Abstract
Objective:
To assess the probability of finding at least one euploid embryo (POE) and euploidy rate in preimplantation genetic screening (PGS) cycles for aneuploidy testing with fluorescent in situ hybridization (FISH) and array comparative genomic hybridization (aCGH) based on the number of retrieved oocytes after adjustment for age.
Design:
Retrospective data analysis.
Setting:
Private in vitro fertilization center in a general hospital, Istanbul, Turkey.
Patient(s):
In this study, 1412 couples with 1739 cycles were included.
Intervention(s):
Embryo biopsy at cleavage stage or trophectoderm biopsy analyzed by FISH or aCGH techniques.
Main outcome measure(s):
Probability of finding at least one euploid embryo for transfer and euploidy rate according to retrieved oocytes after adjustment for age.
Result(s):
Estimated probabilities of finding at least one euploid embryo were plotted based on age and number of retrieved oocytes. When controlled for age, the POE increased with each additional oocyte retrieved (OR 1.11, 95% CI 1.08–1.14), but the euploidy rate did not. When retrieved oocytes were subgrouped, we found that the odds of finding at least one euploidy embryo were highest when 16–20 oocytes were retrieved compared to 1–5 oocytes with both FISH (OR 8.62, 95% CI 4.67–15.93) and aCGH (OR 2.78, 95% CI 1.07–7.21). However, the chances did not increase with the retrieval of more than 20 oocytes.
Conclusion(s):
The estimated probabilities of finding at least one transferrable embryo would enable more accurate counselling of patients based on age and the number of retrieved oocytes. As the highest estimates for finding one euploidy embryo were achieved with 15-20 oocytes, deciding the best protocol for ovarian stimulation to achieve the maximum number of oocytes for retrieval based on the patients age and ovarian reserve would be of crucial importance.
Keywords
Introduction
The ultimate goal of in vitro fertilization (IVF) coupled with preimplantation genetic screening (PGS) for aneuploidy testing is to select at least one euploid embryo for transfer in order to achieve a healthy live birth. Unfortunately, not every couple undergoing PGS can benefit equally from the technique because some patient-specific factors, such as female age and ovarian reserve, may have an effect on the outcome of PGS. Even without additional risk factors, the number of embryos available for transfer is significantly lower in PGS than in regular IVF, increasing the risk of cancellation of transfer simply because of embryo selection in order to eliminate aneuploid embryos. Therefore, finding at least one transferable embryo is the main determinant of clinical success in PGS.
The impact of age on euploidy rate and the outcome of PGS is demonstrated by many studies (Ata et al., 2012; Fragouli et al., 2011; Hellani et al., 2008; Márquez et al., 2000; Munné et al.,1995; Schoolcraft et al., 2010; Verpoest et al., 2009). However, only a few studies have investigated the influence of the number of retrieved oocytes on the outcome of PGS (Santaló et al., 2000; Tur-Kaspa, 2012; Vandervorst et al., 1998). Further information on the probability of finding at least one euploid embryo (POE) according to the ovarian reserve after adjustment for age is needed for improved guidance for patients undergoing PGS. In the present study, we aimed at assessing the probability of finding at least one euploid embryo in PGS cycles for aneuploidy testing and to assess the euploidy rate with fluorescent in situ hybridization (FISH) and array comparative genomic hybridization (aCGH) in order to counsel patients appropriately based on their ovarian reserve and age. In addition, we aimed to determine the optimal number of oocytes which predicts the highest POE with FISH and aCGH after adjusting for age.
Materials and methods
We retrospectively analyzed intracytoplasmic sperm injection cycles where PGS was performed for aneuploidy using either FISH or aCGH in Istanbul Memorial Hospital IVF Center over a 10-year period through years 2003 and 2014. A total of 1739 cycles, 1420 couples with FISH and 319 couples with aCGH, were analyzed from AURA database prepared by our unit.
Institutional review board and ethical committee approval was not considered necessary for this study, as it was a retrospective study involving data analysis and clinical outcomes of well-established treatment cycles carried out in accordance with the IVF guidelines and genetic analysis protocols. Patients were informed about the treatment and procedures and written informed consent was obtained from all patients before starting IVF treatment, embryo biopsy, and genetic analysis and transfer procedures. Patients were informed about the possibility of misdiagnosis and the cancellation of embryo transfer in the absence of euploid embryos.
FISH technique was used until 2011 and 9–11 chromosomes were evaluated. Since 2011, aCGH technology has been used for aneuploidy testing of all chromosomes. Currently aCGH is the preferred technique over FISH, as all the chromosomes can be evaluated accurately. We evaluated the results of 10,456 embryos biopsied and analyzed by FISH and aCGH. Of these embryos, 9203 were evaluated with FISH and 1253 embryos with aCGH. All of the embryos analyzed with FISH were cleavage stage embryos, whereas 520 and 733 embryos analyzed with aCGH were cleavage and blastocyst stage embryos, respectively (Table 1).
Patient and cycle characteristics and resultant euploidy rates according to biopsy day and technique.
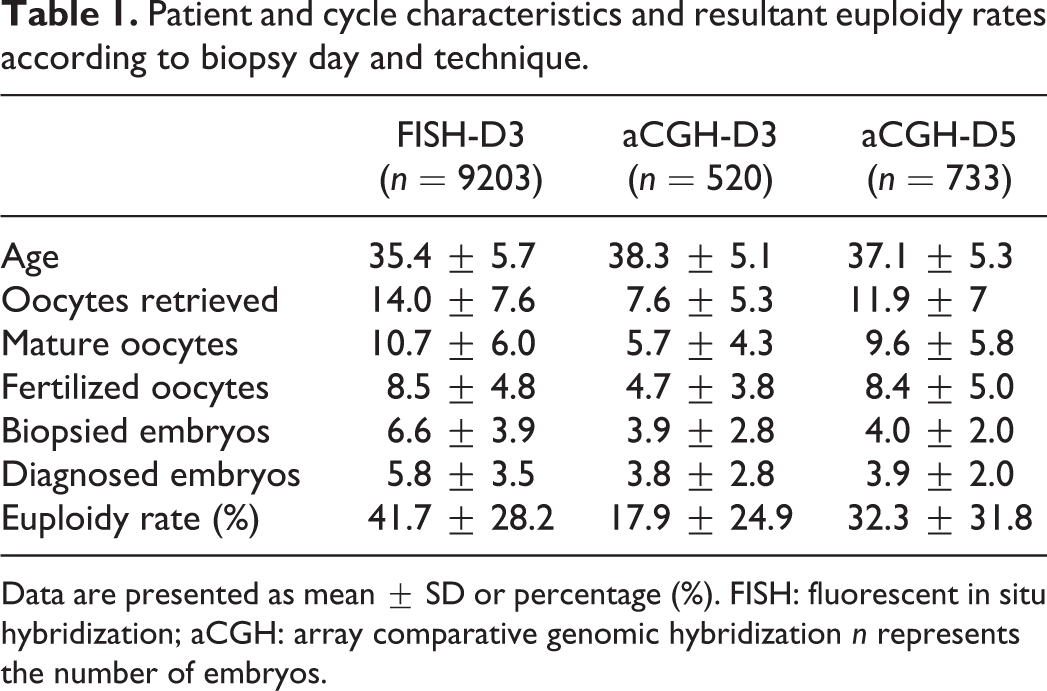
Data are presented as mean ± SD or percentage (%). FISH: fluorescent in situ hybridization; aCGH: array comparative genomic hybridization n represents the number of embryos.
Inclusion criteria
The study cohort included women who underwent PGS for aneuploidy testing with either FISH or aCGH techniques for advanced maternal age, repeated implantation failure, recurrent early spontaneous miscarriages, or a history of abnormal fetal karyotype. Couples having abnormal karyotypes, Y-chromosome microdeletions, and PGS cycles performed for single gene disorders and/or human leukocyte antigen (HLA) typing were not included.
Ovarian stimulation and embryo biopsy
The stimulation protocols have been outlined previously (Kahraman et al., 2011). For ovarian stimulation, gonadotropin-releasing hormone (GnRH
Fluorescent in situ hybridization
FISH technique consisted of the following steps: fixation of biopsied cells, pretreatment and probe application, denaturation and hybridization, stringent wash, and analysis after counterstain application.
Individual blastomeres or trophectoderm cells were fixed with Carnoy’s fixative, as described previously elsewhere(Munné et al.,1996). In the first round of hybridization five chromosomes (13, 16, 18, 21, and 22) and in the second round of hybridization four chromosomes (15, 17, X, and Y) were screened. After the first and second rounds of hybridization, in case of any doubt regarding a chromosome, the nucleus was reevaluated with the use of a new probe set using telomeric- or locus-specific probes specific for that chromosome. In the first round, multivysion PB probe panel (LSI 13, 13q14.1–q14.3, RB1, spectrum red; CEP 16, 16q11.2, D16Z3, spectrum aqua; CEP 18, 18p.11.1–q11.1, D18Z1, spectrum blue; LSI 21, 21q22.13–q22.2, D21S341, D21S342, D21S339, EGR, D21S338, spectrum green; LSI 22, 22q11.2, BCR, spectrum gold) (Vysis, Abbott Inc., Germany) and for the second round 4CC custom probe panel (CEP X, Xp11.1–q11.1, DXZ1, spectrum green; CEP Y, Yq12, DYZ1, spectrum blue; CEP 15, 15p11.1–q11.1, D15Z4, spectrum orange; CEP 17, 17p11.1–q11.1, D17Z1, spectrum aqua) (Vysis, Abbott Inc., Germany) were used. Simultaneous denaturation and hybridizations took place in a humidified chamber (Hybrite, Vysis, Abbott, Germany) and were carried out as described previously by Kahraman and Beyazyurek (2014). The images were stored and processed by image processing software (Isis, Metasystems, Inc. accessed at http://www.metasystems-international.com/isis).
Array CGH
The biopsied samples were placed in microcentrifuge tubes containing 2.5 μl of sterile phosphate-buffered saline (PBS) solution. The contents were centrifuged and spun down. They were either processed immediately or stored at −20°C until they were processed.
The samples were tested using 24Sure™ Cytochip kits (BlueGnome Ltd., Cambridge, UK). In brief, the protocol consisted of the following steps: cell lysis and extraction of DNA, whole-genome amplification (WGA) using degenerate oligonucleotides, fluorescent labeling of samples and reference DNA, combination of differentially labeled products, hybridization onto microarray slides, posthybridization washing, scanning, and analysis of images using the dedicated software. Lysis and WGA were achieved using sureplex amplification kit (BlueGnome, Ltd.) in a vertical laminar flow cabinet according to the manufacturer’s instructions.
Amplified samples and reference DNAs were labeled with Cy3 and Cy5 fluorophores using random primers (BlueGnome Ltd.). An Excel sheet (24 sure V3 planner.xls) was used to divide the two groups that weree labeled differentially. For each different lot of arrays, two sex-matched reference subarrays were included and reference subarrays were located on different slides. The labeling reaction took place in a preheated thermal cycler for 2–16 h at 37°C.
After labeling reaction, Cy3 and Cy5 labeled samples and references were combined with the addition of 25 ml commercially available human COT DNA (BlueGnome Ltd., UK). The volume was reduced to around 3 ml in each tube with the help of a centrifugal evaporator (Lab Conco, Missouri, USA). The labeled DNA were resuspended in dextran sulfate hybridization buffer and denatured at 75°C for 10 min. Eighteen milliliter of this solution was applied to each subarray and hybridized under coverslips in a water bath adjusted at 47°C for 3–16 h. Coverslips were removed gently. High-temperature stringent wash protocol was used to remove unbound DNA (Hybex; Sci Gene, Sunnyvale, CA, USA). A magnetic stirrer was used to continuously mix the wash solution (Velp Scientifika, Italy) to help clear it of the hybridization remnants.
The slides were dried and scanned using a laser scanner (InnoScan, 710; Innopsys, Carbonne, France) and a program (Mapix, Carbonne, France) to store and process the images. The resulting images were stored in tagged image file format and were analyzed by BlueFuse Multi software (BlueGnome Ltd.). Each sample was compared according to both sex references, and gains or losses of individual chromosomes were assessed with the use of 3× SD measurements.
Outcome measures and statistical analysis
As our primary outcome was to assess the POE based on age and number of retrieved oocytes, we first find the clinical determinants affecting the POE and the euploidy rate. The associations between the covariates/predictors (e.g. age, number of retrieved oocytes, body mass index (BMI), gonadotrophin dose, day of transfer) and the outcomes (e.g. euploidy rate and euploidy status) were modeled using linear and logistic binary regression analyses depending on the outcome measures tested. For example, the euploidy rate defined as the rate of POEs per diagnosed embryos was calculated per cycle, and this value has been treated as a continuous variable, whereas the euploidy status of a cycle (the presence of at least one POE) is coded as a binary (1 or 0) outcome measure.
The subsequent cycle of each woman was treated as unique, since each cycle was performed at different time points, which may have an effect on the reproductive aging of ovaries, affecting ovarian reserve and the euploidy rates of oocytes.
Univariate analyses were performed to evaluate the magnitude of effect on euploidy rate and the POE for each covariate. Multivariate analyses were then performed to adjust for all variables that were statistically significantly associated with the POE and euploidy rate to determine the significant predictors after adjustment. Using these models, the graphs were plotted for the POE stratified for age.
As a secondary outcome, we aimed to determine the optimal number of oocytes which predicts the best POE with FISH and aCGH after adjustment for age. For this analysis, we divided the data into five groups according to the number of oocytes retrieved, with group 1 consisting of those with 5 or fewer oocytes, group 2 with 6–10 oocytes, group 3 with 11–15 oocytes, group 4 with 16–20 oocytes, and group 5 with more than 20 oocytes. These subgroups were then introduced into a multivariate logistic regression model which took account of adjustment for age. Each subgroup is compared to the reference category determined as 1–5 retrieved oocytes. The mean probabilities for each oocyte group obtained from this model are shown in a line graph. Analysis was performed by IBM SPSS statistics 20 and graphs were made by Microsoft Excel.
Results
FISH yielded results in 87.4% (8043/9199) of the biopsied embryos. The aCGH yielded results in 96.9% (504/520) of the cleavage and 98.6% (723/733) of the blastocyst stage embryos biopsied. Table 1 shows patient and cycle characteristics according to the biopsy day and technique. The distribution of oocytes and biopsied embryos across all ages is presented in scattered plot graphs for FISH and aCGH (Figure 1). The mean number of retrieved oocytes, biopsied embryos, and resulting euploid embryo numbers are presented in bar graphs for FISH and aCGH (Figure 2).
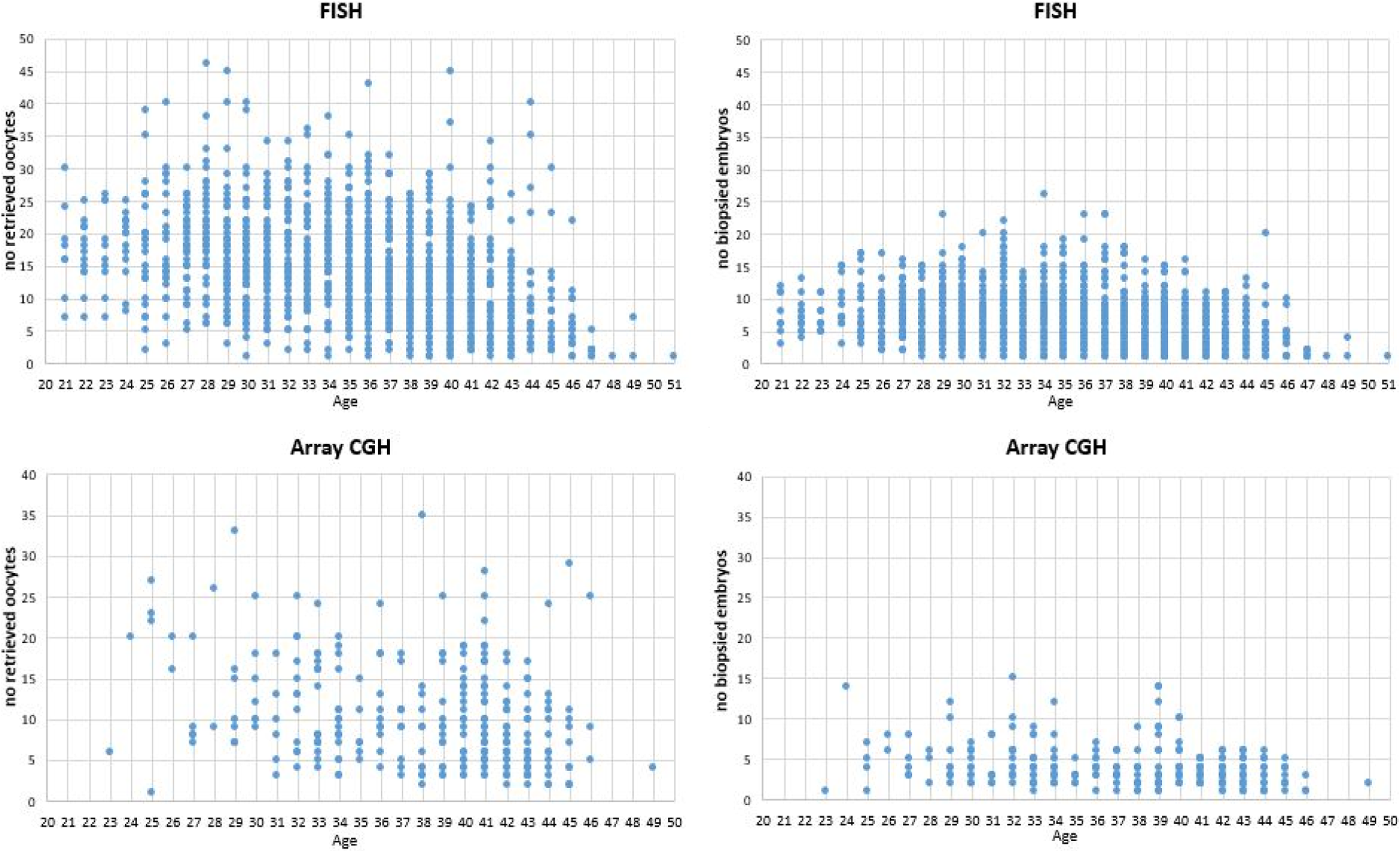
Distribution of oocytes and biopsied embryos across all ages.
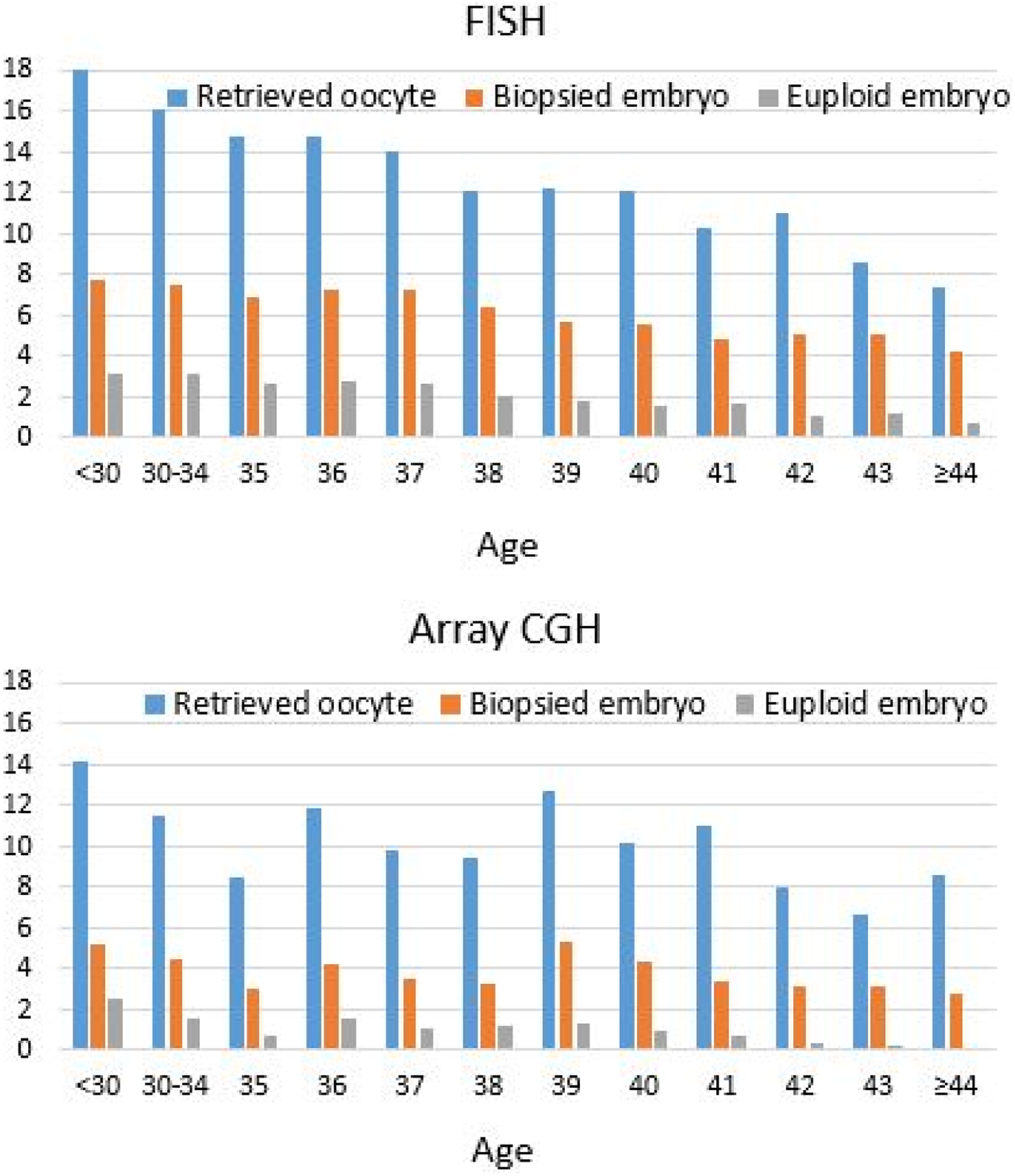
Bar graph showing the distribution of oocytes, biopsied embryos, and resulting euploid embryos in relation to age.
Clinical determinants affecting the POE and euploidy rate
The POE
Univariate logistic regression analysis revealed significant negative correlation with the POE and age and positive correlation with the number of retrieved oocytes. When all of these parameters were entered into the model of multivariate logistic regression, only the age, number of retrieved oocytes, and PGS technique were significant predictors of success in finding one euploid embryo for transfer (Table 2). Among these variables, age was the most significant predictor. When the data were subdivided into FISH and aCGH, the significant predictors for POE were again age and the number of retrieved oocytes for both groups as shown in Table 2. The aCGH data included both day 3 and day 5 biopsies, and the introduction of the biopsy day did not change the estimates.
Multivariate logistic regression analysis showing the clinical determinants of the POFALOEE and linear regression analysis showing the clinical determinants of the euploidy rate for FISH and aCGH.
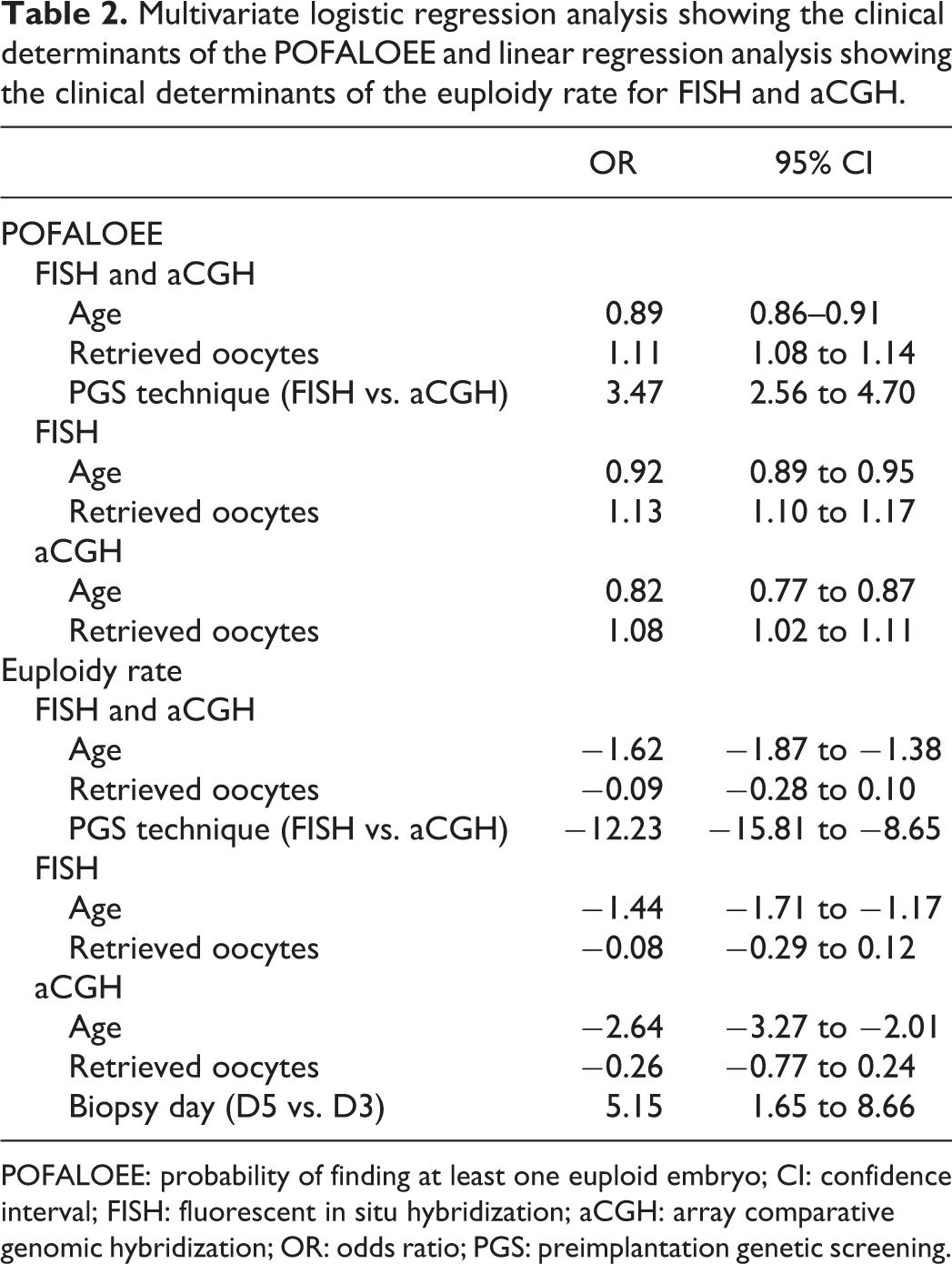
POFALOEE: probability of finding at least one euploid embryo; CI: confidence interval; FISH: fluorescent in situ hybridization; aCGH: array comparative genomic hybridization; OR: odds ratio; PGS: preimplantation genetic screening.
According to the multivariate logistic regression analysis, the odds of having at least one euploid embryo significantly decreased with increasing female age for FISH (odds ratio (OR) 0.92, 95% CI 0.86–0.95) and aCGH (OR 0.82, 95% CI 0.77–0.87), while the odds of having at least one euploid embryo significantly increased with every additional oocyte retrieved for both FISH (OR 1.13, 95% CI 1.10–1.17) and aCGH (OR 1.08, 95% CI 1.02–1.11).
The proportion of cycles with at least one euploid embryo stratified for age is presented in graphs for both FISH and aCGH (Figure 1).
Euploidy rate
Univariate linear regression analysis for the euploidy rate revealed significant negative correlation with age and positive correlation with the number of retrieved oocytes. The PGS technique had a significant effect on the euploidy rate. However, BMI and mean daily gonadotrophin dose did not significantly affect the euploidy rate. With the introduction of age and the number of retrieved oocytes together into the model, age was significantly associated with the euploidy rate, whereas the number of available oocytes was not for either FISH or aCGH. The aCGH data included both day3 and day5 biopsies, and entering biopsy day into the model showed that the biopsy day is a significant predictor of euploidy rate in addition to age (Table 2).
The euploidy rate with FISH and aCGH stratified for age is presented in Figure 3. There was a significant linear decreasing trend for euploidy rate across all ages with both FISH and aCGH. In young patients of less than 35 years, over 52% and 60% of the embryos diagnosed with FISH and aCGH were aneuploid, whereas this rate increased to 82% and 100% for patients over 44 years old, respectively (Figure 3). According to our data, FISH is able to detect 87% of chromosomally abnormal embryos detectable by aCGH in patients at or under 40 years of age. However, in patients over 40 years old, FISH detects 82% of abnormal embryos that are detectable by aCGH (Figure 3).
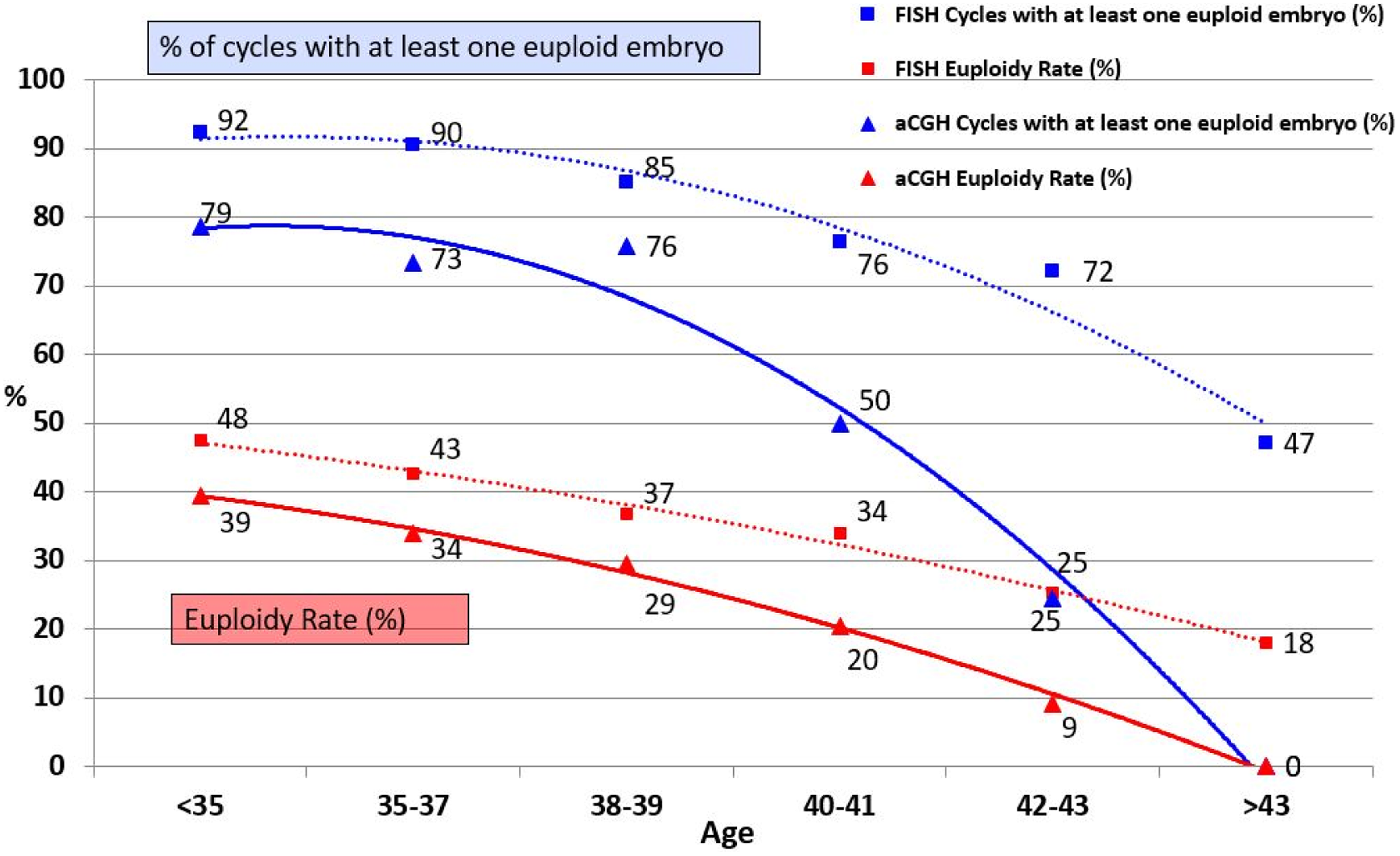
Euploidy rate and proportion of cycles with at least one euploid embryo across age-groups with fluorescent in situ hybridization (FISH) and array comparative genomic hybridization (aCGH).
According to the linear regression analysis, the number of available oocytes was not significantly associated with the euploidy rate with FISH (B −0.084%, 95% CI −0.3 to 0.1, p = 0.423) or aCGH (B −0.545%, 95% CI −1.23 to 0.4, p = 0.118) after adjusting for age. However, for every 1 year increase in female age, the euploidy rate decreased by 1.4% for FISH (95% CI −1.7 to −1.16, p < 0.001) and 2.6% for aCGH (95% CI −3.4 to −1.8, p < 0.001) after adjustment.
POE according to age and number of retrieved oocytes
From the regression models, the probability of finding at least one euploid embryo (y-axis) as a function of age (x-axis) was derived for FISH (Figure 4(a)) and aCGH (Figure 4(b)), with the number of retrieved oocytes fixed at certain values. Specifically, we presented plots to give the POE for 1, 5, 10, and 15 retrieved oocytes.
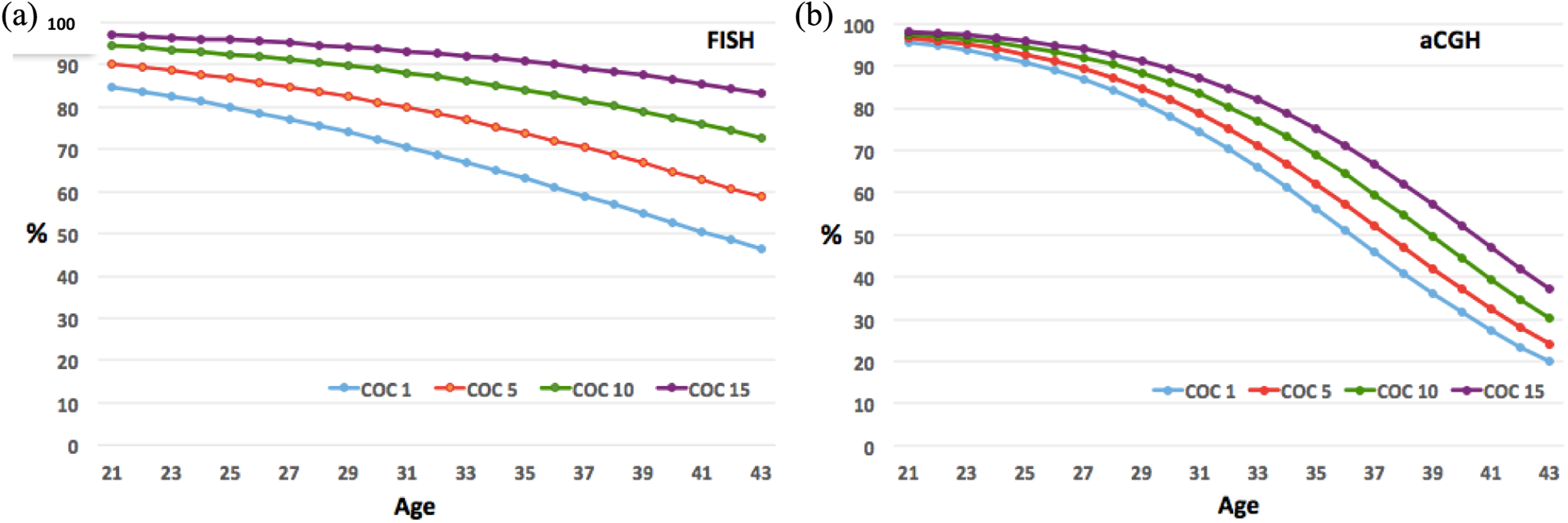
Age-specific probability of finding at least one euploid embryo according to the number of retrieved oocytes assessed with (a) FISH and (b) aCGH. COC: cumulus oocytes complexes (retrieved oocytes); FISH: fluorescent in situ hybridization; aCGH: array comparative genomic hybridization.
For example, the POE for a 35-year-old woman who has 5 retrieved oocytes is 74% and 62% with FISH and aCGH, respectively. If the same woman had 15 retrieved oocytes, her chances of having at least one euploid embryo would increase to 91% and 75% with FISH and aCGH, respectively (Figure 4). In our study, when the possibility of finding at least one euploid embryo for transfer was evaluated according to maternal age, the percentage of cycles with at least one transferable embryo decreased steadily from age 35 (p < 0.001), with a continual decrease until 39 and a steep decline from age 40 onward (p < 0.001) which became even steeper after age 43 (p < 0.001) with aCGH, as shown in Figure 3. In patients over 43 years old, there were no normal embryos for transfer with aCGH, whereas the percentage of cycles with one transferable embryo was 47% with FISH. That’s because of the difference in the detectibility rate of FISH and aCGH which is approximately 15% for patients over 40 and 10% in patients 39 and younger (Figure 3).
Determining the optimal number of oocytes to find at least one transferable euploid embryo
Using multivariate logistic regression analysis, we showed that the odds of finding at least one euploidy embryo was highest when 16–20 oocytes were retrieved compared to 1–5 oocytes with both FISH (OR 8.62, 95% CI 4.67–15.93) and aCGH (OR 2.78, 95% CI 1.07–7.21). With FISH, the POE was significantly higher with each subgroup of retrieved oocytes up until 16–20 oocytes as seen with the increasing ORs compared to the reference group of 1–5 oocytes as shown in Table 3. However, with aCGH, significance for the POE was reached only when 16–20 oocytes were retrieved. Figure 5 shows the mean probabilities with ±2 SD obtained from this analysis for each retrieved oocyte group.
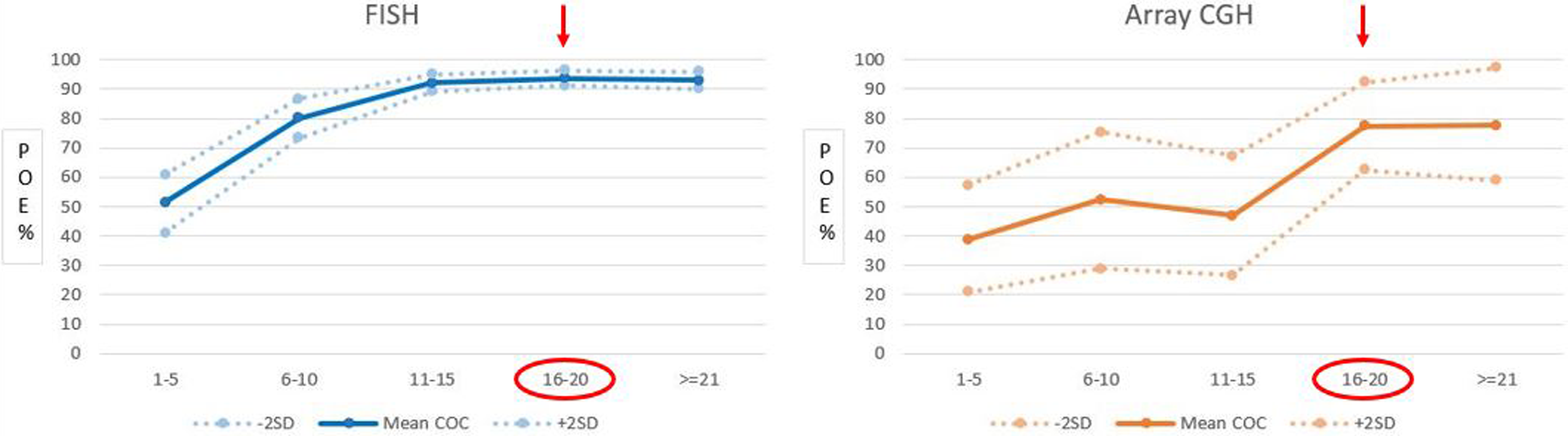
Probability of finding at least one euploid embryo according to each category of oocytes retrieved after adjustment for age with FISH and a CGH. Dots show the mean probabilities of each oocyte subgroups calculated according to logistic regression analysis and the Odds Ratios for each subgroup is given in Table 3. The arrows and circles point the subgroup which gives the highest probability of finding at least one euploid embryo. FISH: fluorescent in situ hybridization; aCGH: array comparative genomic hybridization. POE: Probability of finding at least one euploid embryo.
Multivariate logistic regression analysis showing the impact of age and each category of oocytes retrieved (COC) on the probability of finding at least one euploid embryo.
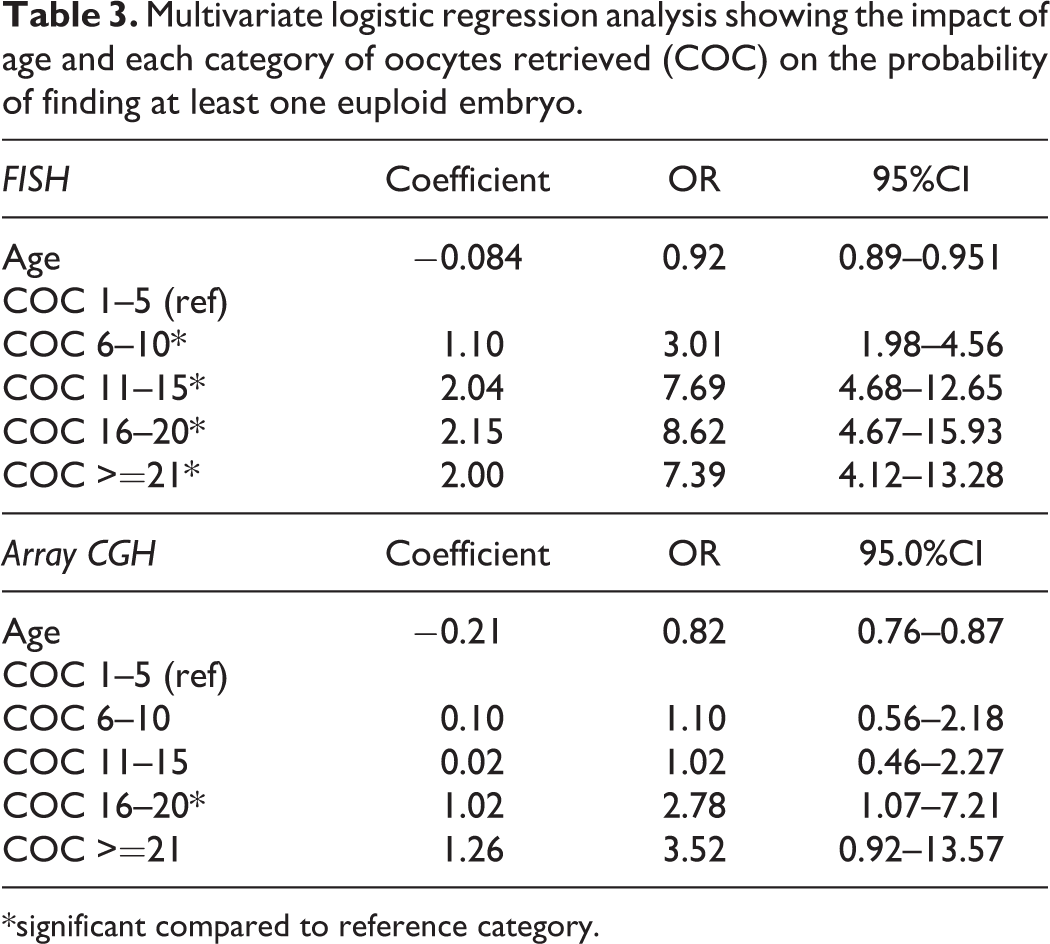
*significant compared to reference category.
As shown in the graph, the mean POE with aCGH for a patient with 1–5 retrieved oocytes is 39%, and it is approximately 50% for 6–15 retrieved oocytes. If the patient had 16–20 retrieved oocytes her chances of reaching transfer would increase 2.8 fold compared to a patient having 1–5 retrieved oocytes regardless of age. However her chances would not increase further after 20 oocytes (Table 3, Figure 5).
With FISH, if a patient has 1–5 retrieved oocytes, then the POE will be 51%, whereas it will significantly increase up to 92% and 94% if she has 11–15 and 16–20 retrieved oocytes, respectively. There will be no further increase in her chance of finding an euploid embryo if she has more than 20 oocytes (Table 3, Figure 5). If a woman has 5 or less oocytes, the chance of finding at least one euploid embryo is significantly decreased with FISH when compared with those with a higher number of retrieved oocytes.
When we subdivided the data into two groups based on age to further analyze the data of patients having ≤5 retrieved oocytes, there was at least one euploid embryo for transfer in 75% (15/20) of the patients younger than 39 years old and in 29.3% (12/41) of the patients aged 39–43 years old with aCGH. In this age-group (39–43 years), in 4 (75%) of the 5 patients with more than 20 oocytes, we were able to find at least one euploid embryo for transfer.
Discussion
In this study, we demonstrate that age is the most significant predictor of success in finding at least one euploid embryo with both FISH and aCGH. The POE and euploidy rate declined with age, regardless of the PGS technique used. When controlled for age, the POE increased with each additional oocyte retrieved. However, the number of available oocytes was not associated with the euploidy rate either with FISH or with aCGH. The biopsy day did not significantly affect the POE, but the euploidy rate was significantly higher with aCGH trophectoderm biopsy compared to cleavage stage biopsy.
According to our data, FISH is able to detect 87% and 82% of aneuploidic embryos that are detectable with aCGH for patients under 40 years old and over 40 years of age, respectively (Figure 1). This is in accordance with a previous report which shows a detectability rate of 80–91% when different number of probes and biopsy days are used (Munne, 2012)
This study shows the importance of complete chromosomal assessment using aCGH especially after age 39. Women over 43 years of age should be counseled regarding the low incidence or the risk of having no euploid embryos for transfer. However, women at the age of 40–43 can benefit from aCGH if they can produce multiple embryos with a relatively higher probability of finding euploid embryos. When we plotted the data after adjusting for age and number of retrieved oocytes, the POE was less than 20% with aCGH in patients older than 40 with one retrieved oocyte and the chances increased up to 50% when the number of retrieved oocytes were 15. If we counsel a 35-year-old normoresponder patient with a chance of retrieving 10–15 oocytes, then the POE is approximately 70–75%. Thus, the study provides us with information for appropriate counseling, which takes into account the age and number of retrieved oocytes (Figure 4).
It seems that women aged between 35 and 43 are more likely to benefit from aCGH if they have numerous suitable embryos from which to select for transfer that is correlated with the number of retrieved oocytes reflecting ovarian reserve. Harton et al. (2013) showed that with the selective transfer of euploid embryos, implantation and pregnancy rates were not significantly different between reproductively younger and older patients up to age 42. In their study, a similar trend of euploidy rate was observed with increasing age.
It is clear that the ovarian reserve as well as female age has significant effects on the probability of finding a suitable embryo for transfer. Therefore, the evaluation of ovarian reserve is of importance for clinical evaluation and counseling. Currently, the most reliable markers of ovarian reserve are anti-Müllerian hormone (AMH) and antral follicle count (AFC; Andersen et al., 2012). In the univariate analysis, although there was a trend (p = 0.07) through a positive correlation with AMH (data not shown) and POE for aCGH, after adjusting for other covariates, the significance disappeared. However, the number of retrieved oocytes, which reflect the AFC, maintained its significance.
Aiming to determine the optimal number of oocytes in finding at least one euploid embryo for transfer, we found that the highest probability of finding at least one euploid embryo could be achieved when 16–20 oocytes were retrieved with both FISH and aCGH, respectively (Figure 1). With more than 15 oocytes, the POE was approximately 78% after aCGH. However, the chance decreased when less oocytes were available. Of course, the chances of retrieving more oocytes decrease with advanced age, but patients with good ovarian reserve are the best candidates for PGS and, according to our data, even with one or two retrieved oocytes there is still a chance for finding one euploid transferable embryo. Therefore, we should not discourage those patients with poor ovarian reserve. In the light of these results, our policy is to share information about the relevant percentages individually with the patients.
Only a few publications have addressed the influence of the number of retrieved oocytes on the outcome of PGS for aneuploidy. Vandervorst et al. (1998) reported a threshold level of nine oocytes below which a significant reduction in reproductive outcome could be expected and suggested the discussion of the option of cycle cancellation with the couple when fewer than nine COCs are expected, and the recommendation of cancellation when this number was <6. In contrast to our study, this study population was very small and consisted of PGD cases for single gene disorders and sex chromosome abnormalities. A study by Tur-Kaspa (2012) showed that the availability of normal embryos and the chances of pregnancy increase with increasing numbers of oocytes retrieved and the chances decrease with a lower number of oocytes, especially in young patients, and they suggested that routine canceling of cycles with an anticipated low number of oocytes is to be reconsidered. In addition to these studies, there is a case report concluding to appropriately weighing the decision to cancel a PGS cycle because even in cases with poor ovarian response there is a chance of pregnancy after PGS. Although there is insufficient data in the literature regarding the optimum number of oocytes for PGS with aneuploidy testing, our results will be useful for appropriately counseling patients undergoing PGS.
The best approach when counseling PGS cases for aneuploidy testing should be first to make a proper evaluation of the ovarian reserve with AMH and AFC and to counsel the couple by giving an estimation of the POE specific to age and ovarian reserve of the woman as shown in Figure 4. We have shown that with aCGH while having one additional oocyte increases the POE 1.1% (Table 2, Figure 4), having more than 15 retrieved oocytes compared to having less than five increases the chances of finding at least one euploidy embryo nearly threefold after adjustment for age (Table 3, Figure 5). Determining the best protocol for ovarian stimulation to achieve the maximum number of oocytes for retrieval based on the patient’s age, basal AFCs, and, if available, previous ovarian responses, is of crucial importance.
PGS gives very many couples the opportunity to have a healthy child. With this study, we report an individualized approach specific to age and number of retrieved oocytes useful for counseling patients undergoing PGS for aneuploidy testing.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
