Abstract
Purpose
Three-dimensional modeling and printing (3DMP) of anatomical structures from cross-sectional imaging data can enhance the understanding of spatial relationships in complex congenital heart defects. Partially due to the substantial financial, material and personnel resources required, 3DMP is not yet universally used. Here, we describe a workflow that addresses and eliminates these drawbacks.
Description
The workflow utilizes the open-source software “3D Slicer” (The Slicer Community) and “Blender” (Blender Foundation) for segmentation and post-editing of datasets. This approach enables the generation of virtual or physical 3D models. The physical models are printed using a standard fused deposition modeling printer.
Evaluation
The financial challenges that likely constrain the wider use of 3DMP are largely addressed by this approach. However, the workflow still requires a considerable amount of time to manually segment the imaging data.
Conclusions
Three-dimensional modeling and printing might improve planning and safety of congenital cardiac surgical treatment. Furthermore, it is a useful tool for education of parents and medical professionals. This workflow increases its suitability for routine use also in regions with low economic resources.
Keywords
Introduction
Three-dimensional modeling and printing (3DMP) can be used in addition to routine imaging to better visualize the anatomic and spatial relationships in complex congenital heart malformations. 1 Three-dimensional modeling and printing aims to transfer two-dimensional cross-sectional images into three-dimensional bodies. These bodies can be expressed as physically printed models made from various materials or as virtual bodies that can be visualized or even manipulated using appropriate hardware and software.
Three-dimensional modeling and printing has been shown to be useful for improving preoperative planning. 2 Furthermore, 3DMP can be successfully used in the perioperative setting and for teaching and training purposes.3,4 Despite its obvious appeal, 3DMP has not entered the clinical diagnostic routine workup in most centers worldwide. 5 This is probably due to several factors: 3DMP is a time-consuming process that requires personnel with specific expertise. Additionally, the costs for appropriate hardware and software are usually high. Contrarily, there are typically no planned resources or funding for 3DMP, whether in hospital environments or insurance reimbursement plans. 6 In order to overcome those obstacles and to increase the feasibility and availability of 3DMP, we aimed to design a workflow which is based exclusively on the use of open-source software and low-cost (consumer) hardware. Using this workflow, virtual and physical 3D-models can be generated.
Description
The different steps of the workflow are shown in Figure 1. In the following figures we show an example of a child with complex transposition of the great arteries (TGA) with ventricular septal defect, left-ventricular outflow tract obstruction, and pulmonary atresia who underwent neonatal palliation with a modified Blalock-Taussig-Thomas shunt. Three-dimensional modeling and printing was conducted in order to evaluate the options for biventricular repair.
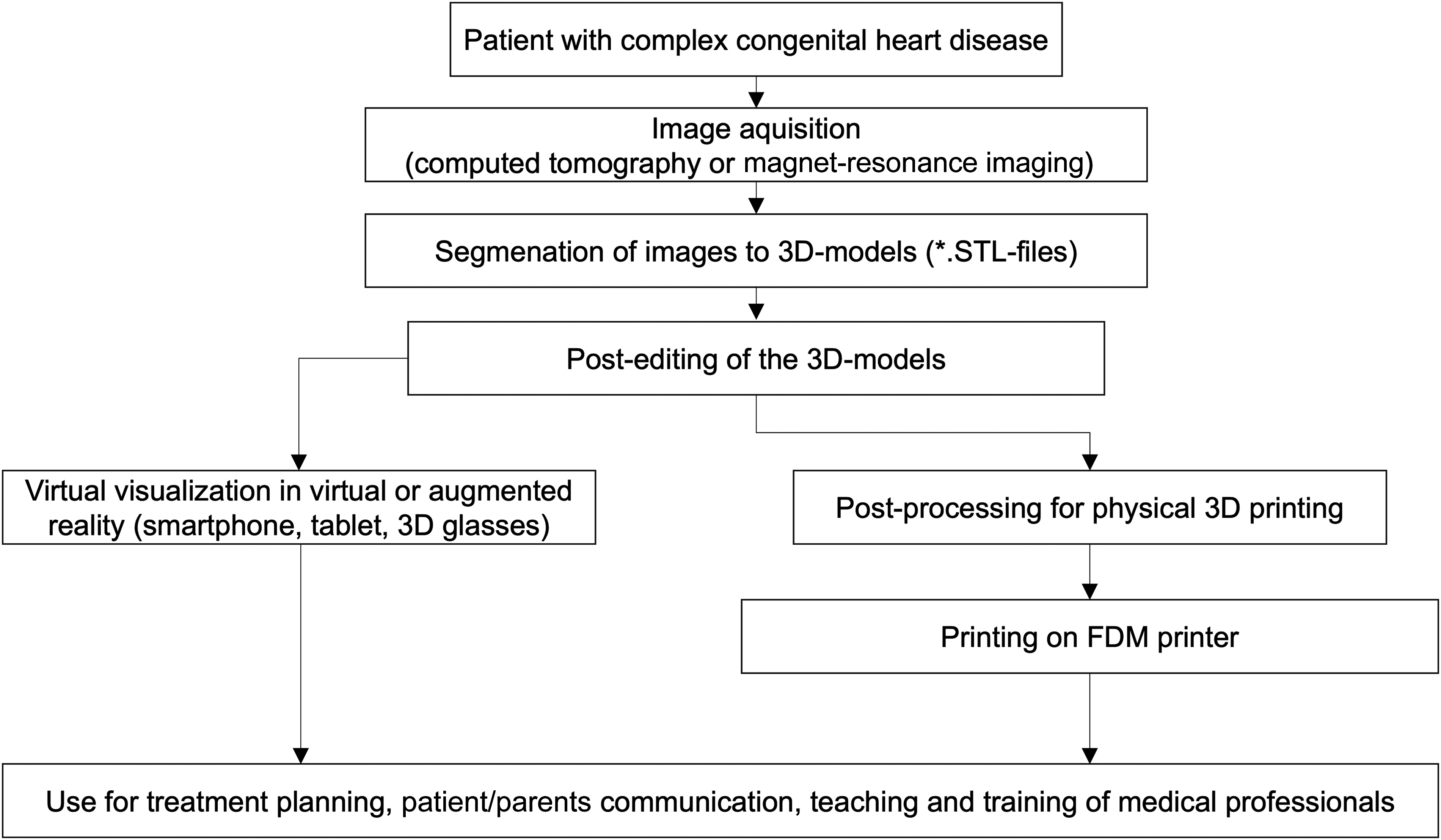
Steps of the workflow to generate and use physical and virtual 3D-models of hearts with congenital malformation. 3D, three-dimensional; FDM, fused deposition modeling; STL, stereolithography.
Image Acquisition
Every cross-sectional image can be potentially used for 3D-reconstruction, although computed tomography (CT) is generally preferred due to its high spatial resolution. 7 In our example, contrast-enhanced images were acquired within the preoperative routine imaging with the layer thickness set at 0.6 mm.
Segmentation
The process of creating a three-dimensional reconstruction of a region of interest (ROI), known as “segmentation,” was done using a “3D Slicer” by “The Slicer Community.” It is a free open-source software that allows viewing of DICOM (Digital Imaging and Communications in Medicine) datasets and, among many other capabilities, the creation of three-dimensional models out of two-dimensional cross-sectional imaging. Along with our aim to create a workflow exclusively based on open-source tools and low-cost hardware, we chose a “3D-slicer” because of its comparable precision in terms of segmentation accuracy to the commonly used cost-intensive “Materialise Mimics” (Materialise NV).8,9
Among the available series of a computed tomography (CT) study, the one with the highest spatial resolution in the X, Y, and Z levels that also subjectively provides a good representation of the ROI is selected for further processing. An area such as the myocardium, the vascular wall, a valve leaflet, or the blood pool is selected in a layer of the dataset (Figure 2A-C). If this process is repeated for multiple consecutive layers, a three-dimensional model of the selected areas can be reconstructed.
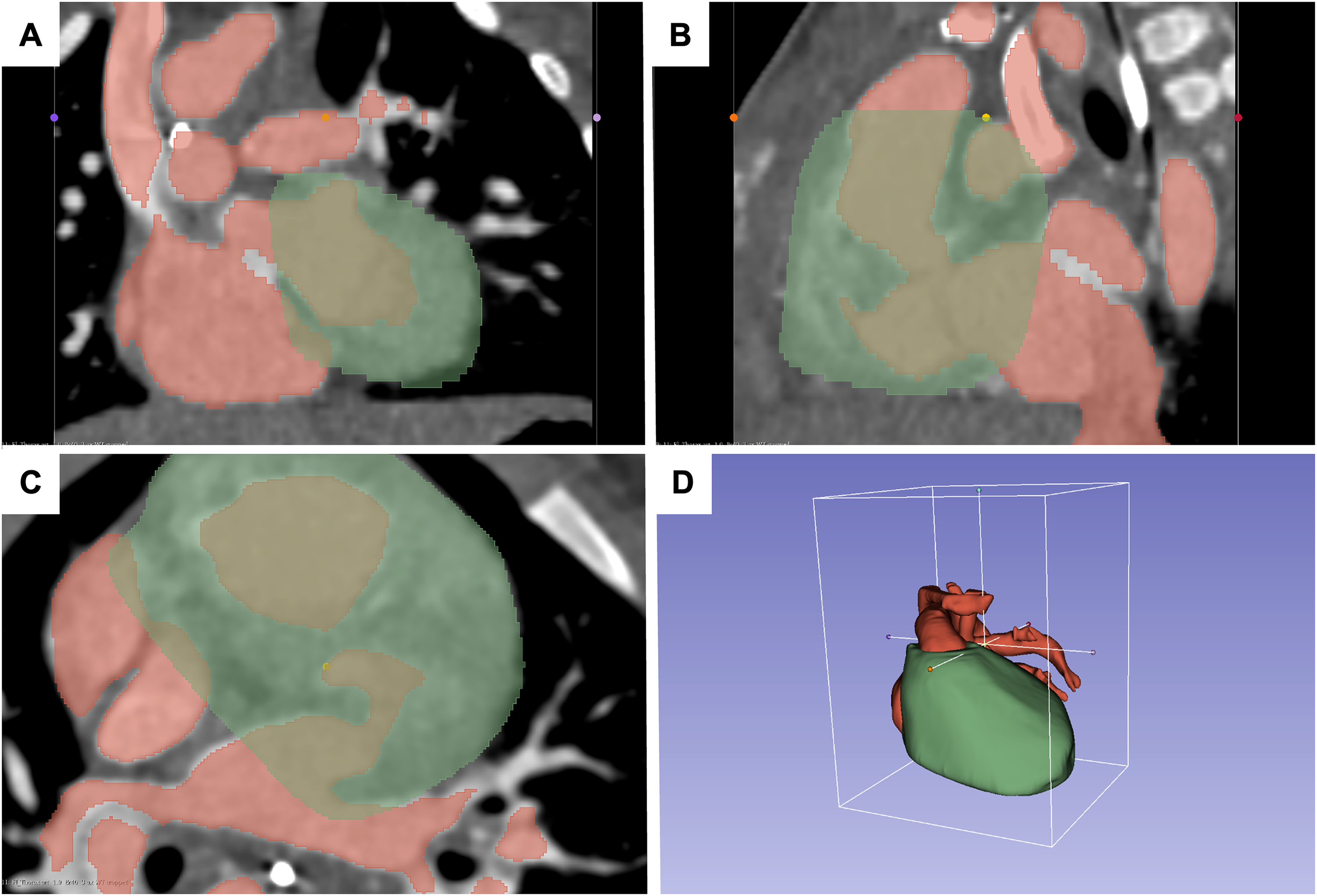
Segmentation of cardiac structures: (A-C) The contrast-enhanced regions of the image are marked (red), and the myocardium is marked (green). (D) This results in a three-dimensional body with segmented red (contrast-enhanced volumes) and green (myocardium) volumes.
While the use of a “3D-Slicer” does not require extensive software-skills, this selection can be time-consuming, when it has to be done manually, layer by layer, which sometimes can be necessary when the ROI is complex (eg, small vessels with incomplete filling of the contrast agent) or when the image quality is challenging. When the image quality is sufficient or the ROI is well-structured (eg, myocardium and contrast-enhanced blood pool within the ventricles), a variety of semiautomatic segmentation tools can speed up this process significantly.
We found the options “threshold,” “grow from seeds,” and “surface cut” useful for our purposes. Once the segmentations are complete, their surfaces can be smoothed using the “smoothing” option (Figure 2D) if necessary, and the segments can then be exported as .stl files.
Post-Editing of the 3D-Models
Further modification is often necessary before display and printing. This includes, among others, removing unwanted structures, creating vascular lumen, smoothing surfaces, and thickening (vessel) walls. “Blender” (Blender foundation) is an open-source software that provides efficient tools while offering a user-friendly interface. To modify the configuration of the .stl file, the options “edit mode,” “object mode,” and “sculpt mode” are useful (Figure 3).
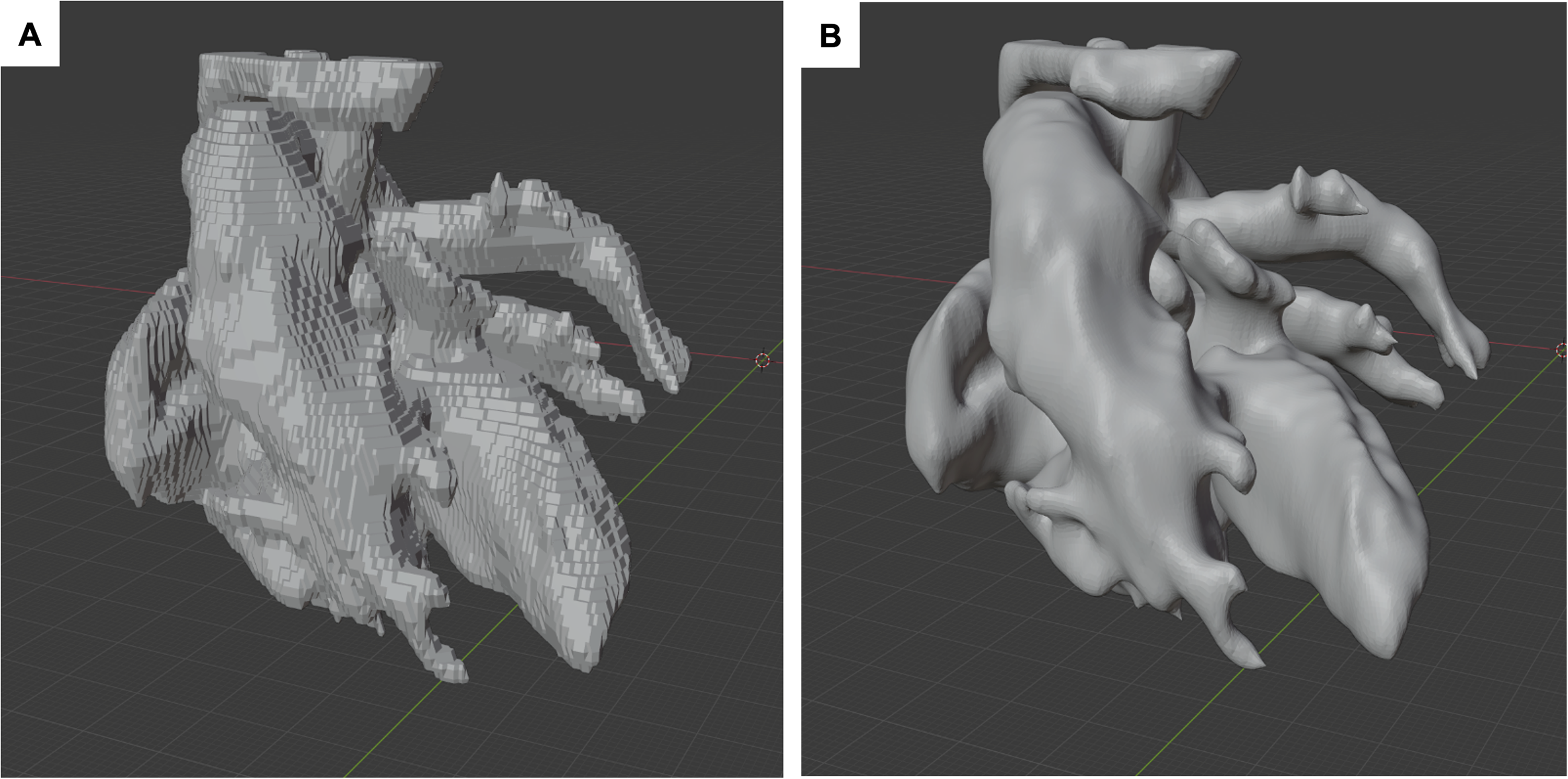
Blood-filled segments of the .stl file before (A) and after (B) smoothening using the “Blender” software.
The completed models can now be sent to a 3D printer or displayed in virtual reality, augmented reality, or mixed reality settings. A convenient way to view and share the 3D models is by using an online viewer such as p3d.in (Brand3D), which allows displaying the models in augmented or virtual reality regardless of the mobile device used.
Processing of Three-Dimensional Datasets to Physical Printouts
Since the regions of interest often lie within the heart (eg, septum, ventricular outflows) we tend to strategically split the model in several parts using “Blender,” allowing a good view on selected structures (Figure 4). To assemble the physical model afterwards, we add connecting mechanisms and then export the parts as .stl files.
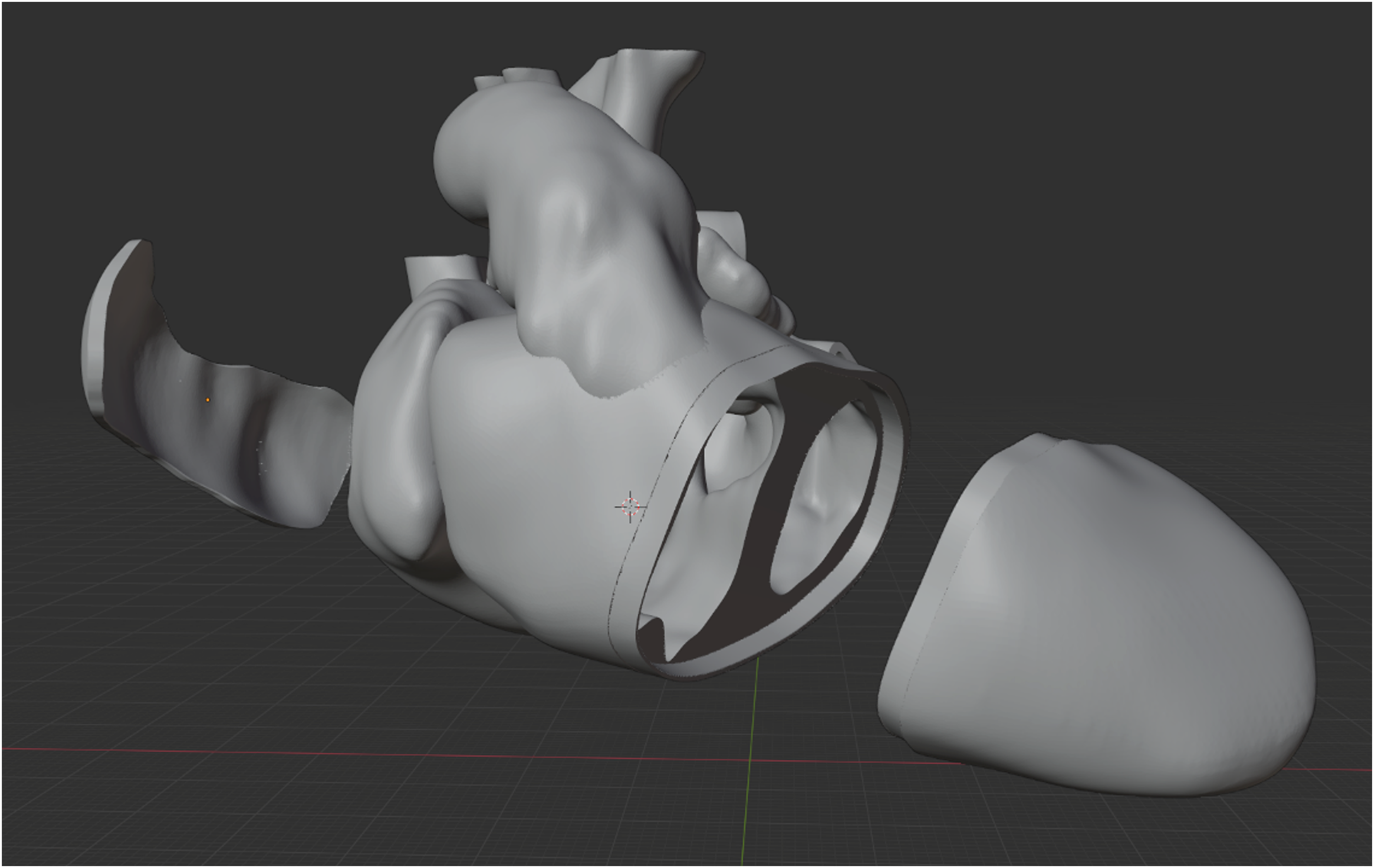

Model with the ventricular mass split transversely in order to achieve a better view of the ventricular septal defect and its spatial relation to the outflow tracts. Additionally, the left atrial roof and right atrial free wall are excised in order to enable transatrial inspection of the ventricular septal defect.
To be processed by a 3D printer, the files need to undergo a procedure called “slicing,” where the model is split into horizontal layers according to the print resolution, and supporting structures are added (Figure 5). The sliced model is then exported as a .gcode file that can be processed by the printer. We found the freeware “Cura” (Ultimaker B.V.) to be an easy-to-use and effective tool for this process. All the aforementioned steps were executed on a commercially available gaming notebook (Lenovo Thinkbook 16p G2, Lenovo; costs: 1.500US4). Sufficient computing power speeds up the process but is not a prerequisite for running the programs mentioned.
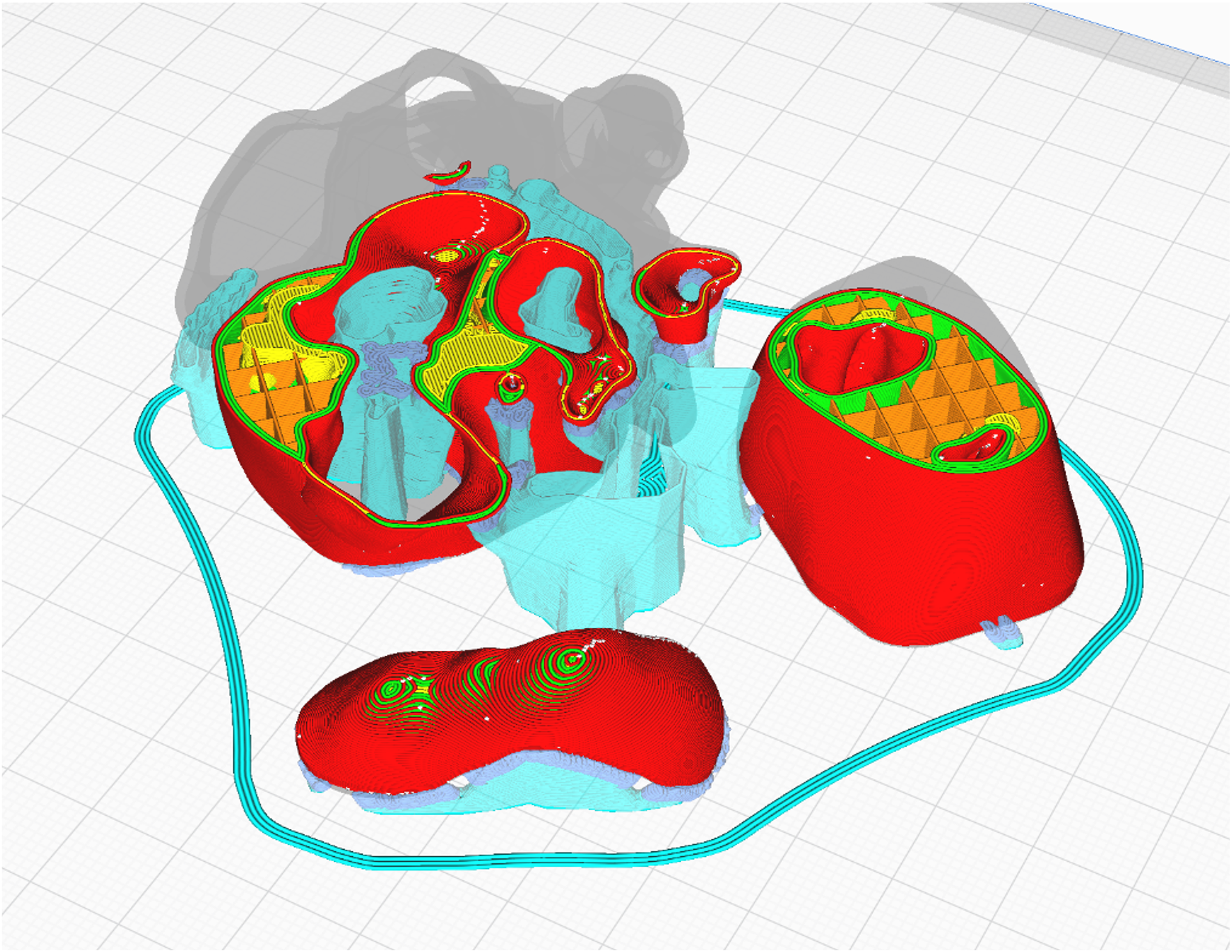
Model after slicing. The parts of the model have been laid out horizontally and have been provided with external (turquoise) and internal (orange) supporting structures to maintain stability of the printed parts during and after the printing process.
We chose a common fused deposition modeling printer from the low-cost segment (i3 Mega, Hongkong Anycubic Technology Co. Limited) with a build volume of 210 × 210 × 205mm for our printouts. Setting the z-resolution, or layer height, to 0.15 mm yields prints of good quality with reasonable printing times. Since we do not require specific physical durability for our models, they do not need to be printed solidly, allowing us to decrease the internal filling to 10%, thereby saving on printing time and material consumption.
As material for rigid 3D models, we chose white polylactic acid, a bioplastic made from organic waste, which is an eco-friendly and less harmful alternative to other printing materials such as acrylonitrile butadiene styrene. 10 White has proven to be a good choice of color, providing good visibility of small and complex anatomical structures (Figure 6).
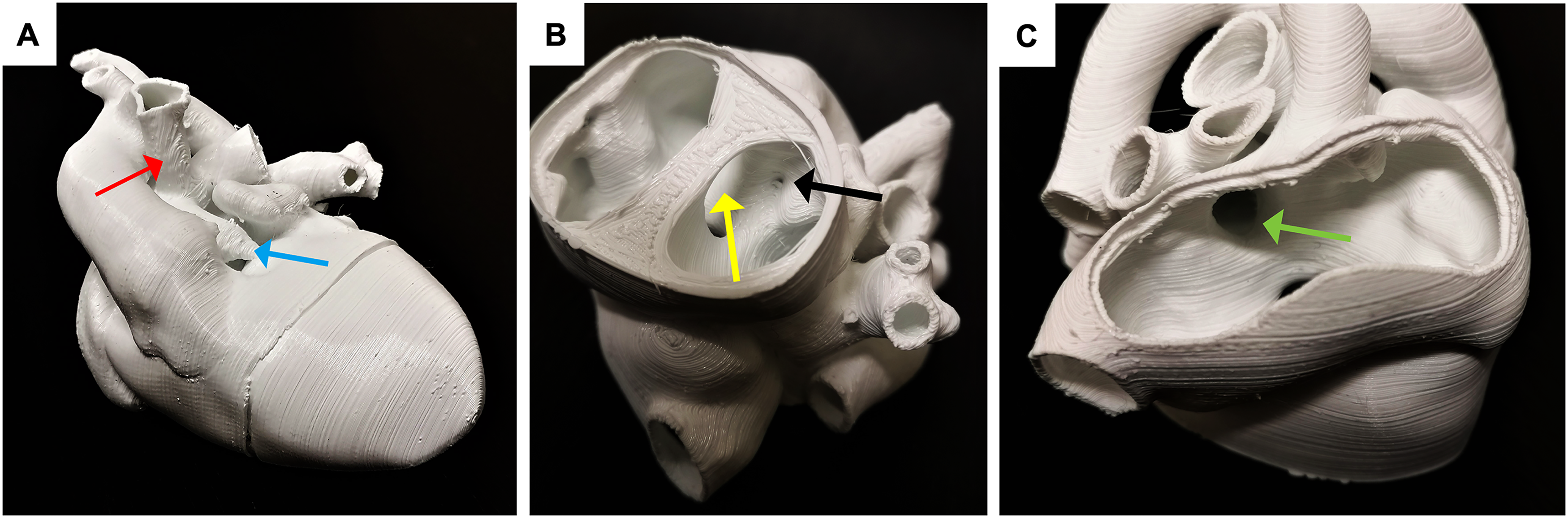
The printed model allows for inspection of the different structures of interest: (A) Hypoplastic pulmonary artery (blue arrow) and the systemic-pulmonary artery shunt (red arrow). (B and C) Frontal view with an axial cut through the ventricles showing the ventricular septa defect (yellow arrow) and the stenotic outflow to the pulmonary artery (black arrow). (C) Dorsal view through the left atrium showing the atrial septal defect (green arrow).
These technologies also enable the production of manipulable 3D models for surgical simulations, although this process is considerably more time-consuming. It involves creating rigid molds of both the outer and inner surfaces of the heart, which are then filled with silicone to produce a soft, flexible model. This method is a cost-effective alternative to directly printing flexible materials, which requires more expensive printers and materials. For example, such printers can cost between $200 000 and $300 000, with each heart model costing approximately $200 to $300. 11
Depending on the required quality of the reconstruction, the quality of the imaging data, the complexity of the pathology, and the required post-processing, the segmentation of one patient specific model using this workflow involves about 1 to 2 h of modeling for an experienced user and 3 to 5 h of printing time. To our best knowledge, this is not more time-consuming than with the use of commercial software. A high-quality print of a model like the one depicted in Figure 6, measuring about 7 × 7 × 6cm and weighing 35 g, takes about 4 h and 30 min to print and has material costs of approximately US$0.70.
Evaluation
The described workflow enables the creation of high-quality anatomical 3D virtual models and prints. The approach emphasizes the use of low-cost and open-source hardware and software, respectively. With this approach, high-quality physical and virtual models of patient-specific anatomies can be produced. The costs for material, software, and hardware use are minimal. However, personnel and organizational challenges persist with this approach. At our institution, the 3DMP process is managed by a pregraduate medical student employed as a student assistant, in collaboration with a congenital cardiac surgeon and a cardiac imager. The surgeon provides the student with guidance on the patient's diagnosis, anatomy, medical history, and specific aspects relevant to the 3D model, such as key areas of interest for the planned surgeries. The medical student carries out the entire process, creating the 3D model with assistance from the cardiac imager when needed. The surgeon and student then review the model together and discuss any necessary modifications. Once finalized, the model is shared with the entire team, either as a printed or virtual version, for further use.
Our experience shows that this organizational approach, where a student assistant operates under close supervision, ensures the production of high-quality 3D models while effectively utilizing personnel and delegating time-consuming tasks away from clinical staff. This makes 3DMP feasible in centers without dedicated reimbursement plans for 3DMPG, like ours, and accessible to a broader community. Alternatively, training dedicated imaging specialists in 3DMP could enhance the quality and sustainability of the process. However, this would require additional resources and reimbursement to support the training and maintenance of such specialized personnel. Taking into consideration that operative planning with a patient-specific 3D-model before an operation led to a significant change in the procedural planning in 47.5% of cases in a study by Valverde et al, and the time needed for training in this technology and 3D-reconstruction in complex cases could be well invested. 2
Another alternative sometimes favored by hospital administrations is outsourcing 3DMP to commercial medical printing companies, which may reduce costs. However, the direct interaction between the clinician and the individual creating the 3D model is often crucial, particularly in cases involving highly complex anatomies or when the quality of the source images is reduced. This interaction can be compromised when external printing companies are involved. Since the segmentation and editing processes still require significant human effort and time, advancements in automated segmentation could further enhance the feasibility of 3DMP in the future. Automatized processes have been described for segmentation of the left ventricle and the aorta of structurally normal hearts. 12 However, the structural complexity and diversity of congenitally malformed hearts have precluded automatized segmentation for congenital heart defects (CHD) to date. The role of deep learning algorithms for automated segmentation of CHD imaging is currently being investigated in experimental settings but its reliability for segmentation of complex CHD has not been developed sufficiently until now. 13
Conclusion
Three-dimensional modeling and printing helps to better understand the anatomical and spatial properties of patients with complex CHD. The production of physical or virtual models preoperatively can improve planning and enhance the safety of congenital cardiac surgical treatments. Additionally, these 3D models can be used for counseling parents and for teaching purposes in medical education. The workflow presented here is easily reproducible and affordable, increasing the suitability of 3DMP for routine use even in regions with limited economic resources.
Footnotes
Abbreviations
Authors' Statement
The parents (legal guardians) of the child whose imaging is shown in this report, gave consent to publication of the image data and the description of the patient's history.
Acknowledgments
The authors thank the Federal state of Baden-Wuerttemberg and the “Studierendenrat” (students’ council) of Heidelberg University for funding the 3D-printing project.
Declarations of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The implementation of the workflow was conducted with funding by the federal state of Baden-Wuerttemberg and the “Studierendenrat” at Heidelberg University Hospital, Heidelberg, Germany.
