Abstract
Angiogenesis, which involves many essential processes, such as human reproduction, organ development, and wound healing, is regulated by multiple signaling pathways. QKCMP is a polypeptide with similar effects to vascular endothelial growth factor (VEGF), which promotes angiogenesis. In this study, zebrafish were treated with different concentrations of QKCMP, and it was found that QKCMP significantly promoted the growth of blood vessels. Human umbilical vein endothelial cells (HUVECs) was then treated with different concentrations of QKCMP, which proved that QKCMP could promote cell proliferation and inhibit cell apoptosis, and thus obtain a complete gene expression matrix. Genes and biological functions or pathways significantly associated with QKCMP were obtained using differential gene expression analysis, weighted gene co-expression network analysis (WGCNA), and enrichment analyses. Among them, genes significantly related to QKCMP are enriched in biological processes (BP) such as vascular formation and development, as well as the main signaling pathway: PI3K/AKT signaling pathway. The proproliferative and antiapoptotic effects of QKCMP on the HUVECs and the induction of cell cycle were then verified using cell counting kit 8 (CCK-8) and flow cytometry. Finally, it was confirmed that QKCMP promotes angiogenesis and rapid endothelialization by stimulating the PI3K-AKT and Hippo signaling pathways using quantitative real-time PCR (qRT-PCR) and western blot (WB).
Introduction
Angiogenesis is required for embryonic development and subsequent physiological homeostasis due to its effect on physiological events such as wound repair, reproduction, and placentation.1,2 Angiogenesis-dependent diseases caused by imbalances in production or activity between antiangiogenic and proangiogenic factors are the etiologies of many diseases, which involve the vascular endothelial growth factor (VEGF), an important endogenous proangiogenic factor. 3 VEGF was initially defined as vascular permeability factor (VPF) because of its vascular permeability, and later possessed its current name as it can promote angiogenesis.4,5 VEGF plays important roles in angiogenesis. It facilitates the regulation of normal and pathological angiogenesis and exerts effects of stimulating mitosis, inhibiting endothelial cell (EC) apoptosis, increasing vascular permeability and promoting cell migration in vivo. 6 In addition, VEGF also contributes to bone repair by accelerating the cartilage-to-bone transition to promote bone healing. 7
VEGF plays essential roles in proangiogenic therapies, which are vital in the context of various diseases or tissue damages. But possesses limited biological function in vivo due to its longevity or activity, thus, alternative materials are needed. 8 VEGF-mimetic peptides exert VEGF-like proangiogenic activity and can be used to present spatially oriented angiogenic cues in collagen scaffolds, which is considered to be the best alternative to VEGF-based therapy.9,10 The VEGF-mimetic peptide QKCMP is a collagen mimetic peptide (CMP) with proangiogenic QK domain through which the QK peptide can bind tightly to collagen.9,11 It has been shown that QKCMP can stimulate early angiogenesis in ECs and has the potential to promote angiogenesis in burn wounds. 11
There are a few reports on the biomedical applications of QKCMP as a new material, mostly focusing on the establishment of stable models that can effectively promote angiogenesis in vivo.8,12,13 In this study, based on the above theoretical knowledge and existing techniques, human umbilical vein endothelial cells (HUVECs) were treated with 0, 50, and 100 µM QKCMP. In vitro experiments were performed to confirm the role of QKCMP in promoting the growth of HUVECs, and the link between QKCMP and possibly related biological pathways was experimentally analyzed to verify the effect of QKCMP on the rapid vascular endothelialization.
Methods and materials
Animal
All zebrafish were raised in fish raising water at 28°C. One day after fertilization (1 dpf), transgenic vascular green fluorescent Fli-1 strain zebrafish was randomly selected and placed in a 6-well plate. Thirty zebrafish were treated in each well (experimental group). Set up Control group, Bevacizumab (Bev) group, Bev + QKCMP high/medium/low dose (100, 50, or 0 μM) group. Inject bevacizumab injection into the yolk sac. After 2 days of treatment at 28°C, 6 zebrafish were randomly selected from each experimental group and photographed under a fluorescence microscope. Data was collected using NIS Elements D 3.20 advanced image processing software to analyze the area and number of subintestinal blood vessels. The statistical analysis results of the above indicators were used to evaluate the induction of zebrafish vascular loss by bevacizumab injection.
Cell culture
HUVECs were cultured in HUVEC-specific medium at 37°C and 5% CO2 in humid air. To explore the effect of QKCMP on the growth of HUVECs, the cells were treated with QKCMP at concentrations of 0, 50, and 100 μM for 5 d, where 0 μM was defined as the Control group without QKCMP, and three samples were replicated in each group. The treated HUVECs continued to be cultured in HUVEC-specific medium. In addition, an Inhibitor group was established in this study, QKCMP-treated HUVECs were treated with PI3K-AKT signaling pathway inhibitors (LY294002, MCE, HY-10108) were then cultured in the medium for 24 h for detection. The morphological changes of HUVECs were observed under a optical microscope and recorded by taking photos.
Quantitative real-time PCR
The total RNA of the HUVECs was extracted using the TRIzol reagent to detect its concentration and quality. The cDNA template was synthesized using the PrimeScript RT kit, followed by qRT-PCR performed on a RT-PCR instrument. The relative expression levels were calculated using the 2−ΔΔCt method. The primer sequences used, with GAPDH as the endogenous control, were as follows: CTGF, Forward primer: TTGCGAAGCTGACCTGGAA, Reverse primer: TGCTGGTGCAGCCAGAAA; YAP, Forward primer: CAACTCCAACCAGCAGCAAC, Reverse primer: TTGGTAACTGGCTACGCAGG; VWF, Forward primer: CCGATGCAGCCTTTTCGGA, Reverse primer: TCCCCAAGATACACGGAGAGG; ITGA10, Forward primer: AACATCACCCACGCCTATTCC, Reverse primer: GTTGGTAGTCACCTAAGTGGC; GAPDH, Forward primer: CTGGGCTACACTGAGCACC, Reverse primer: AAGTGGTCGTTGAGGGCAATG.
Cell proliferation
Cell proliferation was determined through cell counting kit 8 (CCK-8). The HUVECs were inoculated into 96-well plates and co-incubated with CCK-8 reagent at 37°C for 2 h according to the instructions. Subsequently, the cells were treated with QKCMP at concentrations of 0, 50, and 100 μM, and incubated under the same conditions for 0, 12, 24, 48, and 72 h, respectively. The optical density (OD) values at 450 nm were measured with a microplate reader.
Flow cytometry
The apoptosis rate and cell cycle changes of the HUVECs were determined via flow cytometry. For apoptosis, it was assessed with the Annexin V-FITC/propidium iodide (PI) apoptosis detection kit. The cells treated with 0, 50, and 100 µM of QKCMP were collected separately, and stained with Annexin V-FITC/PI after washing with phosphate buffered saline (PBS). Finally, the cells were examined using flow cytometry. For cell cycle, the treated cells collected as described above were centrifuged and washed with PBS. Fixed in 70% methanol at 4°C overnight, the cells were resuspended in the PI staining solution, and incubated for 1 h at room temperature in the dark, followed by detection with flow cytometry.
Western blot
The HUVECs were washed and lysed in PBS, and the supernatant was collected after centrifugation. The protein concentration was determined using a bicinchoninic acid (BCA) kit, and the proteins were separated using sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) after the addition of a loading buffer, which were then transferred to PVDF membranes for WB analysis. The primary antibodies used in this study were all purchased from Abcam, as shown below: anti-p21 (ab109520), anti-p27 (ab32034), anti-cyclin B1 (ab32053), anti-CDK2 (ab32147), anti-CTGF (ab209780), anti-YAP (ab205270), anti-AKT (ab8805), anti-p-AKT (ab38449), and anti-β-actin (ab8227). The protein content was measured after incubating with the secondary antibody coupled with horseradish peroxide for 1 h, with β-actin as the internal reference.
RNA sequencing
Total RNA was extracted from treated HUVECs by using the Trizol kit. After enrichment, mRNA was fragmented and reverse transcribed into cDNA. Double-stranded cDNA fragments were purified and end-repaired for PCR amplification. The sequences were comparatively analyzed to detect the gene integrity and expression, obtaining three groups of complete gene expression matrix.
Differential gene expression analysis
The samples were divided into the Control group (0 µM) and the Treatment groups (50 and 100 µM), and their differences were analyzed using the limma package in R software. Screening thresholds of |log2FC| > 1 and p < 0.05 were used to obtain up- and down-regulated genes with differential expressions.
Weighted gene co-expression network analysis
Weighted gene co-expression network analysis is a method of clustering genes based on sample features. An appropriate soft threshold (β) was selected to construct topological overlap matrices and gene dendrograms. In addition, module colors were established to cluster genes into different color modules based on the correlation between sample features and genes. The gene modules with the greatest correlation were selected for subsequent analyses.
Functional and pathway enrichment analyses
Venn diagrams were used to visualize the intersection of differentially expressed genes (DEGs) and hub genes. Gene ontology (GO) and Kyoto Gene and Gene Encyclopedia (KEGG) enrichment analyses of the obtained overlapping genes were performed using ClusterProfiler software to obtain significant biological functions or approaches. Protein-protein interaction was analyzed using Metascape (https://www.metascape.org), and the biology and signaling pathway enrichment of overlapping genes was analyzed using KEGG, Reactome Gene Sets and WikiPathways. The signal pathway with p < 0.05 was considered to be significant.
Statistical analysis
All calculations were performed using Graphpad Prism 6.01. The expression differences of genes or proteins in the groups with 0, 50, and 100 µM QKCMP were calculated with two-way ANOVA. The statistical differences in apoptosis rates among the the groups (0, 50, and 100 µM) were compared with multiple t-tests. p < 0.05 was considered statistically significant.
Results
QKCMP can promote blood vessel growth and HUVECs proliferation
Compared with the control group, bevacizumab injection significantly reduced angiogenesis in zebrafish, but QKCMP can significantly promote vascular development in zebrafish, and the effect is more significant with increasing concentration (Figure 1(a)). Furthermore, by comparing the area and number of subintestinal blood vessels in each group, it can be concluded that QKCMP can significantly promote the vascular growth of zebrafish (Figure 1(b) and (c)).
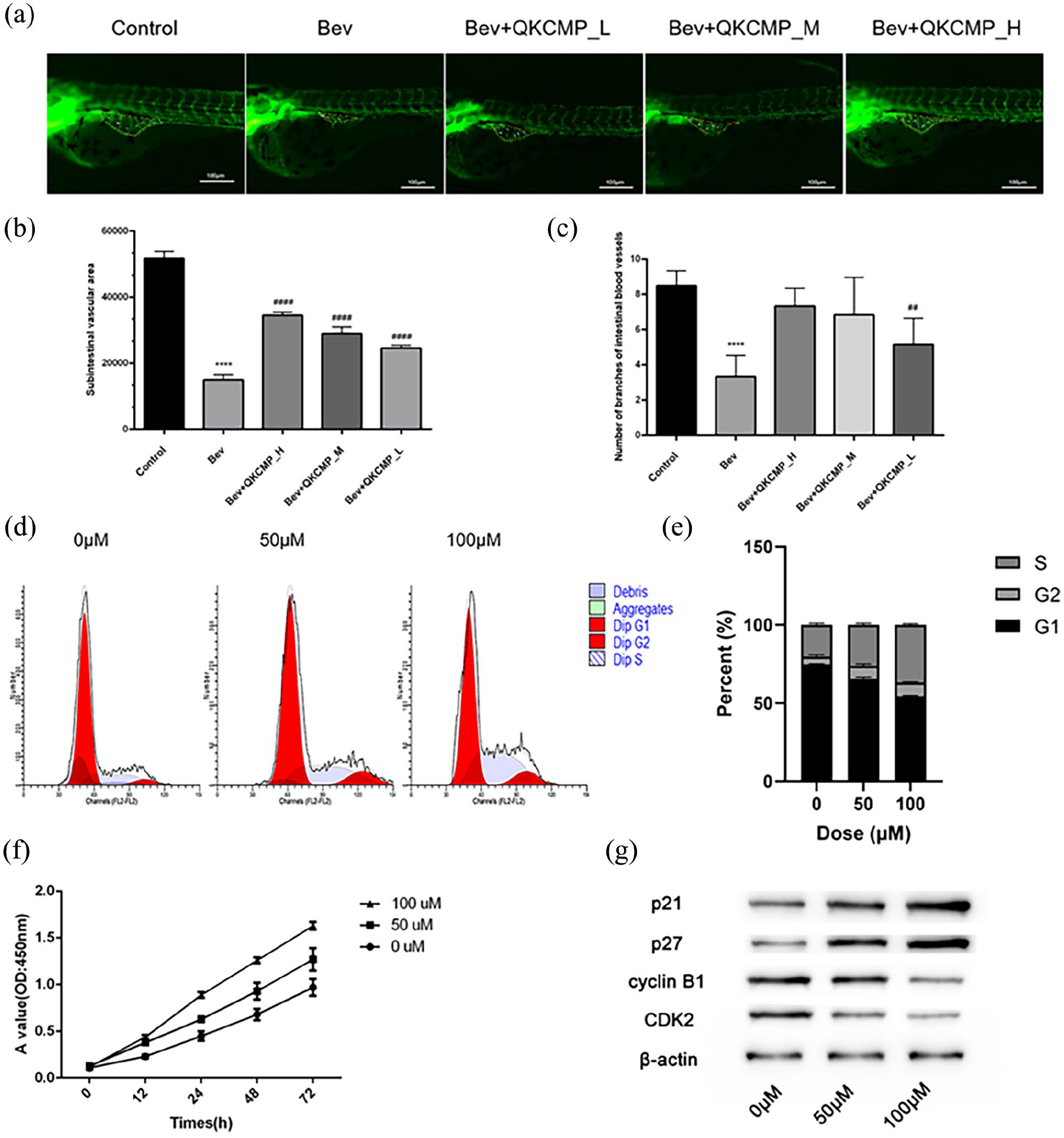
The vasotropic effect of QKCMP was verified in vivo and in vitro. (a) The effects of QKCMP on the embryonic zebrafish vascular development; (b) Subintestinal vascular area; (c) Number of branches of intestinal blood vessels; (d) Cell cycle changes of HUVECs under different concentrations of QKCMP treatment was detected by flow cytometry; (e) The percentage of different periods of the cell cycle in each group; (f) Growth curves of HUVECs under different concentrations of QKCMP treatment; (g) Cell cycle protein changes of HUVECs detected by WB under different concentrations of QKCMP treatment.
CCK-8 and flow cytometry were performed to gain insights into the biological effects of QKCMP on the HUVECs. The findings on cell cycle indicated that the cells mostly arrested in the S and G2 phases (Figure 1(d) and (e)). The CCK-8 results presented that QKCMP had a cell proliferation-promoting effect, which was positively correlated with the QKCMP concentration (Figure 1(f)). The expression of cell cycle pathway-related proteins was detected using WB and it was found that p21 and p27 increased with the QKCMP concentration (Figure 1(g)).
Analysis of hub genes and biological functions
The samples were divided into the Control group (0 µM) and the Treatment groups (50 and 100 µM), and 1575 DEGs were obtained after differential gene expression analysis. The volcano plot in Figure 2(a) displayed the gene expression distribution in the Control and Treatment groups, including 1043 up-regulated genes and 532 down-regulated genes. The heat map in Figure 2(b) showed the expression distribution of genes in the Control and Treatment groups. WGCNA was used to analyze gene modules with a high correlation with the features. Figure 2(c) presented the scale-free fit indices and average connectivity for various soft thresholds. The correlations between the gene modules and features were analyzed, and the findings showed that the “darkgreen” module had higher correlations with 50 µM (R = 0.81, p = 8.7e-3), and the “black” module (R = 0.83, p = 5.3e-3), “grey60” module (R = 0.89, p = 1.1e-3), and “lightcyan” module (R = 0.71, p = 0.03) had the highest correlation with 100 µM (Figure 2(d)). Therefore, these four modules were selected for subsequent analyses.
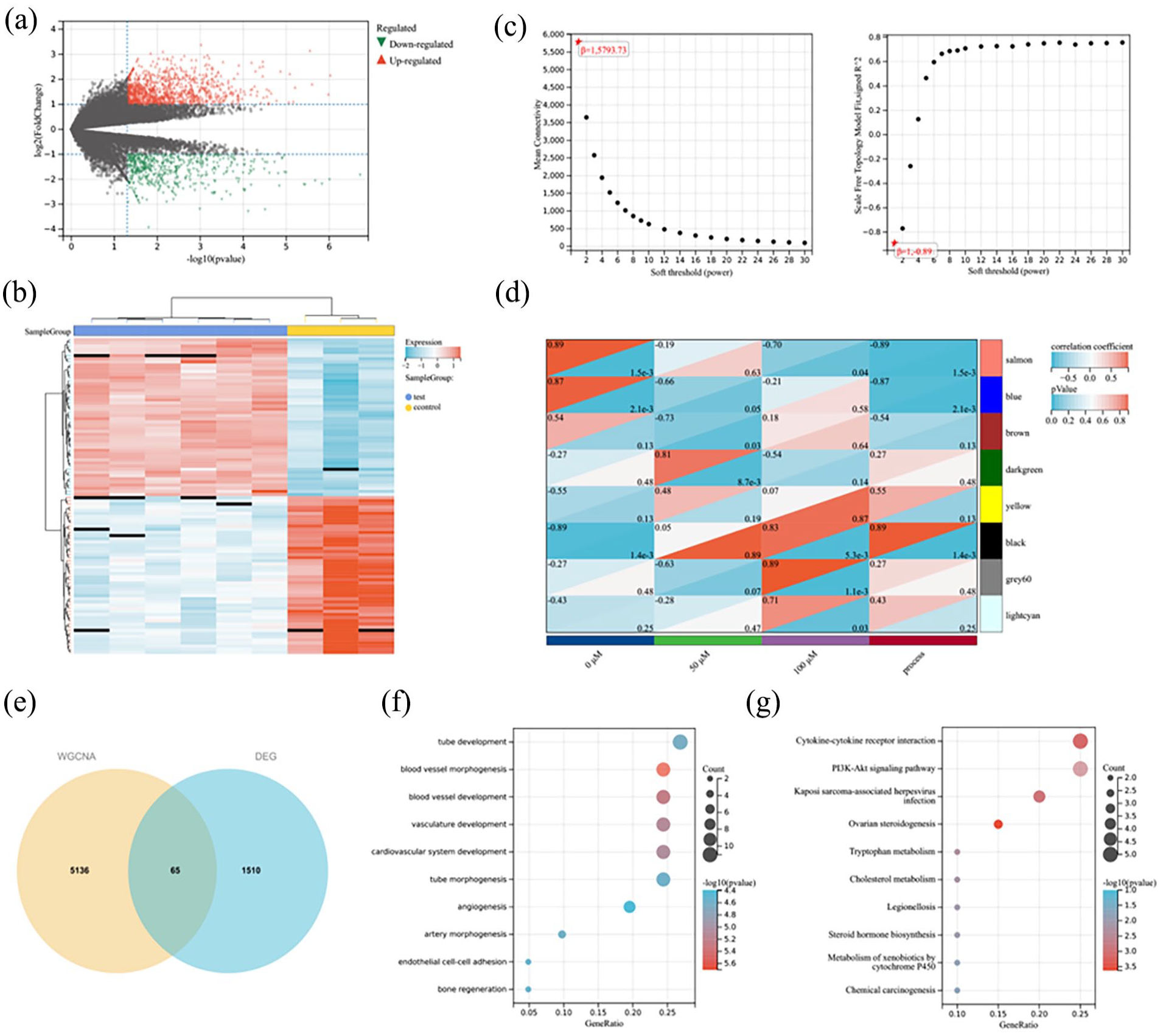
Differential gene expression analysis and WGCNA. (a) Volcano plot, where red (regular triangle) indicated up-regulated genes and green (inverted triangle) indicated down-regulated genes; (b) The heat map showed the expression distribution of DEGs in the Control group and the Treatment group; (c) Scale-free fit indices of soft thresholds and average connectivity; (d) Heat map of gene modules and feature correlations; (e) Venn diagram; (f) BP functional enrichment of overlapping genes; (g) KEGG pathway of the overlapping genes.
The intersection of DEGs and gene modules was identified using a Venn diagram to obtain 65 overlapping genes (Figure 2(e)), which were then subjected to GO term and KEGG pathway enrichment. The results showed that the overlapping genes were mainly enriched in biological processes (BPs), such as Vascular morphogenesis, Vascular development, Cardiovascular system development, and EC adhesion (Figure 2(f)), and disease or metabolic synthesis processes, such as Cytokine-cytokine receptor interactions, PI3K-AKT signaling pathways (Figure 2(g)).
Activation of PI3K-AKT pathway by QKCMP
Analysis of protein-protein interactions between 65 overlapping genes showed that Signaling pathways or biological processes such as blood vessel morphogenesis, Cytokine-cytokine receptor interaction, PI3K-Akt-mTOR-signaling pathway are significantly enriched (Figure 3(a)). In this study, the PI3K-Akt-mTOR-signaling pathway was selected as the key research object, and five DEGs (CSF3, LPAR6, GNG11, ITGA10, VWF) related to this signaling pathway were collected. Subsequently, the expression levels of 5 DEGs in different treatment groups were compared. Their transcription levels increased with the increase of QKCMP concentration, and the expressions of VWF and ITGA10 were significantly different among the three groups (Figure 3(b)). Therefore, qRT-PCR and WB methods were used for further detection. The qRT-PCR results indicated that the expression levels of VWF and ITGA10 were gradually increased with the QKCMP concentration (Figure 3(c)), and the WB assay showed the same results for the protein levels (Figure 3(d)).
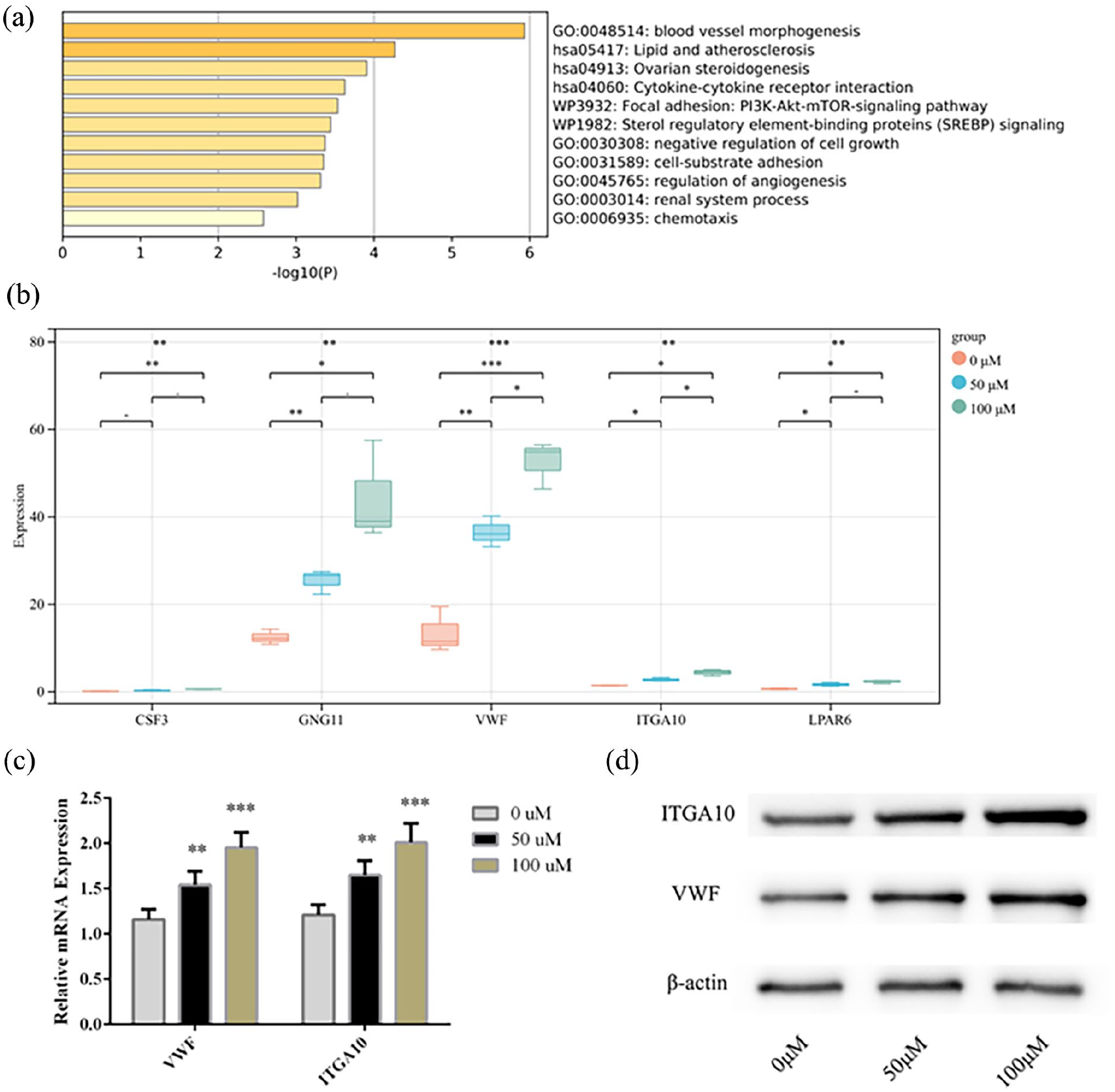
Analysis of PI3K-AKT pathway-related genes and protein levels. (a) Biological function and pathway enrichment analysis of the 65 genes; (b) Transcriptome expression levels of CSF3, LAPR6, GNG11, ITGA10 and VWF in different Treatment groups; (c) Detection of the expression levels of VWF and ITGA10 in HUVECs using qRT-PCR; (d) Detection of the protein levels of VWF and ITGA10 in HUVECs using WB. Compared with the 0 μM group.
The vasotropic effect of QKCMP was related to the PI3K-AKT/Hippo signaling pathway
The mRNA and protein levels of AKT and p-AKT in HUVECs were significantly increased with the QKCMP concentration (Figure 4(a)). After administration of LY294002, HUVEC proliferation was inhibited (Figure 4(b)). These results indicated that the vascular growth-promoting effect of QKCMP was inhibited with the inhibition of PI3K-AKT signaling. The Hippo signaling pathway is another pathway involved in endothelial cell proliferation and angiogenesis. Therefore, this study examined the expression of hub proteins of Hippo signaling pathway in HUVECs. The levels of p-YAP in HUVECs treated with QKCMP were significantly decreased, while the levels of CTGF and YAP were significantly increased with the QKCMP concentration (Figure 4(c) and (d)). This result also proves that QKCMP can activate Hippo signaling pathway.
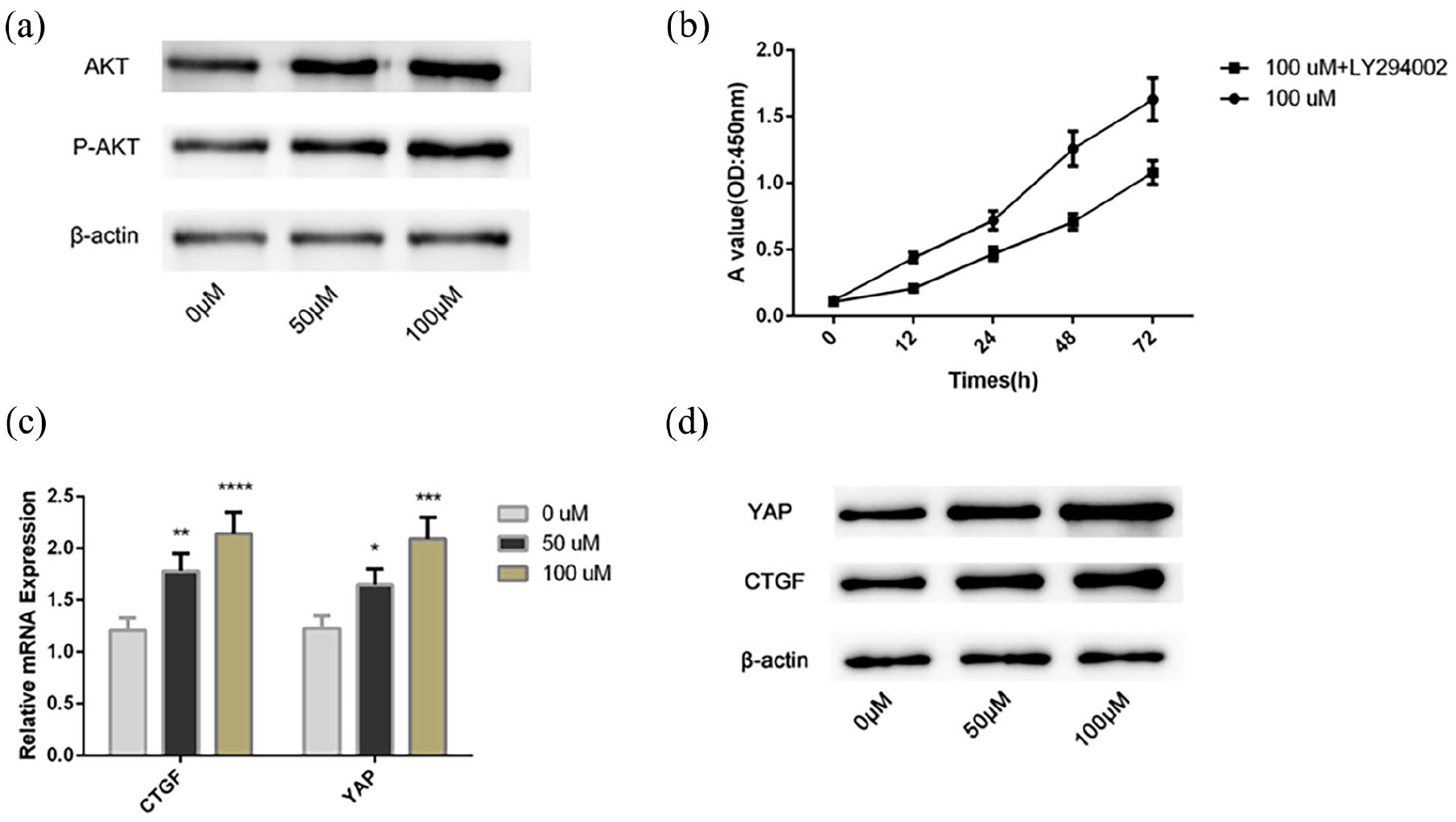
Effects of QKCMP on PI3K-AKT signaling pathway and Hippo signaling pathway. (a) The protein levels of AKT and p-AKT in HUVECs were detected by WB; (b) The proliferation levels of HUVECs after PI3K-AKT inhibitor were detected by CCK-8; (c) The mRNA levels of YAP and CTGF in HUVECs were detected by qRT-PCR; (d) The protein levels of YAP and CTGF in HUVECs were detected by WB; Compared with the 0 μM group.
Discussion
Angiogenesis is involved in important processes, such as human reproduction, organ development, and wound healing, and strictly regulated by various complex networks under normal conditions. 14 This study validated the effect of QKCMP on angiogenesis in vivo and in vitro. Zebrafish experiment showed that QKCMP can significantly promote angiogenesis and development, and with the increase of QKCMP concentration, angiogenesis and development are better. In vitro experiments also confirmed that QKCMP can promote HUVECs proliferation and inhibit apoptosis. In this study, a set of QKCMP-related genes were first obtained using bioinformatics analysis and the related functions and pathways were analyzed through enrichment analyses. The overlapping genes between the DEGs and the WGCNA hub gene modules were mainly enriched in biological functions, such as Angiogenesis and Vascular development, indicating the role of QKCMP in regulating angiogenesis. In addition, these overlapping genes were enriched in Cytokine-cytokine interactions and PI3K-AKT signaling pathways. Subsequently, in vitro experiments confirmed that QKCMP could promote HUVEC proliferation, inhibit apoptosis, and promote HUVEC growth possibly by regulating the PI3K-AKT and Hippo signaling pathways to facilitate rapid vascular endothelialization.
Angiogenesis includes degradation of extracellular matrices, EC migration, cell differentiation and proliferation, microtubule formation, and formation of new capillary sprouts. 15 The growth of vascular ECs includes several processes, such as proliferation, attachment, and migration, and plays an important role in preventing blood clotting. 16 VEGF can promote EC regeneration and angiogenesis. 17 In this study, the promotion of HUVEC proliferation and inhibition of apoptosis by the VEGF-mimetic peptide QKCMP were confirmed through in vitro experiments. Existing findings have showed that high levels of QKCMP could be immobilized on an artificial matrix and implanted in vivo to exert its proangiogenic effects. 11 However, this study focused on its relationship with the PI3K-AKT and Hippo signaling pathways, both of which were involved in regulating cell growth and associated with angiogenesis.18,19
The literatures have revealed that the PI3K-AKT signaling pathway is mostly associated with cancer progression due to its function as a critical signaling pathway to promote tumor development through gene regulation.20,21 Furthermore, angiogenesis and vascular formation are required for tumor growth and metastasis as active ECs in growing tumors release many proangiogenic factors to promote blood vessel growth. 22 VEGF-A, a vital proangiogenic factor, is crucial in embryonic development, wound healing, and angiogenesis. 23 The PI3K-AKT signaling pathway is an essential regulatory pathway for angiogenesis. 24 When VEGF-A is expressed, it can promote angiogenesis and tumor growth by inducing the PI3K-AKT signaling pathway. 25 Nanomaterials can activate the PI3K-AKT signaling pathway by releasing VEGF to affect vascular endothelialization. 26 In this study, two hub proteins associated with PI3K-AKT signaling pathways were used to verify the effect of QKCMP, and it was found that their protein levels were proportional to the QKCMP concentration. Thus, QKCMP can facilitate the regulation of the PI3K-AKT signaling pathway and may promote angiogenesis by activating the PI3K-AKT signaling pathway.
The Hippo signaling pathway, associated with EC proliferation and angiogenesis, 19 is an evolutionarily conserved signaling cascade consisting of an MST-mediated kinase cascade and involved in biological pathways such as regulating cell growth, controlling organ size and regeneration.27,28 Some studies have shown that Hippo-YAP signaling pathway has become a new pathway for vascular development. 29 YAP and CTGF are hub target genes of Hippo signaling pathway, which regulates EC shape and behavior through actin cytoskeleton dynamics. 29 VEGF signaling is associated with the Hippo signaling pathway when delivered to transcriptional programs. 30 The expression of VEGF contributes to the activation of YAP, and YAP-mediated changes in gene expression indirectly promote vascular growth and EC remodeling. 30 In this study, QKCMP with a similar effect to VEGF promoted the expression levels of YAP and CTGF in HUVECs. However, YAP and CTGF levels were significantly decreased, and the proliferation of HUVECs was affected after administration of LY294002. The reason for this phenomenon was probably that LY294002 hinder the effect of QKCMP and inhibit LATS1/2, resulting in the inability of unphosphorylated YAP to transfer into the nucleus and futher regulate the expression of downstream factor CGTF. 31 Therefore, it is clear that QKCMP regulates vascular development through the PI3K-AKT/Hippo signaling pathway.
Although this study found that PI3K/AKT/Hippo signaling may be a potential pathway for QKCMP to promote vascular endothelialization, there are still some limitations. QKCMP, as a simulation peptide of VEGF, has a similar mechanism to VEGF, and further studies should be conducted to compare the similarities and differences between QKCMP and VEGF in function and regulatory mechanism. In addition, the stability and efficacy of QKCMP and its regulatory effect on the PI3K/AKT/Hippo signaling pathway need to be verified in vivo.
Conclusion
Previous studies focused on the application of QKCMP as a new material in biomedicine, while this study focused on analyzing the mechanism of action of QKCMP in promoting vascular endothelialization. This study first confirmed that QKCMP can promote proliferation, inhibit apoptosis and promote vascular growth in a concentration-dependent manner, reflecting the rapid vascular endothelialization effect of QKCMP. Subsequent bioinformatics analysis and in vitro experiments further verified that the promotion effect of QKCMP on vascular endothelialization is achieved by regulating PI3K/AKT/Hippo signaling, and the potential mechanism may be to increase the levels of YAP and CTGF in Hippo signaling pathway by activating PI3K/AKT signaling.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Ningbo Science and Technology Innovation 2025 Major Project (2020Z094, 2022Z125), Key Laboratory of Precision Medicine for Atherosclerotic Diseases of Zhejiang Province (2022E10026), and Ningbo Top Medical and Health Research Program (No. 2022020304).
Ethical approval
The license number of experimental animals is SYXK (Zhejiang) 2022-0004, and the feeding management meets the requirements of international AAALAC certification (certification number: 001458), and the IAUC ethics review number is IAUC-2023-7624-01.
