Abstract
Background:
Breakthrough studies have shown that pluripotent stem cells are present in human breast milk. The expression of pluripotency markers by breast milk cells is heterogeneous, relating to cellular hierarchy, from early-stage multi-lineage stem cells to fully differentiated mammary epithelial cells, as well as weeks of gestation and days of lactation.
Design and methods:
Here, we qualitatively analyze cell marker expression in freshly isolated human breast milk cells, without any manipulation that could influence protein expression. Moreover, we use electron microscopy to investigate cell-cell networks in breast milk for the first time, providing evidence of active intercellular communication between cells expressing different cellular markers.
Results:
The immunocytochemistry results of human breast milk cells showed positive staining in all samples for CD44, CD45, CD133, and Ki67 markers. Variable positivity was present with P63, Tβ4 and CK14 markers. No immunostaining was detected for Wt1, nestin, Nanog, OCT4, SOX2, CK5, and CD34 markers. Cells isolated from human breast milk form intercellular connections, which together create a cell-to-cell communication network.
Conclusions:
Cells freshly isolated form human breast milk, without particular manipulations, show heterogeneous expression of stemness markers. The studied milk staminal cells show “pluripotency” at different stages of differentiation, and are present as single cells or grouped cells. The adjacent cell interactions are evidenced by electron microscopy, which showed the formation of intercellular connections, numerous contact regions, and thin pseudopods.
Keywords
Introduction
Mother’s milk distinguishes mammals from other species and could be considered a new “tissue” or even “temporary organ,” which substitutes for the placenta after childbirth. Mother-offspring biological communication is pronounced by the presence of thousands to millions of cells in the breast milk. 1 Leukocytes constitute less than 2% of mature human milk from a healthy mother and infant, while billions of other cells, ingested every day by the breast-fed infant, are classified as non-immune cells. A breakthrough study showed the presence of stem cells in the human breast milk, including hematopoietic stem cells, mammary stem cells, mesenchymal stem cells, neuro-progenitor cells, and myoepithelial progenitor cells. 2 Amongst these heterogeneous cell populations, 10%–15% of the cells constitute a sub-population with mesenchymal stem cell properties, able to differentiate into distinct cell lineages.3,4
In 2012, Hassiotou et al. 3 reported the expression of pluripotency markers by cell subpopulations in breast milk for the first-time, including POU class 5 homeobox 1 octamer-binding transcription factor 4 (OCT4), sex determining region Y-box 2 (SOX2), and nanog homeobox. 5 Of note, the gene expression levels differed among lactating women, according to the stage of lactation and the degree of breast fullness. 6 Moreover, the varying levels of expression of the pluripotency genes showed the cellular hierarchy, starting from early-stage multi-lineage stem cells, through differentiated progenitor cells, to fully differentiated mammary epithelial cells.3,6
A unique marker selective for mammary gland stem cells (MaSCs) has not been identified in the literature, and several mammary stem cell markers are debatable, likely due to various experimental protocols and culturing methods. 7 However, the limited data of marker functionalities and their expression pattern provide a useful tool for identification and isolation of stem cells. Further, isolated stem cells present in the milk can be cultured to study their pluripotency in various culture conditions. 8 Breast milk cultured stem cells have multiple mesenchymal stem cell surface markers like Cluster of differentiation 44 (CD44), CD29, SCA-1 and are negative for CD33, CD34, CD45, CD73, nestin, vimentin, smooth muscle actin and E-Cadherin, an epithelial to mesenchymal transition marker.4,9 These results are strongly influenced by in vitro conditions, however. 2 Moreover, caution should be taken regarding the analysis and the interpretation of tissue immunostaining and milk flow cytometry data, while the markers currently known to distinguish different mammary cell populations are not definitive. 1
In the last decade, several reports 10 have demonstrated the plasticity of mesenchymal stem cells and generated a great deal of excitement as to their potential for cell-based therapies.11,12 Moreover, fetal and perinatal programming may increase our susceptibility to certain diseases in childhood and adulthood (reference: Fetal programming of COVID-19: May the barker hypothesis explain the susceptibility of a subset of young adults to develop severe disease. European Review for Medical and Pharmacological Sciences 25, 5876–5884). The pluripotency of mesenchymal breast milk cells has been confirmed by their ability to differentiate into adipogenic, chondrogenic, and osteogenic lineages. 13 The presence of different mesenchymal markers may change during lineage maturation induced by specific differentiation in in vitro systems. 14 Noteworthily, different properties of maternal milk samples may be influenced by the presence of distinct subpopulation of milk mesenchimal stem cells. Therefore, the study of mesenchimal cells is very difficult and a better standardization of the system used to isolate or to maintain these cells is mandatory.15,16
The local environment, namely the stem cells niche, is one of the most important elements able to change the stem cell capacity.17,18 Cell adhesion or the interaction between different types of cells is thought to be one of the most important control mechanisms of stem cell activity.19,20 In 2007 Cregan et al. 9 showed that cells isolated from breast milk and cultured in growth medium form colonies containing cells positive for the markers Cytokeratin 5 (CK5) and nestin. In 2015 Sani et al. 21 observed the formation of the colonies, but only some of them grew into larger cell aggregates, while most of the colonies expanded and produced a number of single cells within days. While different studies focused on cell surface marker expression in breast milk cells, less attention has been paid to cell-cell interactions and colony formation.
The aim of this study was to better characterize the qualitative expression of several stem cell markers (CD44, CD133, CD45, CD34, Wilms’ tumor protein 1 (Wt1), nestin, Nanog, OCT4, SOX2, CK5, CK14, P63, thymosin β4 (Tβ4)) and the cell proliferation marker Ki-67 antigen, in samples isolated from human breast milk without any particular manipulation, which could influence the marker expression in vitro. Moreover, we investigated the cell-cell interactions and colony formation among cells expressing different surface markers.
Materials and methods
Breast milk samples recruitment
Breast milk samples were recruited from six healthy volunteers. The experimental analysis were authorized by the Comitato Etico Indipendente Ospedaliero (Università di Cagliari; Prot. PG/2022/795). The samples were collected in the morning with the use of a manual breast pump. The samples were stored at 5°C and delivered to the laboratory within 60 min for the cell isolation process.
Breast milk cells isolation
Breast milk cells were isolated from fresh milk, diluted with an equal volume of Dulbecco’s phosphate-buffered saline (DPBS; pH 7.4; Gibco) and centrifugated at 810g for 20 min at 20°C. The fat layer and liquid were removed, while the cell pellet was washed twice with DPBS. The pellet was fixed with ThinPrep solution (Hologic, Inc. www.hologic.com) and the cell samples were processed for immunocytochemistry analysis with a ThinPrep 5000 Processor.
Immunocytochemistry staining
Antibodies for CD44 (SP37, Ventana 790-4537), CD133 (polyclonal, biorbyt, orb99113, 1:100), CD45 (LCA, 2B11 & PD7/26, Ventana 760-4279), Ki-67 (30-9, Ventana 790-4286), Wt1 (6F-H2, Cell Marque 760-4397), nestin (10C2, Sigma-Aldrich), Nanog (polyclonal, Abcam, ab80892, 1:100), OCT4 (MRQ-10, Cell Marque 760-4621), SOX2 (SP76, Cell Marque 760-4621), CK5 (SP27, Cell Marque 760-4935), CD34 (QBEnd/10, Ventana 790-2927), P63 (4A4, Ventana 790-4509), Thymosin β4 (Tβ4; polyclonal, abcam, ab14334, 1:1000) and CK14 (SP23, Cell Marque 760-4805) were used for immunocytochemistry. Sample preparation was performed using standard validated cytologic protocols (ThinPrep Preserv Cyt solution, ThinPrep 5000 system, https://www.hologic.com). The Ventana automated stains system (http://diagnostics.roche.com) was used for immunocytochemistry staining and a qualitative analysis was performed by three different pathologists comparing the reactivity of different markers in slides of the same sample using an individual scoring system (morphological parameters, % of number of stained cells, area, and intensity of the stain).
Transmission electron microscopy analysis
The samples of breast milk cells (isolated according to method described in 2.2) were fixed for 1 h in a mixture of 1% paraformaldehyde (Electron Microscopy Sciences) and 1.25% glutaraldehyde (Electron Microscopy Sciences) in 0.1 M sodium cacodylate (Electron Microscopy Sciences) buffer pH 7.4. Cells were rinsed in the buffer, postfixed in 1% osmium tetroxide for 1 h and stained in aqueous uranyl acetate 0.25% overnight at 4°C. Cells were processed using embedding epoxy resin standard methods. 22 In detail, cell samples were dehydrated in a graded series of aceton and infiltrated in Epon 812 resin. Cells were finally embedded in pure Epon 812 resin, transferred to flat embedding molds previously filled with embedding medium and polymerized in an oven at 60°C for 24 h.
60–90 nm thick ultrathin sections were cut with an LKB ultratome 8800 ultramicrotome and collected on grids. Sections were stained with uranyl acetate and bismuth subnitrate, washed with distilled water and observed and photographed at a transmission electron microscope (Zeiss Sigma 300; FEG Oberkochen, Germany) operating at 80 kV.
Results
Freshly isolated human breast milk cells were stained with immunocytochemistry, and were positive in all samples for CD44, CD45, CD133, and Ki67 markers (Table 1). Variable positivity was present with P63, Tβ4, and CK14 markers. In our samples, no immunostaining was detected for Wt1, nestin, Nanog, OCT4, SOX2, CK5, and CD34 markers. Of note, Nanog, SOX2, and OCT4 markers of staminal cells are not expressed, most probably due to the transitory form of the studied cells, waiting for more specific signals of stemness.
Immunocytochemistry staining of cell surface markers in freshly isolated human breast milk cells.

A qualitative immunocytochemistry analysis was performed by three different pathologists, comparing the reactivity of different markers in slides of the same sample, using an individual scoring system (morphological parameters, % stained cells, area, and intensity of the stain).
In the next experiment, the freshly isolated human breast milk cell samples were fixed in the ThinPrep solution (after centrifugation), and then immunostained (Figure 1). Compared to flow cytometry analysis, immunocytochemistry gave an advantage of morphological analysis of the cells, and a prominent possibility to observe nuclear staining of selected markers.

(a) Cells isolated from human milk samples, fixed in ThinPrep and stained with hematoxylin and eosin. Single or grouped cells are characterized by heterogeneous morphology. Magnification 10×. (b and c). CD133 positive human milk cells, collected in the first week of lactation, showing different intensity of immunocytochemistry staining. Magnification (b) 10× (c) 20×. (d and e) CD 44 positive and negative milk cells showing different morphology and stain intensity. Magnification 20×. The Ventana automated stains system (http://diagnostics.roche.com) was used for staining of all samples.
In all analyzed samples, we could observe cells of different shape, dimensions, and with different nuclear size. Figure 1(a) shows the presence of single cells, as well as grouped cells, characterized by heterogeneous morphology, while immunocytochemistry staining (Figures 1(b), (c), 2 and 4) presents expression of different markers on the cell surface. Of note, some cells show nuclear positivity.

CD44 and CD45 positive milk cells. CD44 cells shown different intensity of immunocytochemical staining (a and c) In some cases, CD44 positivity was observed in the nucleus and positive and negative cells may be grouped together. (b) CD45 positivity was founded in single (d, f, and h) and in grouped cells (e, g, and i). Negative and positive cells (e, g, and i) coupled with positive or negative vesicles (f and h) may be grouped together. The Ventana automated stains system (http://diagnostics.roche.com). Magnification 40×.
In our experimental conditions, the transmembrane glycoprotein CD44 is one of the most prominent markers for maternal milk mesenchimal staminal cells and shows the putative character of the studied cells. Figure 1(d) and (e) show a high number of CD44 positive cells. These cells are either isolated (Figure 1(d) and (e)) or grouped in colonies with different intensity of marker expression (Figure 2(a)–(c)). Of note, CD44 immunostaining can be also observed in the nucleus of few cells (Figure 1(d) and (e)).
In accordance with previously published data, 4 the percentage of CD44 putative stem cells changed during lactation. In this context, we analyzed the correlation of CD44-positiveness (the most prominent marker in all analyzed samples) and time of lactation and gestation. Our results indicate that the number of the CD44-positive cells increases during gestation, and decreases during lactation (Figure 3). Nevertheless, the quantity of samples is limited, and the preliminary data need further studies, with particular attention to standardized experimental protocols.
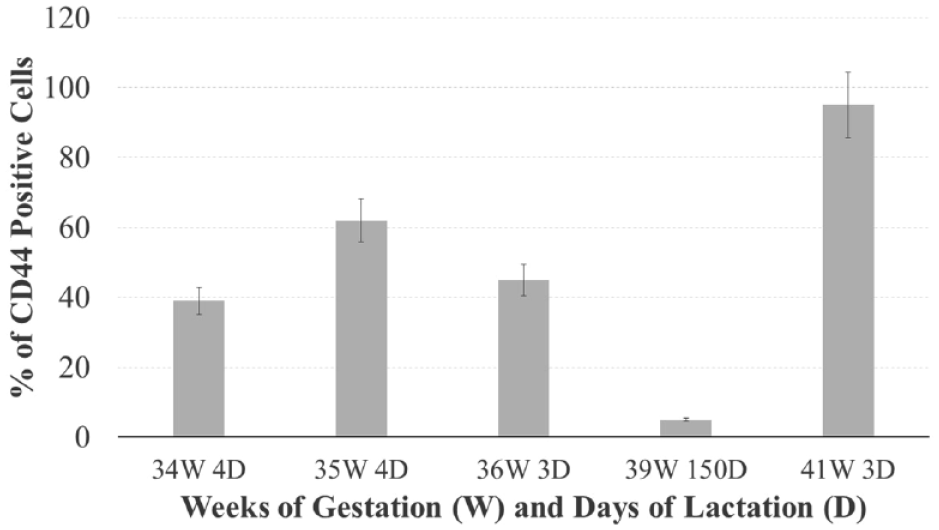
Percentage of CD44 positivity (mean and standard error of three independent experiments staining the same breast milk sample) in several samples of milk with different weeks of gestation (W) and days of lactation (D). The quantitative analysis was made by manual cell counting in optic microscopy (number of squares counted = 12). Statistical comparison between the samples was performed, with 5.0 GraphPad Prism software, by the unpaired t-test. The data were analyzed also with one-way ANOVA as well as the multicomparison Tukey’s test. p values <0.05 was considered significant.
As can be observed in Figure 2(g) to (i), CD45 is present in different cells of the breast milk samples. As for other markers, we also observe a combination of CD45 positive or negative cells, which are single (Figure 2(g)) or grouped with other cells (positive and/or negative; Figure 2(h) and (i)).
The Ki-67 antigen is a nuclear protein, closely associated with cell proliferation, and can be found during all phases of the cell cycle (G1, S, G2, and mitosis), with the exception of G0 phase. For this reason, Ki-67 represents a useful marker to show the growth fraction of a given cell population. The extremely high cell number in the breast milk suggests an intense and dynamic proliferative activity in the mammary gland during lactation. Indeed, Ki-67 immunostaining results positive only in few milk cells (Figure 4) suggests that the remaining negative cells are in a different phase than G0 cell cycle phase.

Ki67 positive cells in different samples of human milk. Ki67 positive cells appear as single cells (a, c, e, f, and h) or grouped cells (b and d). In some cases, immunoreactivity is localized only in the nucleolus (g), and in others amongst several grouped cells where only one is positive (i). The Ventana automated stains system (http://diagnostics.roche.com). Magnification 40×.
Cells isolated from human breast milk form intercellular connections, which together create a cell-to-cell communication network. This phenomenon is clearly underlined by our preliminary ultrastructural investigations (Figure 5). Simultaneously with the presence of well-preserved cytoplasmic organelles and a prominent nucleus, a continuous cytoplasmic membrane appears to be characterized by the presence of numerous cell processes. In the adjacent cells, the plasma membrane seems to play a crucial role in the transmission of intercellular signals. Such phenomenon is evidenced by numerous contact regions established between coupled cells and thin pseudopods-projecting toward adjacent cells (Figure 5(b)–(g)).
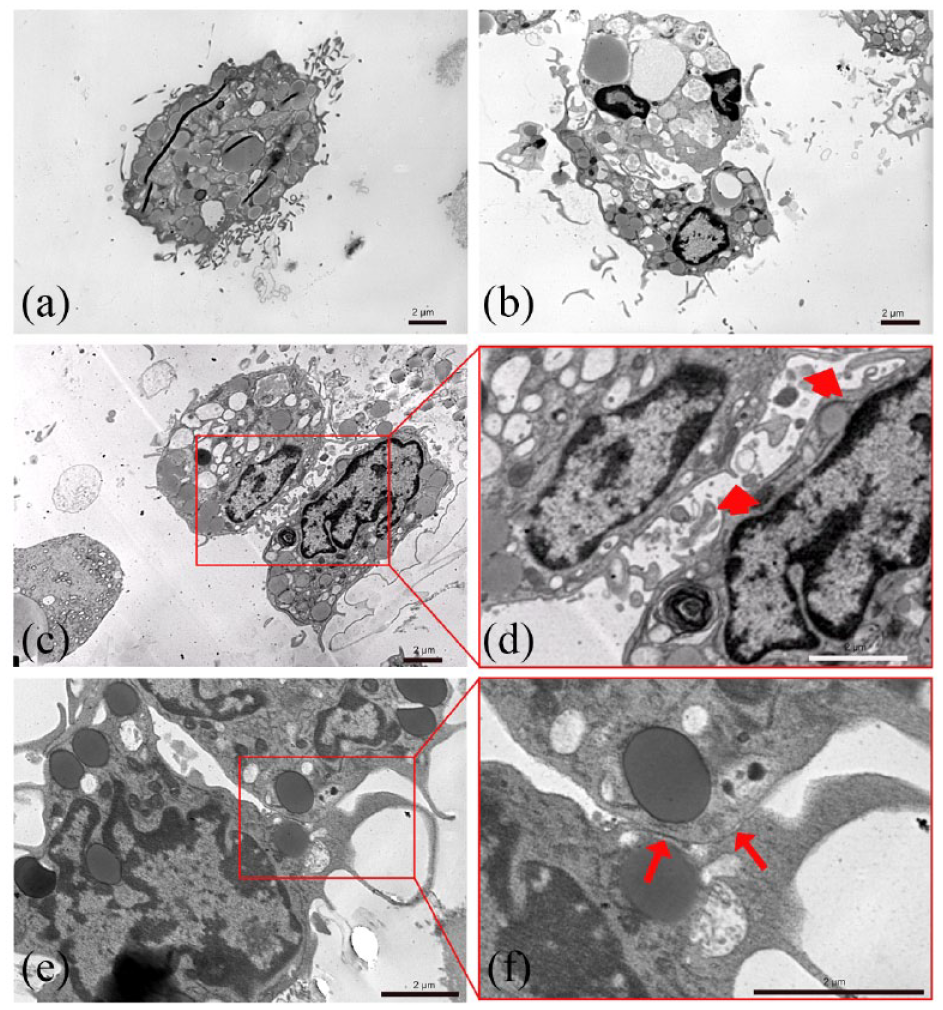
Transmission electron microscopy analysis of human milk cells. The presence of numerous cell processes (a and b), and the presence of numerous thin cell processes projecting toward adjacent cells (c and d) and several contact regions established between coupled cells (e and f) characterize the peculiar morphology of human milk cells.
Discussion
The literature shows that there is large inter-individual variability in progenitor cells in human milk, not only among different studies but also among the women of each single study.6,23 Factors influencing such variations include: breast fullness, the stage of lactation, the health status of the mother/infant, the permeability of the basement membrane, and the development of the breast epithelium. 24 For these reasons, particular attention must be paid to methodological consistency, accuracy, and the reliability of techniques for assessment of milk cell content.25–27
In recent years, numerous clusters of differentiation (CD) markers were used to identify stem cells in human breast milk. Most often, hematopoietic stem cells were immunostained with CD34, CD133, and CD117, while mesenchymal stem cells were stained with CD90, CD105, and CD73. Myoepithelial cells showed reactivity mostly for CD9 and CD44. 28 According to Patki et al., 4 10%–15% of cells isolated from fresh breast milk expressed mesenchymal stem cell (MSC) markers. Further culturing of these cells leads to an increase in the MSC population identified by mesenchymal markers (Smooth muscle actin (SMA), Vimetin, nestin) and surface markers (CD44, CD29, Stem cells antigen-1 (Sca-1)). MSC cells were negative for CD33, CD34, CD45, and CD73 in the studies of Patki et al. 4 Nevertheless, literature data often shows contradictory results. For instance, Fan et al. 2 reported reduced expression of CD34 (1.1% ± 0.15%) and CD133 (2.6% ± 0.79%) in discrepancy with the results of Indumathi et al. 28 whose data showed the expression of CD34 and CD133 as 43.1% ± 2.1%, and 13.07% ± 2.0%, respectively. In the recent literature, different studies of single-cell RNA sequencing reveal sub-populations of mammary epithelial cells with molecular signatures of progenitor and mature states. These cells showed elevated expression for CD44, CD55, aldehyde dehydrogenase 1 family member A3, integrin subunit alpha 6 and others CD49f, and CD14 molecules.8,15,29–31 In this context, our experimental results with immunocytochemistry staining (Table 1) confirm high variability of cell surface markers in human breast milk stem cells, which is most likely influenced by weeks of gestation and days of lactation (Figure 3).
The biological function of the CD133 receptor remains unknown, but alone or in a combination with other markers it is currently used for the isolation of stem cells from numerous tissues. 32 Recently, it was suggested that CD133 is not an organ-specific marker of stem and progenitor cells, but a marker generally expressed by mature luminal ductal epithelial cells. 33 Therefore, it is likely that the increased expression of CD133 in the breast milk cells, found in this study with samples collected in the first period after birth (Figure 1(b) and (c)), is related to the luminal origin of the cells. The subsequent loss of reactivity for CD133 could be instead determined by a greater progressive stemness of these cells. Of note, some cells show nuclear positivity for CD133 (Figure 1(b) and (c)). Such a phenomenon was recently studied by Nunakova et al. 34 and Cantile et al. 35 in cancer cell lines. It was suggested that CD133 in the nucleus may act as transcriptional regulator, but more in depth studies are needed.
CD44 is a cell surface adhesion receptor that is ubiquitously expressed in normal adult and fetal tissues. CD44 interacts with a variety of ligands and can undergo sequential proteolytic processing releasing a CD44 soluble extracellular domain, which is able to translocate from the cytoplasm to the nucleus where it acts as a transcription factor.29,30 For instance, this nuclear translocation occurs under stress conditions (inflammation, oxidative glycolysis, tumor invasion) and takes part of cell survival mechanism. 24 Indeed, we could observe CD44 positivity in the nucleus of few cells, suggesting an active CD44-comunication mechanism. In addition, cleavage of the intracellular domain of CD44 (CD44ICD) activates stemness factors such as Nanog, SOX2 and octamer-binding transcription factor 4 (OCT4), and contributes to the carcinogenesis of breast cancer. 31 In our experimental conditions, we could observe CD44 in the absence of Nanog and SOX2, which could suggest the pre-activation phase of stemness factors in the studied cells.
CD45 is a signaling molecule that regulates a variety of cellular processes including cell growth, differentiation, mitosis, and oncogenic transformation. For these reasons, it was considered a possible candidate for mesenchymal stem cell identification. 36 CD45 was also extensively studied in hematopoietic staminal cells. 33 CD45 is an evolutionary highly conserved receptor protein tyrosinase, which has different natural and artificial extracellular ligands, for instance lectins. 37 In the cell population studied in our experimental conditions (Figure 2(d)–(i)) CD45 marker can be observed in numerous cells, which are present as single cells or colonies. Of note, high positivity of CD44 and CD45 markers—both involved in signaling—could be involved in colony formation and cell-to-cell communication.
The mechanism by which human breast milk derived cells form colonies is poorly described in the literature. To the best of our knowledge, the TEM images of cell colonies (Figure 5) are presented here for the first time. The nanometer resolution images clearly show the contact regions, which were formed before cell isolation. Numerous contact regions, with thin pseudopods—projecting into wide intercellular lacunae and cell surface vesicles—show high activity between neighboring cells with heterogenous morphology. Although the significance of cellular extensions in breast milk cells remains to be ascertained, it is conceivable that such projections could be used to explore and to establish specific contacts with neighboring cells. Our results show that cellular association is not a mere artifact due to sample preparation, but a fundamental process initiating transmission of specific intercellular signals among human breast milk progenitor cells.
Conclusions
Cells freshly isolated form human breast milk, without particular manipulations, show heterogeneous expression of stemness markers. The studied milk staminal cells show “pluripotency” at different stages of differentiation, and are present as single cells or grouped cells. The adjacent cell interactions are evidenced by electron microscopy, which showed the formation of intercellular connections, numerous contact regions, and thin pseudopods.
Footnotes
Acknowledgements
We acknowledge Fondazione Banco di Sardegna for the financial support.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethics statement
These experiments were conducted according to established ethical guidelines, and informed consent obtained from the participants. The experiments were conducted under permission of Comitato Etico Indipendente Ospedaliero (Università di Cagliari; Prot. PG/2022/795).
