Abstract
The Coronavirus 2019 pandemic has strained nearly every aspect of pathology practice, including preanalytic, analytic, and postanalytic processes. Much of the challenges result from high demand for limited severe acute respiratory syndrome coronavirus 2 testing capacity, a resource required to facilitate patient flow throughout the hospital system and society at large. At our institution, this led to unprecedented increases in inquiries from providers to laboratory staff relating to the expected time to result for their patients. The demand was great enough to require redeployment of staff to handle the laboratory call volume. Although these data are available in our laboratory information system, the data do not interface to our electronic health record system. We developed systems using the R statistical programming language that abstract the necessary data regarding severe acute respiratory syndrome coronavirus 2 polymerase chain reaction testing from our lab system in real time, store it, and present it to clinicians for on demand querying. These data have been accessed over 2500 times by over 100 distinct users. Median length of each user session is approximately 4.9 minutes. Because our lab information system does not persistently store tracking information while our system does, we have been able to iteratively recalculate time to result values for each tracking stop as workflows have changed over time. Facility with informatics and programming concepts coupled with clinical understanding have allowed us to swiftly develop and iterate on applications which provide efficiency gains, allowing laboratory resources to focus on generating test results for our patients.
Keywords
Introduction
The coronavirus 2019 (COVID-19) disease caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), first identified in late 2019 in Wuhan, China, quickly reached pandemic levels spreading across the globe. As a result, the virus has caused heretofore unseen levels of stress on health care facilities and society at large. The sheer volume, acuity, and virality of the disease has required strained hospitals to ration limited resources, including beds, lifesaving equipment, diagnostic tools, particularly testing for SARS-CoV-2, as well as health care workers and protective equipment required for their involvement in care. 1
Limitations of SARS-CoV-2 diagnostic testing are distinctly problematic as it is critical for the adequate allocation of many other resources. Patients whose results are pending must remain hospitalized, occupy scarce bedspace, and require care from personnel using limited personal protective equipment. Often those with pending results are health care personnel who must quarantine as they await results from potential exposures. 2
Insufficient availability of SARS-CoV-2 testing has manifested primarily as extended lengths of turnaround time for results. The ramifications of delays in turnaround time include consequences both within and without the hospital setting. In the hospital, this can cause bottlenecks in patient flow 3 and delays in surgical procedures, 4 while in society at large testing is required the safe return to work and school. 5 The intractable nature of the issue has led the federal government to take action to spur efforts to improve this metric, most recently by reducing CMS payments for testing that extends beyond 2 calendar days.
Given the critical implications of results, it is not surprising that the lab receives many inquiries regarding the expected time to result for this testing. In more typical circumstances inquiries related to test status are a large fraction of all questions to laboratory client services departments. In times such as these where results are so critical for the functioning of health care systems and public life in general, the role of disseminating this information critical for patient outcomes, public health, and the functionality of society.
During the course of the pandemic, our infectious disease diagnostic laboratory (IDDL) rapidly expanded the scale of SARS-CoV-2 testing. However, the need for testing correlated with rapid increases in inquiries to the lab from frontline clinicians requesting information regarding the status of testing. Although automation was available to manage the analytical testing for SARS-CoV-2, the process of querying our lab system for a specific test and pulling its status was entirely manual. Here, we describe how we leveraged open-source technology to create a tool that provides this critical information for SARS-CoV-2 polymerase chain reaction (PCR) testing to clinicians in order to facilitate patient flow through the hospital, in particular the emergency department and surgical procedures.
Materials and Methods
Data Architecture
Data were queried from our enterprise data warehouse and our laboratory information system (LIS), parsed, and fed into 2 data marts, one containing SARS-CoV-2 results data and one containing SARS-CoV-2 order status data. This intermediate step simplifies data flow into the dashboards. Data extraction and processing was performed using the R statistical programming language. 6 Data were displayed on dashboards built using the Shiny R package. 7 Dashboards were deployed and data processing automated on an RStudio Connect service (RStudio), which likewise handled security and user authentication. Two apps were created, one for the ED app which went live on April 1, 2020, and another for the anesthesiology group which handles preoperative medical clearance which went live on April 15, 2020. See Figure 1 for architecture of data flow.
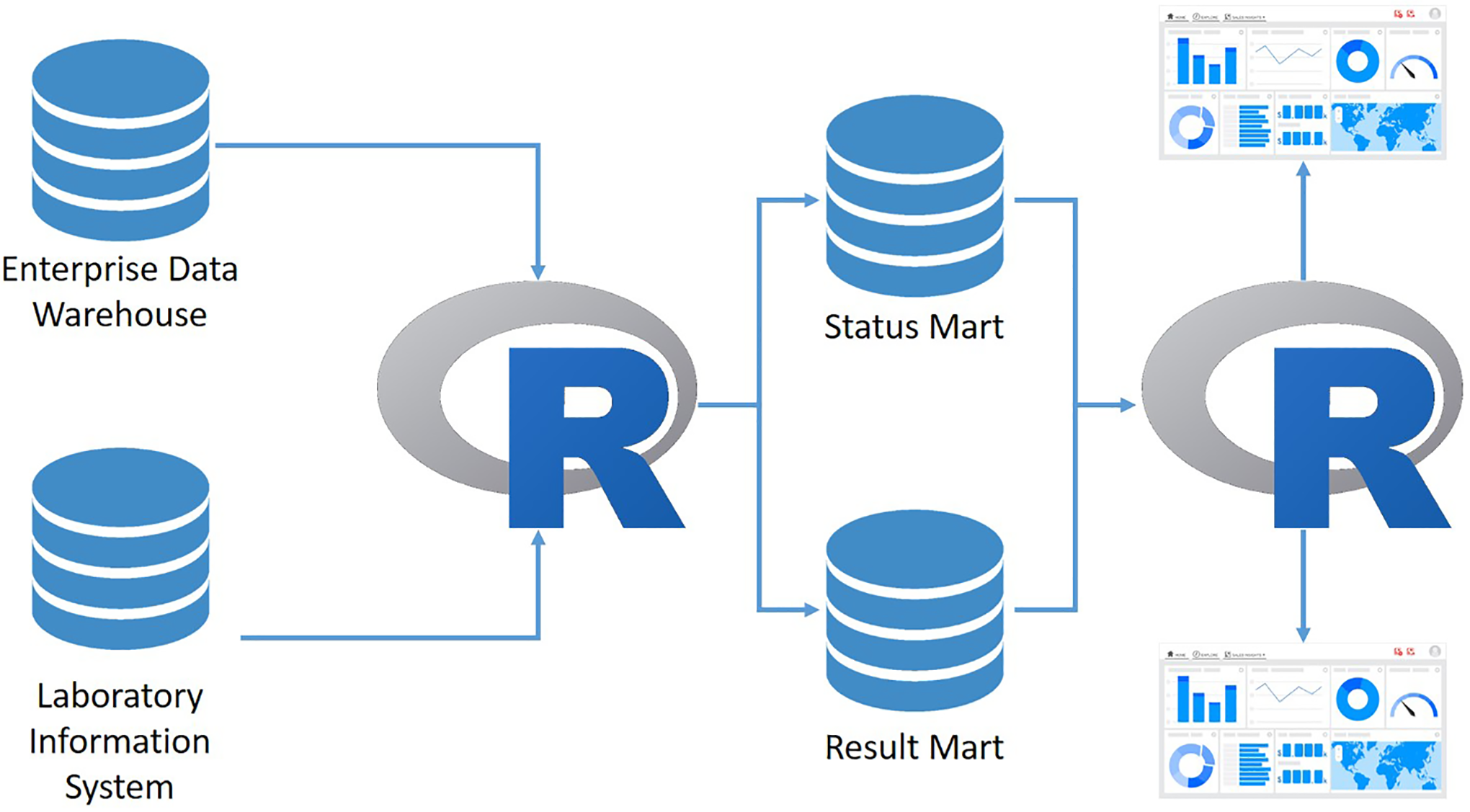
High-level back-end data architecture for 2 apps displaying test order status. Processes written in R move data between the enterprise data warehouse, the lab system, and 2 data marts used to store data. Separate R processes serve dashboards deployed on an RStudio Connect server to users. Database icons represent stores of data, arrows represent data flows.
User Data Acquisition, Processing, and Analysis
User level data were acquired using the shinylogs R package 8 and stored in a sqlite database. Data were accessed, analyzed, and plotted using the R statistical programming language. Raw data were minimally processed prior to analysis; the anesthesia app experienced an error for specific users causing the app to immediately fail upon loading, these entries were removed prior to analysis.
Results
Our electronic health record (EHR) system, Epic (Epic Systems Corporation) provides high-level information regarding the status of a pending lab order, including the time of specimen collection; however, the granularity of the data was insufficient to guide clinical decision-making. Our LIS Soft (SCC Soft Computer) contains far more granular information as to the point in which a specimen is in the analytical workflow; however, interfacing this data to the EHR is not possible. The median turnaround time for the test from time of collection to result is 8.6 hours (interquartile range [IQR] = 6.13-12.5) a time interval sufficient for most clinical purposes but marginal for efficient flow in certain high-acuity clinical areas. The result of this was unprecedented call volumes to the IDDL that proved disruptive to clinical diagnostic work. Although records were not kept throughout the pandemic as to the number of calls received, over several weeks in April these averaged approximately 8 calls per hour. Because staffing was insufficient to handle the quantity of calls, both clinician and staff satisfaction suffered.
We therefore set out to create a custom solution to provide a lab status portal for clinicians to access the status and expected time to result for any given SARS-CoV-2 PCR order. Two apps were created, one specifically for the patients seen in the emergency department and the second for the anesthesia group that facilities preprocedure screening of patients. Both apps used the same data sources and back-end infrastructure (Figure 1), though they differed slightly in front-end functionality as they were tailored for the requirements of each group. For demonstrative purposes, the anesthesia app is presented in Figure 2. The app provides 2 mechanisms for viewing test status (1) through a medical record number lookup tool (Figure 2A) and (2) by browsing a predefined list of patients scheduled for operative procedures within the next 3 days (Figure 2B). In the case of the ED app, this list contained the status for all SARS-CoV-2 tests originating from the ED over the previous day. Focus was placed on ease of use; the app landing page provides direct access to the first mechanism for pulling test status, and the predefined list of patients is easily searchable by common parameters, for example, MRN, date of procedure, and status of test. Test status information is provided as a per patient customized message specifying the expected time range within which the result should be available, dependent on the status of the test within the workflow, which was defined by tracking stops electronically recorded in the LIS. For example, if our LIS indicated that a patient’s sample had been loaded on the PCR instrument at 12
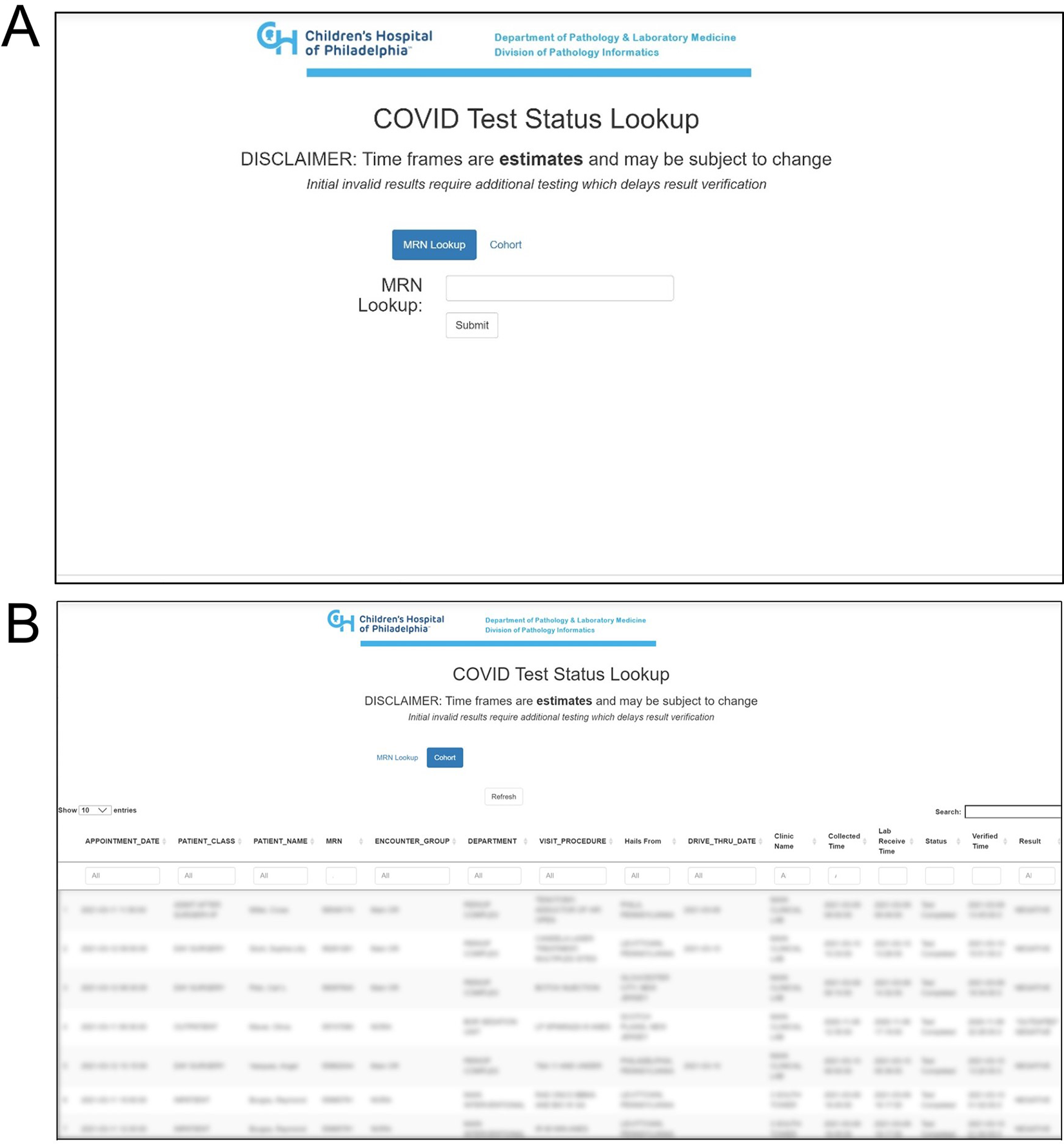
User facing front end of the anesthesia app. Two mechanisms are available for extracting the status of a test. (A) A medical record number look-up tool and (B) a predefined list of patients.
Through 7 months of use, the apps were accessed 2524 times by 107 distinct users. By far the most common user was a nurse, representing 65 of the 107 users. After nurses, lab personnel (n = 21) and physicians (n = 12) were most frequently represented followed by a variety of other administrative staff. Nurses also made the most frequent use of the apps (a disproportionally high 86.2% of all interactions) followed distantly by physicians (6.1%) and lab personnel (5.7%). Use of the app was monitored on a per user session basis. Since going live, relative use of the apps has been essentially constant. Other than high levels of usage of the ED app in April (P < .001 by analysis of variance), no other linear trends or differences in average daily use was identified for either app (Figure 3A). This high level of usage early in the pandemic likely related to overall lack of clarity around testing and hospital processes at that time.
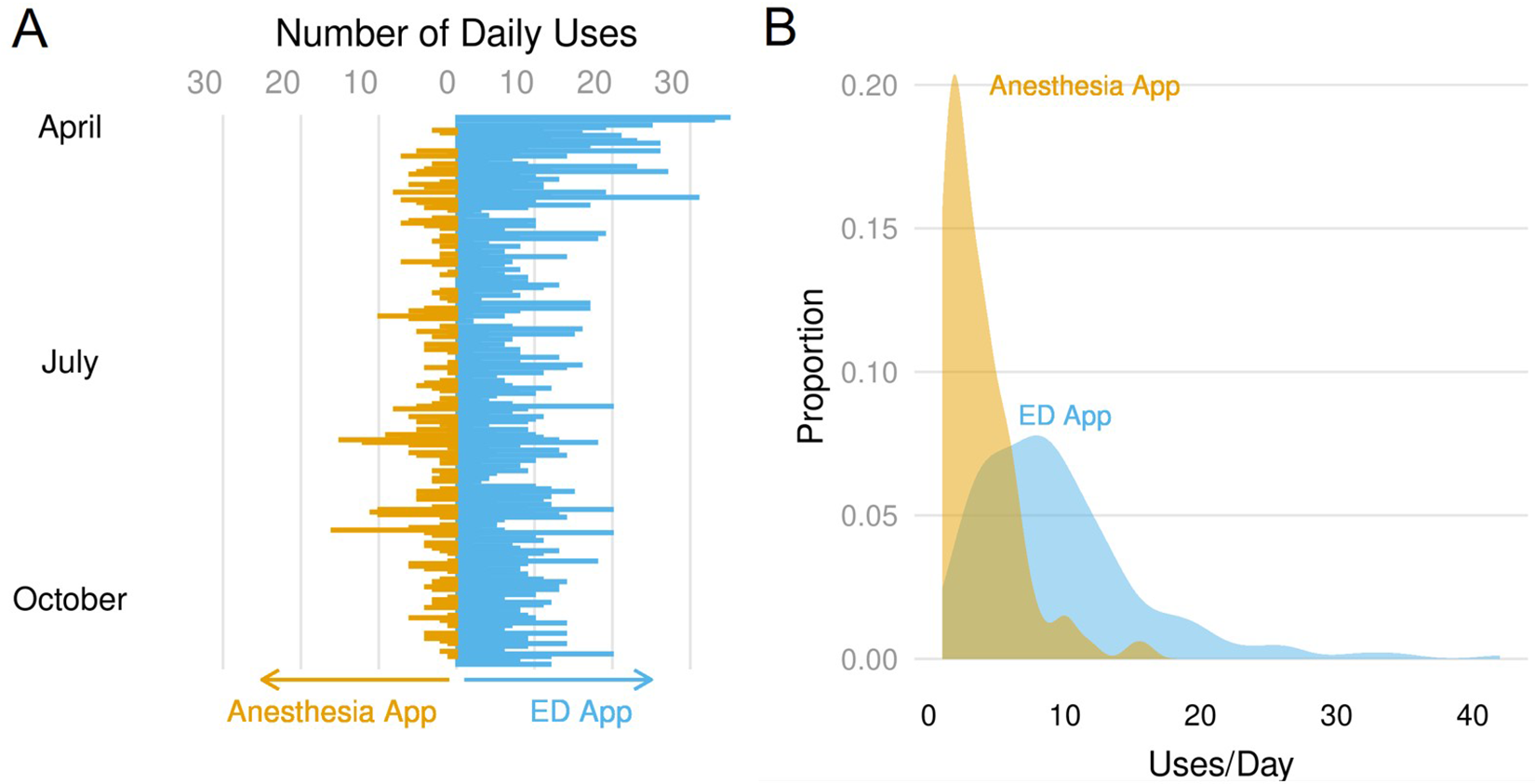
Daily use and distribution of app use over time. A, Counts of daily app use over time. Blue bars represent use of the ED app, while the orange bars represent use of the anesthesia app. The former app went live on April 1, 2020, while the latter went live on April 15, 2020. B, Density plot of distribution of daily use by app. Blue plot represents use of the ED app while the orange plot represents use of the anesthesia app.
Nor was there any statistically significant relationship between daily app use and daily volume of SARS-CoV-2 testing. Regression analysis with daily test volume and app use as independent and dependent variables, respectively, showed R 2 values less than 0.1 and P values >.05 for both apps. These results suggest that app use rate was not dependent on high volumes of test orders.
The apps found quite different patterns of use. Daily use was higher for the ED app (median = 8, IQR = 5-12) than the anesthesia app (median = 3, IQR = 2-5), with a statistically significant difference in median daily interactions (5 uses, P < 3 × 10−16 by Wilcoxon rank-sum test; Figure 3B).
Distinct users were essentially evenly split among the apps with 61 users of the ED app and 53 users of the anesthesia app; however, 2020 interactions came from the former with the remainder from the latter. These data indicate that on a per user basis the ED portal received over 3 times as many visits as the anesthesia app. That said, Pareto analysis demonstrated that while the ED app was used many more times on average per user, the utilization of the anesthesia app was focused in fewer users (Figure 4). Six users accounted for approximately 50% of the interactions with this app, whereas in the ED, use was spread more evenly across a larger quantity of users with 10 users accounting for the same 50% of interactions.
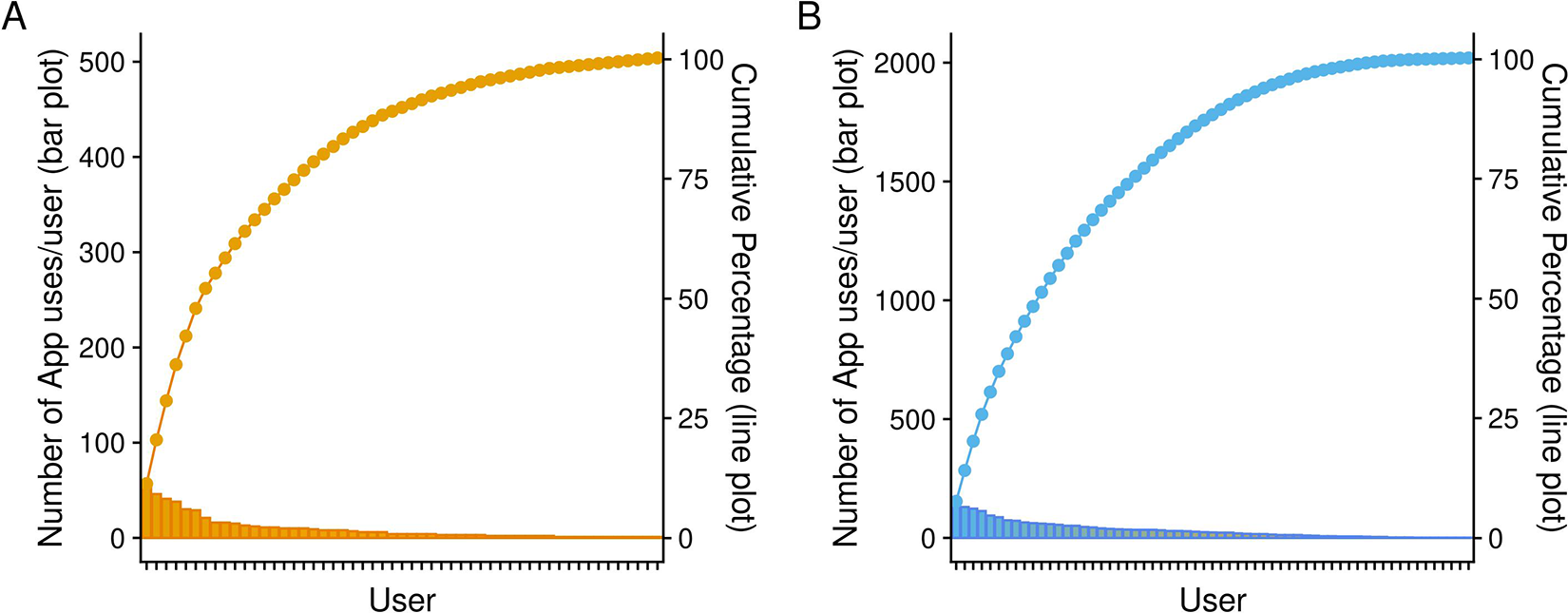
Pareto plots demonstrating distribution of app use by user. Each user of each app is represented on the x-axis. The bar plots represent the number of interactions for a given user and correlate with values on the left y-axis. The line plots represent cumulative contribution of users to the total. A, Number and proportion of interactions with the anesthesia app per user. B, Number and proportion of interactions with the ED app per user.
To further elucidate the patterns of use for the apps, we examined the length of time between app visit and app close for each interaction with each app (Figure 5A). Figure 5B focuses on sessions with time on site below 250 minutes and highlights the difference in pattern of usage where the majority of sessions fall.
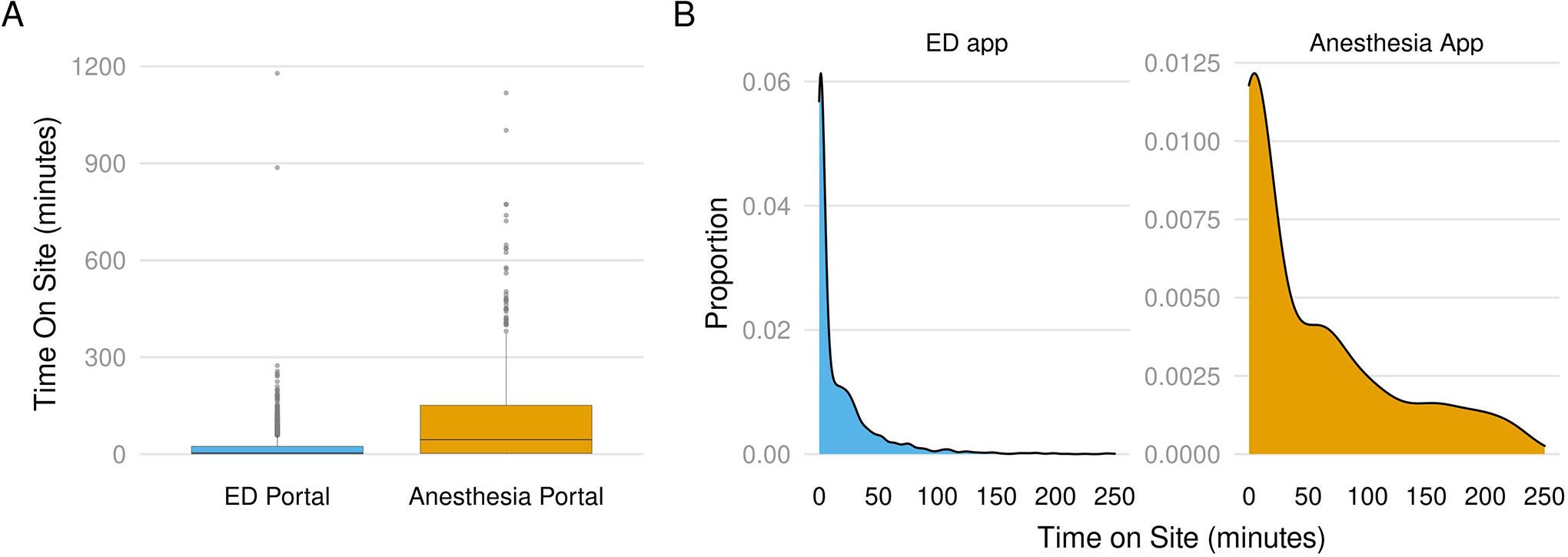
Distribution of time spent using the apps. A, Box plots demonstrating distribution of session time length (in minutes) for each app. Blue box plot represents use of the ED app, while the orange box plot represents use of the anesthesia app. B, Density plots focusing on sessions under 250 minutes. To focus on the bulk of user sessions, which were on the low end of the boxplots, density plots were constructed and the x-axis limited to time intervals below 250 minutes. Blue plot represents use of the ED app, while the orange plot represents use of the anesthesia app.
Although the ED App was used more frequently than the anesthesia app, the interval of time on site/session revealed significantly different patterns of use. The data demonstrate that while the ED app usage showed a predominance of frequent, short sessions (median server time = 3.9 minutes), the anesthesia app showed a wider distribution of session lengths (anesthesia app time, IQR = 2.48-151; ED App time IQR = 0.9-24.1) and a significantly different median interaction length (44.9 minutes, P < 3 × 10−16 by Wilcoxon rank-sum test). Also notable was a long tail of a small number of user sessions that were more lengthy than would be expected in typical use, some stretching beyond 12 hours. These likely represented sessions that were inadvertently left open overnight. Although the data likely do not reflect the typical amount of time required to query status of an individual test, they do provide additional information regarding overall patterns of use in different health care settings.
The consistent use of the apps over an extended period suggests depicted in Figure 3 lends support to the utility of the apps. Further subjective user feedback was collected during the development and optimization of the apps, which was universally positive. Users expressed that the apps were “truly very helpful,” “really awesome,” “saving…from a great deal of redundant work,” and “very helpful to streamline care.”
Discussion
As the COVID-19 pandemic developed in early spring 2020 it quickly became clear that the role of the laboratory in the health care system and in society was evolving. It is often cited that the results from laboratory testing play a part in the large majority of clinical decision-making. As a result of the pandemic, the role of testing for SARS-CoV-2 became an absolute prerequisite for decisions in patient admissions and discharges, scheduling of invasive procedures, and many public health measures governing the integration of exposed and symptomatic individuals back into everyday life. Given the stakes involved, it was therefore predictable that demand for information around testing would increase.
We developed 2 apps to help mitigate this issue by facilitating access to information found within our lab system and publishing it in a secured but accessible location. Our analysis demonstrated significant differences in user behavior when interacting with these apps. An important observation was the significant difference in length of user sessions. Upon discussions with the different user groups, it was revealed that in the ED, the app is used primarily for “grab and go” data pulls, whereas the anesthesia group interacts in a “sit-down” style, uses the app alongside the EHR in working though lists of patients scheduled for procedures in the operating rooms. Implementing a means to monitor usage also proved helpful during the optimization process of the apps. It allowed us to identify champions who could provide feedback on the apps and reflect on outstanding challenges not met by the functionality. Such individuals fall into a class of “Special People” who serve important roles in the success of clinical IT adoption 9 and were very helpful in ensuring that the implementation was a success.
What forced us to undertake this work was the reality that information from our lab system could not be integrated into our EHR. Fundamentally, this was due to limitations in the interfaces, the electronic connections which allow data to flow between these systems. The reliance upon and limitations of interfaces in health care are often cited as bottlenecks interrupting critical data flow needed for decision making. 10 This federated data architecture is often called the “best-of-breed” approach in application selection because it emphasizes the selection of individual systems with optimum functionality. By contrast, the enterprise-wide solution (EWS) emphasizes a single integrated product without the constraints of interfaces. Nevertheless, even in an EWS, the data provided by our apps may not be available if the system itself was not built in a way that makes it accessible. 11
The choice to develop a custom solution for a functional information technology gap is a controversial one. 12 Homegrown applications have to be maintained, bugs must be corrected, enhancements must be prioritized, integrated, and deployed in an orderly fashion. There is risk in that the developer may leave the organization, orphaning the application. Nevertheless, we believe that clinical requirements must be the primary arbiter as to whether to buy, build, or do without and have therefore invested in personnel with informatics expertise and tools to support development of solutions for our department. In the future, as cloud platforms which significantly reduce challenges in app development become commonplace in health care, pathology departments with a trained workforce to take advantage of these resources will be far better equipped to meet the unmet needs of their workflows.
Although we are pleased with the utility that the apps have provided, we have also noted several limitations. A significant drawback of our approach is that it requires the user to leave the EHR and use an independent application. Electronic health records have become to a large degree the hub of clinical workflows, particularly in inpatient care, making this an undesirable limitation on our apps’ usability. The user effort involved in leaving one workspace for another likely weighed against more robust adoption. Various mechanisms exist to support incorporation of third party applications into the EHR, ranging from simple links to incorporation of security, access, and data exchange. Because of the importance of usability on the value of health IT, we will be devoting efforts to laying the groundwork for more robust integration for future apps.
Additionally, while the lab system was able to provide the status of a lab order, the estimation of time to result was a variable that was set manually. These estimates potentially can change over time which would require making continual upgrades to the apps. In practice, we did not find variation in these estimates (data not shown) so this variable did not need to be adjusted. Lastly, this approach was tenable because we had a small number of platforms for SARS-CoV-2 testing. In institutions with many redundant platforms, the complexity of the data may have impeded developing similar applications.
In conclusion, in response to an overwhelming demand for information around SARS-CoV-2 testing, we quickly developed and deployed apps directed at specific high utilizer, clinically critical groups. This was made possible by our department’s investment in informatics and programming expertise within faculty and staff, allowing our medical technologists to focus efforts on clinical testing for our patients.
Footnotes
Acknowledgments
The authors would like to thank Tara Dea, Ari Weintraub, Keri Cohn, Joe Zorc, and Leslie Obermeier for providing useful feedback on the apps. The authors would also like to thank Tim Back, Marianne Henin, and Caroline Burlingame for assisting in querying data. The authors would like to acknowledge Jane Koller and Chris Foehl for volunteering to answer calls from clinicians related to SARS CoV-2 testing. Finally, the authors would like to thank the IDDL staff for tirelessly providing critical results for our patients in trying times.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The article processing fee for this article was funded by an Open Access Award given by the Society of ‘67, which supports the mission of the Association of Pathology Chairs to produce the next generation of outstanding investigators and educational scholars in the field of pathology. This award helps to promote the publication of high-quality original scholarship in Academic Pathology by authors at an early stage of academic development.
