Abstract
Aim:
Various methods investigating the bacterial content causing periodontal abscesses have been applied in studies conducted until today. However, these studies have focused on periodontopathogens. Our study was carried out to research whether different pathogens other than the known periodontopathogens are present in periodontal abscess formation. Therefore, dominant bacterial samples obtained from the periodontal abscess content using the culture-dependent method were identified by 16S rDNA sequencing.
Materials and Methods:
Samples were obtained using a syringe or a periopaper from periodontal abscesses of 20 volunteers who met the research criteria. The three different bacterial colonies that were observed most intensely in each sample were selected and purified, and the isolates obtained were kept until the next characterization. Genomic DNA was isolated from each isolate; 16S rRNA genes were amplified by polymerase chain reaction and identified using DNA sequencing analyses.
Results:
As a result of culture-dependent methods, bacterial species belonging to Streptococcus, Staphylococcus, Neisseria, Actinomyces, Morococcus, Moraxella, and Enterococcus genera were isolated from a total of 60 bacterial isolates, three of which were the most densely growing colonies from each periodontal abscess sample.
Conclusion:
In our study, most of the bacterial species detected were identified for the first time in the bacterial content of periodontal abscesses. In some previously done studies, most of these bacteria species were shown to cause abscesses in different parts of the body. It was concluded that further studies are needed to determine the number and proportion of these bacteria species in total bacterial content to evaluate whether they cause periodontal abscesses.
Introduction
Dentoalveolar abscesses refer to the accumulation of pus localized in the alveolar bone around the dental root. They can spread to adjacent anatomical structures and such a situation can lead to septicemia, cavernous sinus thrombosis, brain abscess, shock, and death if they are not treated early enough. 1 Periodontal abscesses, which are among dentoalveolar abscesses, may be associated with the problems in periodontium (gingiva, periodontal ligaments, alveolar process) such as destruction of alveolar bone. Periodontal abscess is defined as a localized purulent infection affecting the tissues surrounding the periodontal pockets which cause the destruction of supportive dental structures.2, 3
As they are associated with the preexisting periodontitis, the periodontal abscesses are believed to be a complication of periodontitis.
4
It has been suggested that periodontal abscesses are usually the result of untreated chronic periodontitis cases, and the changes in the subgingival microbiota, reduction of the host resistance or both accelerate abscess formation.
2
In some studies in the literature, it is claimed that the microbiology of the abscess originates from periodontopathogenic bacteria. As reported in most studies focusing on this aspect, strains of Aggregatebacter actinomycetemcomitans, Porphyromonas gingivalis, Prevotella intermedia, Fusobacterium nucleatum, Micromonas micros, Campylobacter rectus, Peptostreptococcus micros, Tannerella forsythia, and Capnocytophaga exist prevalently in periodontal abscesses.5, 6 In addition, Peptostreptococcus, Streptococcus milleri (Strep. anginosus and Strep. intermedius), Bacteriodes capillosus, Veillonella, Bacteroides fragilis, and Eikenella corrodens were found to be among other species isolated from periodontal abscesses.
2
Recent human subgingival microbiota studies based on 16S rRNA gene sequences have shown that most existing bacterial species are new species or phylotypes (subspecies etc.).
7
Studies using the similar method revealed that there are a large number of new species in the oral cavity.8, 9 New microbial agents have been identified in previous studies on periodontitis patients. In the literature, it was concluded in a study using DNA-DNA hybridization method that Prevotella tannerae, Filifactor alocis, and Porphyromonas endodontalis could be added to species used for the routine diagnosis of periodontitis-related bacterial flora.
10
Recent literature suggests identification of possible new pathogens, which are thought to be microbiologically almost identifiable. The identification of predominating bacteria in culture whose DNA isolations were registered to GenBank (
Thus, it may be suggested that different pathogens other than the known periodonto-pathogens may be present in periodontal abscess formation or periodontitis. 11 It may also be suggested that bacteria that have not yet been classified as pathogens could be an important factor for periodontal abscess.
In this study, it was aimed to isolate the most densely observed colonies among the culturable aerobic, facultative anaerobic bacteria in periodontal abscess samples and to determine them by molecular methods.
Materials and Methods
This study is an investigation in which bacteria isolation and 16S rDNA sequences are analyzed to determine whether there are different bacterial populations other than the known periodonto-pathogens in periodontal abscesses. This study was carried out between July 2016 and July 2017 on samples that meet the selection criteria in patients with periodontal abscess problem, within the scope of Dr Kubra Karacam’s MD thesis. Details are explained in the subheadings (especially under the section “Laboratorial Procedures for Bacteria Isolations and Identification”).
Research Subjects
Totally 20 volunteer patients who had visited the dental clinic with complaints of localized pain, swelling, and sensitivity, and were seen to have bleeding and suppuration on periodontal probing and diagnosed to have periodontal abscess, were included in the study as subjects.
Inclusion and Exclusion Criteria
Inclusion criteria: Patients with no systemic disease (systemically healthy and having no diabetic problems, tumors, AIDS, nephritic or hepatic diseases/disorders, gastrointestinal problems), and over 18 years old with symptoms/signs of periodontal abscess were included in the study.
Exclusion criteria: Pregnant or lactating patients, those who used antibiotics at least four weeks before recruitment, those who received any periodontal treatment in the last three months until the visit, and those diagnosed with endodontic abscess by radiographic examination and viability tests were excluded from the study.
Ethics and Informed Consent
Ethical approval was obtained from Ethics Committee (2016/13), the informed consent was signed by each participant, and they were informed about the purpose of the study.
Laboratorial Procedures for Bacteria Isolations and Identification
After mucosal decontamination was achieved, periodontal abscess contents were aspirated through a sterile injector and placed in test tubes containing PBS (phosphate buffer saline, pH 7.2) and taken immediately to the laboratory where the experiment would be completed. A serial dilution (dilution series up to 10-6) was prepared by homogenizing gently 0.1 mL abscess sample in 0.9 mL sterile phosphate buffer. Cultures of the samples were incubated for two days at 37ºC in aerobic and anaerobic (anaerobic jar and a kit system [Merck, Mikrobiologie Anaerocult A]) by sowing the cultures in media, which included TSA+Y (Trypticase Soy Agar with Yeast extract: Peptone from casein 15.0 g/L; peptone from soymeal 5.0 g/L; NaCl 5.0 g/L; yeast extract 3 g/L; Agar-agar 15.0 g/L), Colombia Blood Agar (peptone 23.0 g/L; Starch 1.0 g/L; NaCl 5.0 g/L; Agar-agar 13.0 g/L + Sheep Blood after cooled and autoclaved 50 mL/L), BHIA (Brain Heart Infusion Agar: Brain infusion solids 12.5 g/L; Beef heart infusion solids 5 g/L; Proteose peptone 10 g/L; D (+) Glucose 2.0 g/L; NaCl 5.0 g/L; Disodium phosphate 2.5 g/L, Agar-agar 15.0 g/L). All trials were conducted in three repeats. In the first part of the study, the best medium was determined in general examinations and counts. The study continued under Trypticase Soy Broth with Yeast extract (TSBY) medium and aerobic conditions, as there were no major differences in terms of number and species in general observations. On the second day of the incubation, different colonies attracting attention numerically in petri plaques were transported to new media, purified and stored until the next characterization processes at –80 ºC in Nutrient Broth (NB)-containing 16% glycerol. 11
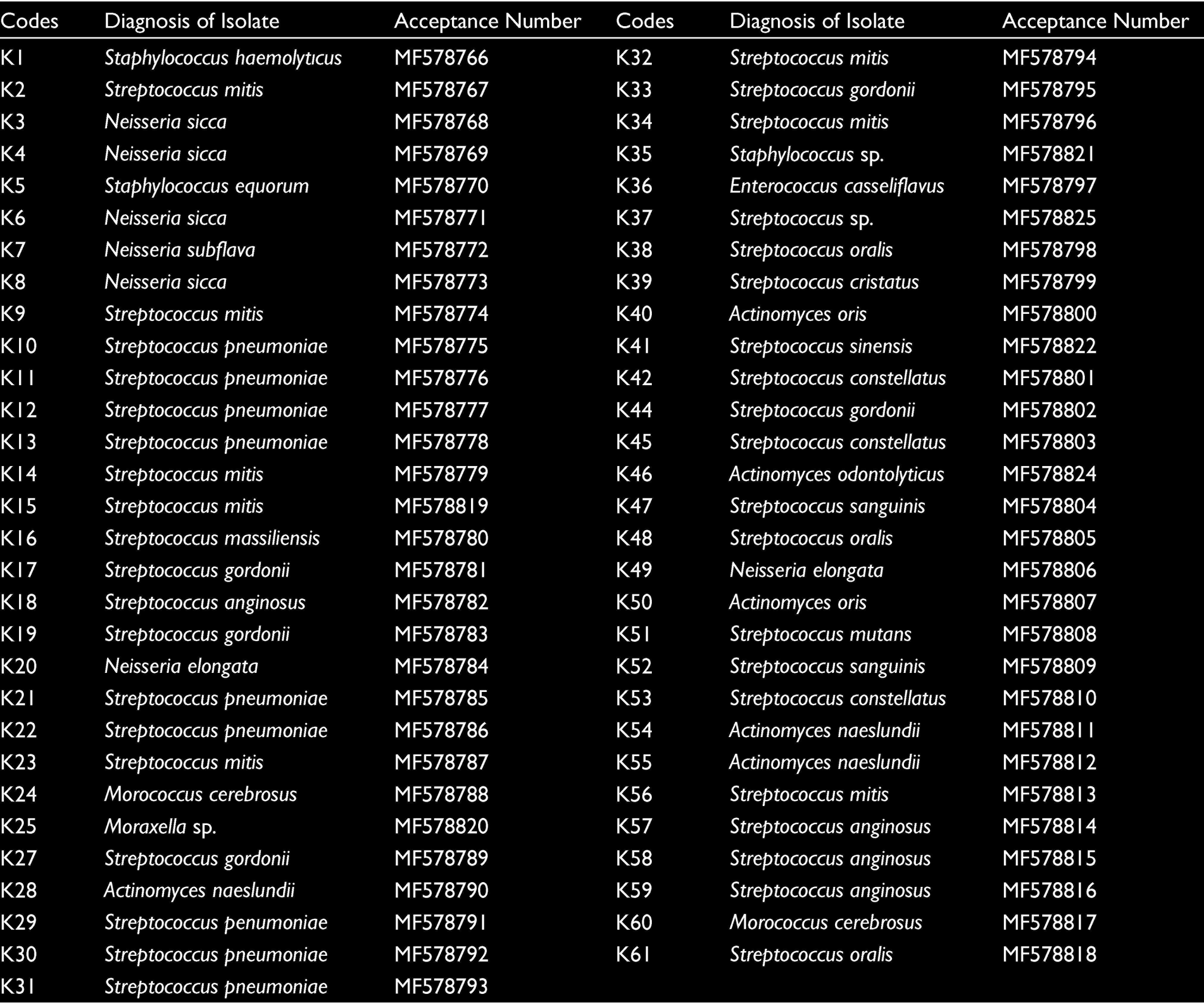
The protocol was applied for genomic DNA extraction of the isolates. For each sample for which PCR would be performed, a 30 μL reaction was prepared. A 29 μL reaction was prepared for one sample with 3 μL 10 × PCR buffer (100 mM Tris – HCl, 500 mM KCI, 15 mM MgCl2, %0.01 gelatin pH: 8.3); 0.6 μL 10 mM dNTP mix; 0.3 μL 50 μM primary 27F (forward 5’- AGA GTT TGA TCC TGG CTC AG -3’); 0.3 μL 50 μM primary 1492R (reverse 5’- GGT TAC CTT GTT ACG ACT T -3’); 1.2 μL Dimethyl Sulfoxide (DMSO); 1.8 μL MgCl2, 0.3 μL 5 unit/μL Taq DNA polymerase, and 21.5 μL sterilized dH2O. Finally, 1 μL of template DNA was added. After an initial denaturation at 95°C for 2 min, the PCR were set as follows: 1 min at 94°C, 1 min at 53°C, and 1 min at 72°C, for 36 cycles, and a final extension at 72°C for 5 min. The samples were analyzed by electrophoresis on a 1% agarose gel and then stained with ethidium bromide (0.5 µg/ml). The PCR product bands were photographed under ultraviolet light. Amplicons were sequenced by Altigen Bio Inc. (Izmir/Turkey). Sequences were compared for similarities with the nucleotide sequences in the National Center for Biotechnology Information (NCBI) library. As the result of the assessments, gene sequences were registered to GenBank by receiving acceptance numbers. 11 Information of the acceptance number is given in Table 1.
Identification of Bacteria and Acceptance Numbers
Results
A total of 61 bacterial isolates, three from each of the periodontal abscess samples taken from each volunteer, were identified and the results are presented in Table 1. These bacteria are Streptococcus, Staphylococcus, Neisseria, Actinomyces, Morococcus, Moraxella, and Enterococcus species.
In the study, the following were found to be the most prevalent species: Streptococcus species (in nine samples Strep. pneumoniae, in eight samples Strep. mitis, in five samples Strep. gordonii, in four samples Strep. anginosus, in two samples Strep. sanguinis, in one sample Strep. massiliensis, in one sample Strep. sp., in one sample Strep. cristatus, in one sample Strep. sinensis, in one sample Strep. mutans), Neisseria species (in four samples N. sicca, in two samples N. elongata, in one sample N. subflava), Actinomyces species (in three samples A. naeslundii, in one sample A. odontolyticus), Staphylococcus species (in one sample Staph. haemolyticus, in one sample Staph. equorum, in one sample Staph. sp.), Morococcus species (in two samples M. cerebrosus), Moraxella sp., and Enterococcus casseliflavus species.
Discussion
In the literature, it has been stated that pathogenic microorganisms can colonize in other parts of the body through bacteremia during periodontal abscess treatment and in rare cases different infections may develop such as endocarditis, meningitis and subdural empyema, atherosclerotic vascular disease, and brain abscess. 12 The treatment of periodontal abscesses is very important, as it can cause systemic infections associated with abscesses as well as tooth loss. The treatment for periodontal abscesses includes drainage of the abscess, irrigation, debridement, and curettage and in worst scenarios, tooth extraction. In a few cases, mechanical therapy does not suffice, thus systemic antibiotic therapy can be given. However, bacterial flora needs to be known, in order to provide an antibiotic molecule with properties of extended inhibitory activity.
The primary reasons for investigating bacterial causes of periodontitis in periodontal abscess are as follows: (a) abscesses usually develop in the periodontal pocket in individuals with periodontitis and these two periodontal diseases are considered to be related with each other, (b) useful and rapid identification techniques are not sufficient and widespread to evaluate a large number of bacterial species in samples obtained from areas with complex microbiota, and (c) there exists a tradition of focusing on a large number of few considered pathogenic species.
13
Studies on abscesses denote that the anatomical region of the infection is related to the polymicrobial etiology and microbiota of the abscesses.
14
Because of the polymicrobial etiology of abscesses, it is a natural situation that periodontitis factors are expected to be determined in periodontal abscess microbial content in studies conducted to date. Indeed, considering the reasons above, in studies investigating the presence of periodontitis factors in periodontal abscesses, the bacterial factors of periodontitis were determined in the periodontal abscess. With the development of new techniques and researches made in this direction, the presence of new bacteria in periodontitis and periodontal abscess that may be a factor in this polymicrobial etiology can be demonstrated.
In a period where newly developing technology has helped the discovery of new possible pathogens, 15 it is likely to expect that cultivable bacteria which have not yet been classified as pathogens may be the causative factors of periodontal abscess. It was aimed in the present study to isolate colony samples growing most prevalently in the media among cultivable aerobes and facultative anaerobe bacteria in periodontal abscess samples, and to identify them using molecular methods. There is no adequate study in the literature which compares present sequences in the GenBank with those detected using regional base sequence analysis and amplified using 16S rDNA PCR.
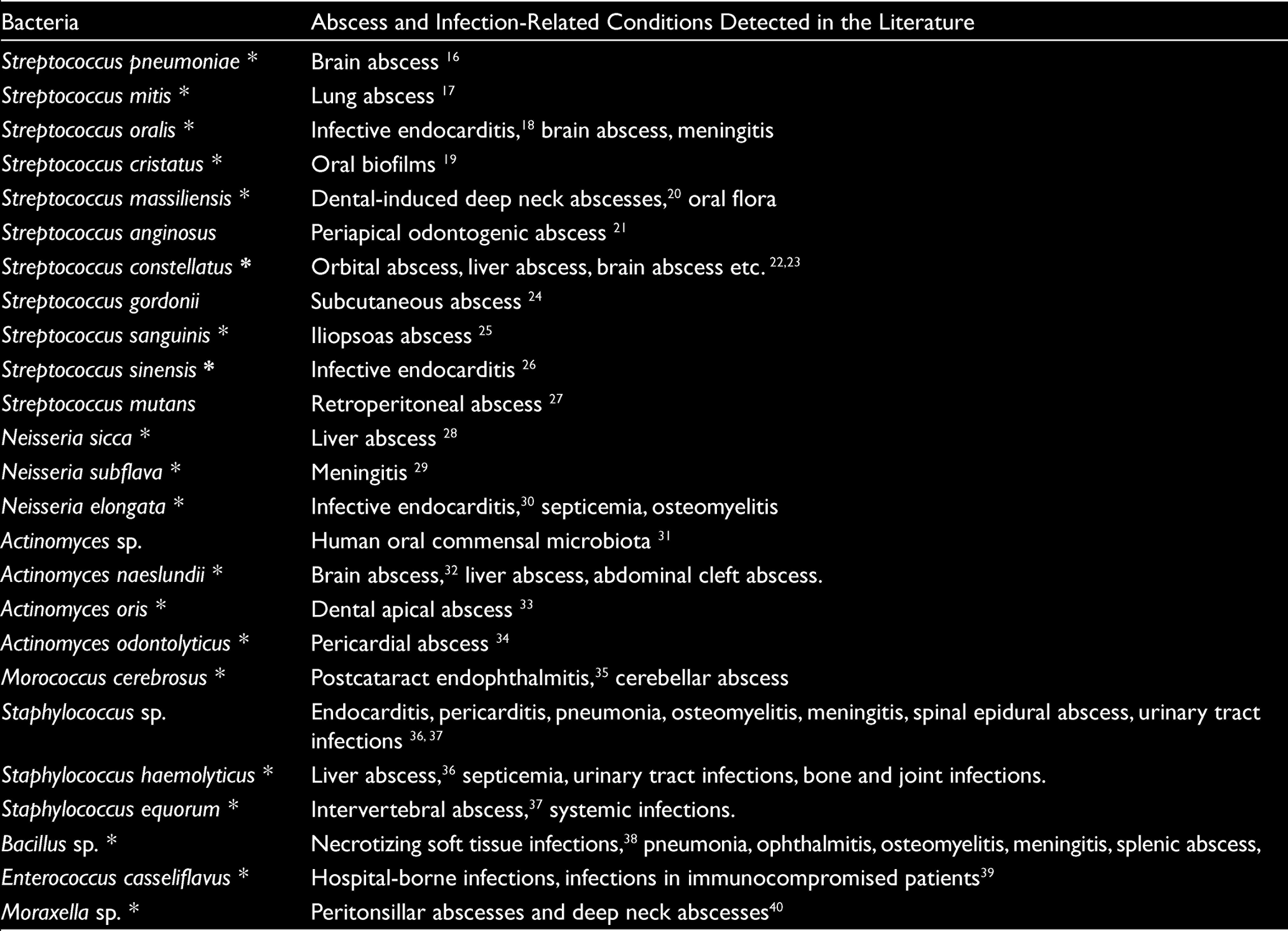
According to the research results, the bacterial species obtained and the potential problems caused by these species in different parts of the body are evaluated in Table 2.
The bacteria identified in the study were associated with infections in parts of the body other than periodontal abscess (e.g., brain abscess: Strep. pneumoniae, Strep. oralis, Strep. constellatus, Strep. sanguinis, A. odontolyticus, Morococcus cerebrosus; Bacillus spp. meningitis: Strep. oralis, N. subflava, Moraxella spp. Bacillus spp. infective endocarditis: Strep. oralis, N. elongata, Staph. haemolyticus, Moraxella spp.; liver abscess: Strep. constellatus, N. sicca, Actinomyces naeslundii, A. odontolyticus, Staph. haemolyticus, and others) and the qualitative existence of most of the bacteria isolated in this study in periodontal abscess samples was different from the existence of periodontal pathogens accepted so far, and this supports our hypothesis that contributing bacteria may also exist.
Infections of the Bacterial Species Obtained in the Study Determined in the Literature
Conclusion
The variety and quantity of bacteria in the environment are of great importance for the maintenance of periodontal health or the emergence of periodontal disease. It is also known that bacteria that are thought to be causative do not always contain the large proportion of microbiota in the first lesions of the disease and that their number increases as the disease lesion develops. Therefore, in order to reveal that the bacteria diagnosed in our study are the cause of the abscess, it is necessary to have information about the ratio between total bacteria rather than their presence in the abscess content. Bacteria determined according to these results may have the potential to reach different parts of the body and cause disease. It is recommended that the treatment of existing periodontal abscesses be studied with much greater precision, considering these conditions.
It can be foreknown that the data obtained with this study will contribute to the literature and shed light on new and comprehensive studies. As a result, it is thought that with each new data obtained, more extensive research is needed and these studies are of great importance not only for periodontology but also for oral and dental health in general, and even systemically.
Footnotes
Acknowledgements
This study is derived from the MD thesis named Identification of Bacteria Isolated from Periodontal Abscess with Culture Assay by 16S PCR and DNA Sequence Analysis.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Review Board Statement
Ethical approval was not needed as this study did not include any living individuals or participants. This was purely In-vitro study.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
