Coker, Cairo, et al. 2021; Li et al. 2021; Sabella et al. 2021) Due to the developmental stage of children’s oral health, it is necessary to study the pathophysiology of HIV-induced inflammation to guide potential interventions (Neuhaus et al. 2010; Goldberg et al. 2015; Starr et al. 2018; Gonçalves et al. 2019; Coker et al. 2020). The dysbiosis of the oral microbiome is not fully restored after effective ART (Li et al. 2020). Meanwhile, ART and HIV infection together contributed to salivary dysbiosis and the changes in the abundance of several bacterial species (Lewy et al. 2019; Coker et al. 2020). Our study provides invaluable baseline information on the oral microbiome of a vulnerable population as CALHIV in a Sub-Saharan country whose HIV was well-controlled.
We used oral swab samples collected by twisting swabs at the cheek. An increasing body of literature has found that the microbiome profile differs when collected from various oral sites (Li et al. 2022). For example, the oral microbiome of the tongue may be distinct from the microbiome profiles of saliva samples, swab samples, or dental plaque. We chose swab samples from CALHIV because the samples collected from the cheek are relatively stable and easier to manage in study protocols for resource-limited settings. Also, we wanted to make sure that younger children would be more cooperative by collecting samples from an easily accessible area with lower gag reflex. The results may not be applicable to different sample collection methods.
We applied a scalable log-ratio LASSO regression model with FLORAL, an open-source computational tool developed by Fei et al. (2024), to enhance feature selection for predicting dental caries status. This method achieved better control of the false discovery rate and improved variable selection based on 10-fold cross-validation and a 2-step model selection process. We did not consider other clinical factors as covariates when estimating the association between microbial pairs and dental caries. In this baseline study, we focused on evaluating the predictive features of paired taxa.
We used the 16S rRNA gene sequencing method, which is a widely adopted approach for bacterial identification and phylogenetic analysis. However, the taxonomic resolution of this method is limited. The 16S rRNA gene is highly conserved among different bacterial species, making it challenging to differentiate taxa at the species or strain level. We analyzed the paired interactions only at the phylum level in the models and compared the composition at genus levels (Appendix Table S3). Additional bias could be introduced during the PCR step of sequencing. Moreover, using only the 16S rRNA method did not allow us to analyze nonbacterial microorganisms, such as fungi and viruses, which could be significant in swab samples from CALHIV undergoing prolonged HIV treatment. Some unknowns in the microbial community may be a result of the limitations of the 16S rRNA resolution. We would need higher-resolution sequencing techniques to further investigate the unknowns and other methods to expand the analysis to include the fungal and viral components of the microbiome.
We have demonstrated the feasibility of conducting oral microbiome studies in resource-limited dental clinics. The protocol involved training local dental officers and assistants to collect oral swab samples and keep them frozen before shipment on dry ice, which may facilitate the expansion of research to include longitudinal oral swab sample collection in dental clinics. In doing so, we can collect samples from both CALHIV and their healthy peers over time to further investigate the impact of HIV infection, ART, as well as oral diseases and malnutrition status on the changes in the oral microbiome among young children with a high burden of oral diseases.
There are some limitations to this study. First, we used convenience sampling to recruit participants from an existing cohort of HIV research. We did not conduct a sample size or power evaluation; instead, everyone who agreed to participate was included. In addition, we lacked a comparison sample of children without HIV infection. Given the limited resources for oral health care, minimal care was provided to these children. We observed a high prevalence of untreated dental caries and similar prevalence rates of untreated caries and risk of caries. In this study, we collected only oral swab samples and did not collect plaque samples, which might have better addressed the risk of dental caries. Furthermore, we did not take radiographs due to the lack of infrastructure. We used the 16S rRNA sequencing technique, which may be accurate at the genus level; however, due to the small sample size, we were able to model associations only at the phylum level. Future studies could use metagenomic sequencing to explore microbial interactions at the species and strain levels. We also had limited data on participant-level factors, such as dietary habits, oral hygiene practices, and socioeconomic status, which are important contributors to oral microbiome diversity and dental caries risk. Finally, as a cross-sectional study with limited sample size, we could address only associations, even with the use of advanced log-ratio LASSO regression.
Conclusions
This is the first study to analyze the oral microbiome and dental caries in a cohort of CALHIV that has received medical care since the first years of life. While this study provides information about the paired feature selection for dental caries in CALHIV on ART using cross-validation, SR1 may interact with other bacteria to increase the risk of dental caries. Future longitudinal microbiome data and samples from multiple oral sites will be essential for understanding and confirming these findings.
Author Contributions
Y. Wang, O. Matangkasombut, G.M. Aldrovandi, A.L. Seminario, contributed to conception, design, data acquisition, analysis, and interpretation, drafted and critically revised the manuscript; A.M. Kemoli, contributed to conception, design, data acquisition and interpretation, critically revised the manuscript; G. John-Stewart, S. Benki-Nugent, J. Slyker, contributed to design, data interpretation, critically revised the manuscript. All authors gave their final approval and agreed to be accountable for all aspects of the work.
Supplemental Material
sj-docx-1-jct-10.1177_23800844241311862 – Supplemental material for Oral Microbiome and Dental Caries in Kenyan Children and Adolescents Living with HIV
Supplemental material, sj-docx-1-jct-10.1177_23800844241311862 for Oral Microbiome and Dental Caries in Kenyan Children and Adolescents Living with HIV by Y. Wang, O. Matangkasombut, A.M. Kemoli, G. John-Stewart, S. Benki-Nugent, J. Slyker, G.M. Aldrovandi and A.L. Seminario in JDR Clinical & Translational Research
Footnotes
Acknowledgements
We thank the patients, parents, and staff of the Kenyan Pediatric Studies for their generosity and willingness to participate in this study. We greatly thank Hu-Friedy Mfg. Co., LLC, for its generosity in donating dental instruments for our oral examinations. We thank Dr. Fan Li and Dr. Nicole Tobin for their technical support.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants NIH/NIDCR K01DE032775, R03DE033631, and NIH/NIMH T32MH080634. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
A supplemental appendix to this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
