Abstract
Positioning single cells on a solid surface is a crucial technique for understanding the cellular functions and cell–cell interactions in cell culture assays. We developed a microfluidic chip for depositing single cells in microwells using a simple micropipette operation. Cells were delivered to microwells by the meniscus motion of liquid interface. The residue deposits of cells were redistributed with air injection, and the isolated single cells were stored in microwells. Different microwell sizes and depths were studied to evaluate the trapping possibility of cells. Medium replacement and cell viability staining with the isolated single cells were achieved in microwells. The chip will serve as a tool for single-cell patterning in an easy-to-use manner.
Introduction
Achieving site-specific placement of individual cells in a cell culture is beneficial for the systematic study of contact and diffusive-mediated communications among neighboring cells 1 –4 . Cell deposition has been demonstrated with laser guidance 5,6 or micromanipulator 7,8 to organize single cells on culture substrates. By using them on a microscope stage, single-cell analysis with onstage incubation for long-term cell culturing can be provided. For example, laser tweezer microscopes 9 –11 are an effective tool to trap individual cells for studying biological mechanisms at the single-cell level. They trap cells by exploiting the optical forces generated by a highly focused laser beam. Currently, cells can be actively printed onto the surface by using laser forward transfer techniques such as matrix-assisted pulsed laser evaporation 12 and inkjet printing 13 .
One facile and simple process to deposit cells on a solid surface is convective sedimentation assembly 14–15 . This process includes convective evaporation for cell redistribution. When a droplet of the cell suspension evaporates on the substrate, the cells in the evaporating portion of the entrained volume are deposited under the meniscus. The deposited cells are pulled into the thin film in front of the meniscus and divided evenly among the entrained volume. A significant amount of the cells in the liquid meniscus will sediment during the deposition process.
In the process of convective evaporation, the surface tension force works at the air–water interface translating on the substrate 16,17 . The translation of the liquid interface can be imposed by sliding a droplet between the 2 glass slides. Prevo and Velev 18 reported a modified convective assembly method that allows rapid and controllable deposition from small volumes of cell suspension. A small liquid body is trapped between 2 plates, and a linear motor pushes the top plate along the long axis of the bottom plate, thereby dragging the meniscus with it. The cell deposition takes place at the edge of a long meniscus of the liquid trapped between 2 plates. The geometry is translationally invariable in the meniscus direction, and there is no redistribution of cells parallel to the meniscus edge.
In this article, we describe a microfluidic cell deposition in which the liquid interface of the cell suspension is manipulated by manual pipetting inside the microfluidic channel. Previously, our group had developed a microfluidic chip for depositing DNA molecules by syringing them through microgrooves 19,20 . This process enabled control over the meniscus motion. Here, we demonstrate an application study of the chip to cell deposition by rapid and simple operation. A microfabricated pattern for isolating single cells is embedded onto the surface of the microfluidic channel. It comprises 2 types of silicone substrates: a microchannel for cell suspension transport and a microwell for cell isolation (Fig. 1). We analyze the cell trapping efficiency for different sizes and depths of the microwells. In addition, we analyze the cell viability for the deposited single cells through medium replacement.
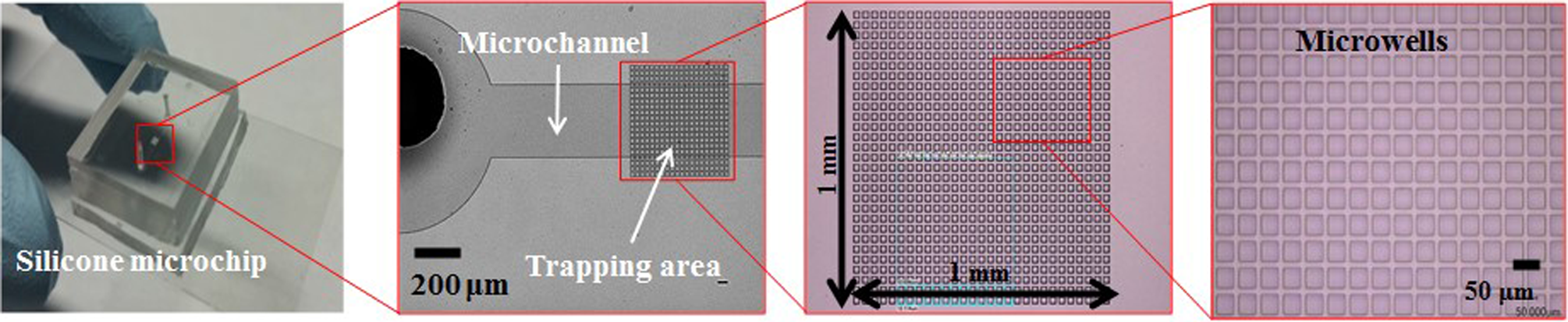
A picture and microscopic images of the microfluidic chip.
Materials and Methods
Cell Sample Preparation
Human non-small cell lung carcinoma cell line NCI-H1299 (American Type Culture Collection, Manassas, VA, USA) was cultured in Roswell Park Memorial Institute (RPMI) medium (Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS; Funakoshi, Tokyo, Japan) and 1% penicillin streptomycin (Thermo Fisher Scientific, Waltham, MA, USA) at 37 °C and 5% CO2. Cells were harvested at 80% confluence by trypsinization and suspended at 1 × 105 cells per milliliter in culture medium for cell deposition experiments. The collected cells were incubated in phosphate-buffered saline with 1 nM calcein-AM (Dojindo Laboratories, Kumamoto, Japan) at 37 °C and 5% CO2. Trypan blue solution, 0.4% (Thermo Fisher Scientific, Waltham, MA, USA), was applied to the deposited single cells for live–dead cell staining.
Fabrication Process
Detailed procedures for the fabrication of a microfluidic chip are as described in Yasaki et al. 19 In summary, a soft lithography technique was used for silicone elastomer polydimethylsiloxane (PDMS) molding. The mold fabrication process for PDMS microstructures was performed according to the SU-8 Data Sheet (Nippon Kayaku, Tokyo, Japan). SU-8 (3025, Nippon Kayaku) was coated on the silicon substrate (3 in., Ferrotec, Tokyo, Japan) by using a spin coater (IF-D7, Mikasa, Tokyo, Japan). After soft baking, this layer was exposed to ultraviolet light through a photomask in order to form patterns by using a mask aligner (M-1S, Mikasa, Tokyo, Japan). After the development, the substrate was washed by rinsing with SU-8 Developer (Nippon Kayaku, Tokyo, Japan) and isopropyl alcohol (Wako Pure Chemical Industries, Tokyo, Japan). A PDMS prepolymer solution containing a mixture of 10:1 mass ratio of PDMS oligomers and a reticular agent from a Sylgard 184 Kit (SILPOT 184 Dow Corning, Toray, Tokyo, Japan) was poured onto the silicon substrate and cured for 2 h at 80 °C. The patterned PDMS plate was cut using a surgical scalpel blade (X-ACTO, Elmer’s Products, Westerville, OH, USA).
Patterns for Microchannel and Microwells
A PDMS microfluidic channel (650-μm width, 0.8-cm length, and 50-μm height) was used as a pattern for cell suspension transport. PDMS microwells for cell trapping were constructed using a total of 841 square patterns in a 1 mm × 1 mm area, varying the size and depth. The side lengths of 25 µm, 30 µm, and 35 µm were prepared. The pattern depths were 26 µm, 36 µm, and 60 µm.
Microfluidic Chip Setting
An inlet hole and an outlet hole were punched as reservoirs of the PDMS microfluidic channel by using a razor sharp stainless steel cutting tip (Harris, UNI CORE, Sigma-Aldrich, St. Louis, MO, USA). The size of the inlet and outlet holes was 1 mm in diameter. The flow path of the microfluidic channel was overlaid on the square pattern of the microwells. The surfaces of the PDMS structures were bonded by exposure to a vacuum plasma spray for 90 s. The plasma treatment was performed using soft plasma etching equipment (SEDE-PFA, Meiwafosis, Osaka, Japan).
Cell Deposition and Observation
An air displacement micropipette (PIPETMAN P-20, Gilson, Middleton, WI, USA) was used to introduce cell suspension (5 μL) into the PDMS microfluidic channel. The trapped cells in the PDMS microwells were observed using a cooled CCD camera connected to an all-in-one fluorescence microscope (BZ-X700, Keyence, Osaka, Japan). The excitation wavelength was selected using a BZ-X filter GFP (470/40, Keyence, Osaka, Japan). The deposited single cells were directly counted from the obtained fluorescence images.
Results
Deposition Process of Cells
The procedure from the start of the 2 bonding PDMS structures to the cell deposition is illustrated in Fig. 2A. The cell suspension was injected into the microchannel, enabling slow flow by single pipetting. After 15 min, the cells fell down from the microwells due to gravity of incubation. The overspilled cells were pushed out and redistributed to the microwells by pipetting of air into the microchannel. The side views depicted in Fig. 2B are based on the sequential microscopic monitoring. Figure 2C shows a sample of the deposited and isolated cells.
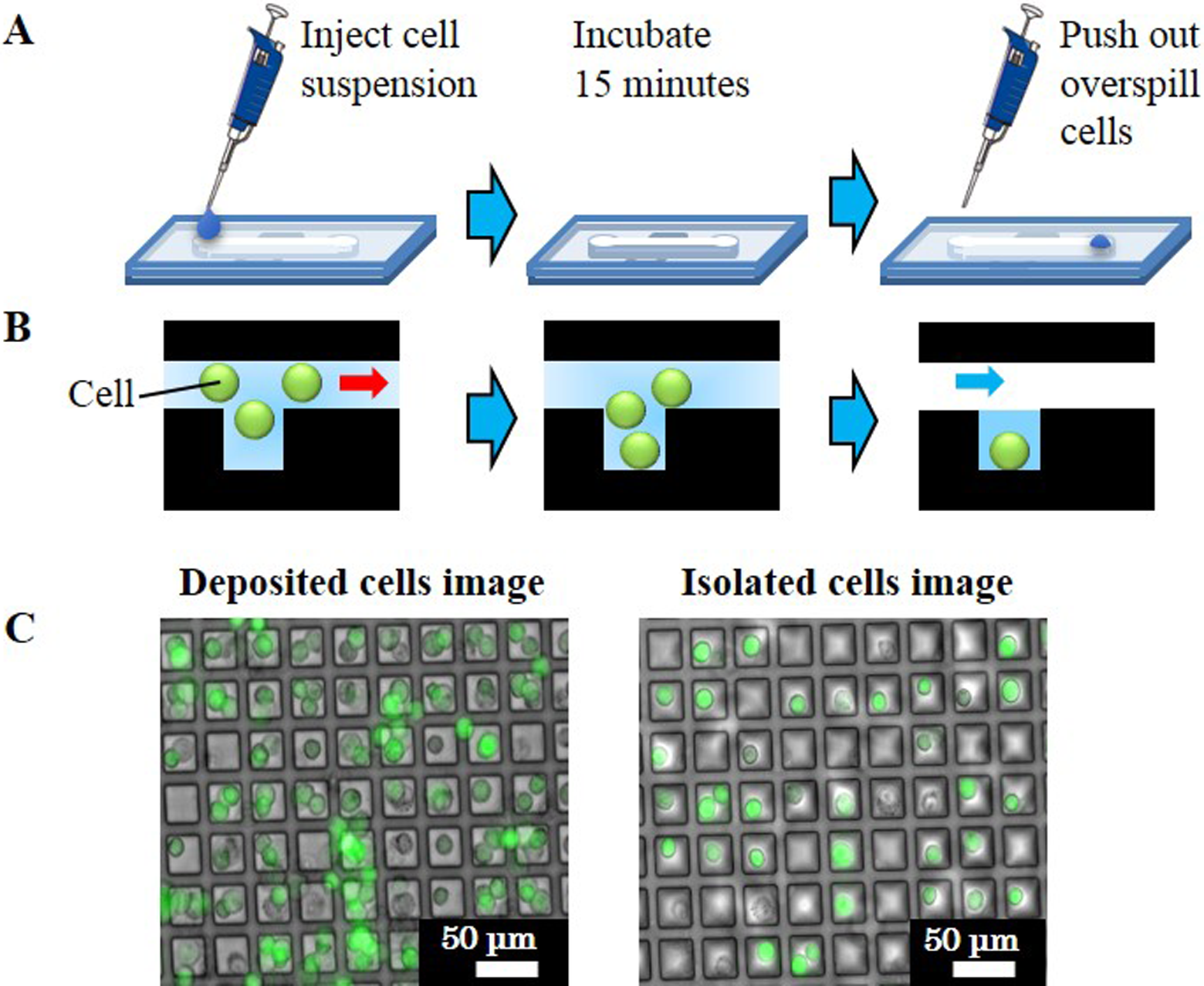
Operation of cell deposition microchip. (A) Experimental procedure with micropipette. (B) Schematic of single-cell trapping in a microwell. (C) Microscopic fluorescence images of cells in microwells.
Cell Trapping Efficiency Analysis
We defined simple metrics 19,20 to allow the comparison of a cell trapping efficiency in different sizes and depths of the microwells. The trapping possibility of cells was calculated as a cell trapping efficiency, where the number of microwells with cells was divided by the total number of microwells on the surface of the microfluidic channel. The cell trapping efficiency was taken to be the number averaged value. The microwells in the center area (325 µm in width) of the microfluidic channel were counted for the calculation of the cell trapping efficiency. By using values from 3 replication tests, the cell trapping efficiency was obtained as a mean standard deviation (SD). The number of isolated single cells in a microwell was clearly imaged (Fig. 3A). The cell trapping efficiencies in different microwell sizes and depths in the microfluidic channel are shown in Fig. 3B and C. The highest cell trapping efficiency of single cell was obtained at the size and depth of the microwells for 25 μm (40 ± 2%) and 26 μm (57 ± 6%), respectively. Under such conditions, over 500 cells were successfully deposited and isolated simultaneously at each occurrence.
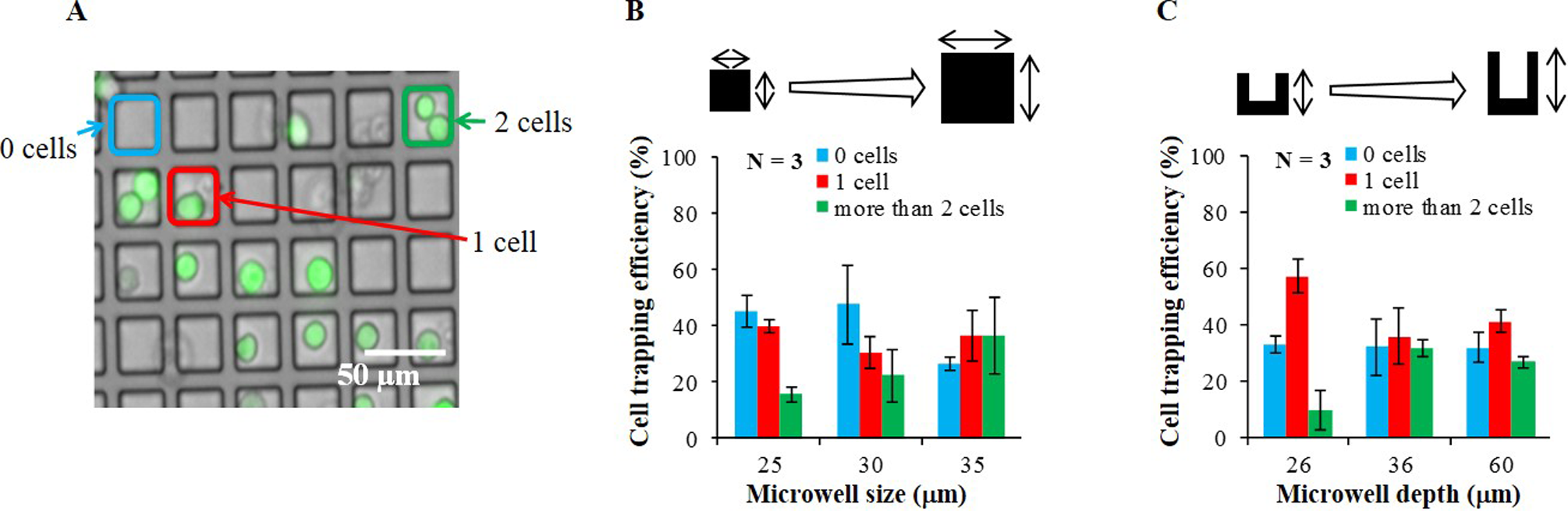
Results for trapping possibility analysis. (A) A microscopic fluorescence image for counting cell numbers in a microwell. (B) Cell trapping efficiency for different size of microwells. Depth of microwell was fixed at 36 μm. (C) Cell trapping efficiency for different depth of microwells. Size of microwell was fixed at 30 μm.
Medium Replacement and Cell Viability Staining
The changes in cell trapping efficiency of the isolated cells by medium replacement inside a pattern of microwells (size = 30 μm, depth = 26 μm) are shown in Fig. 4A. Before medium replacement, the cell trapping efficiency of more than 1 cell was 67 ± 6%. After medium replacement, a large part of the isolated cells remained inside the microwells, and the cell trapping efficiency of more than 1 cell was 50 ± 13%. A microscopic image of cell viability testing for the isolated single cells inside microwells is shown in Fig. 4B. The trypan blue staining revealed that over 90% of cells were viable.
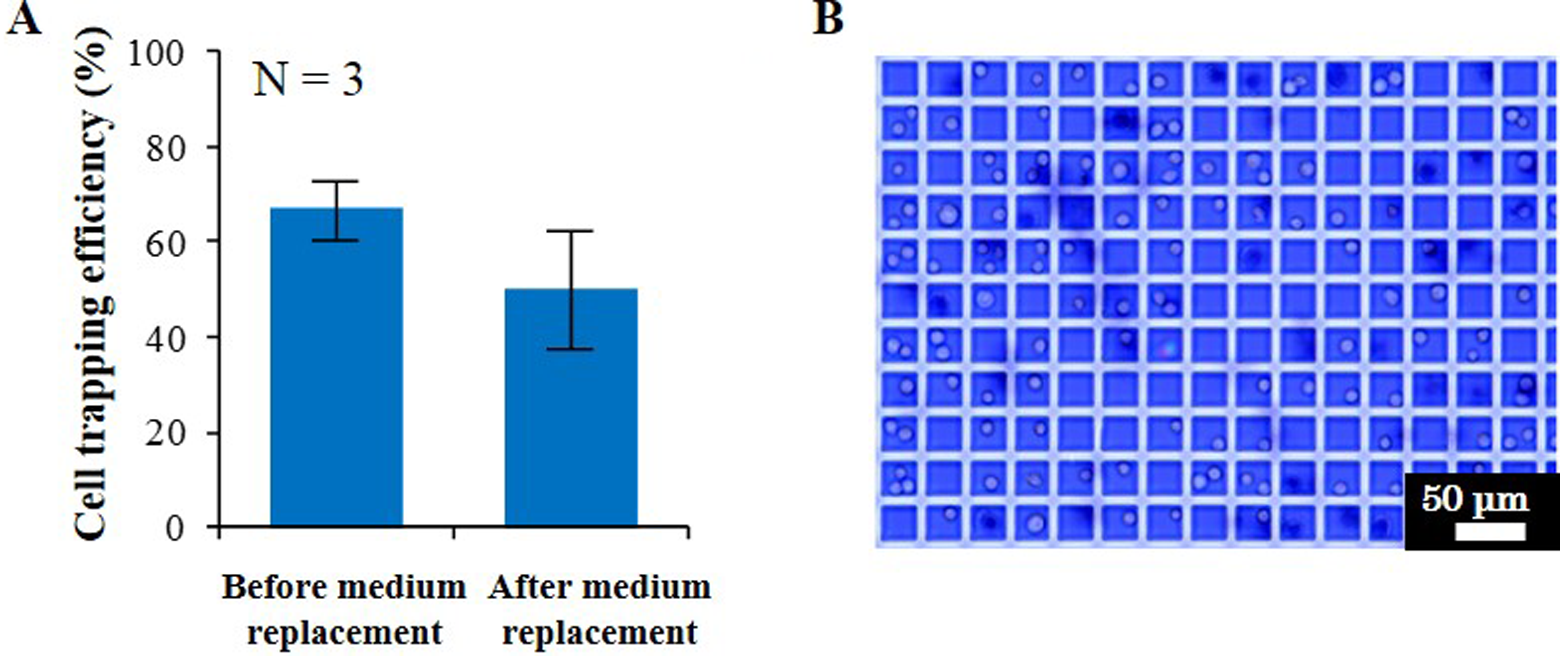
Results for cell viability testing through medium replacement. (A) Cell trapping efficiency for more than 1 cell before and after medium replacement. (B) A microscopic image of trypan blue staining for the isolated single cells.
Discussion
When the cell suspension approached the microwells in the fluidic channel, some of the cells near the liquid interface were induced to the cavity. At that moment, the contact line of the liquid interface passed through the cavity and the moving liquid interface trapped the cells in the cavity (Fig. 2B). The air injection washed the residue deposits of cells away, and the isolated cells were stored in the cavity with the medium (Fig. 2C). When the size and depth of the microwells were increased, the cell trapping efficiency of a single cell was slightly decreased; in contrast, the cell trapping efficiency of more than 2 cells was certainly increased (Fig. 3B and C). This is likely due to the spherical size (16 μm) of the single cell.
In this article, simultaneous cell deposition and isolation in a PDMS microfluidic chip were demonstrated. In the course of micropipette control over liquid interface motion, cells were trapped in the cavities of the microwells by 2-step pipetting with cell suspension and air injection. The trapping possibility of single cells was highly influenced by the size and depth of the cavity. The isolated single cells were captured alive and remained inside the microwells through the medium replacement.
Footnotes
Acknowledgments
This research is supported by the Center of Innovation Program from Japan Science and Technology Agency, JST, and JSPS KAKENHI Grant Number 26750146.
Ethical Approval
This study was approved by our institutional review board.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Center of Innovation Program from Japan Science and Technology Agency, JST, and JSPS KAKENHI Grant Number 26750146.
