Abstract
Vanishing twin syndrome (VTS) refers to the spontaneous reduction of a fetus during multiple pregnancies, resulting in its invisibility during gestation. Vanishing twin syndrome is commonly identified through ultrasonographic examination in early pregnancy; nonetheless, the genetic causes of VTS are rarely detected. This report aimed to investigate the feasibility of genetic testing for VTS using low-coverage whole-genome sequencing and short tandem repeat (STR) analysis. We report a 39-year-old woman who underwent in vitro fertilization and conceived dichorionic diamniotic twins. The heartbeat of 1 fetus ceased at 11 weeks, leading to a diagnosis of VTS, whereas the surviving fetus developed normally and was delivered at full term. Placental examination revealed a rudimentary gestational sac adjacent to the placenta that correlated with the vanished fetus. Chorionic tissues were collected from the placenta of the live-born infant and the rudimentary gestational sac that was considered to have originated from the vanished fetus. Genetic testing of chorionic tissues from the gestational sac and placenta showed that both fetuses were monozygotic twins with a normal 46, XY male karyotype. However, a triallelic pattern at the Penta E locus was identified in the vanished fetus on STR analysis, which was a unique genetic characteristic. This report highlights the feasibility of genetic testing for VTS, despite the unclear relationship between the identified genetic pattern and the vanished fetus.
Keywords
Introduction
Vanishing twin syndrome (VTS) refers to the spontaneous reduction of the fetus during multiple pregnancies, resulting in its subsequent invisibility during gestation. 1 Vanishing twin syndrome occurs in approximately 53% of pregnancies with 3 or more gestational sacs and 36% of twin pregnancies, 2 with the majority of cases being identified through ultrasonographic examination in early pregnancy. Risk factors for VTS include advanced maternal age, chronic maternal diseases, multiple pregnancies, use of assisted reproductive technology, small placentas or other anatomical abnormalities in the placenta, and teratogenic factors.3 -6 Despite the identification of these risk factors, ascertaining the direct cause of VTS remains challenging because of the lack of utilization of appropriate diagnostic methods. Clarifying the etiology of VTS may be the key to the identification of a select number of cases, which may influence the obstetric outcomes and the subsequent reproductive outcomes.
Chromosomal analysis of the product of conception (POC) is instrumental in the investigation of the cause of pregnancy loss (PL). Karyotyping is recognized as the traditional gold standard for POC analysis; nevertheless, karyotyping has limitations owing to its reliance on cell culture, including culture failure and maternal tissue contamination from poor sample quality. 7 Moreover, cases such as VTS in which chorionic tissues remain in the uterus for an extended period, karyotyping is impractical owing to the loss of cell viability. The use of next-generation sequencing (NGS) has become widespread in genetic testing, 8 because of its ability to directly assess genomic DNA (gDNA) from samples, obviating the need for cell culture and enabling genetic testing even with deteriorated samples. Thus, NGS is increasingly being utilized in the chromosomal testing of miscarriages.8,9 In addition, recent reports using NGS, specifically whole-exome or whole-genome sequencing, suggested that a higher proportion of miscarriages and stillbirths with normal chromosomes might be attributed to genetic abnormalities undetected by karyotyping than that was previously expected.10,11 Furthermore, discerning the genetic differences between tissues is crucial to avoid maternal cell contamination (MCC) when conducting POC testing. Tissue individualization is enabled by comparing the short tandem repeat (STR) allele patterns of individual gDNA, 12 and STR analysis is beneficial for confirming the individuality of zygotes in twin pregnancies, aiding in distinguishing monozygotic from dizygotic twins. 13
In our report, we present a case of VTS in which NGS was utilized to detect copy number variants (CNVs), and STR analysis was conducted on chorionic tissues obtained from a live-born infant and a vanished fetus that were retained for 8 months, a duration scarcely reported in the literature, underscoring the innovative aspect of our work.
Case Report
Sample Collection
The chorionic tissues were rinsed with saline and collected under a stereomicroscope to prevent contamination from maternal tissues. Approximately 10 mg of chorionic tissues was collected. Maternal blood samples were collected in Cell-Free DNA BCT tubes (Streck, La Vista, NE, USA) and subsequently preserved via flash freezing in liquid nitrogen, followed by storage at −80°C.
Genomic DNA Extraction and Genetic Analysis
NGS
Genomic DNA was extracted from the chorionic tissues and maternal blood using NucleoSpin Tissue and NucleoSpin Blood, respectively (Takara Bio Inc., Shiga, Japan). The quality of each extracted gDNA sample was evaluated using Quant dsDNA BR Assay Kits (Invitrogen, Cergy Pontoise, France) and agarose gel electrophoresis to confirm the presence and integrity of gDNA. For the NGS analysis, whole-genome amplification (WGA) of gDNA from the chorionic tissues was performed using the Embgenix PGT-A Kit (beta version; Takara Bio Inc.), followed by DNA library construction and lcWGS using the MiSeq System (Illumina, Inc., San Diego, California). Sequencing data were analyzed using Embgenix analysis software version 0.32 (Takara Bio Inc.) to detect CNVs.
STR analysis
Quantitative fluorescence polymerase chain reaction was conducted on gDNA from the chorionic tissues and maternal blood using the PowerPlex 16 System (Promega Biosciences, LLC, San Luis Obispo, California). Short tandem repeat typing was performed using a capillary sequencer. Allelic patterns of STRs from both chorionic tissues and maternal blood samples were detected and compared to identify consistencies/discrepancies and to confirm or exclude the possibility of MCC.
Case Presentation
A 39-year-old woman (gravida 2, para 1) conceived through in vitro fertilization with double embryo transfer and was initially diagnosed with dichorionic diamniotic twins in another hospital. At the initial diagnosis, the crown-rump length (CRL) of the first viable fetus was 19.0 mm, whereas the CRL of the second viable fetus was 19.7 mm. The patient was referred to our hospital for perinatal management. Upon presentation at our hospital, VTS was diagnosed following cardiac arrest in 1 fetus at 11 weeks of gestation; the first viable fetus had a CRL of 43.4 mm, whereas the second non-viable fetus had a CRL of 20.0 mm (Figure 1). At 20 and 28 weeks of gestation, the surviving fetus underwent fetal screening, which showed no abnormalities and an adequate amniotic fluid index. The non-viable fetus was no longer detectable on ultrasonography at 34 weeks of gestation. The pregnancy progressed without complications, and the patient vaginally delivered a healthy male infant at 40 weeks of gestation after spontaneous labor. The live-born infant weighed 3058 g, with an Apgar score of 8/9. The placenta was normal in size, and an approximately 5-cm rudimentary gestational sac was found at the edge of the amniotic membrane, which was presumed to have originated from the vanished fetus (Figure 2).
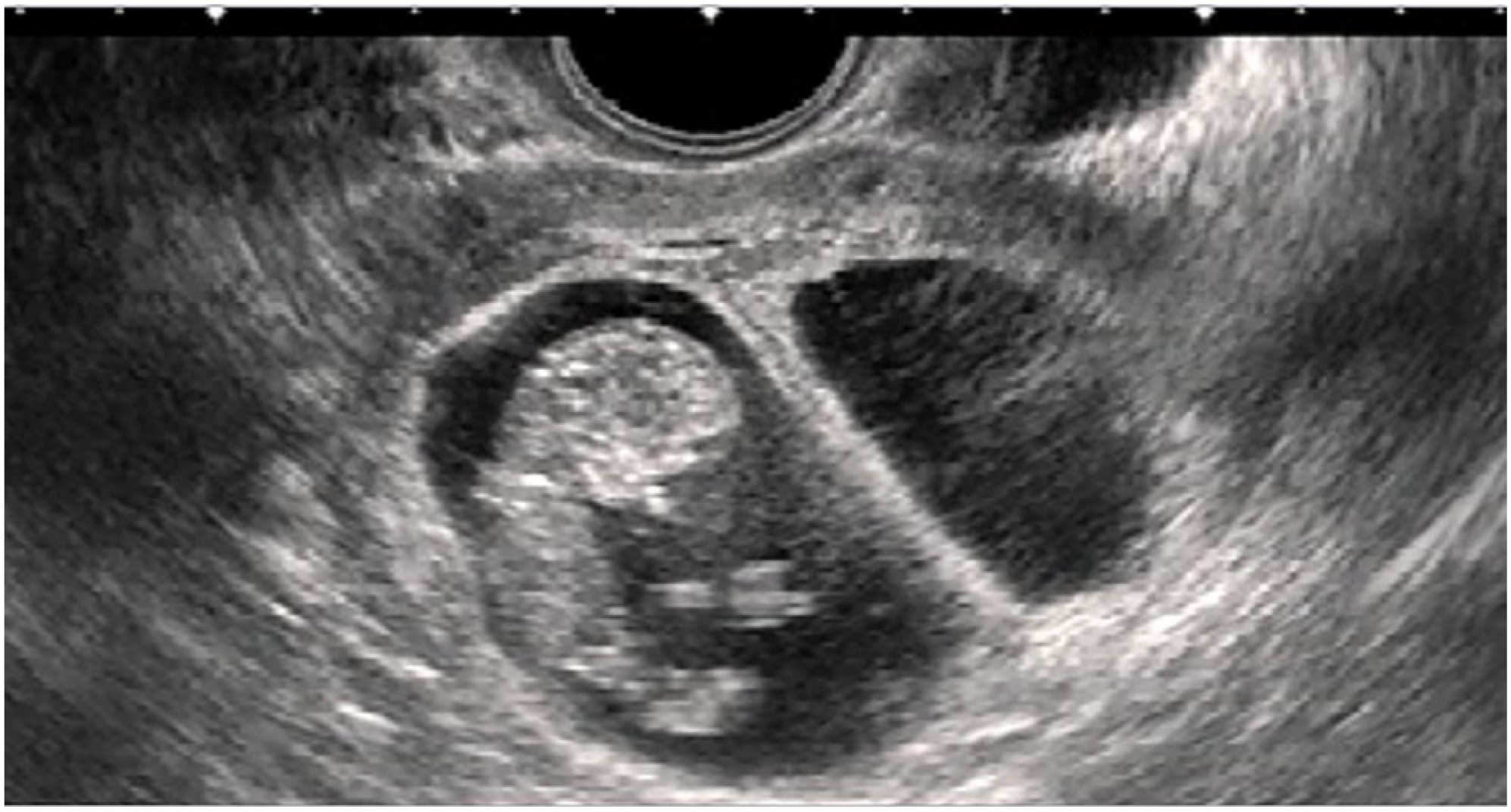
Diagnosed with dichorionic diamniotic twins and VTS at 11 weeks’ gestation, with the first fetus having a CRL of 43.4 mm, and the vanished fetus.

Rudimentary GS at the edge of the amniotic membrane, which was thought to have originated from the vanished fetus. Both chorionic tissues were confirmed and corrected under a microscope.
Chorionic tissues were collected from both (1) the placenta of the live-born infant and (2) the rudimentary gestational sac that was considered to have originated from the vanished fetus; these were confirmed under a microscope (Figure 2). For gDNA extraction and STR analysis, maternal blood was collected on the day of delivery. Genomic DNA was extracted from the chorionic tissues and maternal blood. Agarose gel electrophoresis revealed fragmented DNA in the vanished fetus. Chromosomal analysis with lcWGS of both chorionic tissues indicated no chromosomal aneuploidy, mosaicism, or structural abnormalities in the CNV plot; furthermore, both samples showed a normal 46, XY male karyotype (Figure 3). Maternal and chorionic tissue STR allele typing was conducted and concordance rates were assessed, confirming a very low possibility of MCC. The chorionic tissues matched 15 out of 16 STR markers, indicating that the twins were monozygotic. A unique triallelic pattern at the Penta E locus was identified in the vanished fetus, with alleles 9, 14, and 15 in the rudimentary gestational sac compared with alleles 14 and 15 in the placenta of the live-born infant as well as alleles 9 and 15 in the maternal blood (Figure 4).
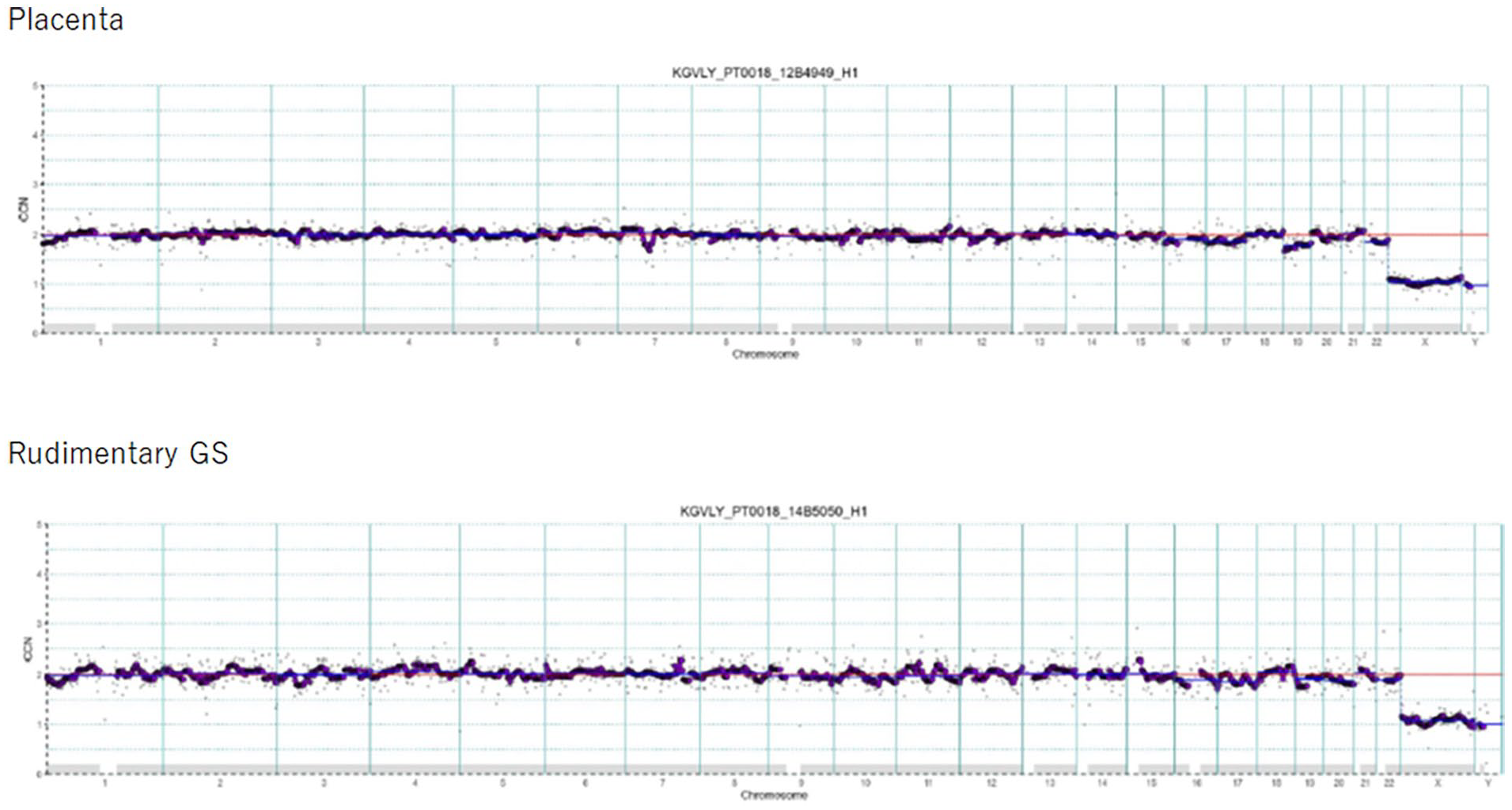
lcWGS revealed no evidence of chromosomal aneuploidy, mosaicism, or structural abnormalities in the CNV plot, and both chorionic tissues showed a normal 46, XY male type.

Concordance in 15 out of 16 STR markers suggested that the twins were monozygotic. A unique triallelic pattern at the Penta E locus was observed in the vanished fetus, with alleles 9, 14, and 15, compared with alleles 14 and 15 in the placenta of the live-born infant, and alleles 9 and 15 in the maternal blood.
Discussion
We demonstrated the potential of genetic diagnosis in VTS by successfully performing CNV analysis through lcWGS and STR analysis, which clarified the genetic profile of monozygotic twins, including a distinctive triallelic pattern at the Penta E locus in the vanished twin.
Couples who have experienced VTS are often concerned regarding the well-being of the surviving twin. When VTS occurs before the second trimester, the obstetric outcomes of the surviving twin are similar to those of the original singleton. In contrast, VTS occurring after the second trimester is significantly associated with reduced gestational duration, decreased birth weight, and increased incidence of neurological disorders.14,15 In this case, the live-born infant was delivered at full term, without any abnormalities. Although understanding the etiology of VTS may provide insights into its potential impact on perinatal outcomes in certain cases, the specific cause of VTS often remains unclear, and comprehensive analyses of vanished fetuses are limited.
While numerous factors can cause PL, more than half of the cases are associated with fetal chromosomal abnormalities.10,16 The European Society of Human Reproduction and Embryology guidelines for recurrent PL have emphasized the limitations of traditional karyotyping and highlighted the importance of future research on the potential of NGS techniques. 17 Karyotyping of POC samples requires cell culture, which often fails (5%–38%) because of factors such as contaminated bacteria, fetal demise, macerated tissues, and preferential growth of maternal cells, resulting in a low diagnostic yield.18,19 Moreover, the quality of karyotyping varies among samples and laboratories and relies on the expertise of technicians and cytogeneticists.20 -24 Next-generation sequencing is an effective alternative approach to genomic testing because it does not require cell culture and allows for both qualitative and quantitative analyses. 8 Next-generation sequencing also enables the testing of very old samples through gDNA extraction. 25 While the quantity and quality of nucleic acid samples are key factors that can limit research, 26 genomic testing can be successfully conducted even with poor-quality samples by employing techniques such as WGA and NGS. 27 Short tandem repeat analysis can show other information such as MCC. These features are beyond the sensitivity of conventional methods such as karyotyping. 28 Previous large-scale studies that included 1745 patients with PL reported that 50.4% of POCs were karyotypically abnormal. However, the application of genome-wide haplotyping to a subset previously identified, through traditional karyotyping, as normal revealed additional chromosomal aberrations, which eventually increased the rate of chromosomal abnormalities in PL to 67.8%. 28 Chromosomal and genetic analyses of POC can pinpoint the cause of PL and offer vital clinical insights for future pregnancies, contributing to a reduction in the need for further costly and complex maternal investigations to identify other causes of recurrent PL.29,30 If a balanced translocation is suspected based on the results of POC analysis in either parent, preimplantation genetic testing is suggested. 31 In addition, if POC chromosomes exhibit a normal karyotype, it could guide the identification of other factors contributing to PL. 29 In this case, a sac from the vanished fetus was found adjacent to the amniotic membrane. The tissues, which were retained within the uterine cavity for approximately 8 months, exhibited features of fibrosis under microscopic examination, rendering cell culture and subsequent karyotyping unfeasible. The WGA process ensured that the DNA quality was sufficient for testing despite significant degraded DNA in the tissues of the vanished fetus. Our findings demonstrate that POC analysis using lcWGS is feasible even when the tissues remain in the uterine cavity for an extended period. However, lcWGS was used specifically to detect CNVs in our study. This method allows for the identification of CNVs that might not be detected by traditional karyotyping, and it does not have the same high yield of detecting other genetic variants, such as those identified through whole-exome or whole-genome sequencing.
In this case, the concordance of 15 out of 16 STR markers, excluding the Penta E locus, in the chorionic tissues of the live-born infant and vanished fetus indicated that the twins were monozygotic. In addition, only the discordance in 1 STR marker between the twins suggested an extremely low possibility of tissue contamination between the chorionic tissues. A discrepancy was observed in the triallelic pattern at the Penta E locus in the vanished fetuses. The non-matching STR profiles of the chorionic tissues and maternal cells served as definitive evidence to exclude the possibility of MCC. The triallelic pattern can be classified into 2 types: type 1 arises from somatic variants, whereas type 2 results from chromosomal rearrangements. 32 Type 1 consists of 3 alleles with varying signal intensities, as observed in this case in which alleles of different strengths were present. Short tandem repeat analysis revealed that both the mother and the vanished fetus had allele 9 at the Penta E locus. As for the other 15 STR markers, excluding Penta E, distinct genetic loci were observed between the mother and the vanished fetus, indicating genetic dissimilarity. Furthermore, the signal intensity of allele 9 at the Penta E locus differed between the mother and the vanished fetus. These findings suggest that the presence of allele 9 in the vanished fetus was not due to MCC but was potentially attributed to a somatic variant. Therefore, the evidence points to a scenario in which despite the predominance of cells with repeat sequences of alleles 14 and 15, a mosaic condition may exist, comprising mutant cells with repeat sequences of allele 9 arising from somatic variants.
The Penta E locus is located on chromosome 15q26.2. 33 Upon examination using lcWGS, no aneuploidy was observed in this chromosomal region, and the possibility of a chromosomal mosaic pattern was deemed unlikely. The contribution of this variant to PL remains uncertain, as a triallelic pattern has been reported in healthy individuals.34,35 In contrast, when a variant occurs in the chromosome 15q26.2 region, it may affect the expression of genes in the same region such as NR2F2 and SPATA8, in which region defects or variants can reportedly cause cardiac and renal diseases. Despite no previous reports on the triallelic pattern of Penta E related to PL in the database, the potential association between the triallelic pattern and PL cannot be definitively excluded. 36 The study limitations include (1) the uncertainty in identifying the exact cause of VTS and (2) limited sample; therefore, the importance of genetic testing could not be established.
Conclusions
Chromosomal analysis was successfully performed with lcWGS to detect CNVs and STR analysis in a case of VTS that remained in utero for 8 months. While the exact cause of VTS was not identified in this report, the feasibility of genetic and chromosomal diagnoses was demonstrated. Widespread use of genetic testing and newer testing methods in larger cohorts of VTS may lead to a deeper understanding of this syndrome.
Footnotes
Acknowledgements
The authors thank Takara Bio Inc. for their invaluable technical support and expertise throughout the course of this research. Their dedication to excellence and collaborative spirit has been instrumental in overcoming challenges and achieving meaningful results.
Author Contributions
H.H. and M.H. designed the study, developed the key conceptual ideas, outlined the proof, and conducted data collection. S.T., T.G., and T.M. aided in interpreting the results and worked on the manuscript. T.T. and T.K. supervised the project. H.H. wrote the manuscript with support from T.T. and T.K. All authors discussed the results and commented on the manuscript.
Data Availability
All data generated or analyzed during this study are included in this article. Further enquiries can be directed to the corresponding author.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethics Approval
Ethical approval to report this case was obtained from Institutional Review Board of Osaka University Hospital, approval number (accession no. 21321).
Informed Consent
Written informed consent was obtained from the patient for their anonymized information to be published in this article.
