Abstract
Eating disorders (EDs) are complex and debilitating conditions. Prior efforts to predict outcomes (onset, prognosis, treatment response) have yielded inconsistent findings. Machine-learning (ML) techniques have shown promise to improve outcome prediction, but a systematic literature synthesis is missing. We conducted a systematic scoping review to summarize extant literature on ML applications in ED-outcome-prediction research, identifying 75 studies. ML has mostly been used to predict ED diagnostic status (k = 45); other studies have predicted escalation of ED risk and symptoms (k = 13), treatment outcomes (k = 12), and ED onset (k = 6). Decision trees, random forest, and support-vector machines were the most common models used. Although many studies reported moderate to high predictive performance, the benefits of ML over traditional statistical techniques remains unclear in light of inconsistent findings. We make several recommendations for future research (i.e., integrating multiple data types, external validation) to encourage continued progress in this developing field.
Eating disorders (EDs) are complex, chronic conditions associated with a number of adverse outcomes, including high comorbidity and mortality, impairment in psychological and social functioning, and poor quality of life (Klump et al., 2009; van Hoeken & Hoek, 2020). A host of barriers (i.e., cost, limited therapist availability, stigma) contribute to low uptake of treatment; approximately three quarters of people with EDs are not accessing appropriate intervention (Ali et al., 2017) or are reporting significant delays (5 years on average) between symptom onset and treatment seeking (Hamilton et al., 2021). This is concerning given that recovery without treatment is uncommon and that the likelihood of full recovery is reduced the longer that symptoms persist (Fernández-Aranda et al., 2021; Wonderlich et al., 2012). Despite the development of numerous evidence-based treatments, treatment outcomes also remain suboptimal and variable; fewer than 50% achieve full symptom remission (Linardon & Wade, 2018). Many studies have sought to improve ED outcomes through exploring factors associated with ED risk, onset, and treatment outcomes, yet accurate prediction of outcomes has proven challenging; studies have often yielded unreliable predictors that lack clinical utility (Linardon et al., 2017; McClure et al., 2024).
Increasingly, machine-learning (ML) techniques are being applied to optimize diagnosis, prognosis, and treatment-outcome predictions in the field of psychiatry, and promising results have emerged (Dwyer et al., 2018). Although the term “machine learning” was coined by Arthur Samuel in the 1950s, the use of ML in psychiatric research has been steadily growing over the past 2 decades as data availability and computational resources have increased (Z. S. Chen et al., 2022). ML is a collection of data-driven computational techniques that learn patterns in data, enabling underlying data structures to be identified and accurate predictions of future outcomes to be made (Jordan & Mitchell, 2015). Unlike traditional statistical techniques (i.e., linear regression, logistic regression), which perform most optimally when only a small number of linearly related variables are used, ML techniques can model more complex (nonlinear, interactions), high-dimensional (i.e., large number of variables) data while enabling several different data types (i.e., survey data, imaging data, social media data) to be considered in a single model. These capabilities provide greater opportunity to capture the diversity and complexity inherent in psychiatric phenomena, potentially leading to predictions that are more accurate, parsimonious, and generalizable. ML techniques are often more suited to modelling relationships that are generalizable to single cases, potentially allowing more accurate predictions at the individual patient level (Bzdok & Meyer-Lindenberg, 2018; Dwyer et al., 2018). This stands in contrast with traditional techniques that often examine group-level effects, producing findings that can have limited validity when applied to individuals (Dwyer et al., 2018). Consequently, researchers have touted the potential for ML to usher in an era of precision psychiatry, in which prevention, diagnosis, and treatment of mental-health conditions are based on the unique biological, psychological, and social characteristics of each individual (Bzdok & Meyer-Lindenberg, 2018).
Although ML offers significant potential for improving outcome prediction, there are several challenges associated with ML applications that are worth noting. These include the need for large data sets to adequately train and validate models, difficulties with interpretation of complex models involving many predictors, and the computational resources and expertise required to perform ML analyses (Bzdok & Meyer-Lindenberg, 2018; Dwyer et al., 2018). These complexities can not only make the development of robust models challenging but can also cause difficulties with translation of ML models into clinical practice (Steyerberg et al., 2013). Thus, it is crucial that researchers consider the potential limitations of applying ML techniques alongside its anticipated benefits and the clinical utility of models (i.e., data availability, generalizability, clinical interpretability). To help facilitate this, adherence to best practice guidelines on ML model development and reporting is recommended (i.e., TRIPOD statement, Collins et al., 2024; PROGRESS, Steyerberg et al., 2013).
Broadly, ML techniques can be grouped into two main categories: supervised learning and unsupervised learning. Supervised-learning techniques are applied to data sets in which the input data (i.e., predictors, features) and output data (i.e., outcomes, response variable) are labeled (Wang, 2021). These techniques learn patterns and relationships between variables to accurately classify or predict a prespecified outcome (i.e., predicting ED onset). In contrast, unsupervised-learning techniques identify patterns and relationships in unlabeled data (i.e., input data are provided without corresponding output data). They are commonly used to identify meaningful clusters or groups in data in which the underlying structure may be previously unknown (i.e., identifying clusters of ED subtypes in data; Wang, 2021).
ML techniques have been applied across various mental-health conditions, demonstrating promise in areas including detection and diagnosis, treatment and support, and clinical and research administration (Shatte et al., 2019). For example, supervised-ML techniques have been used with data derived from thousands of electronic health records (EHRs) to predict future suicide attempts with high accuracy (Walsh et al., 2017). The use of neuroimaging data with ML has shown promise in enhancing early detection and diagnosis of schizophrenia (Koutsouleris et al., 2021; Skåtun et al., 2017), and wearable-sensor data have been used to accurately predict the presence and severity of depression (Tazawa et al., 2020). Unsupervised techniques have also been used to improve understanding of diagnostic heterogeneity—that is, variability in symptoms and treatment responses in a specific diagnosis—by uncovering clinical subtypes of diagnoses that may show similar etiologies, prognoses, and treatment responses (i.e., Amoretti et al., 2021; Chekroud et al., 2017; Kung et al., 2022; Pelin et al., 2021). Several studies have also compared ML and traditional techniques; ML has demonstrated enhanced performance for predicting treatment outcomes in depression (Kessler et al., 2016), obsessive compulsive disorder (Lenhard et al., 2018), and psychosis (Koutsouleris et al., 2016). Together, these studies provide preliminary evidence of the potential utility of using ML to enhance psychiatric research.
Although ML techniques have been applied more widely in the fields of depression and schizophrenia, there has been a growing number of studies testing ML in ED research that have yielded encouraging results (Fardouly et al., 2022). For example, ML has shown promise in predicting ED diagnoses using data from surveys (i.e., Linardon et al., 2020), social media posts (i.e., Abuhassan et al., 2023), and neuroimaging scans (i.e., Cerasa et al., 2015). Preliminary evidence has also been found for predicting ED risk and onset (Krug et al., 2021; Mitchison et al., 2023), illness course (Haynos et al., 2021), and treatment response (Espel-Huynh et al., 2021), and ML models have achieved high predictive performance. Several studies have also investigated whether ML outperforms traditional techniques by comparing approaches; some have found that ML enhanced predictive performance (Haynos et al., 2021), and others have found comparable results (Espel-Huynh et al., 2021; Krug et al., 2021). Thus, enhancing understanding of the conditions under which ML performs optimally in this context is critical for maximizing its potential benefits and, ultimately, improving understanding of the complexities of EDs.
Furthermore, the clinical benefits of constructing accurate and robust ML models are substantial. By integrating ML into clinical workflows, health-care providers could offer more precise, timely, and individualized care, ultimately improving patient outcomes and optimizing resource allocation (Lee et al., 2021). For example, ML models processing large amounts of patient intake data may aid in generating differential diagnoses, enabling clinicians to promptly identify conditions aligned with a patient’s symptoms and history. Likewise, ML may be used to predict treatment responsiveness, guiding the selection of appropriate therapies. However, before ML models can be integrated into clinical practice, they must undergo rigorous testing and external validation to ensure they reliably contribute to improved patient outcomes (Steyerberg et al., 2013). Ongoing monitoring and regular updates to these models are also essential to ensure ongoing adherence to current clinical standards and adaptation to emerging data trends (Steyerberg et al., 2013).
Considering the exciting potential ML offers for enhancing diagnosis, prognosis, and treatment-outcome prediction and the increasing number of studies using ML in this field, a systematic scoping review mapping this emerging body of literature is timely. Although previous narrative reviews have provided a preliminary overview of ML applications in the ED field (Fardouly et al., 2022; Ghosh et al., 2024), narrative reviews do not follow predefined, systematic methods and may be subject to selective reporting, making their findings less comprehensive and more prone to bias (Sarkar & Bhatia, 2021). In contrast, a systematic scoping review follows more rigorous, transparent, and replicable methods to comprehensively identify, map, and synthesize the available evidence, ensuring minimized bias and enhanced reproducibility (Peterson et al., 2017). This approach allows for a more structured synthesis of trends, helping to identify common methodologies, gaps in the literature, and directions for future research, which is essential for advancing the field.
Although a systematic review has been conducted on ML and natural language processing for detecting EDs (Merhbene et al., 2024), its focus was limited to a specific application of ML in the field (text-based detection of EDs) rather than the broader landscape of ML applications for ED-outcome prediction. In contrast, our review provides a more comprehensive examination of how ML has been used to predict ED outcomes, encompassing a wider range of methodologies, data types, and clinical applications. This broader scope allows for a more complete understanding of ML’s potential and limitations in ED research, offering insights that are critical to guiding future work in the field.
Thus, we conducted a systematic scoping review aiming to locate, examine, and summarize the existing literature on the application of ML in ED-outcome-prediction research. Specifically, we aimed to understand (a) which ML techniques have been used and in what context, (b) whether there is evidence of ML demonstrating superior performance over traditional statistical techniques, and (c) whether models have been externally validated in new samples.
Method
A systematic-scoping-review methodology was selected given the purpose was to explore, locate, and summarize how ML has been applied in ED-outcome-prediction research. Scoping reviews aim to map the available literature on a given topic, providing an overview of the research conducted to date while allowing key concepts and gaps in knowledge to be identified (Arksey & O’Malley, 2005). A scoping review was also selected given anticipated heterogeneity across study designs, samples, assessment instruments, and so on. The Arksey and O’Malley (2005) methodological framework for scoping reviews and the PRISMA Extension for Scoping Reviews (Tricco et al., 2018) were used to guide the methodology of this review.
Information sources and search strategy
Six online databases (Medline, PsycINFO, Web of Science, EMBASE, Cochrane Database, and ProQuest Dissertations and Theses Global) were searched in February 2023 (updated in April 2024). Terms related to EDs and ML were combined and searched for in the title and abstract (for search terms, see the Supplemental Material available online). Terms related to ML were adapted from prior reviews conducted in the broader health field (Christodoulou et al., 2019; Morgenstern et al., 2020). The reference list of a relevant narrative review (Fardouly et al., 2022) was hand searched to identify any additional studies.
Selection of sources of evidence
Studies were included if they (a) reported on a method or application of an ML technique; (b) evaluated the ML technique’s performance for predicting ED risk, diagnosis, and/or prognosis either naturally or in the context of an intervention; (c) made an original contribution to the literature (i.e., reviews were excluded); and (d) was available in English language. Studies that applied only logistic regression were excluded because this was not regarded as a ML technique (see Christodoulou et al., 2019). Given that we were interested in the prediction of ED risk, onset, and diagnosis, no restrictions were placed on sample type (i.e., nonclinical and clinical samples were included). All study designs were considered for inclusion. Titles and abstracts were screened for relevance by Z. McClure. Full texts were sourced for potentially relevant studies and were reviewed to determine whether full inclusion criteria were met (performed by Z. McClure, J. Linardon, M. Fuller-Tyszkiewicz).
Data extraction
Data were extracted using a template developed for this study. Data extracted included authors; design; study aim; outcome; sample characteristics; data type; predictor variables/features; name of ML techniques used, including key details to help ascertain the predictive performance of the techniques, such as comparison with more traditional techniques (i.e., logistic regression, linear regression); number of outcome events in training set; validation method used; and key findings. Data were extracted by Z. McClure in consultation with J. Linardon.
Risk of bias
For studies comparing ML models with a traditional technique, we assessed for risk of bias using a tool developed by Christodoulou et al. (2019). Each criteria assessed methodological issues of model development and aspects that may compromise comparisons between approaches. Five signaling items are used to indicate potential bias: (a) unclear or biased validation of model performance, (b) difference in whether data-driven variables selection was performed (yes/no) before applying traditional and ML algorithms, (c) difference in handling of continuous variables before applying traditional and ML algorithms, (d) different predictors considered for traditional and ML algorithms, and (e) whether corrections for imbalanced outcomes were used only for traditional algorithms or only for ML algorithms. Each item was scored as “no” (not present), “unclear,” or “yes” (present). Risk of bias was considered high if at least one item was scored unclear or yes. For further detail on items, see Table S1 in the Supplemental Material.
Synthesis of results
Studies were synthesized according to four broad themes that were developed through discussion among authors following the data-extraction process. The themes are (a) “predicting ED onset,” studies that predicted onset of an ED in previously asymptomatic or at-risk samples; (b) “predicting ED risk and symptoms,” studies that predicted ED-related behaviors and cognitions (i.e., binge-eating episodes, weight/shape concerns, severity) in clinical and nonclinical samples (in the absence of predicting diagnostic categories); (c) “predicting ED diagnosis,” studies that aimed to predict the presence versus absence of an ED or to classify different ED subtypes (i.e., anorexia nervosa [AN] vs. bulimia nervosa [BN]); and (d) “prediction of treatment outcome,” studies that aimed to predict responsiveness to a particular intervention program.
Results
Search results
Seventy-five studies were included (for PRISMA flowchart, see Fig. 1; for studies, see Tables S2–S5 in the Supplemental Material). Publication year ranged from 1998 to 2024; 84% were published from 2015 onward (see Fig. S1 in the Supplemental Material). A synthesis of findings is presented for each theme below.
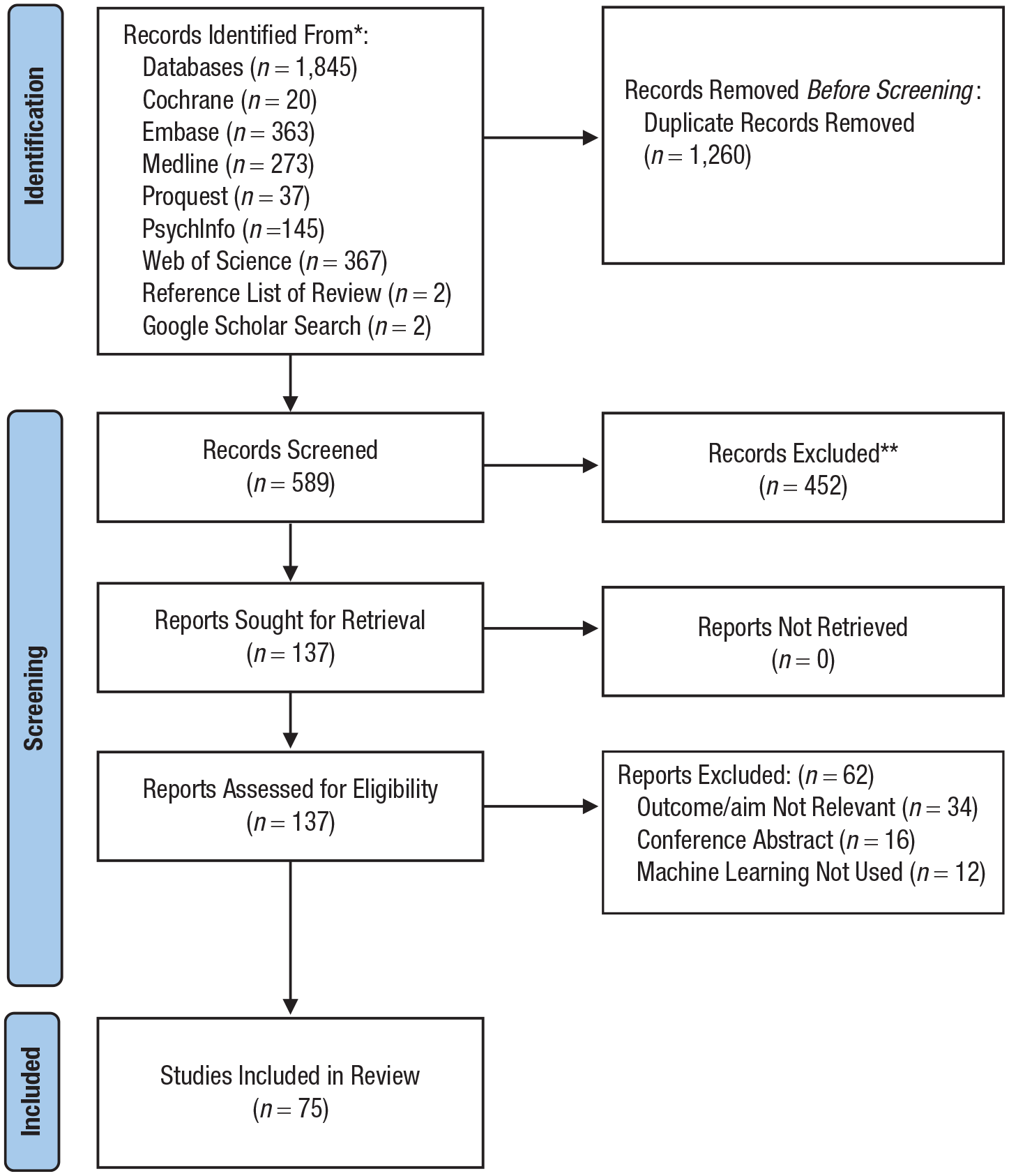
PRISMA flowchart.
Predicting ED onset
Study characteristics
Six studies used ML models to predict ED onset (see Table 1). Sample size ranged from 206 to 1,297 (Mdn = 920). Follow-up time ranged from 10 months to 8 years. All but one (Mitchison et al., 2023) used a classification and regression tree (CART) to detect interactions between variables and identify important predictors. None reported conducting external validation of models.
Aggregated Study Characteristics of Studies Predicting ED Onset (k = 6)
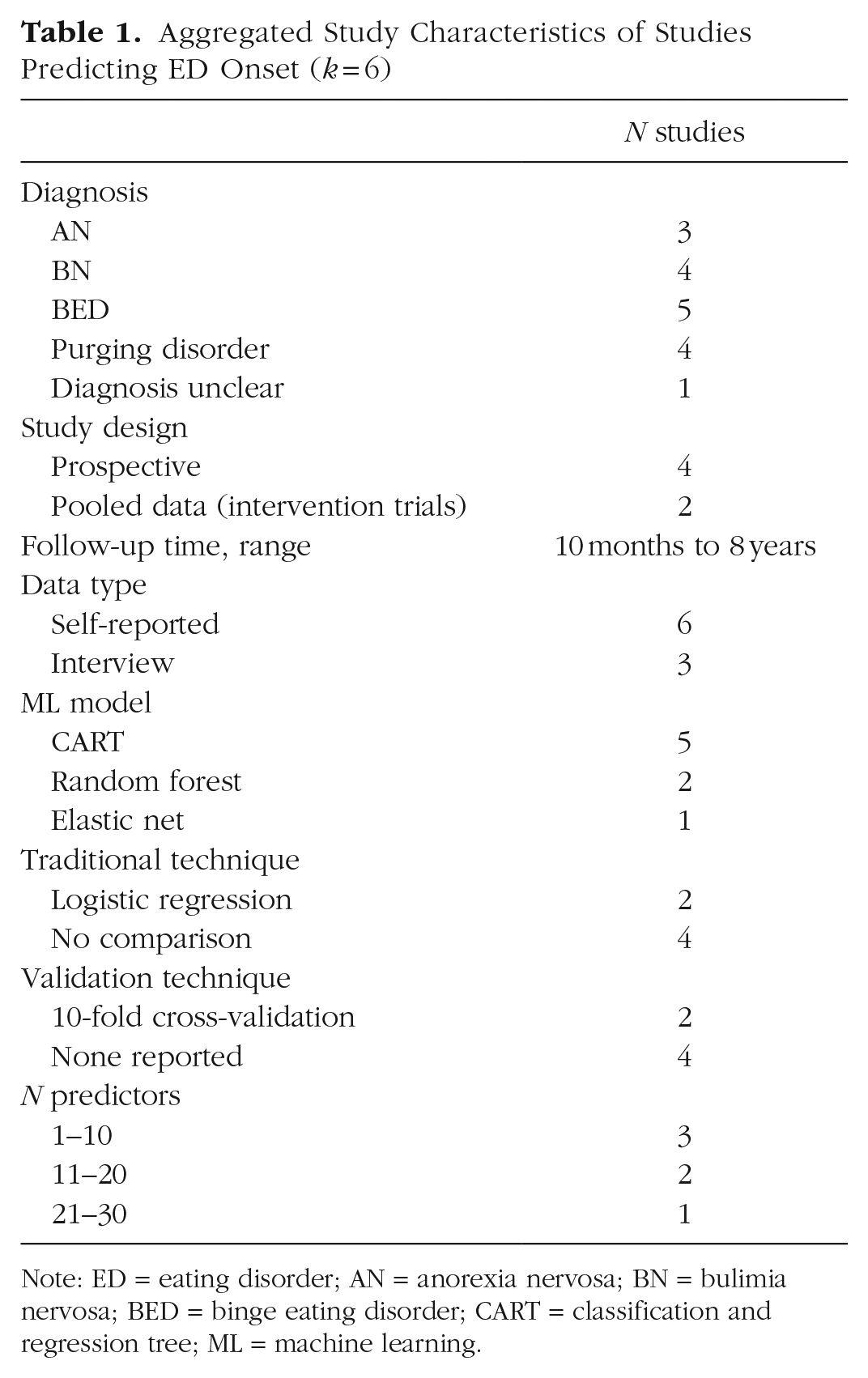
Note: ED = eating disorder; AN = anorexia nervosa; BN = bulimia nervosa; BED = binge eating disorder; CART = classification and regression tree; ML = machine learning.
Study findings
Four studies reported classification accuracy (76%–98.2%; Mehl et al., 2019; Mitchison et al., 2023; Stice & Desjardins, 2018; Stice et al., 2002), and one reported the aggregated out-of-bag error rate (Allen et al., 2016; 0.4% error rate for classifying onset in boys, 13.8% error in girls). Two implemented a traditional approach alongside an ML technique, enabling comparison (for studies comparing approaches, see Table S6 in the Supplemental Material). Allen et al. (2016) tested CART and logistic regression. Using six predictors, CART found sex was the most important predictor of ED onset such that adolescent girls displayed higher risk than boys (n = 1,297). A nonlinear interaction was also found between weight and eating concerns and onset for boys and girls, and an interaction between moderate weight and eating concerns and externalizing problems was found for girls. The logistic-regression model failed to identify any two-way interactions. In contrast, Mitchison et al. (2023) found comparable performance between an ML technique and traditional technique (n = 687). Using six predictors, elastic net and logistic regression performed similarly at predicting ED onset over 1 year (both models obtained an area under the curve [AUC] of .75), demonstrating moderate predictive performance.
Predicting ED risk and symptoms
Study characteristics
Thirteen studies used ML models to predict ED risk and symptoms (see Table 2). Studies aimed to predict the occurrence of specific ED behaviors, cognitions, or general ED risk. Sample size ranged from 13 to 11,620 (Mdn = 371). Most recruited nonclinical samples and used self-reported data as input. Several ML models were tested; CART and random forest were the most common. Only one reported conducting external validation (X. Chen et al., 2023). For ease of readability, findings were synthesized according to sample type and by data type for nonclinical samples.
Aggregated Study Characteristics of Studies Predicting ED Risk and Symptoms (k = 13)
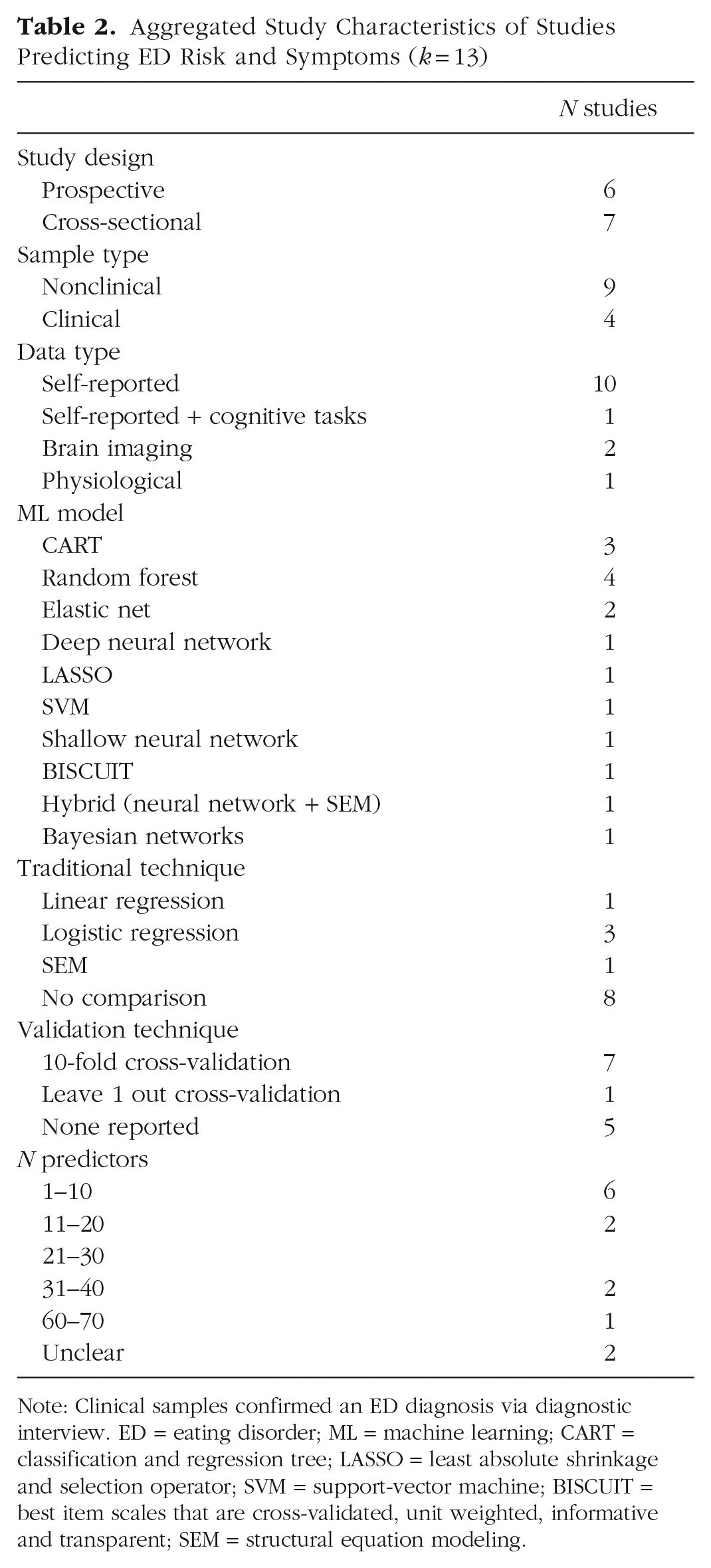

Note: Clinical samples confirmed an ED diagnosis via diagnostic interview. ED = eating disorder; ML = machine learning; CART = classification and regression tree; LASSO = least absolute shrinkage and selection operator; SVM = support-vector machine; BISCUIT = best item scales that are cross-validated, unit weighted, informative and transparent; SEM = structural equation modeling.
Clinical samples
Four studies (all prospective) predicted symptoms in a clinical sample. Three used self-reported ecological momentary assessments (EMAs) as input obtained via smartphone (Arend et al., 2023; Levinson et al., 2022) or a blood-glucose-monitoring device (Presseller et al., 2024). Presseller et al. (2024) found a random-forest model could distinguish eating and noneating episodes in binge eating disorder (BED) and BN with high accuracy (82%), and Arend et al. (2023) reported good model performance for predicting binge eating in women with BN and BED (AUC = .80, best items scale that is cross-validated, unit-weighted, informative, and transparent algorithm; for explanation of algorithm, see Table S3 in the Supplemental Material). Only Levinson et al. (2022) employed ML models (CART, random forest, support-vector machine [SVM], shallow neural network) alongside a traditional technique (logistic regression; n = 60). A shallow-neural-network model performed best, predicting behavioral symptoms with high accuracy (> 75%) across several time points. However, performance metrics were reported only for the shallow-neural-network model, so the degree to which it outperformed logistic regression is unclear.
Haynos et al. (2021) demonstrated superior performance of ML models (elastic net and random forest) compared with logistic regression for predicting symptom trajectories over 2 years in transdiagnostic women (n = 415). When researchers used more than 30 baseline predictors (self-report and interview), ML models achieved higher mean AUCs across outcomes, including ED diagnostic status, presence of binge eating, compensatory behaviors, and underweight body mass index (elastic net: range = 0.61–0.93; random forest: range = 0.62–0.92; logistic regression: range = 0.47–0.83).
Nonclinical sample
Self-reported data
Seven studies recruiting a nonclinical sample used self-reported data as input to predict symptoms. Least absolute shrinkage and selection operator (LASSO) or Bayesian networks were used to identify ED risk factors (Bercht & Costa, 2023; Han & Zhang, 2021). Two used CART to predict orthorexia nervosa risk (Dell’Osso et al., 2018) or general ED risk (Ren et al., 2022), correctly classifying 67% to 88% of high-risk participants. Three other studies predicting ED symptoms compared ML with a traditional technique (Kheirollahpour et al., 2020; Liang et al., 2022; Mitchison et al., 2023). Kheirollahpour et al. (2020) found a hybrid ML model (R2 = .552) outperformed structural equation modeling (SEM; R2 increased by 27%) for predicting eating-behavior patterns using three predictors (n = 340). In contrast, Liang et al. (2022) found no performance benefit between random forest, deep neural network, and linear regression for predicting body-image disturbances (n = 11,620) using four predictors. Mitchison et al. (2023) also reported similar performance between elastic net (AUC = .64) and logistic regression (AUC = .62) for predicting symptom persistence using three predictors (n = 276).
Brain-imaging data
Two studies used ML models to explore brain structure and function as predictors of symptom variation. X. Chen et al. (2023) sampled Chinese students and used connectome-based predictive modeling (n = 660). Although models failed to predict binge/purge behavior, connectivity between networks involved in cognitive control, reward sensitivity, and visual perception predicted increased body-image concerns (significant correlations between actual and predicted scores). Findings were replicated in an independent sample (n = 821). X. Chen et al. (2022) also reported good model performance of a linear SVM for predicting binge eating using brain structure and function indexes in primary school children (n = 76).
Predicting ED diagnosis
Study characteristics
Forty-five articles applied ML to classify individuals with an ED or differentiate ED subtypes (see Table 3). Many studies focused on classifying AN versus healthy control subjects (k = 18). Sample size ranged from 30 to 1,165,000 (Mdn = 423). Several data types were used as input; text from social media profiles were the most common. Most studies undertaking text classification implemented more than one ML technique, proposing custom models combining natural-language-processing techniques (i.e., feature extraction) with supervised-learning models (i.e., SVM, random forest) or deep-learning models (i.e., neural networks, transformer). Studies using other data types mostly implemented one ML technique; SVM was the most common. External validation was reported in one study (Burstein et al., 2023). Findings are synthesized below according to the data type used as model input, with emphasis on noteworthy studies (i.e., large sample size) and/or those that implemented ML alongside traditional techniques.
Aggregated Study Characteristics of Studies Predicting ED Diagnostic Categories (k = 45)

Note: ED = eating disorder; ML = machine learning; CART = classification and regression tree; LASSO = least absolute shrinkage and selection operator; SVM = support-vector machine.
The value of v was not specified.
Textual data
Sixteen studies used textual data as input. Most used data from posts on social media platforms Twitter and Reddit (k = 14) and aimed to classify ED cases versus healthy control subjects. Most reported F1 scores, ranging from poor to excellent performance across studies (F1s = 0.43–0.98). One noteworthy study is Abuhassan et al. (2023), who collected a large sample of Twitter biographies from 1,165,000 users and classified them into five groups: individuals with an ED, health-care professionals, communicators, health-care professional communicators, and other. They implemented a deep-learning model based on bidirectional encoder representations (BERT) and long short-term memory (LSTM), achieving a high classification accuracy (98.37%). Four studies compared the performance of ML models with logistic regression (Fano et al., 2019; Noguero et al., 2023; Ramirez-Cifuentes et al., 2018; Uban et al., 2021; ns = 177–1,288). All but one (Noguero et al., 2023) found that an ML model performed better. The best performing models were a multilayer perceptron with GloVe vectors (F1 = 0.78; logistic regression: F1 = 0.56), SVM (F1 = 0.85; logistic regression: F1 = 0.76), and LSTM hierarchical attention network (F1 = 0.61; logistic regression: F1 = 0.49).
Self-reported data
Studies using self-reported data as input were cross-sectional (k = 8) or retrospective (k = 2) or used data from a longitudinal and case-control study design (k = 1). ML models generally achieved high accuracy (> 70%) for classifying ED cases or differentiating subtypes. Three studies compared ML models with a traditional approach (Krug et al., 2021; Orru et al., 2021; Sandoval-Araujo et al., 2024). Orru et al. (2021) used four predictors, and the others used 43 to 51 predictors. One reported comparable performance between ML models and logistic regression (Krug et al., 2021), although they noted that ML achieved similar performance using fewer predictors, leading to a more parsimonious model (AUCs = .69–.82). In contrast, Sandoval-Araujo et al. (2024) found ML models (CART: AUC = .81; random forest: AUC = .79) outperformed logistic regression (AUC = .62) for classifying AN versus atypical AN, whereas Orru et al. (2021) found ML models (naive Bayes, SVM, random forest; AUCs = .81–.90) were superior to logistic regression (AUC = .77) for classifying ED cases versus healthy control subjects. Overall, ML models tended to perform better at distinguishing between ED cases and healthy control subjects compared with distinguishing between ED subtypes (i.e., AN vs BN).
Brain-imaging data
Seven studies investigated using ML (SVM: k = 5; LASSO: k = 1; linear relevance vector: k = 1) with brain-imaging data to identify biomarkers of EDs that may distinguish EDs from healthy control subjects (ns = 30–658; Arold et al., 2023; Cerasa et al., 2015; Cyr et al., 2018; Lavagnino et al., 2015, 2018; Weygandt et al., 2012; Zheng et al., 2023). Studies were primarily interested in exploring whether patterns in brain structure or function could be used to classify people at the individual patient level, and most reported high accuracy (> 70%). One study that examined brain-activation patterns involved in food-cue processing was also able to distinguish BN from BED cases with high accuracy (84%; Weygandt et al., 2012). Another found that features extracted from diffusion tensor imaging classified AN versus BN cases with good performance (AUC = .79; Zheng et al., 2023).
A case study (Strigo et al., 2017) used brain-activation patterns during anticipation and experience of high pain conditions to recommend a diagnosis of either AN, gastrointestinal problems (GIP), or depression. An SVM classifier trained on samples with recovered AN, GIP, or depression was implemented with a 15-year-old female with overlapping ED, depressive, and gastrointestinal symptoms. The model (accuracy of 56% when trained on diagnostic samples) classified the subject into the gastrointestinal group. These findings were corroborated by a second model using participant self-reported behavioral measures (84% when trained on diagnostic samples).
Physiological data
Studies using physiological data as input categorized individuals with AN versus healthy control subjects using eye-gazing tracking in response to a body-image-related visual-scanning task (Liu et al., 2021), concentrations of several trace elements (Zhao et al., 2004), and genotypic and phenotypic data (Guo et al., 2016). One classified AN versus BN cases with moderate performance (AUC = .72) using several metabolic indices (Dönmez et al., 2023). Two studies used electroencephalography data to accurately classify people with BED versus without BED (Raab et al., 2020; accuracy = 81.25%) and AN cases versus healthy control subjects (Karavia et al., 2024; accuracy = 75%–85%). Guo et al. (2016) was the only study to compare ML with a traditional approach, finding no benefit of SVM (AUC = .69) over logistic regression (AUC = .69) for classifying AN cases (n = 4,402).
Predicting treatment outcome
Study characteristics
Twelve studies predicted response to ED treatment (see Table 4). Most were predicting treatment outcomes to inpatient treatment and used self-reported data as input. A range of ML models were implemented; random forest was the most common. Sample size ranged from 36 to 826 (Mdn = 262). No study reported conducting external validation.
Aggregated Study Characteristics of Studies Predicting ED Treatment Outcome (k = 12)

Note: ED = eating disorder; ML = machine learning; RCT = randomized controlled trial; CART = classification and regression tree; LASSO = least absolute shrinkage and selection operator; SVM = support-vector machine.
Inpatient treatment
Studies show promise for using ML models to predict inpatient treatment response. CART and random forest were useful for identifying important predictors of good treatment outcome versus poor treatment outcome in BN (Hannöver et al., 2002; n = 630, model misclassification rate = 18%) and treatment dropout in a transdiagnostic sample (Todisco et al., 2023; n = 420). Four studies used supervised-learning techniques to classify people into different treatment-response categories; studies reported good performance (accuracy > 75%; Espel-Huynh et al., 2021; Espel-Huynh & Lowe, 2019; Ioannidis et al., 2020). Only one compared ML (SVM, k nearest neighbor) with a traditional approach (Espel-Huynh et al., 2021; n = 333), classifying participants into three different response trajectories (rapid, gradual, low symptom static). All models were trained using either 80 predictors or three predictors. SVM (radial) with three predictors was the best performing ML model (AUC = .94) but did not significantly outperform logistic regression (AUC = .93).
Digital intervention
Two studies applied ML models to predict response to ED digital interventions. von Brachel et al. (2014) used LASSO to identify important predictors of dropout to an online motivational program for women with an ED. Linardon et al. (2022) compared predictive performance of several ML models with linear regression for predicting engagement and symptom-level change (n = 826). ML models did not significantly outperform linear regression using 36 self-reported baseline predictors (across models for engagement outcomes: AUCs = .48–.52; for symptom change: R2s = .15–.40). However, predictive performance improved considerably across models (except SVM radial) for dropout (AUCs = .92–.99) and adherence outcomes (AUCs = .62–.93) when intervention-usage-pattern variables were added as input (although ML had comparable performance with traditional regression).
Other treatment types
Other treatment types included general outpatient treatment for EDs (Svendsen et al., 2023), pharmacological treatment for BED (Goyal et al., 2022), and behavioral weight loss and stepped care treatment for BED (Forrest et al., 2021). Although Svendsen et al. (2023) did not compare ML performance with a traditional approach, their model performed better than chance at predicting treatment nonresponse, demonstrating high precision (positive predictive value = 70%–71%) and sensitivity (78%–95%). Note that a synthetic data set was used to train the model. Two other studies predicting likelihood of placebo response (Goyal et al., 2022; n = 189) and reduction in binge eating, ED psychopathology, and weight loss (Forrest et al., 2021; n = 191) in adults with BED compared ML techniques with traditional techniques. Only Goyal et al. (2022) found better performance of an ML model (Gaussian naive Bayes; accuracy = 72%, specificity = 88%, sensitivity = 63%) compared with logistic regression (accuracy = 66%, specificity = 53%, sensitivity = 73%).
Risk of bias assessment
Of 19 studies that implemented a traditional technique alongside an ML model, seven were assessed as having a high risk of bias (see Table S7 in the Supplemental Material). Five of these reported greater predictive performance of ML models. The most frequently unmet criterion was unclear or biased validation performance (Criterion 1; 7/19 studies scored yes or unclear), often because of insufficient reporting on whether all model-building steps, such as feature selection and hyperparameter tuning, were repeated in each validation fold.
Discussion
This scoping review located, examined, and summarized existing literature on the application of ML in ED-outcome-prediction research. Specifically, we aimed to investigate (a) which ML techniques have been used and the contexts in which they have been applied, (b) whether there is evidence of ML outperforming traditional techniques, and (c) the extent to which models have been externally validated. We included 75 studies. Below, we summarize several key findings and trends in the literature pertaining to the three key aims of the review.
Summary of key findings
Synthesis of studies identified four broad domains in which ML techniques have been applied. These included predicting (a) onset of an ED (k = 6), (b) presence or severity of symptoms and risk factors (k = 13), (c) ED diagnostic categories (i.e., ED vs. healthy control subjects, AN vs. BN; k = 45), and (d) outcomes during treatment/intervention (k = 12).
We found that there were several different types of data used as input to predict outcomes, yet studies mostly relied on one type of data to generate prediction models. The most common data type used was self-reported data; most studies using this data type reported moderate to high predictive performance across the four domains. Other types of data used for predictive models—particularly when the goal was to predict diagnostic categories—were textual data, brain-imaging data, and physiological data. For example, studies implemented natural-language-processing techniques with ML models to analyze and classify text; many demonstrated the potential to detect ED cases through content posted on an individual’s social media profile (e.g., Aragón et al., 2020; Benitez-Andrades et al., 2023; Fano et al., 2019). Several studies also showed preliminary evidence for using brain-imaging data with ML models to identify biomarkers that can accurately classify ED cases or predict symptom variation (e.g., Cerasa et al., 2015; X. Chen et al., 2023; Lavagnino et al., 2015, 2018).
Although a number of ML techniques were tested across studies, the most common were CART, random forest, and SVM. Several studies implementing CART suggested that interpretability may be a key advantage of this model; studies highlighted its utility for identifying cut points associated with ED-risk/diagnostic categories and interactions between predictors (e.g., Allen et al., 2016; Hannöver et al., 2002; Linardon et al., 2020; Ren et al., 2022; Stice et al., 2011). Although authors were overall optimistic about the performance of ML models, external validation of findings was reported in only two studies (Burstein et al., 2023; X. Chen et al., 2023), which limits understanding of model robustness and out-of-sample generalizability.
There is emerging evidence supporting the potential for ML to outperform traditional techniques, although findings are mixed. In particular, 12 of 19 (63%) studies comparing approaches reported higher predictive performance of ML models compared with traditional techniques. However, five of these studies were assessed as being high risk of bias, indicating that findings should be interpreted with caution given that some reported advantages may be influenced by methodological limitations. Although most of these indicated enhanced performance through observing greater predictive accuracy or classification ability, one noted superior performance because of ML producing a more parsimonious model (but similar AUC to logistic regression; Krug et al., 2021). Likewise, another found that a ML model (CART) was more useful for detecting significant, nonlinear interactions compared with logistic regression (Allen et al., 2016). It appears that ML may improve text-classification tasks given that three studies found ML performed better at classifying ED cases based on social media content (Fano et al., 2019; Ramirez-Cifuentes et al., 2018; Uban et al., 2021). However, note that the traditional approach still performed relatively well at text-classification tasks, indicating that it may be worth considering whether the incremental benefits of ML outweigh the potential complexities that may come with implementing ML models in this context. This consideration is particularly relevant for text-classification tasks, which often rely on more complex ensemble or deep-learning methods (i.e., LSTM). These models can introduce additional challenges related to interpretability, transparency, computational demand, and clinical implementation, which must be weighed against their potential performance benefits (Miotto et al., 2018).
In contrast, studies finding comparable performance suggest there are circumstances when simpler traditional techniques perform just as well. This may be the case in studies with smaller samples, limited number and type of predictors, and predictors that are either weakly or linearly related to the outcome (Chekroud et al., 2021). Note, however, that some studies that had a large number of predictors (i.e., 80 predictors; Espel-Huynh et al., 2021) or a large sample size (Liang et al., 2022) found minimal benefits, suggesting that multiple factors may determine the relative benefits of ML. However, given that studies comparing approaches were limited and heterogenous, it is difficult to know for certain which specific contexts ML techniques show superior performance in this field. Further studies comparing approaches in similar contexts (i.e., similar samples, number and type of predictors, and outcomes) and under conditions hypothesized to optimize ML performance (i.e., diverse data types, larger samples) are needed to provide a more robust understanding of when ML techniques should be used over traditional approaches.
Literature gaps and future directions
Although the application of ML in predicting ED outcomes is growing and has been met with great enthusiasm, this synthesis identified a number of important literature gaps and directions for future research. We focus on four key gaps.
First, given that research has primarily applied ML to predicting ED diagnosis, knowledge of how ML techniques may be used to facilitate more personalized support, predict the evolving nature and complexities of EDs over time, and forecast responsiveness to treatment is limited. This is an unfortunate oversight and may explain why even the field’s best treatments (i.e., cognitive-behavior therapy) produce modest outcomes (Linardon & Wade, 2018). Future research should seek to explore how ML techniques can be leveraged to deliver more personalized treatments plans that are tailored to an individual’s unique symptom profile and are administered at critical moments. For example, future studies may investigate how ML can be used to deliver such support through just-in-time adaptive interventions (JITAIs). JITAIs offer timely support through technological means (i.e., smartphones) using real-time analysis of passive smartphone or sensor data (i.e., usage patterns, social media interaction, location, heart rate, movement) or low-burden self-reporting (i.e., EMA; Juarascio et al., 2018; Nahum-Shani et al., 2018). ML may enhance JITAIs by predicting the optimal timing and type of intervention to deliver to individuals in moments of need (i.e., high risk for displaying ED behaviors). ML-enhanced JITAIs are emerging across the health field, demonstrating promise in areas including weight-loss interventions (Forman et al., 2019) and alcohol-consumption reduction (Bae et al., 2018). Although some studies used ML models with EMA (Arend et al., 2023; Levinson et al., 2022) and sensor data (Lekkas et al., 2023) to predict ED symptoms, it may be beneficial to also explore how predictive models can be used with these data to deliver JITAIs, providing timely support to people during high-risk moments.
Second, although across studies there were several different types of data used as input (i.e., self-report, neuroimaging, text, physiological), studies rarely implemented more than one data type in a single model. The ability to integrate and process complex and varied data sources is a key advantage of ML and may improve prediction by enabling novel interactions and associations to be discovered between a diverse set of input features (Iniesta et al., 2016). An example of this was seen in Linardon et al. (2022) when ML model performance significantly improved after including usage-pattern variables alongside an initial set of self-reported baseline predictors for predicting responsiveness to ED digital interventions. In clinical settings, it may also be useful to explore whether combining routinely collected clinical data with health-service-use data improves prediction of therapeutic outcomes given that this information is readily accessible in clinical contexts. However, it is important to consider how increasing model complexity may also limit interpretability. Future research should prioritize investigating how diverse types of data sources may be integrated in models to enhance prediction while also evaluating the feasibility of applying these models in clinical settings.
Third, many studies had relatively small sample sizes in the context of ML (n < 1,000) and did not conduct external validation of models on new, independent samples. ML models generally require larger samples to optimize model performance and mitigate the risk of overfitting (Yarkoni & Westfall, 2017). Overfitting occurs when the model captures noise in the training data rather than the true underlying patterns, causing the model to perform well on the training data but fail to generalize to new data (Yarkoni & Westfall, 2017). Prior recommendations suggest avoiding using ML models with fewer than several hundred observations (Poldrack et al., 2020). Furthermore, it is also critical that future studies conduct external validation of their models on new, independent samples because only two studies reported doing so (Burstein et al., 2023; X. Chen et al., 2023). External validation is crucial for mitigating the risk of overfitting and determining whether a model is generalizable to new data that span different clinical settings, populations, and subgroups (Riley et al., 2016). Without this, it remains unclear whether results would replicate in conditions outside the samples in which the models were developed. This is particularly pertinent given that many studies developed models on samples consisting of predominantly White, educated female participants. The potential for racial, ethnic, and gender biases in ML models is a well-recognized concern in the development, validation, and implementation of ML models (Hooker, 2021). Training and testing of models in populations that are traditionally underrepresented in ED research is critical to establish more robust and generalizable models.
To facilitate the collection of large, heterogeneous (e.g., self-report, neuroimaging, social media) data sets, we recommend cross-institutional and international collaborations among researchers in the ED field. An example of cross-country collaboration can be seen by Krug et al. (2021), who collaborated to collect a large clinical sample (n = 1,402) across six different centers in Europe. Data collection may also be facilitated by establishing clinical registries. For example, in Australia, the TrEAT registry—a clinical registry for EDs—includes data from more than 10 clinics and provides a valuable resource for researchers (University of Technology Sydney, n.d.). Such registries may enable the application of ML techniques to address complex questions and understand clinical variability in real-world settings. In addition, EHRs and passive data collected via smartphones (i.e., social media interactions, communication patterns, usage patterns) are increasingly being used to develop risk signatures across other psychiatric disorders (Chekroud et al., 2021). Such data also provide the opportunity to collect large amounts of information across a variety of domains and time points while being low effort for participants.
Fourth, once models have undergone external validation to establish model generalizability, the clinical application of models should be assessed. Specifically, it is important to understand the feasibility of implementing ML models into real-world settings and their actual clinical value. This includes evaluating the usability of the model by clinicians, the incremental utility of the model compared with current clinical practice (i.e., whether the accuracy of ML models for identifying ED cases improved on clinical diagnostic accuracy), and the cost-effectiveness of implementing the model into clinical workflows (Cearns et al., 2019; Steyerberg et al., 2013). To achieve this, external validation should be followed by clinical-impact studies that evaluate the real-world effectiveness of ML-based tools for improving patient outcomes and optimizing management of care (Steyerberg et al., 2013). It is also crucial to conduct research aimed at understanding clinicians’ perspectives, concerns, and potential barriers to using ML in practice. Ongoing collaboration between researchers and clinicians is required to facilitate acceptability of ML-based systems and ensure compatibility with clinical practice and guidelines (Dwyer et al., 2018).
Limitations of this review
A limitation of this review is the lack of preregistration. Because of the broad and exploratory nature of this scoping review, it was difficult to predetermine rigid parameters for the research questions, methodology, and data synthesis. Although this approach allowed for greater flexibility, enabling our methods to evolve with data, preregistration may have enhanced transparency and reduced the risk of bias. However, we sought to mitigate this by adhering to established scoping-review frameworks and ensuring a systematic and replicable approach to study selection and data extraction.
Conclusion
In conclusion, research applying ML to ED-outcome prediction is rapidly emerging, with 75 studies identified in this scoping review. Although the utility of ML is promising in this field, the available research largely contains proof-of-concept studies that simply demonstrate feasibility of these advanced computational approaches. It is evident from this review that larger-scale validation studies are required to establish more robust and generalizable findings.
Supplemental Material
sj-docx-1-cpx-10.1177_21677026251340348 – Supplemental material for Machine-Learning Applications in Eating-Disorder-Outcome Prediction: A Systematic Scoping Review
Supplemental material, sj-docx-1-cpx-10.1177_21677026251340348 for Machine-Learning Applications in Eating-Disorder-Outcome Prediction: A Systematic Scoping Review by Zoe McClure, Matthew Fuller-Tyszkiewicz, Mariel Messer and Jake Linardon in Clinical Psychological Science
Footnotes
Acknowledgements
Z. McClure wishes to acknowledge the support of the Australian government provided through an Australian Government Research Training Program Scholarship.
Transparency
Action Editor: Kelsie T. Forbush
Editor: Jennifer L. Tackett
Author Contributions
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
