Abstract
Introduction
A role for bacteria and other microbes has long been suspected in the chronic inflammatory sinonasal diseases. Recent studies utilizing culture-independent, sequence-based identification have demonstrated aberrant shifts in the sinus microbiota of chronic rhinosinusitis subjects, compared with ostensibly healthy controls. Examining how such microbiota shifts occur and the potential for physician-prescribed interventions to influence microbiota dynamics are the topics of the current article.
Methods
The nasal cavity microbiota of 5 subjects was serially examined over an 8-week period using pan-bacterial 16S rRNA gene sequencing. Four of the subjects were administered topical mometasone furoate spray, while 1 subject underwent a mupirocin decolonization procedure in anticipation of orthopedic surgery.
Results
Measures of microbial diversity were unaffected by intranasal treatment in 2 patients and were markedly increased in the remaining 3. The increase in microbial diversity was related to clearance of Moraxella spp. and a simultaneous increase in members of the phylum Actinobacteria. Both effects persisted at least 2 weeks beyond cessation of treatment. Transient changes in the relative abundance of several bacterial genera, including Staphylococcus and Priopionibacteria, were also observed during treatment.
Conclusions
The effects of intranasal steroids on the sinonasal microbiome are poorly understood, despite their widespread use in treating chronic sinonasal inflammatory disorders. In this longitudinal study, administration of intranasal mometasone furoate or mupirocin resulted in shifts in microbial diversity that persisted to some degree following treatment cessation. Further characterization of these effects as well as elucidation of the mechanism(s) underlying these changes is needed.
Introduction
Chronic inflammatory diseases of the nasal cavity and paranasal sinuses, including chronic rhinitis and chronic rhinosinusitis (CRS), are highly prevalent and problematic disorders with poorly understood etiologies. Chronic rhinitis affects up to 20% of the population globally and consists of a heterogeneous group of suspected etiologies that result in a common end point of symptoms derived from chronic inflammation.1,2 Chronic rhinitis is broadly classified into allergic rhinitis (AR), infectious rhinitis, and nonallergic rhinitis (NAR).3,4 CRS is a chronic inflammatory disorder of the paranasal sinuses, which likely represents a heterogeneous group of disorders resulting in a common end point of signs and symptoms resulting from chronic inflammation. 5 Prolonged medical therapies such as saline rinses, corticosteroids, antihistamines, and antibiotics are the mainstays of therapy for such inflammatory sinonasal disorders.
Management of chronic rhinitis and rhinosinusitis has generally required the use of intranasal corticosteroids (INS). INS use is considered first-line therapy for AR, and although less effective in NAR, it is frequently utilized as part of the regimen introduced by the physician. 6 Many potential mechanisms for INS modulation of host inflammation were proposed early after its introduction into clinical practice, directly targeting both epithelial cells and inflammatory cells; these include modulation of mast cells, TH2 cells, eosinophils, and the arachidonic acid pathway.7,8 However, despite 20 years of clinical practice and research, the mechanism(s) of action of INS in chronic sinonasal disorders remains incompletely understood. 9 In addition to immunomodulatory mechanisms, certain INS formulations manifest antimicrobial properties, for example, against Staphylococcus aureus biofilms and planktonic Streptococcus.10,11 The potential for a direct antimicrobial effect is supported by the finding of reduced bacterial recovery in sinus samples obtained during surgery among CRS patients receiving INS treatment. 12 The rare occurrence of mucosal candidiasis with INS use also supports the hypothesis of drug-related microbial community shifts, analogous to antibiotic-associated vaginal yeast infection. 13 Currently, 7 different INS formulations are available in the United States, and their different additives may also convey antimicrobial properties or other unknown effects. 14
Commensal microbes are critical determinants of human health,15–20 and imbalances in human-associated microbial communities (“dysbiosis”) are strongly associated with several human diseases (obesity, inflammatory bowel disease, vaginosis, S. aureus carriage).21–24 In fact, our initial studies and those of others have suggested that community diversity may be a sign of health in the sinuses.25–27 However, the origins and etiological significance of these alterations remain unclear due, in part, to limited understanding of both the temporospatial organization of microbial communities and of the host and environmental determinants that shape them. An individual’s homeostatic “core microbiome” is relatively stable over time but exhibits a quick response to external perturbations and may shift to a new baseline when stressed, resulting in alterations in bacterial community dynamics and host–microbe homeostasis.28–31 Although the human microbiome has been the focus of much research, the basic ecological principles that govern human–microbiota interactions require further investigation. 32 Despite the rich theoretical foundation of community ecology 33 developed primarily from studies of plant and animal communities, basic understanding of how the composition of a microbial community affects its temporal stability, spatial organization, resistance, and resilience to perturbation is generally lacking.34,35
Utilization of INS or other medications and the resulting bacterial community shifts may temporarily, or even permanently, alter microbial composition and function, potentially eliciting a number of unseen and pathogenic effects.36,37 As the equilibrium between commensals and putative pathogens is hypothesized to determine an individual’s susceptibility to disease, in this study, we tested the hypothesis that INS use could profoundly alter the commensal and pathogen burden either by direct or indirect mechanisms.
Materials and Methods
Study Design and Population
A prospective, institutional review board-approved study (Colorado Multiple Institutional Review Board protocol number 11-1134) was performed on 5 adult volunteers. Four otherwise healthy adult males, aged 25 to 40, with a known diagnosis of chronic noninfectious rhinitis underwent a minimum of 1-month washout period without the use of any medical therapies for rhinitis and then were given mometasone furoate nasal spray daily for 1 month. The fifth one was a female subject, aged 25, who did not have any sinonasal disorder but underwent twice daily topical mupirocin therapy to the anterior nares as a standard preoperative therapy for a planned orthopedic surgery. Subjects underwent weekly sampling of the anterior nares in the classic well-described fashion using a sterile swab 24 for 2 weeks prior to therapy, during 4 weeks of treatment, and for at least 2 weeks after cessation of therapy. The anterior nares were chosen as the nasal sampling site owing to the anterior distribution of nasal topical medications, comparison ability across many other published studies, keen interest in medical implications of S. aureus carriage, and its spatial relevance for nasal cavity inflammatory disorders such as rhinitis. Baseline time points were used to confirm microbiome stability of this bodily subsite as has been shown in prior Human Microbiome Project studies.
Sample Collection
All swabs were collected from subjects during the same season using CultureSwabs (BD Diagnostics, Franklin Lakes, NJ) in the standard fashion, rotating at least 5 full turns until fully saturated. CultureSwabs were placed on ice immediately following collection and frozen at −80°C until DNA extraction.
16S Amplicon Library Construction
A phenol:cholorform bead-beating method was used to extract total genomic DNA from the swab heads. Swab heads were placed in sterile microcentrifuge tubes that contained 500 μl of buffer B (142.9 mM NaCl, 142.9 mM Tris-Cl [pH 8.0], 14.3 mM EDTA, and 5.7% SDS) and 500 μl of 1:1 phenol:chloroform.38,39 A total of ∼0.25 g of 0.1-mm zirconia/silica beads was added to the suspension, and samples were mechanically disrupted in a Mini Beadbeater-16 (Biospec Products, Bartlesville, OK) at the highest setting for 2 min. The samples were centrifuged (>14,000 × g) for 5 min, and the aqueous layer was reextracted 2 more times with an equal volume of phenol:chloroform and once with an equal volume of chloroform. DNAs were precipitated by the addition of 0.5-volume ammonium acetate (7.5 M) and 1 volume of 100% isopropanol, incubated at −80°C for 10 min, then centrifuged (>14,000 × g; 25 min). Nucleic acid pellets were washed with 250 μl of 70% ethanol and centrifugation (>14,000 × g; 5 min), dried in air, then resuspended in 30 μl of sterile 1× Tris-EDTA (pH 8.0), and stored at −80°C until polymerase chain reaction (PCR) processing. All DNA extraction and PCR steps were performed in a high-efficiency particulate air -filtered laminar flow hood that was decontaminated by ultraviolet light.
Amplicons of the 16S rRNA gene (∼500 b.p.; primers 27FYM + 3 40 and 515R41,42) were generated via broad-range PCR using 5′-barcoded reverse primers. PCR yields were normalized using a SequalPrep™ kit (Invitrogen, Carlsbad, CA), pooled, lyophilized, and gel purified, as previously described. 25,43 Pooled amplicons were provided to the Center for Applied Genomics at the University of Toronto for pyrosequencing on a 454/Roche Life Sciences GS-FLX instrument using Titanium chemistry (Roche Life Sciences, Indianapolis, IN). Sequences were demultiplexed and screened for nucleotide quality (bases at 5′ and 3′ ends with mean Q < 20 over a 10 nucleotides window were discarded), ambiguous bases (sequences with >1 N were discarded), and minimum length (sequences <200 nucleotides were discarded). Chimera screening was performed by the tool ChimeraSlayer. 44 Genus-level taxonomic calls were produced by the Ribosomal Database Project (RDP) classifier, which performs naive Bayesian taxonomic classification versus a training set.45,46 Species-level taxonomy precision was obtained via BLAST against a database of sequences obtained from Silva104 tagged as isolates. 47 Species-level results required that the candidate sequence overlaps the database sequence by at least 95% with at least 99% sequence identity and that the Silva-derived taxonomy of the database hit matches the RDP classifier genus-level taxonomy result.48,49
Statistical Analysis
The R (v3.0.3, cran.r-project.org) 50 and Explicet (v2.10.5, www.explicet.org) 51 software packages were used for data display, analysis of results, and figure generation. Principal coordinates analysis (PCA) applied the R prcmp command to a covariance matrix of species abundance data that was first center log transformed to correct for the compositional nature of the data set. 52 A small value (1/total sequences per sample) was added to each count value prior to log transformation. 52 Correlations between principal component axis scores and microbial abundances were assessed by Spearman rank correlation test. Standard alpha-diversity indices (Good’s coverage, Chao1, Shannon diversity, and evenness) 53 were estimated with rarefaction (n = 50 sequences/sample) and replicate resampling (n = 1000 replicates) using Explicet. 51
Results
This prospective study of the effects of INS on nasal microbial biodiversity included 4 adult volunteers with a known diagnosis of chronic rhinitis. Each subject underwent a minimum washout period of 1 month without the use of any medical therapies for rhinitis and then were given mometasone furoate monohydrate nasal spray (Nasonex®, Merck) administered as two 50 μg sprays in each nostril once daily for 1 month (total dose 200 μg per day). 54 A fifth subject did not have any sinonasal disorder but underwent twice daily topical mupirocin therapy to the anterior nares as a standard preoperative therapy for a planned orthopedic surgery. All subjects underwent weekly sampling of the anterior nares by sterile swab as has been previously described 24 for 2 weeks prior to therapy, during 4 weeks of treatment, and for at least 2 weeks after cessation of therapy.
Broad-range 16S rRNA gene PCR and pyrosequencing were successful in 30 of the 32 (94%) samples from INS-treated subjects (swabs taken 2 weeks prior to and at week 4 of treatment from subject A were negative). No differences in PCR or sequence yield were noted before, during, or following treatment in these subjects, suggesting that the steroid nasal spray was not broadly bactericidal. In contrast, 4 of the 8 (50%) swabs obtained from the mupirocin-treated subject were PCR negative, specifically those from 2 week-pretreatment, and treatment weeks 1, 2, and 4. From those samples with positive PCR, we generated a median of 1030 16S rRNA gene sequences per sample and Good’s coverage indices ranging from 91% to 99%, indicating that sequence coverage was adequate.
As expected from previous studies,24,29,48,55–58 nasal microbiotas were dominated by several genera of the phyla Actinobacteria, Firmicutes, and Proteobacteria (Figure 1; Supplemental Figure 1), including Corynebacterium (29.5% of sequences), Staphylococcus (14.4%), Propionibacterium (13.1%), Moraxella (6.8%), Gordonia (3.4%), and Streptococcus (3.4%). Furthermore, substantial variation in baseline nasal microbiotas was evident across the 5 study participants (Figures 1 to 3).
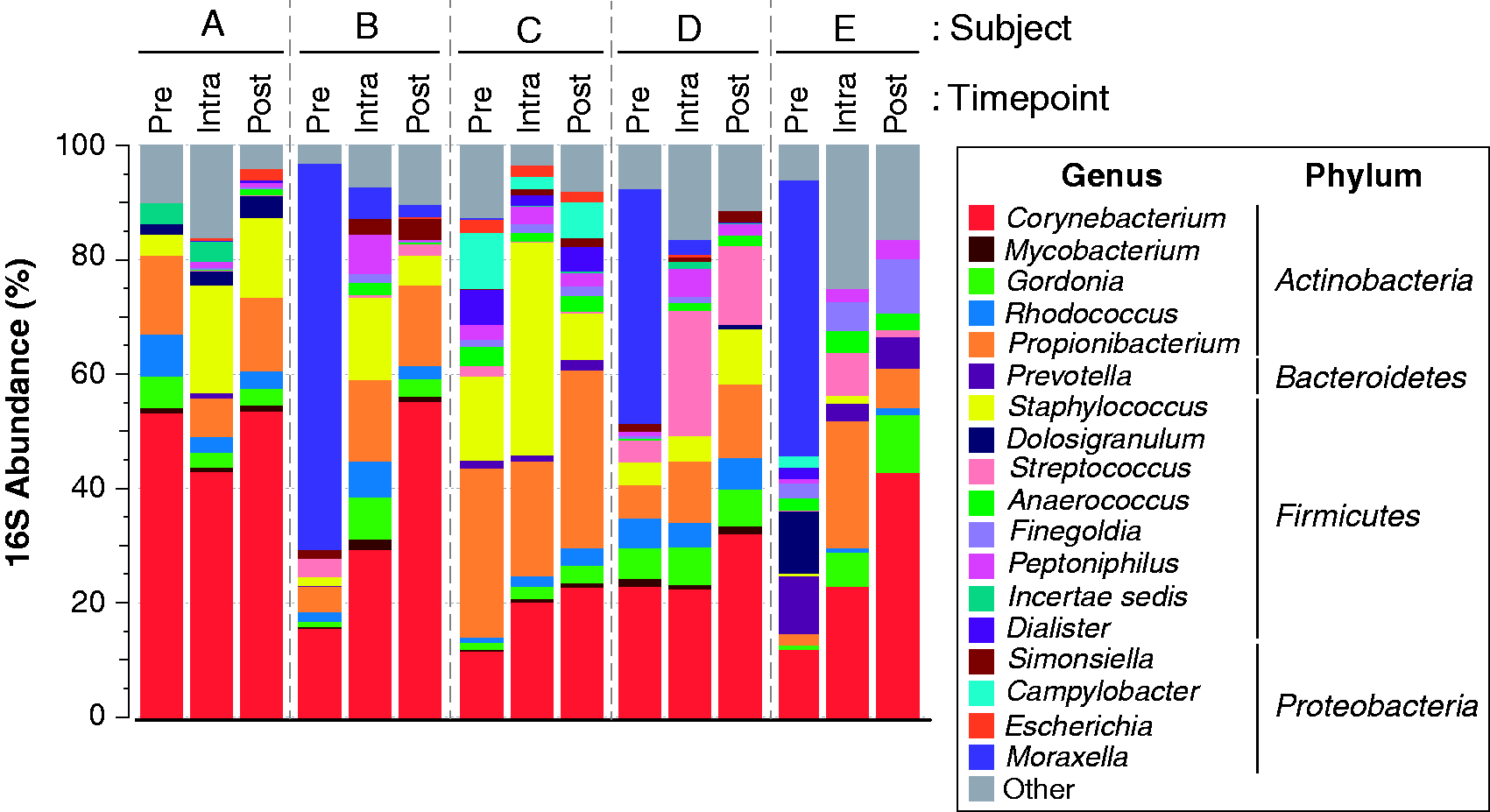
Changes in nasal microbiota accompanying INS and mupirocin treatment. Relative abundances of nasal bacterial genera are summarized as mean values for pre-, intra-, and posttreatment samples. For simplicity of display, only genera with mean relative abundances greater than 0.5% across all specimens are displayed; rare taxa are aggregated into the “Other” category. The legend displays the color coding of genera along with the phyla to which these taxa belong (ie, Actinobacteria, Bacteroidetes, Firmicutes, and Proteobacteria). Subjects A to D were treated with intranasal steroids, while subject E was treated with topical mupirocin. Pre: pretreatment time points. Intra: intratreatment time points. Post: posttreatment time points.
Upon initiation of treatment with either INS or mupirocin, the nasal microbiota of each subject shifted in composition and biodiversity (Figures 1 to 3). For example, PCA results of species-level data showed clear separation between pretreatment and intratreatment samples among INS-treated individuals (Figure 2(a)); these differences appeared to persist even after treatment cessation. Because intra- and posttreatment samples were shifted relative to pretreatment sample along both principal coordinates axes 1 (PC1) and 2 (PC2), we next performed Spearman correlation tests between PC1, PC2, and PC3 scores and the 20 most highly abundant species-level taxa (Figure 2(b)). Significant positive and negative correlations were identified in this analysis. For example, several Staphylococcus spp. species were positively correlated with PC1 scores, whereas Moraxella spp. were negatively correlated with PC1. These results suggest that despite differences in baseline microbiota, INS treatment generally increased the relative abundances of several taxa, most notably the staphylococci, while suppressing other taxa such as Moraxella spp. and streptococci. In contrast, the effects of mupirocin treatment were not evident along PC1 and PC2, indicating that nasal microbiota respond differently to this treatment, compared with INS.
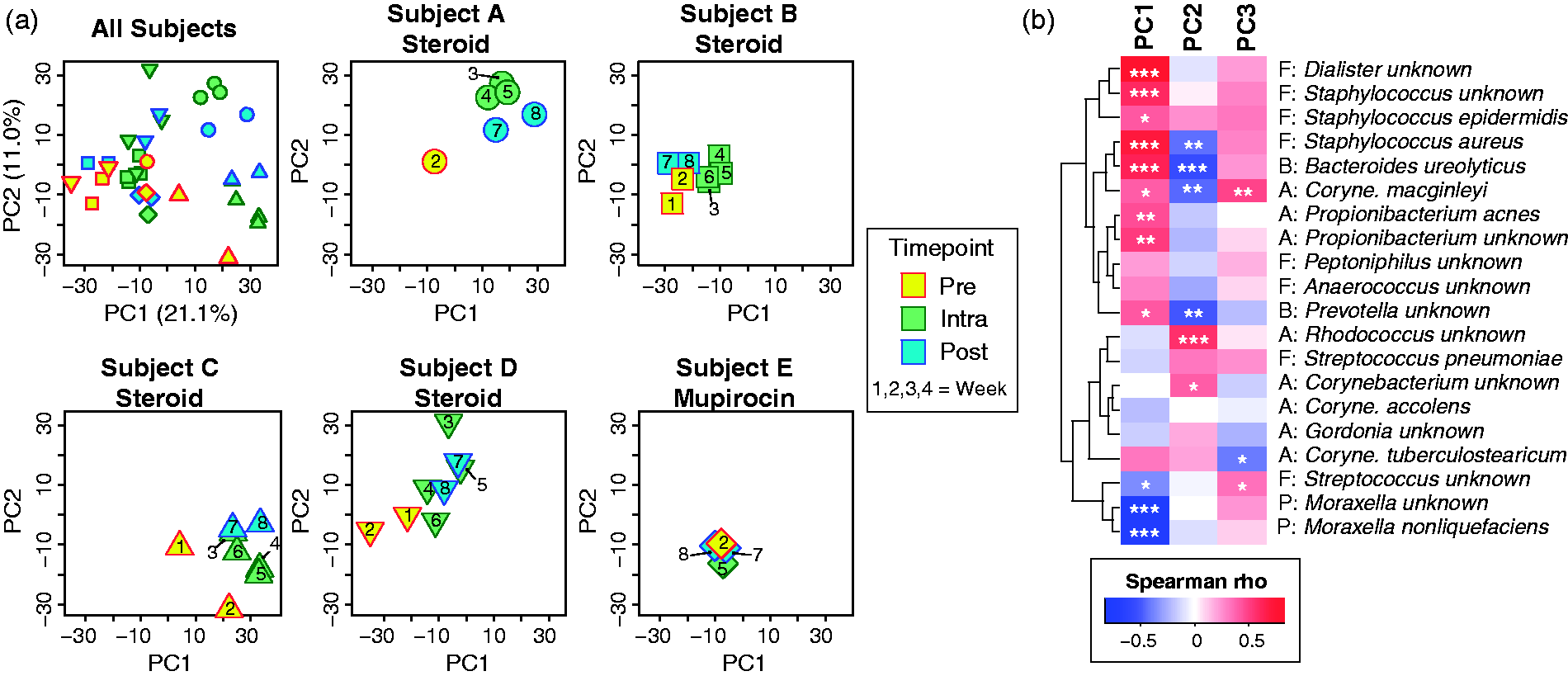
Principal coordinates analysis (PCA). PCA was performed on species-level data for all subjects and time points. PC1, PC2, and PC3 accounted for 21.1%, 11.0%, and 10.3% of the overall variance, respectively. (a) Plots of samples along principal components axes 1 and 2 for all subjects/time points together along with separate plots for each individual. Subjects are distinguished by different symbol shapes, while time points are color coded by treatment phase (pre-, intra-, and posttreatment) and marked by time of sampling, in weeks (missing time points represents samples from which 16S rRNA gene PCR was not successful). (b) Correlations between PC1, PC2, and PC3 scores and most abundant bacterial species/genera. The heat map is color coded by Spearman correlation coefficient (rho), while P values are indicated by symbols: *P < .05. **P < .01. ***P < .001. PC1: principal coordinates axis 1; PC2: principal coordinates axis 2.
The dynamic changes occurring in Staphylococcus, Moraxella, and other genera during the course of treatment are illustrated by examining the relative abundances of genera across the study time course (Figure 3). Some alterations, such as increased Staphylococcus, appeared transiently and resolved once treatment ceased. The genera Corynebacteria and Gordonia exhibited a consistent and stable increase in relative abundance with treatment that remained at least 2 weeks beyond cessation of treatment. In contrast, the diminished levels of Moraxella that occurred during treatment also lasted throughout the posttreatment period. Remaining genera did not exhibit consistent or notable changes in relative abundance in response to treatment.

Temporal variation in microbial diversity. These panels display the relative abundances of select bacterial genera (the 5 most highly abundant) and alpha-diversity indices across the study. Samples obtained at weeks 1 and 2 were pretreatment, those collected at weeks 3 to 6 were during treatment (indicated by shaded box), while those collected at weeks 7 and 8 were posttreatment. Subjects A to D were treated with intranasal steroids, while subject E was treated with topical mupirocin.
In addition to changes in relative abundances of taxa, 2 of the INS-treated subjects (A and E) exhibited sharp, but transient increases in alpha-diversity indices (ie., richness, complexity; Figure 3) immediately following initiation of treatment. In the remaining subjects, these indices were either unchanged (richness, complexity) or trended downward (evenness) over the course of treatment.
Discussion
Data from this prospective, longitudinal survey of human subjects using mometasone furoate nasal spray demonstrate specific shifts in the types and relative abundances of bacteria inhabiting the anterior nares. Changes in the nares microbiota were observed not only during the use of the steroid spray but for several weeks after cessation, indicating another potential mechanism for symptomatic benefit beyond their classic direct anti-inflammatory actions.
In general, healthy adults harbor anatomical site-specific core microbiota that directly influences immunity and epithelial barrier function.15,59–64
However, a subject’s baseline microbiome at a particular body subsite, for example, nasal cavity, may be subject to perturbation from external stimuli, such as a viral infection, alteration in the local microenvironment (eg, pH), and, as observed in this pilot study, by medication administration.28–31 The response to a given perturbation could result in at least a transient shift away from the stable core followed by return to the baseline state (ie, resilience; Figure 4). Study of stability and resilience in the gastrointestinal tract has demonstrated that humans may have a small degree of random “drift” from the baseline, and although individuals typically exhibit stability and resilience, they do so to varying degrees. 66 The factors that govern stability and resilience are unclear, and it has been hypothesized that shifts from the baseline microbiota create periods of susceptibility to disease or pathophysiology. The overall stability of the nasal cavity microbiome has recently been investigated;24,29,64,67,68 however, its response to perturbation and resilience is not well defined. We have shown in a prior study of sinus microbiota in CRS that antibiotic use is associated with loss of microbial diversity and may allow for S. aureus dominance. 25 Subsequently, we examined changes in the sinus microbiota after surgery with perioperative antibiotics and observed significant early changes with several subjects returning to their preoperative microbial state by 6 weeks following surgery and antibiotic treatment. 30 Although further elaboration of these findings is merited, taken together, these data suggest that the potential exists for physician-administered therapies to effect changes in the upper airways microbiota. As first-line medical therapy for chronic rhinitis, the mechanisms of action of intranasal steroid formulations should be more thoroughly understood but unfortunately remain largely unknown. 9

Microbiome perturbation by host or external stressors can lead to a variable extent of disturbance in which return to a healthy baseline may occur (resilience). Reprinted from Vickery and Ramakrishnan 65 with permission from Elsevier.
Specific pathogens such as S. aureus contribute to recalcitrance in the most challenging subset of chronic rhinitis and CRS patients and in a broader medical context confer significant risk to general health. 69 Pathogen colonization has been theorized as a potential risk factor for acute disease, particularly in the setting of airway dysbiosis, 70 as communities of nonpathogenic microbes colonizing the upper airway surface may limit the capacity of pathogens to proliferate through direct commensal–pathogen interactions (“pathogen exclusion”) or indirect mediation of mucosal immunity. 71 INS use, based on our preliminary data, alters commensal microbiota and may coincidentally decrease pathogen presence and abundance and therefore may serve as a complementary method to eradicate colonizing pathogens.
Interestingly, our results suggest that mupirocin and intranasal mometasone furoate manifest similar effects on nares microbial diversity. Preoperative decolonization of S. aureus carriers with mupirocin decreases S. aureus surgical site infections when administered as a part of a bundle of preoperative interventions,72–74 and current guidelines recommend its use in select patients undergoing certain cardiac and orthopedic procedures. 75 However, the degree of sinonasal clearance of S. aureus by mupirocin and the further effects of mupirocin on the diversity of sinonasal microbiota have not been previously examined. Although the absence of S. aureus carriage by the subject treated with mupirocin precludes us from addressing these questions directly in this study, it may be that the clearance of S. aureus by mupirocin is similar to that observed for Moraxella, with clearance of 1 species followed by the expansion of niche-occupying commensal microbes to fill the void. Furthermore, the results obtained from this individual highlight the potential for iatrogenesis with mupirocin use, as our data suggest that its administration results in unintended consequences for the sinonasal microbiota. As with intranasal steroids, we must continue to use this medication in a thoughtful manner as we seek to further understand its on-target and off-target effects.
Studies to date of the sinonasal microbiota have been limited by their cross-sectional study design, which this study overcomes despite its limited sample size. Our findings warrant further study within a larger cohort, over a longer period of time, and with the addition of a crossover arm. Cross-sectional sinonasal microbiome studies to date are limited by large intersubject variation and the numerous confounding variables that may exist in human subjects with rhinosinusitis or atopic disease. To overcome this challenge, prospective longitudinal sampling was utilized in this pilot study and is recommended for future studies of the human microbiome in health and disease.76,77 In addition, mechanistic studies are required to better understand the observed effect on the sinonasal microbiota, as these results would likely have broader implications, given the widespread use of both topical and oral corticosteroids to treat myriad conditions, coupled with a lack of understanding of the effects on the microbiome at local and distant sites of administration.
Conclusion
INS are a mainstay of treatment for inflammatory sinonasal disorders, although multiple formulations exist whose mechanisms of action are poorly understood. Here, we demonstrate their potential effects on the sinonasal microbiome. Further characterization of these effects in larger cohorts with longer periods of observation is indicated.
Supplemental Material
Supplemental Figure - Supplemental material for Determinants of the Nasal Microbiome: Pilot Study of Effects of Intranasal Medication Use
Supplemental material, Supplemental Figure for Determinants of the Nasal Microbiome: Pilot Study of Effects of Intranasal Medication Use by Vijay R. Ramakrishnan, Justin Holt, Leah F. Nelson, Diana Ir, Charles E. Robertson and Daniel N. Frank in Allergy & Rhinologys
Footnotes
Author's Note
The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by the Department of Otolaryngology, University of Colorado, National Institute on Deafness and Other Communication Disorders of the National Institutes of Health under Award Number K23DC014747 (V.R.R.) and by Flight Attendants Medical Research Institute grant CIA13006 (D.N.F.).
Ethical Approval
This study was approved by our institutional review board.
Statement of Human and Animal Rights
This article contains research using human subjects.
Statement of Informed Consent
Informed consent was obtained from all subjects in this study.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
