Abstract
Background: Neonatal sepsis is the third leading cause of neonatal mortality, behind prematurity and intrapartum-related complications. The main objectives of this study are to assess the proportion of sepsis in preterm newborns and identify the etiologic agents and their antibiotic sensitivity patterns. Methods: A longitudinal observational study was done from July 2016 to May 2018. Whenever clinical diagnosis of sepsis was made, blood cultures and antibiotic susceptibility tests were done. Result: We did 690 blood cultures, 255 (36.9%) showing bacterial growth. The most commonly isolated bacteria were Klebsiella species 78 (36.6%), Coagulase negative Staphylococcus 42 (19.7%) and Staphylococcus aureus 39 (18.3%). Gram-positive bacteria showed high resistance to penicillin (98.9%) and ceftriaxone (91.3%) whereas Gram-negative bacteria were highly resistant to gentamicin (83.2%) and ceftriaxone (83.2%). Conclusion: Resistance to the more commonly used antibiotics such as ampicillin and gentamycin was very high, necessitating reconsideration of the empiric use of these antibiotics.
Background
Every year, 2.6 million neonates die globally; three fourths of these deaths occur in the first weeks of life, and almost all (99%) occur in low- and middle-income countries (LMICs).1,2 Neonatal sepsis is the third leading cause of neonatal mortality, only behind prematurity and intrapartum-related complications (or birth asphyxia). 3 The World Health Organization (WHO) estimates that one million deaths per year are due to neonatal sepsis and that 42% of these deaths occur in the 1st week of life. 2 In addition, the survivors of neonatal sepsis are vulnerable to short and long-term neuro-developmental morbidity.4-6
Neonatal sepsis is a life-threatening condition and needs immediate empirical antimicrobial therapy to reduce mortality. It is important to choose an antibiotic regimen that covers the most common pathogens. 7
Blood culture remains the gold standard for diagnosis of neonatal sepsis, despite its low sensitivity which may be due to small volume of the blood sample, or the use of empirical antibiotics prior to sampling. 8 Neonatal sepsis is classically defined as the presence of symptoms of sepsis in the neonatal period combined with bacteriological isolation of an infectious agent from blood or cerebrospinal fluid (CSF). 9 Early-onset neonatal sepsis refers to the presence of a confirmed infection in the blood or CSF of patients younger than 3 days of life, and late-onset neonatal sepsis refers to the onset of such infections between 3 and 28 days. 10
The type of organisms causing sepsis varies from one region to another and changes over time even in the same place.11,12 This variation is often attributed to the changing pattern of antibiotic use and changes in lifestyle.13,14
In developing countries, clinically diagnosed sepsis occurs in 49-170 newborns per 1000 live births, whereas culture-proven sepsis occurs in 16 per 1000 live births. 15 The continuous evolution of drug resistance of pathogens causing neonatal sepsis is a major problem. Notably, methicillin resistant S. aureus (MRSA) and extended spectrum beta lactamase (ESBL) producing bacteria and multidrug resistant (MDR) Gram-negative organisms represent the principal setbacks to fighting infections. Most Gram-negative bacteria are now resistant to ampicillin and many are also becoming resistant to gentamicin.16,17
Although antimicrobial treatment is the cornerstone of the treatment of neonatal sepsis, due to the lack of local data on bacterial profiles and antimicrobial resistance patterns, there is a lack of clear consensus on the choice of antimicrobials for many settings in LMICs. Therefore, clinicians in LMICs are limited to use the data from the other settings, mostly from developed countries, to inform their treatment decisions.
Objectives
The primary objective of this study was to assess the proportion of clinical and bacteriologically confirmed sepsis among preterm newborns admitted to selected hospitals in Ethiopia.
The secondary objectives of this study include:
To determine common organisms causing sepsis in preterm newborns
To determine antimicrobial susceptibility of the identified microorganisms
Methodology
Study Design
A multi-center longitudinal observational study design was used which included all preterm babies (<37 weeks of gestation) who were either delivered in or were referred to the study hospitals before 7 days of life from July 2016 to May 2018. Preterm newborns admitted at >7 days of life were excluded from the study as the original study used Ballard score (which is accurate before 7 days of life) to accurately estimate the gestational age of the babies for inclusion into the study. Thus, for this analysis, we included those neonates who were admitted to a NICU at <7 days. The preterm babies admitted to the NICUs were identified based on the study eligibility criteria by trained nurses. After screening, written informed consent was obtained from parents before enrolling the babies. The maternal medical history and physical examination including gestational age estimation was done by trained study physicians. Once a clinical diagnosis of sepsis was made (based on clinical signs including temperature instability, respiratory distress, decreased age appropriate neonatal reflexes, altered mentation, abnormal body movement or poor feeding), patients often underwent necessary laboratory investigations including complete blood count (CBC), blood culture, and CSF analysis and culture to establish the diagnosis of sepsis. Other investigations were also done as clinically indicated. 18
Study Settings
The study was conducted in five hospitals in Ethiopia: Tikur Anbessa Specialized Hospital (TASH), St. Paul’s Millennium Medical College (SPMMC) and Ghandi Memorial Hospital, located in Addis Ababa, and Jimma University Medical Centre (JUMC) and Gondar University Hospital (GUH), located in the southwestern and northwestern parts of the country, respectively.
Ethical Approval
Addis Ababa University and all participating institutions provided ethical approval of the study. All parents were provided written informed consent prior to their infants’ participation in the study.
Laboratory Analysis
Sample Collection, Isolation and Identification of Bacteria
At all hospitals, a minimum of 1ml of blood was drawn aseptically from the neonates with suspected sepsis using a butterfly needle. At Jimma and St. Paul Hospitals, the blood was added into BD BACTEC Peds Plus blood culture bottle and gently mixed. The blood culture bottle was then incubated in the BD BACTECTM FX40 machine which was inspected daily for a total of 5 days. When an alert was observed, the BACTEC bottle was taken from the machine, and sub-cultured on MacConkey Blood agar (Oxoid, England), and Chocolate Agar (candle jar) media and incubated at 35-37°C for 18-24 hours aerobically. At TASH and GUH, the blood specimens were collected and inoculated on brain heart infusion broth and inspected for five days. At all hospitals, further bacterial identification was performed based on the American Society for Microbiology (ASM) identification guidelines.
Antimicrobial Susceptibility Testing
Antibiotic susceptibility testing was carried out using the Kirby Bauer disc diffusion technique using Muller Hinton agar (Oxoid, England). Antibiotic discs were selected based on CLSI guidelines. Accordingly, penicillin, cefoxitin, ampicillin, erythromycin, clindamycin, trimethoprim-sulphamethoxazole, oxacillin, amoxicillin clavulanic acid, ceftazidime, gentamicin, ciprofloxacin, and chloramphenicol were used. All the antimicrobials used for the study were obtained from Oxoid Ltd., Basingstoke, Hampshire, UK. The drug susceptibility results were measured using calipers and based on zone of inhibition; the results were reported as resistant, intermediate and susceptible to the treating clinicians so that they could modify the treatment accordingly.
Multi drug resistance (MDR) was defined as bacterial strains resistant to at least one agent in three or more antimicrobial classes.
Quality Control
Data were checked and reviewed regularly for any incompleteness or outliers and any issues were rectified by the site principal investigators before entry into the database. Reference strains of Escherichia coli ATCC 25922 and S. aureus ATCC 25923 were used for media and antibiotic disk performance checks.
Data Analysis and Interpretation
Data were collected using Case Report Forms (CRF) that were formulated and reviewed by various physicians in their respective field of expertise. The data were entered into a computer using the data management system (DMS) developed for this study. Finally, electronic data were transferred, cleaned, merged to one master data and stored centrally using the Structured Query Language database that was later analyzed in SPSS version 20. Frequencies and cross tabulations were used to summarize descriptive statistics. Descriptive statistics were also used to present antimicrobial susceptibility patterns.
Results
Clinical Characteristics of the Participants
Among all preterm newborns (n = 4,919) enrolled into the study, 2,003 had a clinical diagnosis of sepsis, making the overall proportion of clinical sepsis 40.7%. Of those diagnosed with clinical sepsis, 1,807 (90.2%) had early onset neonatal sepsis (EONS) and 196 (9.7%) had late onset neonatal sepsis (LONS) (Figure 1).
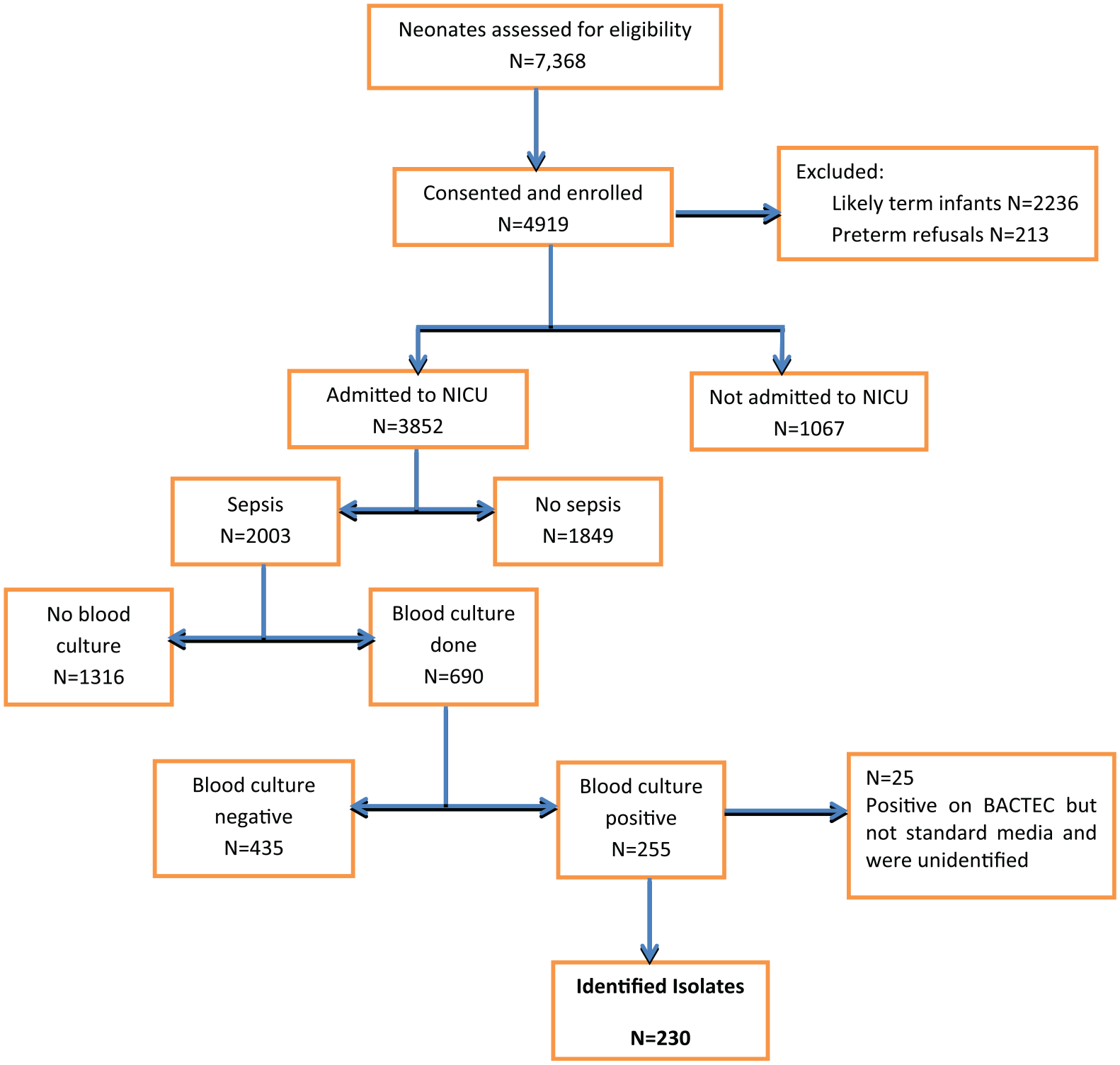
Consort flow diagram.
The majority of those with clinical sepsis were male (53%) and 905 (45%) were female. There were more early (<34 weeks) preterm newborns with a diagnosis of clinical sepsis, 1,071 (53.4%) than late preterm newborns (34-36 weeks), 932 (46.5%). Most were delivered in health institutions (1893,94.5%) either in the study hospitals or health centers.
Of the study participants with neonatal sepsis, only one-third, 690 (34%) had blood cultures done, of which 255 (36.9%) showed bacterial growth (Table 1). Nineteen different bacterial pathogens were identified. Gram-negative bacterial species were the most commonly isolated organisms 134 (58.2%) compared to Gram-positive organisms 90 (39.1%). The most commonly isolated bacteria were Klebsiella species (spp.) 78 (33.9%) followed by Coagulase negative Staphylococcus (CoNS) 42 (18.2%), S. aureus 39 (16.9%) and E. coli 12 (5.2%).
Bacteria Identified in Preterm Newborns Admitted with a Diagnosis of Sepsis to Selected Hospitals in Ethiopia.
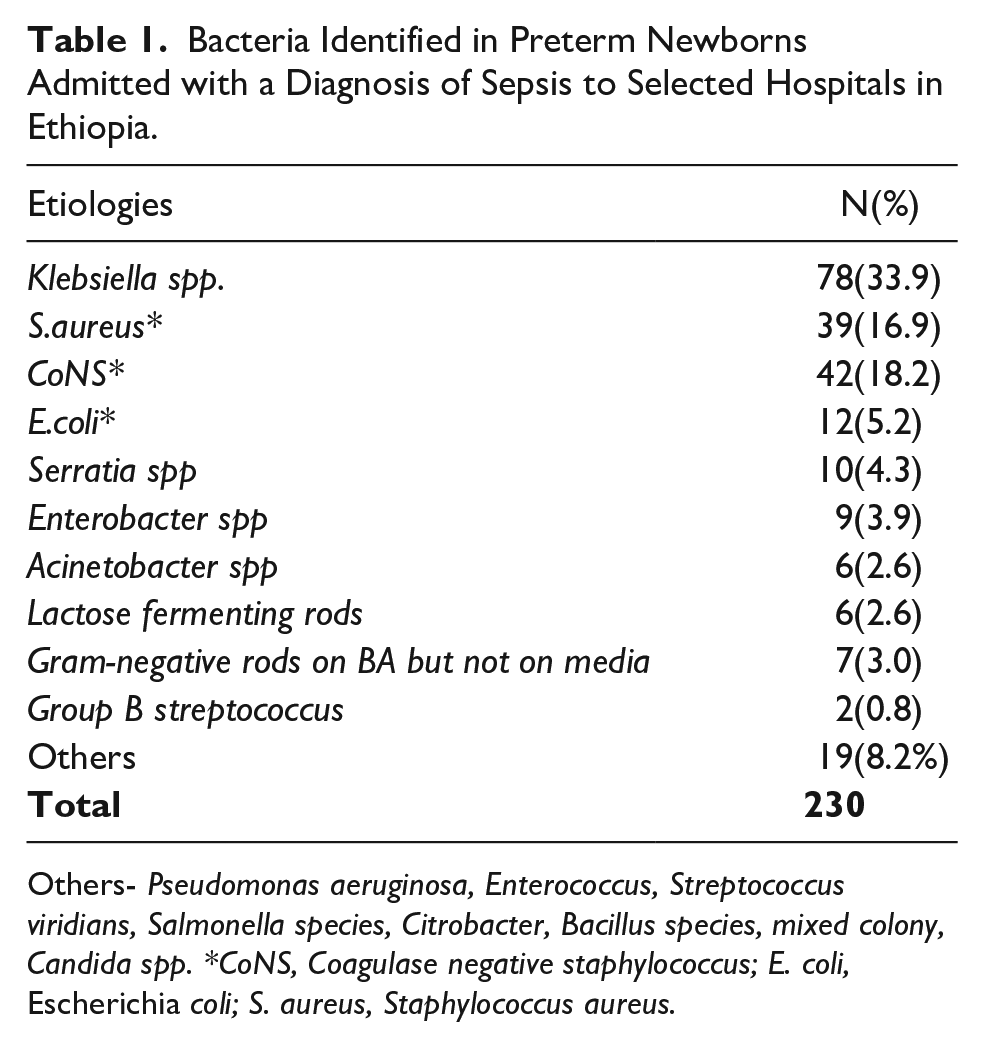
Others- Pseudomonas aeruginosa, Enterococcus, Streptococcus viridians, Salmonella species, Citrobacter, Bacillus species, mixed colony, Candida spp. *CoNS, Coagulase negative staphylococcus; E. coli, Escherichia coli; S. aureus, Staphylococcus aureus.
Gram-positive bacteria showed high resistance to penicillin (98.9%), amoxicillin-clavulanic acid (79.3%) and ceftriaxone (91.3%). S. aureus was found to be highly resistant to Oxacillin(85.3%). Better sensitivity to ciprofloxacin was observed among the Gram-positive bacteria (59%) (Table 3). Gram-negative bacteria were highly resistant to the first- and second-line empiric antibiotics such as gentamicin (83.2%) and 3rd generation cephalosporins such as ceftazidime (88%) and ceftriaxone (83.2%). The best sensitivity was observed to ciprofloxacin (65%) (Table 2).
Antibiotic Resistance Patterns of Isolated Gram-Negative Bacteriapathogens.

Note: spp. species, CN, gentamicin; AMP, ampicillin; CAZ, ceftazidime; AUG, amoxicillin clavulanic acid; CHL, chloramphenicol; TE, tetracycline; CRO, ceftriaxone; CIP, ciprofloxacin; SXT, sulfamethoxazole-trimethoprim.
Antibiotic Resistance Patterns of Isolated Gram-Positive Bacterial Pathogens.

Note: CoNS, Coagulase-negative staphylococci; GBS, group B streptococcus; P, penicillin; FOX, cefoxitin; ERY, erythromycin; CLN, clindamycin; SXT, sulfamethoxazole-trimethoprim; TE, tetracycline; CRO, ceftriaxone; CIP, ciprofloxacin; CN, gentamicin; OX, oxacillin; NA not applicable.
Multidrug Resistance
Multidrug Resistance (MDR) was observed in 199 isolates (91.3%). Klebsiella spp.(92%) and S. aureus (92.3%) were found to be predominantly MDR (Table 4).
Frequency Distribution of Non-MDR and MDR Patterns of Isolated Bacteria.

Discussion
The overall culture positivity rate from patients with symptoms of neonatal septicemia was 36.9%, which was slightly lower than results previously reported from Gondar (46.6%) and Addis Ababa (44.7%). 19 Higher rates of positive results were reported in Indonesia (65.3%), Georgia (63%) and Yemen (57%).20-22
On the other hand, the present study culture positivity rate was higher than observed in studies from Nepal (20.3%) and Gondar (32.1%).10,23 In contrast, much lower rates of positive results were reported from Iran (5.6%), Kuwait (8.7%), and Saudi Arabia (5%).24-26This variation could be due to methodological differences, time of blood culture collection and prior antibiotic administration, all of which might affect the culture positivity rate.
In our study, the predominant isolates causing sepsis were Gram-negative organisms with Klebsiella spp. accounting for 78 (33.9%) followed by Gram-positives like CoNS 42 (18.2%) and S. aureus 39 (16.9%). There was a similar report from Addis Ababa where Klebsiella spp. (39.2%) and S. aureus (22.2%) were the most commonly isolated organisms. A report from Sudan showed that Klebsiella (71.1%), S. aureus (15.8%), E. coli (5.3%), Gram-positive cocci (2.6%) and Serratia marcescens (5.3%) were associated with sepsis in newborns. 27 In Nigeria as well, neonatal infections were due to Gram-positive and Gram-negative organisms such as S. aureus (24.6%), CoNS (24.6%) and Klebsiella spp. (16.4%). 28
The results of the current study agree with the reports from most developing countries but differ from those reports from Kuwait where the most frequent organisms isolated were Gram-positive organisms like Staphylococcus epidermidis (34%), Streptococcus viridians (28%) and Candida species (14%). A report from the United Kingdom showed CoNS and GBS being organisms frequently seen in EONS, whereas CoNS and E. coli were most frequently associated with LONS. A study from the USA also showed that Gram-positive bacteria such as GBS were more predominant etiologies of sepsis. 25
As shown in this report and reports from other developing countries, most pathogens isolated in the hospital setting before 72 hours of life are similar to those isolated afterward. It is highly likely that unclean delivery practices are associated with infections with nosocomial agents very early in life 29 given that the deliveries of our patients occurred in health facilities (95%).
Klebsiella spp. was the predominant isolate in the present study, representing 33.9% of the total isolates, of which K. pneumoniae was the predominant bacteria identified. Similar findings reported from Georgia, USA showed K. pneumoniae accounting for 29% and Yemen 36.7%. 22
A very high prevalence of bacterial causes of neonatal sepsis and an alarmingly high level of antibiotic resistance in the isolates were observed in the current study. The overall MDR prevalence was 91.3% and it has been demonstrated that bacteria like Klebsiella spp. (92%), S. aureus (92.7%) and CoNS (100%) were found to be mostly MDR. This frequency is higher than that found from a study done in Gondar where the overall MDR prevalence was 65% and S. aureus (61.2%) and K. pneumoniae (74%) were found to be the principal MDR strains. 10 In Addis Ababa, Gram-negative bacteria showed high-level resistance to ampicillin, ceftriaxone, cephalothin, chloramphenicol, and gentamicin. Multidrug resistance was observed in 45.7% and 84.2% of Gram-positive and Gram-negative bacteria respectively. 19
One of the major limitations of this study is that only about one-third of infants with a clinical diagnosis of sepsis had a blood culture done. Reasons for this gap included failure to order blood culture by the hospital staff as a routine investigation for sepsis and difficulty of obtaining blood encountered in some of the newborns. The other limitation is the different methods of culture used at the different study sites which affects the blood culture positivity, as conventional blood culture suffers from poor sensitivity due to the lack of use of nutritionally enriching substances, inability to neutralize the inherent antimicrobial components in blood and/or in patients receiving antimicrobial treatment.
Although advances in neonatal care have improved survival and reduced complications in preterm infants, sepsis still contributes significantly to mortality and morbidity among very-low-birth-weight infants in neonatal intensive care units. Bacterial isolates from neonatal sepsis show variation from place to place, and indicates that there is a need for continuous monitoring of causative organisms and their drug susceptibility patterns at a local level.
Conclusion and Recommendations
A higher proportion of bacterial isolates was observed in this study. Klebsiella spp, CoNS and S. aureus were the predominant pathogens associated with sepsis. The overall prevalence of MDR was 91.3% which was high for most of the isolates. Ciprofloxacin was the most effective drug against both Gram-positive and Gram-negative bacteria. Even though there is a high rate of resistance seen in this report for the commonly prescribed antibiotics, further research is needed at the community level for revision of treatment guidelines.
Footnotes
Acknowledgements
We would like to thank all the parents/care takers of the patients who participated on the study. We would also like to thank the following individuals for their contribution to this paper: Asrat Demtse, Abayneh Girma, Sara Ayinalem, Netsanet Workneh, Riccardo E. Pfister and all the data managers.
Abbreviations
CoNS, coagulase negative Staphylococcus; CSF, Cerebrospinal fluid; CBC, complete blood cell count; EONS, Early onset neonatal sepsis; ESBL, extended spectrum beta lactamase; LMICs, low- and middle-income countries; MDR, Multidrug resistant; MRSA, methicillin resistant Staphylococcus aureus; Spp., Species; WHO, World Health Organization.
Authors’ contributions
BE: Contributed to conception and design; contributed to acquisition, analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy.
LMM: Contributed to conception and design; contributed to acquisition, analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy.
MB: Contributed to acquisition, analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy.
MG: Contributed to acquisition, analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy.
AA: Contributed to acquisition, analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy.
AM: Contributed to acquisition, analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy
BW: Contributed to acquisition, analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy.
RLG: Contributed to analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy.
EMM: Contributed to analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy.
GM: Contributed to acquisition, analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy.
MA: Contributed to acquisition, analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy.
SS: Contributed to acquisition, analysis, and interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring integrity and accuracy.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by grants from the Bill & Melinda Gates Foundation.
Ethics Approval and Consent to Participate
This study was approved by the ethical review committees of participating institutions (Addis Ababa University (AAU IRB no is AAUMF 03-008), St. Paul’s Millennium Medical College, Jimma University and the University of Gondar, Ethiopia). Informed consent was obtained from all participants.
Availability of Data and Materials
Study data will be made available at completion of the study.
