Abstract
Objective
To investigate the effects of a probiotic drink containing
Methods
Caries-free young adults were administered a probiotic drink containing
Results
Out of samples from 10 included participants, 256 814 sequences passed through quality control, clustered into 170–234 different ‘species-level’ phylotypes.
Conclusion
Yakult intake changed the abundance of some bacteria related to caries, suggesting that the change of composition may be beneficial to oral health, while the overall microbiota structure remained unaffected.
Introduction
Dental caries, or tooth decay, is a complex biofilm disease involving a net loss of minerals from the teeth due to prolonged periods of low pH in the mouth. 1 With negative impacts on quality of life, dental caries also adversely affects children’s school attendance and academic performance, 2 and is one of the main reasons for hospitalization of children in some high-income countries. 3 Efforts to prevent dental caries include water fluoridation, fluoride toothpastes, fluoride varnishes, sealants and/or interim therapeutic restorations, and silver diamine fluoride, each of which has shown varying levels of efficacy in clinical trials. 2 Although the prevalence of dental caries has declined in most industrialized countries since the 1970s, 4 dental caries remains the most common non-communicable disease worldwide. 3 Thus, new methods for caries prevention remain in urgent demand.
Probiotic therapy has been introduced for the maintenance of oral health,
5
but studies on the same probiotic strain have presented conflicting results.6–8
To date, there are no reliable reports of species and genotypic diversity in the oral cavity following consumption of Yakult. Therefore, the present study was conducted to explore the effects of
Patients and methods
Study population
This self-controlled case study recruited healthy students aged 20–22 years, who were attending Sun Yat-Sen University, between November 2016 and December 2016. Students were recruited through advertisement and inclusion criteria were self-reported healthy status and complete dentition. The oral cavity of each subject was checked by an experienced dentist (XH) and individuals with the following conditions were excluded: less than 28 teeth; clinically detectable signs of dental caries (white spot lesion or carious cavity), periodontal disease (gingival bleeding, deep periodontal pocket, attachment loss and tooth mobility) or other pathologies (such as sinus tract, blisters and swelling); presence of dental restorations, crowns or orthodontic appliances; current smokers or smoker co-habitants; current participation in another clinical study; use of antibiotics or probiotic products within 3 months prior to examination. 10 Subjects with current, or history of, diseases associated with oral microbiota, such as diabetes mellitus, rheumatoid arthritis, inflammatory bowel disease and atherosclerosis,11,12 were also excluded.
The study was reviewed and approved by the Ethics Committee of the Guanghua School and Hospital of Stomatology, Sun Yat-sen University (KQEC-2019-01), and all participants provided written informed consent.
Study design and sampling procedure
For each subject, supragingival dental plaque was collected from the surfaces of four first molars on the morning of day 1, namely timepoint A, at least 24 h after the last tooth brushing and 12 h after the last food intake. Subjects were then instructed to drink 100 ml of Yakult (Yakult Sdn Bhd, Selangor Darul Ehsan, Malaysia) three times on day 1, one bottle immediately following sampling, one after lunch and one after dinner, by swishing the beverage in the mouth for about 20 s to prolong contact time with the teeth before swallowing. They were further instructed to drink one bottle of Yakult after lunch daily from day 2 to day 29 according to the method above. Supragingival dental plaque was collected on the morning of day 2 and day 30, namely timepoints B and C (Figure 1). During the study, subjects were instructed to maintain their regular oral hygiene regimen except for the 24 h preceding sample collections on day 1, day 2 and day 30. 10 All oral samples were collected at the Guanghua Institute of Stomatological Research, Sun Yat-Sen University, and were immersed in 4 ml of sterilized 0.9% saline solution and stored at –80°C prior to DNA extraction.
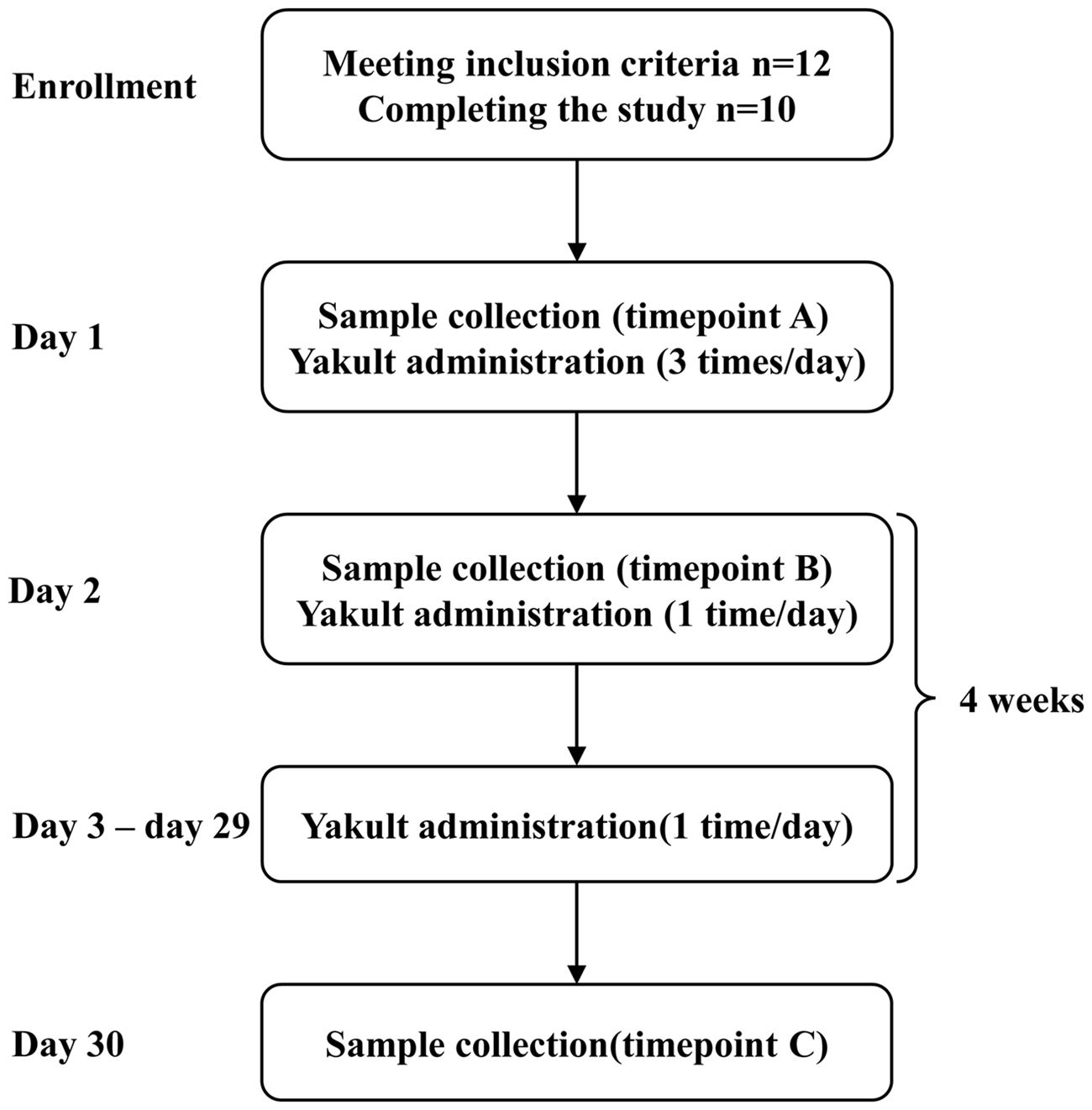
Flow chart showing study design.
Bottles of Yakult were purchased from general stores in Guangzhou, and stored at 4–6°C prior to consumption, according to the manufacturer's instruction. Each bottle of Yakult (100 ml) was purported by the manufacturer to contain 1.2 × 1010 colony-forming units (cfu) of
DNA extraction and polymerase chain reaction
DNA was extracted from plaque samples using a Powersoil® DNA isolation Kit (MO BIO Laboratories, QIAGEN GmbH, Hilden, Germany) according to the manufacturer’s instructions. Polymerase chain reaction (PCR) was used to generate amplicon libraries of the small-subunit (16S) ribosomal (r)RNA gene V4 and V5 hypervariable regions for each individual sample. PCR was performed using universal primers (forward, 5′-GTGACAGCAGCCGCGGTAA-3′ and reverse, 5′-CCGTCAATTTATTTAAGTTT-3′ [HonorTech Co., Beijing, China]) with 30 barcodes, which helped to assign sequences to different samples. The amplification mix contained 30 ng of DNA sample, 2 µl of forward primer (10 µM), 2 µl of reverse primer (10 µM), 4 µl dNTPs (2.5 mM), 5 µl 10 × Pyrobest™ Buffer (TaKaRa, Tokyo, Japan), 0.3 µl Pyrobest™ DNA Polymerase (2.5 U/μl; Code, DR005A; TaKaRa) and 36.7 µl ddH2O in a total volume of 50 µl. Cycling conditions were an initial denaturation at 95°C for 5 min, 30 cycles of denaturation at 95°C for 30 s, annealing at 56°C for 30 s, and elongation at 72°C for 40 s, followed by a final extension for 10 min at 72°C, Samples were then held at 4°C. Each sample was amplified using one specific barcoded primer. To assess quality, the PCR product for each sample was subjected to electrophoresis using a 3% agarose gel and visualized using a UVP BioDoc-IT2 UV Imaging System (Analytik Jena, Upland, CA, USA). DNA fragments were excised from the gel and further purified using QIAquick Gel Extraction Kit (250) (Cat No./ID: 28706; QIAGEN GmbH). Qubit 2.0 (Thermo Fisher Scientific, Waltham, MA, USA) was used to determine the concentration of PCR products. Finally, an amplicon library was constructed and pyrosequencing of the 16S PCR-amplicons was performed using the Illumina® MiSeq System (Illumina, San Diego, CA, USA) according to the manufacturer’s instructions.
Sequence processing
To minimize the effects of low-quality data, raw data generated from pyrosequencing was preprocessed before further analysis. First, raw data was split into individual samples according to the barcode. Then, paired reads obtained from dual terminal sequencing were assembled into a sequence according to their overlapping relationship using Connecting Overlapped Pair-End (COPE) software, version V1.2.3.3. 13 The length of the overlap sequence was 19 base pairs (bp) and the matching rate was set at 98%. The barcode and primer sequence were then removed by an in-house script. No mismatch was allowed for barcode while one bp of mismatch was allowed for primer. Spliced tags were pooled according to their lengths and chimeric sequences were removed. Finally, a statistical table of Operational Taxonomic Unit (OTU) classification level was obtained according to the data statistics. Raw sequences were deposited in the NCBI Sequence Read Archive (https://www.ncbi.nlm.nih.gov/sra) with accession numbers SRR8497113–SRR8497142.
OTU assignment and taxonomic classification
Following optimization, spliced tags were clustered into OTUs of less than 97% similarity using USEARCH software, version 8.0.1623 (https://www.drive5.com/usearch/) and the abundance of each OTU was obtained. Singleton reads, defined as reads with an abundance of 1, were removed from clustering. Sequences in the same OTU were considered to be from the same assumed taxon. Representative sequences, defined as sequences that appeared most frequently in the OTUs, were screened by UPARSE (http://www.drive5.com/uparse/) and analysed using RDP software, version 2.2 (http://pyro.cme.msu.edu/). The SILVA database, version 123 (http://tax4fun.gobics.de/Tax4Fun/ReferenceData/SILVA123.zip) was used for taxonomic classification (with similarity threshold set at 90%) and phylogenesis. Alpha diversity was evaluated using number of observed OTUs, Chao1 Index, Shannon Diversity Index and Phylogenetic Diversity whole tree analysis.
Statistical analyses
Data are presented as
Results
Twelve healthy students of Sun Yat-Sen University, aged 20-22 years, were recruited into the study. A total of 10 subjects (named 1–10) completed the full study and were included in the final analyses.
Thirty samples, 10 for each timepoint (A1–10, B1–10, C1–10) were collected and analysed. Barcoded 16S rDNA amplicon sequencing using the Illumina MiSeq platform yielded a total of 978 091 raw reads (OTU Sequences), resulting in a dataset of 256 814 reads (sample size). The number of reads per sample ranged from 5 891–12 571, with an average of 8 560. Richness and diversity analyses were based on OTUs, and clustering the unique sequences into OTUs at a 3% genetic distance resulted in 170–234 different ‘species level’ phylotypes per sample. For all of the samples analysed, the number of OTUs detected (125–234) was very close to the total number of OTUs estimated by Chao1 richness indicator (144–278). Using Good’s method, the average coverage level was approximately 99.8% in all samples, indicating that approximately two new phylotypes would be expected for every 1 000 additional sequenced reads. This level of coverage suggested that the identified 16S rDNA sequences represented the majority of bacterial types present in the current study samples.
Rarefaction curves were analysed for individual samples (Figure 2a) and for samples pooled for each of the three timepoints (Figure 2b). Rarefaction curves among the three timepoint groups displayed similar richness of species, with no statistically significant between-group differences (

Rarefaction curves for all samples comprising 10 healthy subjects at three timepoints, namely A (before intervention with Yakult, day 1), B (day 2) and C (one day following end of intervention, day 30): (a) rarefaction curves for all individual samples showing number of observed species (alpha diversity) plotted against increasing subsampled sequence sets for each sample. A trend towards a plateau was first observed following approximately 20 000 reads, suggesting this was the minimum number of reads needed per sample to accurately represent microbial diversity; (b) rarefaction curves for samples pooled for each of the three timepoints showing number of observed species (alpha diversity) plotted against increasing sequences for each pooled sample. Similar patterns of increasing species diversity were observed between individual and pooled sample sets, indicating similar phylogenetic diversity.
To determine whether microbiota from the three timepoint groups could be distinguished from each other, PCA was employed (Figure 3). This technique extracted the important information from a multivariate dataset and visualized them in an eliminated dimension way. No apparent clustering trend was observed, indicating that the overall structure of the plaque microbiota was not affected by probiotic intake.

Principal component analysis (PCA) of dental plaque microbiota in 10 healthy subjects at the three study timepoints, namely A (before intervention with Yakult, day 1), B (day 2) and C (one day following end of intervention, day 30). No clustering trend was observed in PCA space. PC1, 2, 3, principal component 1, 2, 3; Var, variance.
The effects on microbiota structure of different timepoints, subjects and Yakult consumption versus baseline control (namely ‘by timepoint’, ‘by subject’ and ‘by consumption’; Figure 4) were assessed using ANOSIM, based on Bray–Curtis dissimilarity. The resulting R values were all larger than zero (Figure 4), suggesting that dissimilarities between timepoints, between consuming Yakult and baseline control, and between subjects, tended to outweigh the corresponding within-group dissimilarities. However, only the differences between and within subjects were statistically significant (

Analysis of similarity based on Bray–Curtis dissimilarity to investigate the effects of timepoint, Yakult consumption (versus baseline, day 1) and subject, on dental plaque microbiota in 10 healthy subjects: ***
Taxonomy-based characterization of dental plaque microbiota was performed by identifying and quantifying bacterial taxa against reference databases using mothur software (https://www.mothur.org/). Plaque microbial communities were then taxonomically profiled, summarizing the relative abundances of microbial clades at different taxonomic levels (Figure 5, showing phylum and genus levels only). All sequences were found distributed in 23 phyla, 48 classes, 82 orders, 113 families, 184 genera and 253 species. The five most abundant phyla were
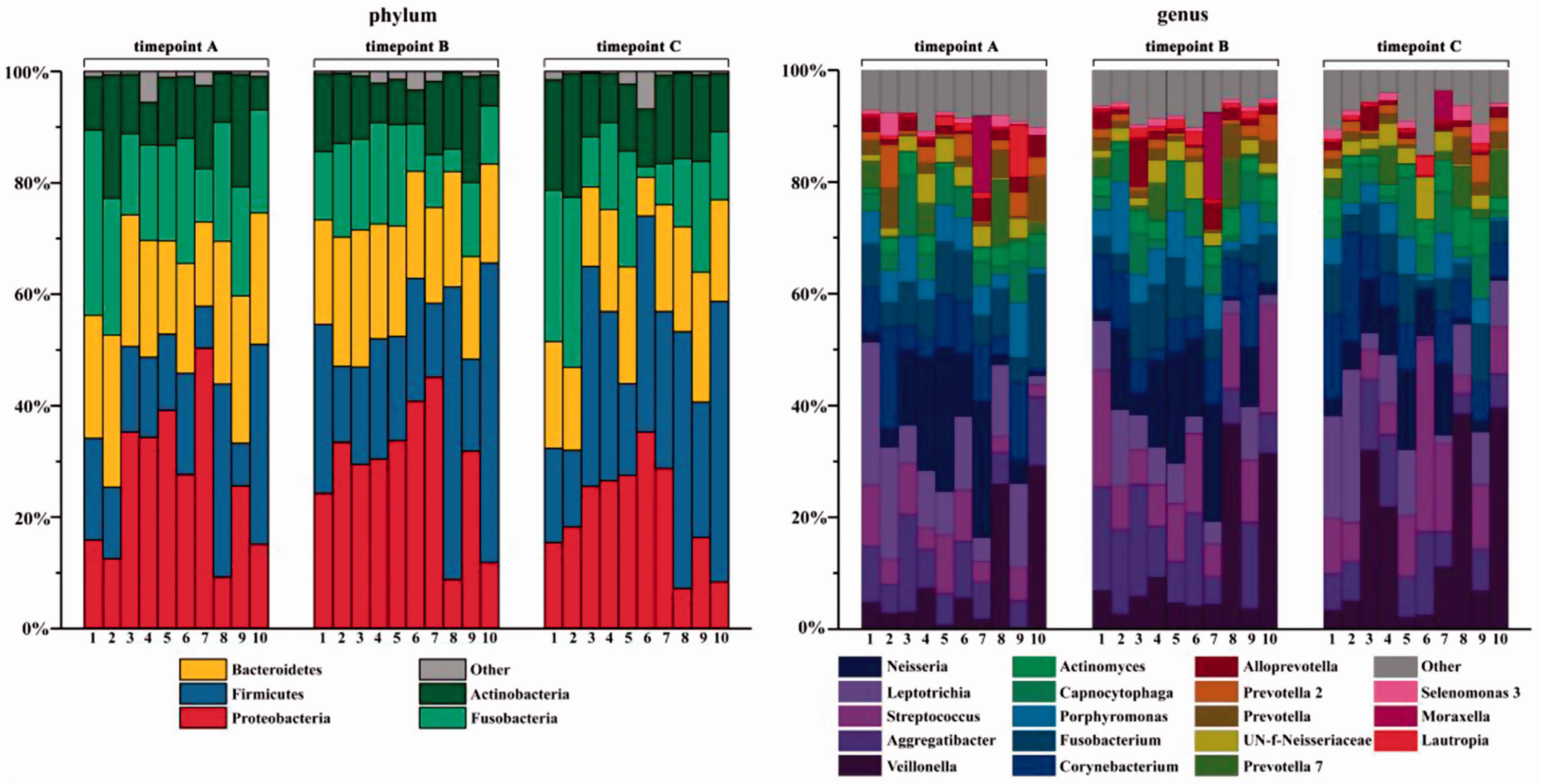
Taxonomic composition of dental plaque samples from 10 healthy subjects at different timepoints (A, before intervention with Yakult [day 1]; B, day 2 during intervention; and C, one day following end of intervention [day 30]), displayed as relative abundance according to phylum and genus. At phylum level, relative abundances highlighted the major roles of
To study changes in the composition of dental plaque microbiota following Yakult intake, the plaque microbiota was compared between the three timepoints (Figure 6). Among five dominant phyla,
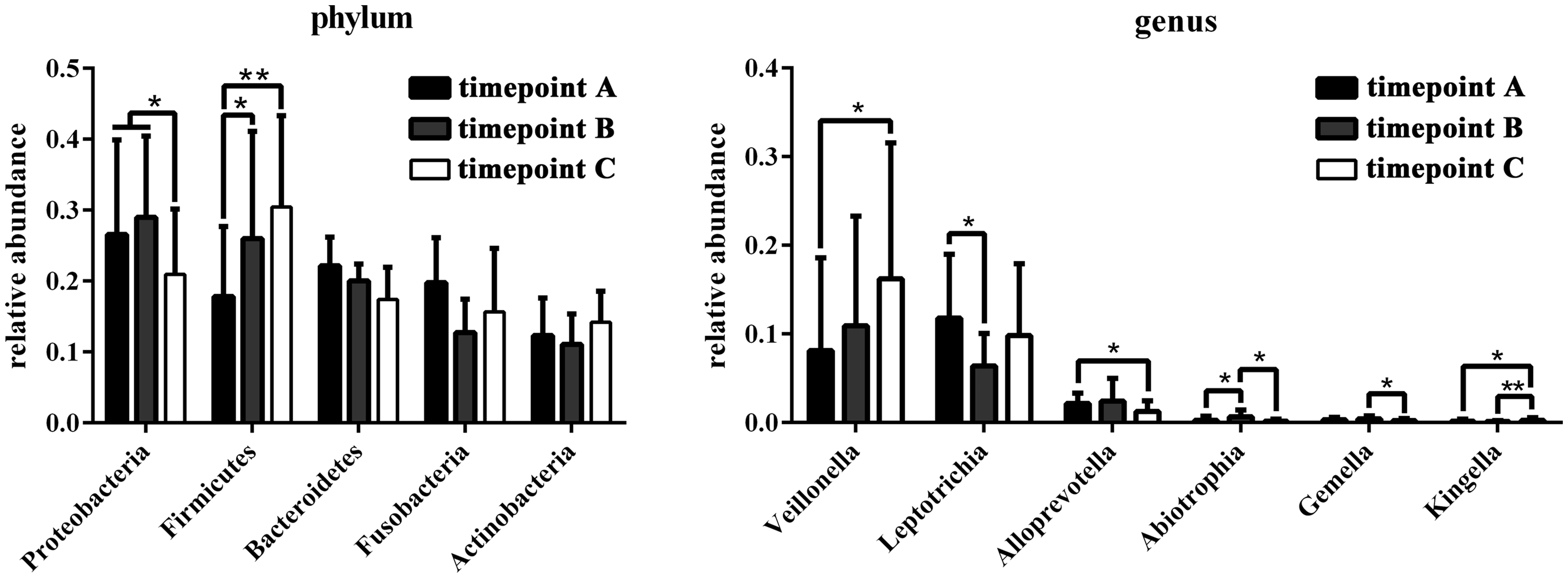
Comparison of bacterial phylum and genus profiles between three timepoint groups of dental plaque samples from 10 healthy subjects: Timepoint A, before intervention with Yakult (day 1); timepoint B (day 2 during intervention); and timepoint C, one day following end of intervention (day 30). Data presented as mean ± SD relative abundance of dominant taxa (>1% for phylum and > 0.1% for genus) between three timepoint groups (*
Discussion
Modulation of resident microbial communities by consumption of probiotics has become an appealing means for host health promotion.
14
Yakult, a well-known probiotic beverage, contains a single live bacterial species,
The oral cavity has also been a target site for testing various strains of probiotics.10,14 In the present study, it was hypothesized that Yakult consumption may change the structure and composition of dental plaque microbiota, which was assessed using 16S rRNA sequencing. Data analyses revealed changes in the abundance of some bacteria that have been linked with the presence or absence of caries, while the overall microbiota structure remained unaffected. In particular, changes were noted in some genera associated with caries, including
Many studies have investigated the effects of
Since the majority (>90%) of microbial species cannot be readily cultured using current laboratory culture techniques,
36
conventional plate culture and colony counting are insufficient in microbiota analysis. In order to gain a deeper insight into the plaque microbiota, the present study engaged 16S rRNA sequencing. As the most common sequencing approach to analyse microbiota, it has been used to compile most of the data collated by the Human Microbiome Project.
36
The major phyla found in the present study samples were
As the first study to investigate the effects of
In conclusion, Yakult intake was found to change the abundance of some bacteria related to caries, while the overall microbiota structure remained unaffected. The compositional change may be beneficial for oral health, but further analyses of biofilm acidogenicity and other enzymatic activity changes are needed to verify whether continued intake of probiotics for longer periods may have a significant impact on dental plaque microbiota.
Footnotes
Declaration of conflicting interest
The authors declare that there is no conflict of interest.
Funding
This work was supported by the National Natural Science Funding of China (grant No. 11772361).
