Abstract
The mechanical properties of Co–Cr–Mo (CCM) alloys are advantageous in various biomedical applications. However, because of their bioinert surface, CCM alloys exhibit poor endothelial cell attachment properties; thus, problems of biocompatibility remain. In this study, we aimed to improve the biocompatibility of the CCM alloy surface using solid-binding peptides. We selected peptides with high binding affinity for cast CCM alloy surfaces through in vitro evolution by the phage display method. The peptides were functionalized on the CCM alloy surfaces by simple immersion in the peptide solution. The peptide bound to both cast and 3D-printed CCMs with the same affinity. The peptides linked to the amino acid motif that promotes cell adhesion, and improved the attachment of endothelial cells on the 3D-printed CCM in serum and serum-free conditions. Hence, CCM-binding peptides are attractive tools for constructing a biofunctional surface on CCM-based biodevices.
Keywords
Introduction
Co–Cr–Mo (CCM) alloys have been widely used in medical devices, such as coronary stents, dental implants, and artificial hip joints.1–3 However, because of their high melting point and hardness, fabrication of devices with these alloys by conventional processes is typically expensive and time-consuming. 4 Recent developments in three-dimensional (3D) metal printing technology could overcome the above-mentioned limitations. Products generated through such methods tend to exhibit better mechanical properties than cast CCM alloys and also satisfy the type 5 criteria of ISO22764. 5 CCM alloys also exhibit better corrosion resistance; however, their surface is bioinert. Previously, researchers found that 3D-printed CCM and cast CCM have similar cell attachment properties but that these are drastically lower than that of titanium. 6 To address this issue, surface modification can be used as it allows the construction of a bio-surface that improves the tissue compatibility of CCM.
Modification with biofunctional molecules regulates the function of artificial solid surfaces, such as metallic biomaterials, 7 toward cells and extracellular matrix (ECM). However, modification of metallic surface with ECM proteins is difficult because solid surfaces sometimes induce conformational changes in the proteins leading to protein inactivation. Hence, short peptides derived from the functional site of ECM are useful. The Arg-Gly-Asp (RGD) peptide is the most known peptide motif derived from fibronectin. It acts on the cellular scaffold via integrin, a cell membrane protein. RGD-modified metal surfaces offer a better environment for the attachment of endothelial cells.8,9 Furthermore, the Arg-Glu-Asp-Val (REDV) peptide was found to be a specific ligand for endothelial cells derived from fibronectin, improving biocompatibility of CCM-based coronary stents.10–12
Several techniques, including physical, chemical,13,14 and electrochemical methods,15–17 have been employed to immobilize biofunctional molecules on the metal surface. In this study, we have utilized the material-specific binding properties of solid-binding peptides (SBPs).18–22 SBPs are identified by the in vitro evolution technique using the phage display method. They are attractive molecular linkers for immobilizing biofunctional molecules, because SBPs can be conjugated to other proteins and peptides, either chemically or through molecular cloning. The conjugates of SBP with functional proteins or other biofunctional molecules are beneficial since their biological activity is generally retained despite the conjugation. 22 These constructs are likely to be non-toxic and are stable both chemically and physiologically. Moreover, they present a rigid molecular orientation on the surface of the metal. 18 In addition, surface treatments to functionalize SBPs are usually simple. The material surface can be functionalized by immersion in the peptide solution. Thus, the conjugation of biofunctional molecules to SBP would provide a one-step process for constructing a bio-interface. However, SBPs for CCM alloys have not been identified so far.
The aim of this study was to select and identify the CCM alloy binding peptide by in vitro evolution using the phage display method. Peptides were selected from a random peptide pool, and their binding property to cast CCM alloy was tested. A peptide that could bind to 3D-printed CCM was also identified from this peptide library. Then, the peptides were conjugated to RGD peptide to improve cell adhesion. The fusion peptides were applied to functionalize 3D-printed CCM alloy and to demonstrate improvement in the adhesion of endothelial cells.
Methods
Selection of CCM-binding peptides
A PhD-12 phage display peptide library kit (New England Biolabs (NEB), Beverly, MA, USA) was employed for isolation of CCM-binding peptides (BPs). The CCM alloy (ASTM F75) disk (∅10 × 1.0 mm) surface was polished with a 400-grit polishing paper and sonicated in acetone for 15 min three times, then irradiated using an UV/ozone cleaner (PC-440; Meiwafosis, Tokyo, Japan) for 30 min to remove any extraneous material from the surface. The panning procedure was performed according to the phage display manual. The phages eluted from the CCM surface at pH 2.2 were collected and amplified. Then, their DNAs were isolated and sequenced using the 96 gIII sequencing primer (NEB).
Characterization of the binding ability of CCM-binding peptides
To evaluate the binding ability of CCM-binding peptides, we used the N-terminal biotinylated peptides (Thermo Fisher Scientific, Waltham, MA, USA). CCM disks were immersed in 1 μg/mL peptide solutions for 1 h at room temperature. After immersion, the disks were washed with phosphate buffered saline (PBS) pH7.2. Then, the disks were immersed in 0.1 μg/mL streptavidin-horseradish peroxidase (HRP) (Thermo Fisher Scientific) for 1 h at room temperature. Finally, the disks were washed with PBS pH7.2, and were immersed in peroxidase assay solution containing 0.08 mg/mL of 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid) (ABTS) (Tokyo Chemical Industry, Tokyo, Japan) and 0.17 (v/v)% H2O2 in 50 mM sodium citrate buffer pH 4.0. After 30 min of incubation, the absorbance of the solution was measured at 415 nm using a microplate reader (MTP-880Lab; Corona Electric, Ibaraki, Japan).
Evaluation of the binding ability of peptides to 3D-printed CCM
The 3D-printed CCM was fabricated by selective laser melting process as reported in a previous study. 6 EOS MP1 powder (Co-28Cr-6Mo; EOS), which has the same content as ASTM F75, was used in this study. The surface was cleaned with an UV/ozone cleaner (PC-440; Meiwafosis Co., Ltd.) for 30 min before the evaluation of the binding ability of CCM BPs. The binding strength was evaluated by the method described above, and compared with that of the cast CCM alloy.
Cell culture
Bovine pulmonary artery endothelial cells (CPAE) were obtained from RIKEN Bio Resource Research Center (Tsukuba, Japan). CPAE were cultured in minimum essential media α (MEMα) (Wako Pure Chemical Industries, Osaka, Japan) containing 20% fetal bovine serum (FBS) (Biosera, Sussex, UK) and 100 unit/mL penicillin and 100 μg/mL streptomycin (Nacalai Tesque, Kyoto, Japan) in a 5% CO2 incubator at 37°C.
CPAE attachment activity test
CCM disks were polished using #1200 silicon carbide (SiC) polishing paper to obtain similar surface roughness among the disks, because surface roughness leads to changes in the cellular function.23–25 The cell adhesive peptide Arg-Gly-Asp-Ser (RGDS) was fused at the C-terminal of CCM-BP04, and was named CCM-RGD. 3D-printed CCM was sterilized and immersed in 1 g/mL CCM RGD peptide for 30 min. CPAE were seeded at a cell density of 3,000 cells/cm2 onto the 3D-printed CCM, and cultured for 4 h. After cultivation, the surface was washed with PBS to remove non-adhered cells. Then, the cells on the surface were fixed with 70% ethanol for 5 min and then stained with 0.5% crystal violet (Wako Pure Chemical Industries). Adhered cells were captured with a digital microscope system (Keyence, Osaka, Japan), and were counted under a microscope (Wraymer, Osaka, Japan). Cell attachment activity was calculated as the ratio of the number of adhered cells to the seeding number of cells.
Statistical analysis
For the peptide binding ability test, the experiments were performed in triplicate. Statistical differences were determined by one-way ANOVA analysis with the Dunnett’s test for multiple comparisons. For the CPAE attachment activity test, average and standard division were calculated from triplicate quantitative analysis of five disks. Statistical differences were determined by Student’s t-test. A p < 0.05 was considered significant.
Results
We utilized a phage displayed library that provided linear 12-mer peptides on the surface of M13 phage with a diversity of 4.1 × 1015 for the screening of CCM-binding peptides. After four rounds of panning procedure toward CCM alloys, 85 clones were randomly picked from the phage pool. They were then sequenced. Characterization results of the CCM-binding peptides are listed in Table 1. Although the selected peptides had different sequences, they contained more positively charged amino acid residues than negatively charged amino acid residues.
Amino acid sequence of peptides used in this study.

Calculated by subtracting the number of basic residues (R, H, and K) from the number of acidic residues (D and E).
Hydropathicity index was calculated using ProtParam tool.
Gold-binding peptide from Brown et al. 26
Quantification of peptide binding to CCM surface was determined by HRP-activity using ABTS chromogenic substrate. To evaluate the binding affinity of the selected peptide, biotinylated peptides were used. After incubating the biotinylated CCM-binding peptides on alloy, streptavidin fused HRP was added, then free streptavidin fused HRP was removed from the surface. The ABTS solution changes the color to blue-green, in which absorbance maximum is 415 nm, by HRP activity. Therefore, CCM BPs binding ability was exhibited as absorbance at 415 nm.
Figure 1 shows the binding affinity of CCM-binding peptides to CCM, Ti, and SUS 316L stainless steel. CCM BP-04 and -08 bound to CCM disks in significantly higher amounts than the control peptide (GBP). CCM BP-02, -04, -06, and -08 exhibited significantly higher binding affinity to CCM than to Ti and SUS 316L. CCM BP-04 showed the highest specific binding affinity to CCM. Therefore, BP-04 was selected for evaluation of binding to 3D-printed CCM.

CCM BPs binding to CCM (black bar), Ti (gray bar), and SUS 316L stainless steel (white bar). The results are presented as the mean ± SD (n = 3).
Figure 2 compares the binding affinities of the peptides onto the 3D-printed CCM. CCM BP-05 showed the lowest binding affinity for cast CCM alloy, and was not statistically different from the negative control. Moreover, CCM BP-04 showed four times higher binding affinity to 3D-printed CCM than to CCM BP-05. Hence, CCM-RGD peptide was synthesized using CCM BP-04. Figure 3 shows the scanning electron microscopy (SEM) images of the cast CCM and of the 3D-printed CCM, which were polished with 400-grid and 1200-grid polishing paper, respectively. The surface topography and roughness were different between as-cast CCM and 3D-printed CCM; however, the CCM BP-04 binding ability was similar.
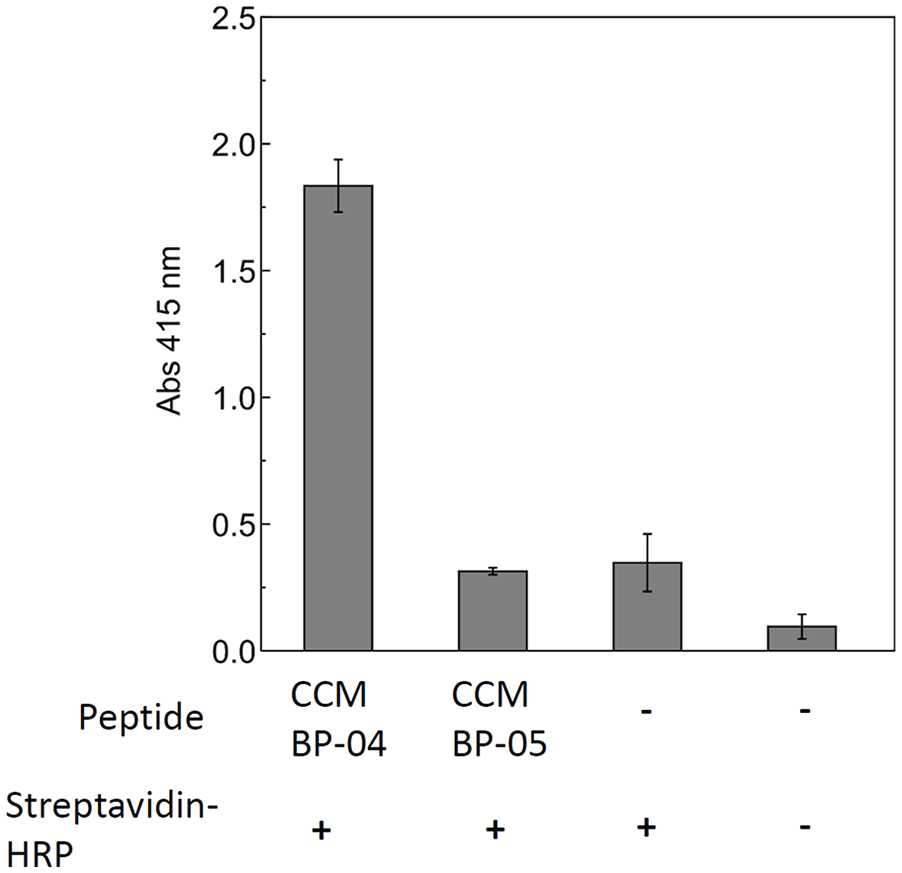
CCM BPs binding to the as-prepared 3D-printed CCM disk. The results are presented as the mean ± SD (n = 3).

SEM images of the CCMs. (a) The as-cast CCM surface that was polished with a #400-grid polishing paper and used in the peptide-binding test. (b) The 3D-printed CCM that was polished with a #1200-grid polishing paper and used in the endothelial cells attachment test.
Figure 4(a) shows the effect of CCM-RGD peptide functionalization on the attachment of endothelial cells. The peptide induced higher cell attachment activity in the culture medium with and without serum. Figure 4(b) shows the images of cells on the 3D-printed CCM surface. In a serum-containing medium, there was no difference in cell shape between the peptide-immobilized CCM and the control CCM. However, in serum-free conditions, the peptide-immobilized CCM displayed higher cell attachment and greater cell spread. The cells exhibited adhesion failure on the control CCM in serum-free conditions, as shown by the arrows in Figure 3(b). Therefore, the CCM-RGD peptide had a considerable effect on cell adhesion and spread even in the serum-free condition.
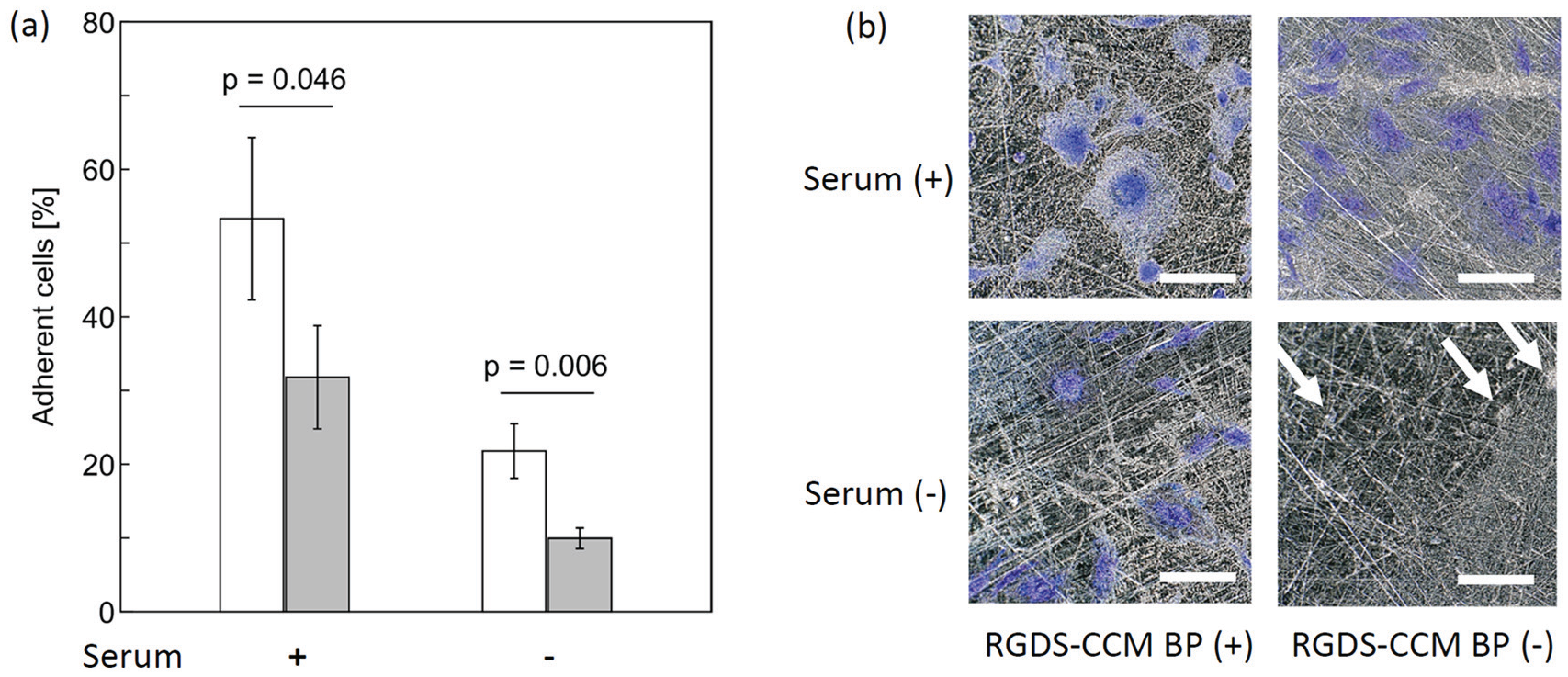
Endothelial cell adhesion on 3D-printed CCM. (a) Percentage of adherent cells; white bars represent cells attached on CCM coated with CCM-RGD peptide and gray bars represent that on bare CCM. (b) Phase contrast images of cells on the substrate. Scale bar (white) = 100 μm.
Discussion
CCM BP-04 and BP-08 showed higher specific binding to CCM than other CCM BPs (Figure 1). To better understand this phenomenon, the binding constants of CCM-RGD peptide in the serum-containing medium will be determined in the next step of this study. Although the binding mechanism of CCM BPs is still unclear, our results show that BP-04 and BP-08, which had similar charge and pI, exhibited higher binding affinity to CCM than other peptides (Table 1). Therefore, the charge and/or pI value are possibly important for the binding of CCM BPs to CCM. Interestingly, the binding strength of BP-04 to both cast CCM and 3D-printed CCM was similar. The fact that the SLM 3D printing process produces materials with better mechanical properties than those of cast CCM, 5 suggests that the process would be of no effect to protein adsorption because the cell attachment to 3D-printed CCM was similar to that to as-cast products. 6 Hence, BP-04 could recognize 3D-printed CCM as cast CCM.
The cell attachment activity of CCMs was drastically lower than that of Ti, 6 because the surface of CCM is bioinert.27,28 The RGDS cell adhesion motif induces integrin-mediated cell adhesion. 29 CCM-RGD peptide bound specifically to 3D-printed CCM and could provide a scaffolding function onto the surface for integrin-mediated cell adhesion. In the serum-containing medium, the cells could attach onto 3D-printed CCM with or without the peptide because the serum components also provided the scaffolding function to the cells. However, CCM-RGD peptide induced significantly higher cell attachment in the serum-containing condition. Although there were concerns that peptides might detach due to competitive adsorption from elements present in the serum, the serum was found to exhibit scaffolding function for the 3D-printed CCM during the cell attachment and spreading processes. Understandably, the peptide effect was markedly less pronounced in the serum-free condition than that in the serum-containing medium.
The conjugation of RGDS with other biomolecules could greatly enhance the functionality of the surface modification.30–32 In fact, a fused peptide construct, containing both RGDS and laminin-derived YIGSR (Tyr-Ile-Gly-Ser-Arg) motifs, 33 exhibited 10-fold higher adhesion to endothelial cells than that to SMCs.9–11 In addition, RGD-functionalized biomaterials surface promotes not only cell adhesion but also various in vitro and in vivo phenomena, such as proliferation, 34 differentiation,34–36, endothelialization, 37 and wound healing. 38 The CCM BPs linked to these peptides could be effectively functionalized on the CCM surface and provided biocompatibility. However, the stability of CCM BP binding layer on CCM alloy is unclear. To demonstrate the stability, long-term cellular events such as proliferation and differentiation need to investigate. While the immobilization process with biomolecules usually requires multiple steps, the CCM BPs could be functionalized by a simple one-step immobilization method for coating biofunctional molecules on CCM alloys.
Conclusions
In this study, a CCM BP peptide with a high binding affinity to cast CCM and 3D-printed CCM was selected by the phage display method. The CCM-RGD peptide, a fusion peptide of CCM BP and RGDS cell adhesive peptide, enhanced the adhesion of endothelial cells on the surface of 3D-printed CCM in serum and serum-free conditions. The coating process was simple. Collectively, our results show that the CCM BPs would be an attractive tool for constructing biofunctional CCM-based biodevices.
Footnotes
Acknowledgements
The authors would like to thank Ms. Satomi Kohara, Ms. Satomi Magae, and Ms. Tishiko Ishizuka for their technical assistance with the experiments. We are deeply grateful to Dr. Tetsushi Taguchi (NIMS, Japan) for providing alloy disks and technical suggestions. Part of this work was conducted at the NIMS Molecule & Material Synthesis Platform, supported by the Nanotechnology Platform Program of the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan. We would like to thank Editage (![]() ) for English language editing.
) for English language editing.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
