Abstract
Background:
Although highly active antiretroviral therapy has improved the quality of life among HIV-infected people in India, the emergence of drug resistance along with the limited access and affordability to routine monitoring continues to be a challenge worldwide.
Methods:
The frequency and patterns of HIV-1 drug-resistance mutations among the first-line failing HIV-infected patients attending a hospital in Salem, Tamil Nadu, India, were genotypically analyzed using the online Stanford HIV Database.
Results:
Of the study patients followed up for 6 months, 23 patients failed first-line therapy and the mutation of I135R/T/V/X, L178 I/M, M184V/I, D67N, K70R, and K103N was most common. Phylogenetic analysis revealed that most of these patients belonged to HIV subtype C.
Conclusion:
The study documents the frequency of nucleoside reverse transcriptase inhibitor and nonnucleoside reverse transcriptase inhibitor mutations that are prevalent in the first-line failing HIV-infected patients of South Indian region and adds up to the data for developing future algorithms to study the drug-resistance mutations of HIV subtype C. Thus, the results of the study call for the need for rational approach for selecting and for frequent viral monitoring to be performed to detect failure, followed by genotyping.
Introduction
HIV continues to be a serious public health issue, with over 78 million people infected and having claimed around 39 million people’s lives since the start of epidemic. 1 Although recent reports demonstrate a steady declining trend in adult HIV prevalence in India, the HIV epidemic in India is concentrated among high-risk groups and is heterogeneous in its distribution. The total number of people living with HIV/AIDS in India was estimated at around 2 million in 2011, reflecting a general adult population prevalence of less than 1%. 2
The advent of highly active antiretroviral therapy (HAART) and increased access have dramatically brought down the morbidity and mortality associated with HIV/AIDS, 3 and as of June 2014, about 13.6 million people living with HIV had access to antiretroviral therapy (ART). However, the success of HAART depends on the susceptibility of the virus to the drugs and optimal adherence to the therapy, failure of which could result in the emergence of HIV drug resistance. 4 With the factors such as excessive variability and the high evolution rate of HIV favoring the development of resistance, 5 in the context of HAART scale-up, emergence of HIV drug resistance (DR) is unavoidable. 6 Apart from bringing down the efficacy, thereby hampering the benefits of HAART, HIV drug resistance could also lead to the spread of resistant strains among the population. 7 This is particularly an issue of serious concern in developing countries where the HIV-infected patients have limited access to the second-line drugs.
While HIV subtype B predominates in North America and Western Europe, subtype A is most prevalent in west and central Africa, and subtype C is the major subtype in sub-Saharan Africa and Southeast Asia. 8 While studies on HIV-1 subtypes indicate that HIV-1 subtypes A and C are the most prevalent subtypes globally, most of the data on the genetic mechanisms of HIV-1 drug resistance and interpretations are based on the study of subtype B viruses. However, HIV-1 subtype B viruses account for only about 12% of global HIV-1 infections. 9 An increasing number of observational studies, in vitro and in vivo, suggest that the currently available protease (PR) and reverse-transcriptase (RT) inhibitors are as active against non-B viruses as they are against subtype B viruses. 10,11 However, fewer data are available on the genetic mechanisms of drug resistance in non-B viruses, and some in vitro and in vivo observations also suggest that various subtypes may respond differently to certain antiretroviral (ARV) drugs. 12,13
In country like India, where non-B type viruses predominate, extensive studies of resistance mutations among patients exhibiting therapeutic failure are essential for updating of the existing algorithms. With drug resistance an emerging problem, ART in resource-limited settings requires a paradigm shift as monitoring strategies including HIV viral load and drug resistance testing are often not available or not affordable for the patients. In the backdrop of the limited studies on HIV drug resistance among subtype C-infected patients experiencing therapeutic failure, we aimed to study the patterns of drug-resistance mutations (DRMs) and to study the various mutational interactions selected in South Indian HIV-1-infected patients experiencing virological failure (VF).
Methods
Study Design and Study Population
The study was conducted in Mohan Kumaramangalam Medical College Hospital, Salem, Tamil Nadu. In this prospective study, the participants included all HIV-infected patients visiting Integrated Counseling and Testing Centre and ART clinic, who met the following criteria: confirmed HIV-positive status per National AIDS Control Organisation (NACO) guidelines, >18 years old, with no previous exposure to ARV drugs by self-report and consented to participate in the study. Pregnant and lactating women were excluded from the study. During the period between March 2010 and March 2014, a total of 250 chronic HIV-1-infected individuals were enrolled, and of these, 37 dropped out for certain reasons as lost to follow-up and death, and the remaining 213 were followed up for a period of 6 months. Of these followed patients, based on the plasma viral load, 23 patients with VF were selected for genotyping analysis (Figure 1). The study received ethical clearance from the Directorate of Medical Education and from the Government Mohan Kumaramangalam Hospital. Written informed consent was obtained from all study participants before enrollment.

Phylogenetic tree of RT region of pol gene from the strains of the study participants.
Whole blood specimens were collected in 10 mL K3 EDTA tubes (Becton Dickinson, New Jersey) patients. Three milliliters of whole blood were used for CD4 count estimation, and the remaining were centrifuged within 6 hours of sample collection at 400 g for 10 minutes to separate plasma. The plasma specimens were stored as multiple aliquots at −75°C until the time of testing. A total of 23 plasma specimen aliquots from HIV-reactive ART clients were shipped at −75°C to YRG Centre for AIDS Research and Education, Chennai, for genotypic study. Detailed patient case history and demographics were collected from patient records of the hospitals, including the clinical follow-up within 6 months and ARV regimens.
Drug-Resistance Genotyping
The HIV genotyping assay was performed using the home brew methodology as described elsewhere. 14 Viral RNA was extracted from plasma (QIAamp viral RNA extraction kit, Qiagen Inc, Valencia, CA, USA) and reverse transcribed to complementary DNA (cDNA). The RT and PR regions of polymerase (pol) gene were amplified from cDNA by single or with nested polymerase chain reaction (PCR) using relevant primers. The PCR products were column purified and sequenced on an automated 3100 Avant genetic analyzer (Applied Biosystems, Foster City, California). Both forward and reverse sequences were assembled and manually edited with the Seqscape Version 2.5 multiple sequence alignment software and exported as a FASTA format consensus RT sequence. On July 5, 2014, the consensus sequences of each specimen were assessed for drug resistance using the online Stanford Drug Resistance Database available at http://hivdb6.stanford.edu/asi/deployed/hiv_central.pl?program=hivdb&action=showSequenceForm.
Phylogenetic Analysis
The phylogenetic tree of the RT region of the pol gene was constructed using Molecular Evolutionary Genetic Analysis software, version 6.0, using neighbor-joining method (Kimura 2-parameter) and boot strap values of 500. Reference strains belonging to subtypes A, B, C, and Circulating Recombinant Forms (CRFs) were used to compare the diversity of the strains. All the data were documented on a specially designed paper form and subsequently transferred to Microsoft Excel spreadsheet. All the entries were checked for possible keyboard error(s) at the entry level. The electronic data were exported into the STATA software v11 for statistical analysis. Baseline clinical and biological characters of the study participants were summarized as frequency (%) for the categorical variables and mean ± standard deviation or median (interquartile range) for quantitative variables. Prevalence of DRMs was computed with 95% confidence interval.
Results
Demographic Characteristics of the Study Population
Of the 250 study participants enrolled (median age 32 years; median CD4 count 222 cells/mm3), after dropouts due to lost to follow-up and death (n = 37), 213 were followed-up for 6 months. All participants self-reported good adherence and were in the median age of 35.4 years, and 67% of them were males. Ninety-five percent of individuals had contracted HIV-1 infection by heterosexual transmission and only 1 individual by blood transfusion (Table 1).
Baseline Characteristics of HIV-1-Positive Participants Treated with First-Line ARV Regimens in Salem, Tamil Nadu, India.a
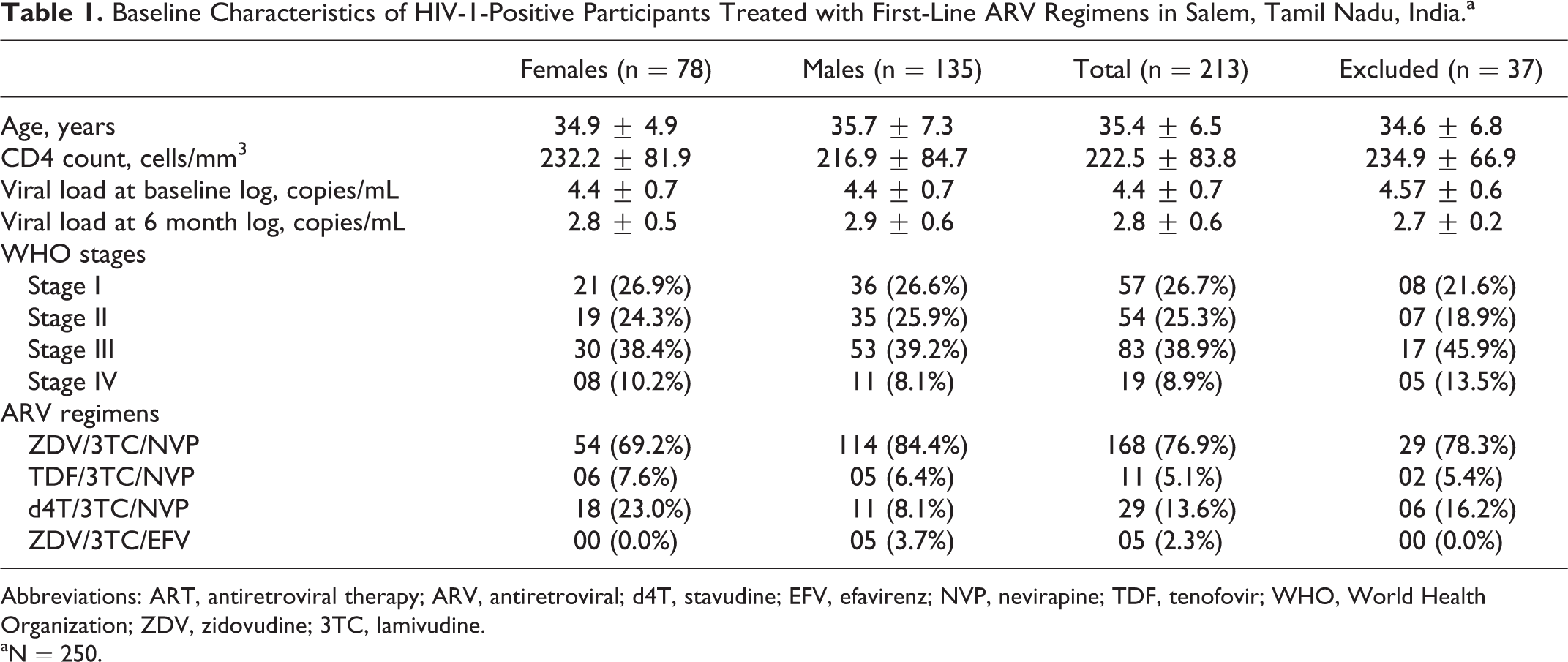
Abbreviations: ART, antiretroviral therapy; ARV, antiretroviral; d4T, stavudine; EFV, efavirenz; NVP, nevirapine; TDF, tenofovir; WHO, World Health Organization; ZDV, zidovudine; 3TC, lamivudine.
aN = 250.
Of the 213 participants followed, based on the plasma viral load data, 37 cases with VF were identified. Of these, specimens from 23 participants could be successfully amplified at the RT region of the pol gene. Among the 23 isolates, 21 isolates were found to belong to subtype C and 2 isolates observed as subtype B (Figure 1).
The most commonly prescribed first-line regimen was the combination of tenofovir (TDF), lamivudine (3TC), and efavirenz (EFV) or nevirapine (NVP). The other first-line regimen was zidovudine (ZDV) or stavudine (d4T) instead of TDF. Of the 213 study participants, 23 (9.26%) patients experienced the VF with the significantly lower CD4 count at 6 months (Table 2).
Consolidated Demographic Characteristics of Participants with Virological Failure.a

Abbreviations: ARV, antiretroviral; d4 T, stavudine; EFV, efavirenz; NVP, nevirapine; PVL, plasma viral load; TDF, tenofovir; ZDV, zidovudine; 3TC, lamivudine.
an = 23.
Pattern of Mutations in HIV-1
Of the study participants, any one of the known RT drug resistance–associated mutations was found in 87% (20 of 23). Any one of nucleoside reverse transcriptase inhibitor (NRTI)-associated mutations was found in 87% (20 of 23), of which M184 V was predominant with 78.2% (18/23), followed by T215Y (52%). Thymidine analogue mutations (TAMs) were observed in 78% (18 of 23) of the study participants with T215Y being the most common mutation (52%; 12 of 23), followed by D67N (48%; 11 of 23). Fifty percent of these belonged to TAM-1 and the remaining to TAM-2 group. The frequency of other NRTI mutations and distributions of TAMs are shown in Table 3.
Frequencies of NRTI and NNRTI Resistance Mutations Observed in the Study Group.
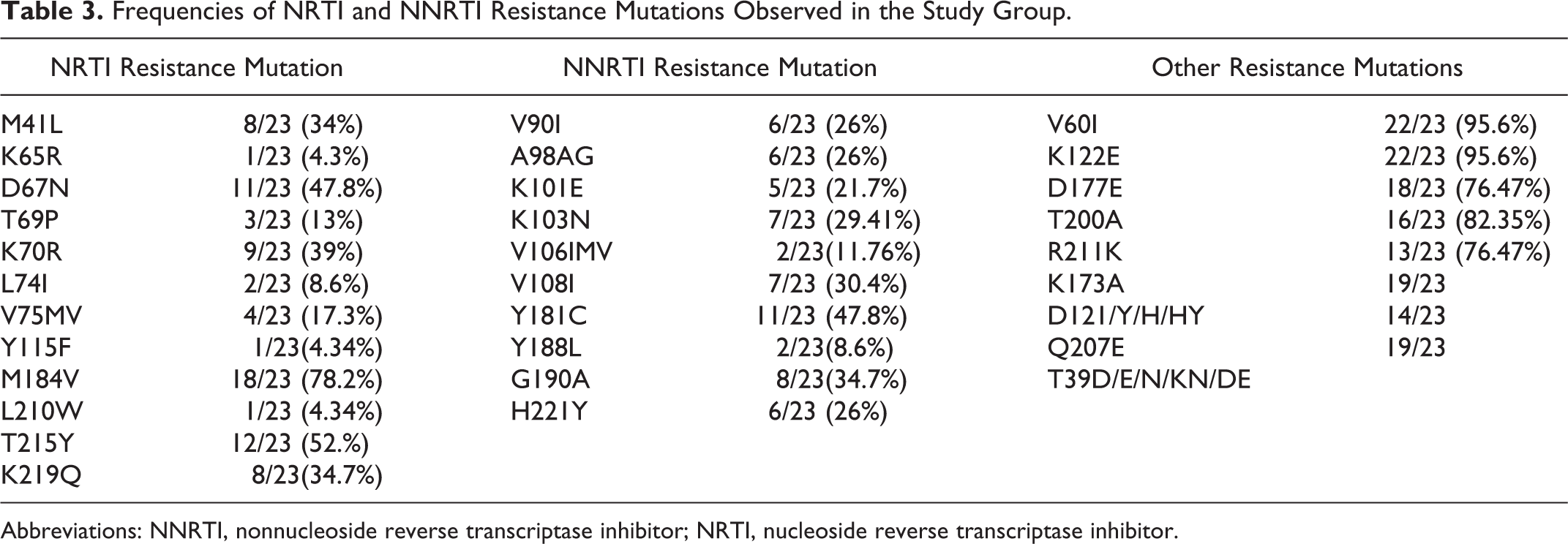
Abbreviations: NNRTI, nonnucleoside reverse transcriptase inhibitor; NRTI, nucleoside reverse transcriptase inhibitor.
Of the study participants, any one of the nonnucleoside reverse transcriptase inhibitor (NNRTI)-associated mutations was observed in 91% (21 of 23), of which Y181C (48%) was the predominant followed by G190A (35%). Other NNRTI mutations that were observed are presented in Table 3. NVP seemed to select for a wider range of mutations when compared to EFV. The patient who developed K65R was on ZDV-based regimen and coselection of K65R along with M184V was observed. Y115F was also found to be coselected. None of the patients had multidrug-resistance mutations.
Of the patients who developed NNRTI-associated mutations, 19 and 3 patients were in NVP- and EFV-based regimens, respectively. The combination of M184 V with Y181C was the predominant NRTI hypersusceptibility mutation (HSM), followed by M184V and of the NNRTI HSMs, only T215Y was observed (12 of 23). The other newer mutations observed in the study include V60I, K122E, T200A, D177E, R211K, and K173A.
Discussion
HIV drug resistance is one of the major hurdles for achieving and maintaining of successful viral suppression. Most data on the genetic mechanisms of HIV-1 drug resistance are from studies of subtype B viruses, the predominant subtype in the North American and Europe. Several of studies suggest that the currently available PR and RT inhibitors are as active against non-B viruses as they are against subtype B viruses. 15 The mutations that cause drug resistance in subtype B viruses appear to be the main mutations that cause drug resistance in non-B viruses as well. 16 However, the spectrum of mutations at some drug-resistance positions differs among subtypes.
The results of this study document the DRMs among the first-line ART failure patients from Southern India and reemphasizes the significance of viral load monitoring genotyping for rational selection of regimens. The DR genotyping results of isolates from the patients with VF in the current study demonstrated high level of mutations known to cause resistance to reverse transcriptase inhibitors. Of the 23 cases, one or more DR mutations were observed in the RT region of isolates.
Studies have demonstrated that NRTI mutations like T215Y might lead to viral hypersusceptibility to the NNRTIs in NRTI-treated individuals and that the presence of these mutations may improve subsequent virologic response to NNRTI-containing regimens in NNRTI-naive individuals. 17,18 The most prevalent mutations observed in the study confer high-level resistance to NRTIs (especially ZDV, in case of M184V mutation) and NNRTIs (specifically NVP and EFV, in case of K103N/S mutation and Y181C/Y/I/V mutation).
These results corroborate with the reports from systematic reviews of studies on HIV drug resistance from resource-limited settings. 19,20 Mutations at positions 41 (M41L) and 65 (K65 R) are found to confer resistance to NRTI drugs such as ZDV, d4T, 3TC, emtricitabine (FTC), and TDF, which are the most commonly used drugs in India. 21 Moreover, M41L confers immediate to high-level resistance to ZDV and d4T when existing along with T215Y. The study demonstrates the prevalence of these mutations and that caution needs to be given while selecting initial regimen.
The mutation patterns observed in this study were similar to those observed from earlier studies done in subtype B. As observed in previous studies from India and Kenya, M184V was the predominant mutation. Although most of the patients were on 3TC as first-line drug, majority of these patients failed, probably due to the selection of M184V mutation. Since most of the study participants were exposed to thymidine NRTIs and ZDV, this explains the number of NRTI-related mutations observed in this study. However, the most frequent mutations in RT were associated with ZDV and NNRTI exposure, which confirms the low genetic barrier of these drugs. 22
Studies have shown that non-subtype B viruses might be more prone to rapid emergence of NNRTI resistance because of background polymorphisms in the RT gene. 23 After NVP use, Y181C is the most frequent mutation in the HIV-1 RT gene, but this does not confer high-level cross-resistance to EFV. In contrast, the K103N mutation is relatively uncommon after NVP use but confers high-level EFV cross-resistance. K103N is studied to occur in over 30% of patients with VF while receiving NVP and reduces susceptibility to NVP by 50-fold. The mutation also occurs in over 50% of patients failing EFV and reduces susceptibility to EFV by 25-fold. K103N alone is sufficient to prevent a virological response to EFV. A98G mutation observed in 26% of the study patients is known to confer low-level, approximately 2-fold, resistance to NVP with uncertain effects on virological responses to the drug.
The increased proportion of K103N + Y181C mutations suggests that there is no impact of NRTI exposure on accumulation of NNRTI resistance mutation with sequential use of NNRTIs. Efavirenz exerting specific selective pressure could probably be the reason for this phenomenon. Also, as observed earlier, HIV strains with both K103N and Y181C mutations together had high prevalence of G190A. 24 Unlike reported earlier, 25 –27 NNRTI mutations with the highest prevalence among the first-line failure was 181C followed by G190A. It is well reported that G190A causes high-level resistance to NVP and intermediate-level resistance to EFV.
As observed in a study from Chennai, 27 TAM pathways 1 and 2 were evenly distributed, which suggests that patients who had a history of mono/dual exposure have very few drug sequencing options. This equal accumulation of both TAM pathways is associated with increasing levels of NRTI cross-resistance. Among the TAM-1 pathway, the combination of M41L and T215Y was commonly observed, while the combination of M41L and L210W was the least observed as reported earlier for subtype C. 28 Lower prevalence of L210W in subtype C compared to B might probably be due to the difference in the encoding of leucine at this position as described elsewhere. 29,30 Coexistence of both pathways was observed to be lower than previously reported from South Africa, 28 showing that this combination could be unique to subtype C and this genotype is usually not found in subtype B.
On analyzing the incidence of TAMs with respect to the thymidine analogues used, the emergence was higher in patients on first-line triple therapy including ZDV than those on D4T. Similar findings have been reported earlier in France. 31 Several studies have shown that thymidine analogue mutations are associated with NNRTI hypersusceptibility, wherein there is an increased susceptibility to a particular drug, that is, lower amounts of drug are needed to inhibit replication of virus when compared with wild-type virus. 32 Patients with such mutations have been observed to have better outcomes on NNRTI-based ART. Although at least 23 different RT codons have been associated with NNRTI hypersusceptibility, the strongest effect is seen with mutations at codons 215 (T215Y), which was the predominantly observed mutation in the present study.
The genetic variability of HIV-1 is a major challenge to the development of an effective vaccine and long-acting treatment. Our observation based on the phylogenetic analysis shows that over 91% of the study population were infected with subtype C of HIV-1. Previous studies from other parts of the country also corroborate the predominance of HIV-1 subtype strains in India. 14,33 Interestingly, along with subtype C, a couple of subtype B strains, related to North American and European viruses, were found in the study population as reported earlier. 33 –35 Although majority of the studies globally have used env gene for HIV-1 subtyping, the HIV-1 pol gene has also been used for subtyping in regions where there is less diverse HIV-1 strains circulating, hence for developing countries like India which mainly have subtype C, can use pol gene for subtyping. This method has been used mostly for European strains as well as in other countries. 36
Limitations of this study include the minimal sample size and the design wherein only a representative group of patients are chosen for genotyping analysis. Also, the findings of this study should be interpreted in light of the other limitations, namely, of the study participants followed, only the 23 cases of VF were chosen for genotyping based on their PVLs and that blips in viral load cannot be ruled out. It is well reported that occasionally, viral blips can occur even during effective treatment. 37 Also, it is to be noted that the study participants may not have achieved virological suppression in the first place, even after being on ART for more than 6 months, which might possibly be attributed to the effect of persisting HIV-1 primary or transmitted resistant mutations. 38
Transmission of drug-resistant HIV strains has been well documented and has been associated with suboptimal virologic response to initial ART. 39,40 The likelihood that a patient will acquire drug-resistant virus is related to the prevalence of drug resistance in persons engaging in high-risk behaviors in the community. In India, primary resistance has been documented in a few studies on naive patients, 14,41 and with increasing ART access, problems of primary drug resistance could increase in future and thus screening patients prior to initiation of HAART might be essential.
However, it is to be noted that several studies have demonstrated viral resuppression in spite of the presence of major DRMs. 42,43 A recent study from India had reported patients with controlled viremia in the presence of multiple major NRTI- and NRTI-associated mutations, without change in therapy. 44 Based on these results, these studies go on to suggest that work has to be done in understanding how the DRMs affect the viral fitness, drug susceptibility, and the therapeutic responses. 45
Conclusion
The genotypic results revealed that the pattern of mutations was based on the Stanford database where it only refers to the HIV subtype B, whereas most of the isolates in this study were of subtype C. Thus, the results of the study calls for the need for rational approach for selecting drugs such as using NRTI drugs with high potential for cross-resistance be used later rather than earlier stage and frequent viral monitoring be performed to detect failure, followed by genotyping. The importance of genotyping for identification of mutations in the RT gene of HIV-1 isolates pertaining to drug resistance from first-line therapy failures has reemphasized the importance of a regimen selection prior to the start of second-line therapy for the better efficacy of the treatment and the need for well-designed and long-term longitudinal studies.
Sequence Data
Nucleotide and amino acid RT sequences of the 23 study patients were submitted to GenBank (KM260546, 1786571, 1786562, 1786551,1786167, 1785656, 1785650, 1785332, 1785329, 1785326, 1785091, 1784957, 1784934, 1784933, 1784931, 1787203, 1786593).
Footnotes
Acknowledgments
The authors would like to express their sincere gratitude to the study participants, without whom this study would not have been possible.
Authors’ Note
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
