Abstract
Objectives
Honey total polyphenolic fraction (HTPF) is reported to have anti-disease potential, however the role of HTPF in the regulation of microRNAs (miRNAs) has never been investigated. This study was undertaken to investigate the potential of HTPF against inflammation via regulation of miRNAs in pancreatic islets of Langerhans.
Methods
Pancreatic islets were isolated from C57BL/6 mice and HTPF was purified from honey. Bioinformatics algorithms were used to determine miRNA target genes. Expression of miRNA and mRNA was determined using their specific taqman assays. Pairing between miRNA and 3′ untranslated region (3′UTR) of mRNA was confirmed using luciferase reporter clone containing the 3′UTR of mRNA sequences and results were verified by transfection of mouse pancreatic β-cell line Min6 with miRNA inhibitors.
Results
The data showed that mmu-miR-26a-5p is a direct regulator of cyclooxygenase-2 (COX-2) expression and HTPF inhibits COX-2 expression or prostaglandin E2 (PGE2) production via up-regulating mmu-miR-26a-5p expression. Transfection of islets with anti-miR-26a-5p significantly enhanced COX-2 expression and PGE2 production (
Conclusion
HTPF suppresses glucose-induced PGE2 production and activation of NF-κB via negative regulation of COX-2 and mmu-miR26a-5p. These novel pharmacological actions of HTPF on glucose-stimulated pancreatic islets provide new suggestions that HTPF or HTPF-derived compounds inhibit glucose induced inflammation in pancreas by up-regulating the expression of microRNAs.
Introduction
Type 1 diabetes (T1D) is an organ-specific disorder caused by autoimmune destruction insulin-producing beta cells in Langerhans of the pancreatic islets. 1 Due to ethical and practical issues, the current understanding of destructiveness of pancreatic islets and the process to understand diabetic pancreatic pathology is limited. Small number of studies conducted on the autopsies and biopsies from T1D patients, indicate that beta-cell destruction and islet infiltration is performed through lymphocytes, macrophages, natural killers and dendritic cells and by the overexpression of a few immune system associated genes in T1D patients.1,2 To explain the pathology more clearly and move forward in our understanding of the process leading to islets destruction is to have detailed information occurring at the molecular level. However, it is believed that the underlying the destruction process of pancreatic islets is multifactorial and its etiology still needs to be fully studied. 3 Molecular evidences clearly show that the onset of T1D and the destruction process of islets is related to the overproduction of several pro-inflammatory cytokines and inflammatory mediators including cyclooxygenase-2 (COX-2, also known as PTGS2).4,5 It is important to know that usually COX-2 is expressed at low levels in most of the cells and tissues, but it is well known to induce a wide range of inflammatory stimuli such as cytokines, chemokine and other inflammatory mediators.4–6 In diabetic pancreatic islets, COX-2 is expressed constitutively and dominantly and its excessive induction is well known to increase the production of prostaglandin E2 (PGE2),7,8 therefore, along with COX-2, PGE2 is considered as one of a possible therapeutic targets for treatment of T1D. MicroRNAs (miRNAs) are small non-coding RNAs known to regulate mRNA genes by pairing with their complementary sequences at 3’ untranslated regions (3’UTR) at the post-transcriptional level and it is speculated that miRNAs regulate more than 70% of all human protein-coding genes and participate in the regulation of almost all cellular events studied so far.9–11 Now it is well established that miRNAs play an important role in the pathogenesis of T1D,12–14 but their role in the destruction of diabetic islets remains largely unexplored.
Honey has been used as a traditional sweetener for many decades but its potential as a therapeutics has also been tested in recent years.15,16 Numerous studies demonstrated its therapeutic values as it is a rich source of antioxidants, polyphenols and flavonoids. 17 Studies showed that it has potential to improve lipid profile and reduces the risk of coronary heart diseases.17–19 It has potential also as a tropical medicine as it shows healing of wounds, burns, diabetic foot, ulcers and also healing of psoriatic and herpes lesions.19–21 Interestingly, honey and its polyphenolic contents have been reported to possess antioxidant, anti-inflammatory, antimicrobial and anti-cancer activities.22–27 Most importantly, it is also now well documented that the glucose induced induction of COX-2 or PGE2 mediated inflammatory activity in human islets suggested one of the major routes through which hyperglycemia contributes to the destruction of pancreatic beta cells. 28 In view of these, we hypothesized that the polyphenolic fraction of honey inhibits the glucose-induced COX-2 or PGE2 mediated inflammatory activities by regulating the expression of microRNAs in pancreas, the main organ affected by hyperglycemia. To test this hypothesis, pancreatic islets were isolated from normal C57BL/6 black mice and treated with honey total polyphenolic fraction (HTPF) and glucose. Bioinformatics algorithms were used to screen 3’UTR of COX-2 mRNA for the identification of seed matched sequences complementary to miRNAs sequences. The role of specific miRNA in regulation of inflammatory activity was determined by transfection of islets with specific miRNA inhibitor. Our novel data showed HTPF inhibits the glucose-induced COX-2 expression and PGE2 production via upregulating the expression of microRNA mmu-miR26a-5p in mouse pancreatic islets. These investigations further verified in pancreatic beta cell lines Min-6 using transfection with reporter constructs and anti-miRNAs. These are novel findings have never been investigated before and may be of value in the design of novel therapies for the reducing inflammation or treatment of hyperglycemic patients.
Methods
Extraction of honey polyphenols
Natural honey was collected from Egypt and total polyphenols were extracted as described previously.
29
Briefly, the solid-phase extraction of polyphenols was performed using Amberlite XAD-2 column (catalog # 10357, Sigma-Aldrich Inc., St. Louis, MO., USA). The sugar contents from honey were removed by an acidification of the column using hydrochloric acid solution and the polyphenolic fraction from the column was eluted using methanol solution and the collected fraction of polyphenols was then dried using a Rotavapor and re-suspected in minimum volumes of ethanol and extensively dialyzed against PBS (10 m
Pancreatic islets isolation and pancreatic β-cell line Min6 culture
With the ‘Institutional Review Board’ Ethical approval (IRB approval #19-09-02) from the Deanship of Scientific Research, Qassim University, KSA, mouse pancreatic islets were isolated in accordance of the World Medical Association Guidelines for use of laboratory Animals. Islets from 3-6 months old C57BL/6 mice were isolated as described by the published method. 14 Briefly, the mice were euthanized, and collagenase P solution, 1 mg/mL (catalog # 11215809103, Roche, Indianapolis, IN, USA) was injected into the bile duct to inflate the pancreas, followed by removal and incubation at 37°C for digestion until the pancreas formed a milky solution with only a few clumps. After digestion, islets were purified in density gradient Histopaque solution (Sigma-Aldrich). Islets were manually selected, washed in HEPES buffer, and cultured overnight in RPMI 1640 complete medium supplemented with 15% heat-inactivated fetal bovine serum (FBS). Whereas, the mouse pancreatic beta cell line Min6 31 was cultured in DMEM (catalog # SLM-120-B, Millipore, Darmstadt, Germany) supplemented with 15% FBS as described previously. 32
Treatment of pancreatic islets and pancreatic β-cell line Min6 with honey polyphenolic fraction and glucose
Purified islets were plated in 24-well plates (30-40 islets per well), allowed 24 h recovery after purification, washed twice and transferred to complete RPMI 1640 complete medium to stabilize basal gene expression and serum-starved for 12 h as described previously.
14
Whereas pancreatic Min6 cells (70–80% confluent) were also serum-starved for 12 h in DMEM culture medium as described previously.32,33 Both serum-starved islets and Min6 cells were washed twice with PBS and pretreated with honey polyphenolic fraction (HTPF, 20–40 μg/mL) for 2 h and stimulated with glucose (0–25 m
Transfection of islets and Min6 cells with miRNA inhibitors
Freshly isolated mouse pancreatic islets or Min6 cells were transfected with anti-miRNAs (100 nM; Qiagen, Hilden, Germany or Ambion, Foster City, CA, USA) in replicates using HiPerfect Transfection Reagent (Qiagen) without disrupting of islets structure in accordance to the manufacturers’ instructions with some modifications.14,34 Briefly, freshly isolated intact islets (∼50) or Min6 cells (70–80% confluent) were coated in culture plates and allowed to recover for 24–72 h and were transfected with the miRNA inhibitors, or miRNA controls. After 72 h post-transfection, islets or Min6 cells were pretreated with honey polyphenolic fraction (20–40 μg/mL) for 2 h or stimulated with glucose (0–25 m
Luciferase reporter assays
Luciferase reporter assays were performed using luciferase assay kit (Promega, Madison, Wisconsin, USA) as described previously. 10 Briefly, a luciferase reporter vector containing the entire 3′untranslated region of PTGS2 (or COX-2) mRNA (# ENST00000367468) and empty vector containing only luciferase gene and its constitutively active promoter (SwitchGear Genomics, Menlo Park, California, USA) were used in the reporter assays. Pancreatic beta cell line Min6 were co-transfected with 100 ng reporter plasmid, 50 and 100 nM selected anti-miRNAs (Qiagen) or negative control miRNAs (Qiagen) using HiPerfect Transfection Reagent (Qiagen) in a 96-well plate. Min6 cells were cultured 24 h after transfection and treated with HTPF (20–40 μg/mL) for 2 h and dual luciferase activity was measured. Each experiment was performed in triplicate and repeated three times using different sets of Min6 cells.
MicroRNA isolation, first strand cDNA synthesis and TaqMan assays
Total RNA containing miRNA fraction was isolated from non-transfected or transfected pancreatic islets and Min6 cells using mirVana miRNA isolation kit (Ambion, Foster City, CA, USA) as described by the manufacturers and published procedure.35,36 Total RNA (0.5–1.0 μg) was reverse-transcribed using Super Script First Strand cDNA synthesis kit (Applied Biosystems) and desired mRNAs and miRNAs expressions were quantified by TaqMan Gene Expression Assays (Applied Biosystems) using the Step One Real Time PCR System (Applied Biosystems). GAPDH/snoRNA202 expression was used as an endogenous control. Relative expression levels were analyzed using double delta CT method as described previously. 37
Western immunoblotting for protein expression
Protein expression in non-transfected and transfected pancreatic islets or Min6 cells was determined by western blotting as described previously.38,39 Briefly, the total cell lysates were prepared and were loaded on 10% resolving SDS-PAGE with 4% stacking and blots were probed with primary antibodies specific for the COX-2 (catalog # D5H5; Cell Signaling Technology Beverley, MA, USA) and β-actin (catalog # 8457; Cell Signaling Technology).
Prostaglandin E2 ELISA
Release of prostaglandin E2 (PGE2) in the culture of non-transfected or transfected pancreatic islets and Min6 cells was determined using PGE2-specifc ELISA kit (Cayman Chemicals, Ann Arbor, MI, USA) according to the instructions of the manufacturer.
Nuclear factor-kappa B activity assays
Nuclear levels of NF-κBp65 and NF-κBp50 in treated or untreated Min6 cells were determined in the nuclear extracts using a highly sensitive Transcription Factor ELISA Kit according to the instructions of the manufacturer (Abcam, MA, USA).
Statements
The study was performed in accordance of ARRIVE guidelines (http://www.nc3rs.org.uk/arrive-guidelines). Moreover, we also confirm that all methods were conducted in accordance with our institutional guidelines and regulations for the use of animals.
Statistical analysis
The data are expressed as the mean±SD unless stated otherwise and were collected from at least three independent experiments. One-way ANOVA of variance followed by Tukey-Kramer multiple comparisons test, and two-way ANOVA of variance followed by Bonferroni post hoc comparisons test.
Results
Bioinformatics algorithms for the detection of conserved seed matched sequences of microRNAs in 3′UTR of COX-2 (or Ptgs2) mRNA
First we scanned 3’-untranslated (3’UTR) of mouse COX-2 mRNA for the determination of conserved seed matched sequences of microRNAs using TargetScan bioinformatics algorithm. TargetScan algorithm showed that the 3’UTR of mouse COX-2 (ENST00000367468.4) contains 2487 nucleotide bases and has seven conserved sites for different microRNAs namely miR-26a-5p, miR-103-3p, miR-107-3p, miR-101-3p, miR-143-3p and miR-144-3p (Figure 1(a)). The details of the predicted consequential pairing of target region of 3′UTR of COX-2 with these highly conserved miRNAs are given in Figure 1(b). Interestingly, results also analyzed that the microRNAs miR-26-5p, miR-103-3p/miR-107-3p, miR-101a-3p, miR-143-3p and miR-144-3p seed sequence conserved for mouse 3’UTR of COX-2 mRNA is also conserved for other species such as human, chimp, rhesus, squirrel, rat, rabbit, pig, cow, cat, dog and brown bat (Figure 1(c)). In order to validate these results, we further analyzed the whole 3’UTR of COX-2 mRNA for the sequence complementary to mmu-miR26a-5p by another computer based bioinformatics approach miRBase, which uses different algorithm methods. Using miRBase algorithm, the results showed that the stem-loop sequence of mmu-miR26a-5p showed similar complementary sequences in the same two positions of 3’UTR of mouse COX-2 mRNA (Supplemental data Figure 2). Furthermore, another bioinformatics approach miRanda database (miRDB) also confirmed the occurrence of complementary sequences of mmu-miR26a-5p in the 3’UTR of COX-2 mRNA (Supplemental data Figure 3). All three algorithms TargetScan, miRBase and miRanda databases identified sequences conserved in the 3′UTR of COX-2 are complementary to microRNA-26a-5p seed sequence. TargetScan bioinformatics algorithm for the prediction of seed matched sequence of conserved mouse microRNAs in 3′UTR of COX-2 (or PTGS2) mRNA. (a) 3′UTR of mouse PTGS2 mRNA shows conserved sites for miR-26a-5p, miR-103-3p, miR-107-3p, miR-101-3p, miR-143-3p and miR-144-3p. (b) Duplex prediction of mmu-miR-26a-5p, mmu-miR-103-3p, mmu-miR-107-3p, mmu-miR-101a-3p.1, mmu-miR-143-3p or mmu-miR-144-3p with the seed sequence in the 3′UTR of mouse PTGS2. (c) Cross-species conservation of the miR-26a-5p, miR-103-3p, miR-107-3p, miR-101-3p, miR-143-3p or miR-144-3p seed sequence in the 3′UTR of PTGS2 mRNA. The sequences showing in red boarders are the locations of potential seed-matched sequence for PTGS2 mRNA.
Expression of COX-2, mmu-miR-26a-5p, mmu-miR-103-3p, mmu-miR-107-3p, mmu-miR-101a-3p, mmu-miR-143-3p and mmu-miR-144-3p in un-stimulated and glucose-stimulated mouse pancreatic islets
Treatment of primary islets with 25 m Expression of COX-2 and expression of microRNAs conserved in 3′UTR of COX-2 mRNA in glucose stimulated mouse pancreatic islets. (a) COX-2 mRNA expression in primary mouse pancreatic islets stimulated with glucose (2.0 or 25 m
Validation of mmu-miR-26a-5p binding in the 3′UTR of COX-2 mRNA
To test whether the 3′UTR of COX-2 is a functional target of the bioinformatics predicted mmu-miRNA-26a-5p, we used a luciferase reporter clone containing entire 3′UTR of COX-2 mRNA including the predicted the seed-matched binding sites and luciferase assay was performed in mouse pancreatic beta cell line Min6. As shown in Figure 2(d) and (a) significant concentration-dependent increase in luciferase activity in Min6 pancreatic beta cells co-transfected with COX-2 reporter and anti-miR-26a-5p (
Honey TPF up-regulates microRNA-26a-5p expression and down-regulates COX-2 expressions in pancreatic islets
Purified islets after 72 h culture in RPMI 1640 complete medium were pretreated with honey total phenolic fraction (Honey TPF, 20–40 μg/mL) for 2 h and stimulated with glucose (25 m Honey polyphenolic fraction up-regulates mmu-miR-26a-5p expression and down-regulates COX-2 expression in glucose-stimulated mouse pancreatic islets. (a) Effect of honey total polyphenolic fraction (Honey TRF) on glucose-induced down-regulation of mmu-miR-26a-5p in mouse pancreatic islets quantified by TaqMan assays using real-time PCR. *
Involvement of Honey TPF in the inverse co-regulation of mmu-miR-26a-5p and COX-2 was further verified by transfection of mouse pancreatic islets with anti-miR-26a-5p used as miRNA inhibitor. Mouse pancreatic islets were transfected with 100 nM anti-miR-26a-5p or anti-miR-control and were then treated with Honey TPF (20–40 μg/ml). As shown in Figure 4(a), transfection of islets with anti-miR-26a-5p significantly suppressed the mmu-miR-26a-5p expression compared with non-transfected islets ( Honey polyphenolic fraction up-regulates the mmu-miR-26a-5p expression and down-regulates COX-2 expression in anti-miR-26a-5p-transfected mouse pancreatic islets. (a) Effect of honey total polyphenolic fraction (Honey TRF) on mmu-miR-26a-5p expression in pancreatic islets transfected with anti-miR-26a-5p. *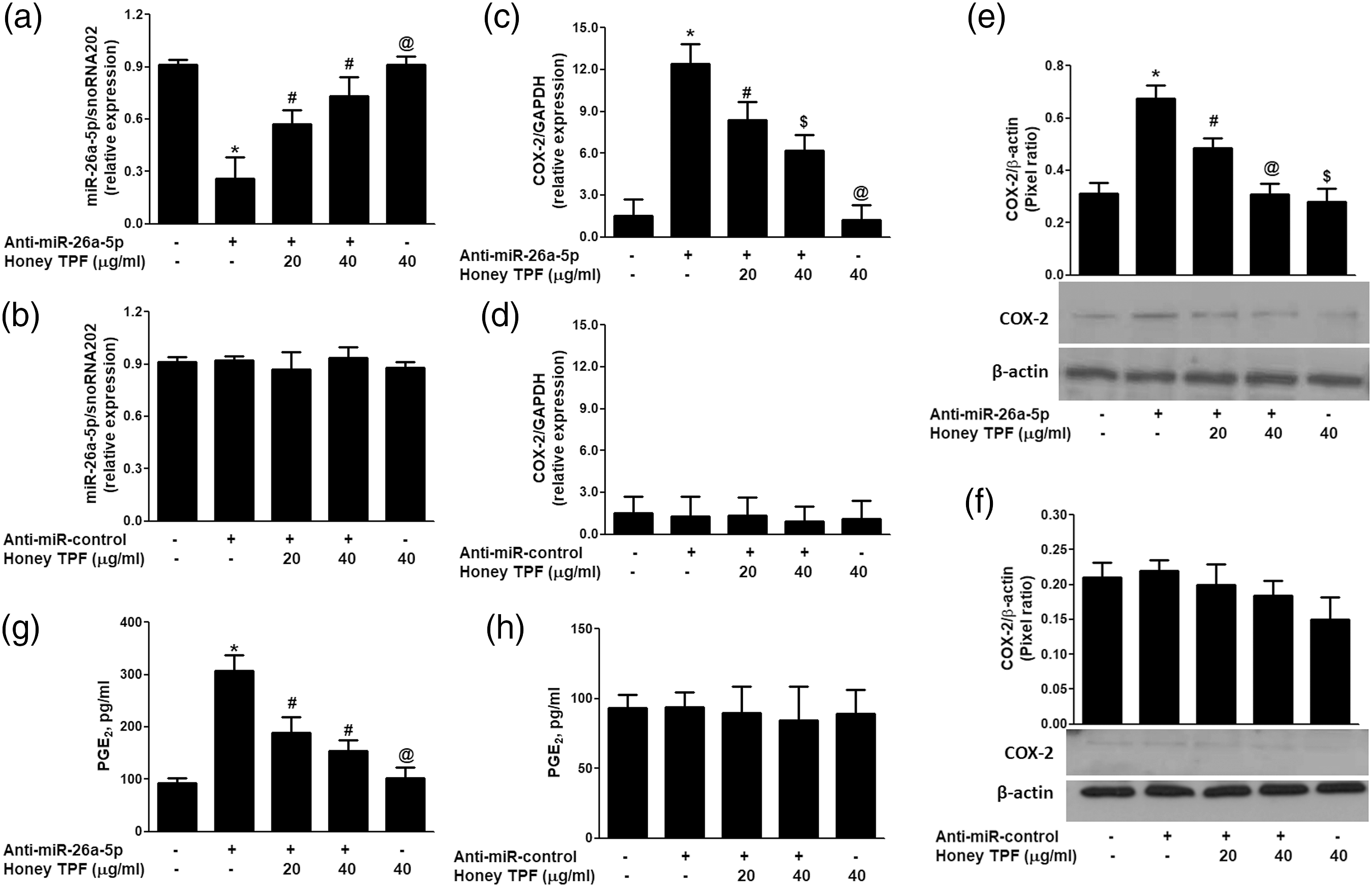
Honey TPF up-regulates microRNA-26a-5p expression and down-regulates PGE2 production in pancreatic islets
We also determine whether inhibition of COX-2 expression also inhibits PGE2 production in the same sets of mouse pancreatic islets transfected with anti-miR26a-5p or anti-miR-control, the culture medium was quantified for PGE2 secretion. Transfection of islets with anti-miR-26a-5p markedly enhanced the PGE2 production (
Effect of honey TPF on negative co-regulation of mmu-miR-26a-5p and COX-2 and the involvement of 3′UTR of COX-2 mRNA
Involvement of honey TPF in the negative co-regulation of mmu-miR-26a-5p and COX-2 was confirmed by a luciferase reporter assays using a reporter clone containing entire 3′UTR of COX-2 mRNA in pancreatic beta cell line Min6. The results showed a significant increase in relative luciferase activity (RLA) in Min6 beta cells co-transfected with 3′UTR COX-2 reporter and anti-miR-26a-5p ( Honey polyphenolic fraction directly up-regulates the mmu-miR-26a-5p expression and down-regulates COX-2 expression via 3′UTR of COX-2 mRNA sequence in mouse pancreatic beta cell line Min6. (a) Effect of honey total polyphenolic fraction (Honey TRF) on the luciferase activity in Min6 cells co-transfected with the reporter vector and anti-miR-26a-5p. @
Honey TPF inhibits glucose-induced NF-κB signaling via mmu-miR-26a-5p in mouse pancreatic β-cells
In order to investigate whether honey TPF inhibits the glucose-induced downstream signaling via microRNA modulation, we transfected pancreatic beta cells Min6 with anti-miR-26a-5p and then treated with honey TPF for 2 h and stimulated with glucose for 30 min and the levels of nuclear NF-κBp65 and NF-κBp50 were determined by the highly sensitive NF-κB p65/p50 Transcription Factor Binding Assay kit (Abcam). Our data showed that the transfection of MIN6 cells with anti-miR-26a-5p or MIN6 cells treated with glucose alone significantly increased the nuclear levels of NF-κBp65 ( Honey polyphenolic fraction inhibits glucose induced NF-κB signaling events through mmu-miR-26a-5p in mouse pancreatic beta cell line Min6 cells. (a) Effect of Honey TRF inhibits glucose stimulated NF-κB p65 activation in Min6 cells transfected with anti-miR-26a-5p. #
Discussion
This is the first study that shows that the polyphenolic fraction of natural honey inhibits COX-2 mRNA/protein expression and PGE2 production via upregulating microRNA mmu-miR-26a-5p expression in pancreatic islets. In recent years, scientists from all over the globe actively investigating the role of microRNAs in diabetes and several novel miRNAs were discovered and their roles in regulation of genes relevant to the pathogenesis of diabetes were identified.40,41 In the last two decades, lots of research on prostaglandin pharmacology and physiology was conducted but the most important was based on the identification of COX-2, as it expresses predominantly in numerous pathological conditions and now it’s well known for inducing inflammation in almost all major tissues including pancreatic islets.4,42 It is well known that PGE2 is generated via activation of COX-2, which has now been considered as a key player in the induction of pro-inflammatory pathways not only in diabetes but also in several metabolic disorders. 43 Importantly, developed pharmacological inhibitors of COX-2 show promising therapeutic effects against the onset and progression of a number of inflammatory diseases. 44 Although diabetes is predominant in every population but the treatment is still limited to a few classes of drugs and none of the treatment reverts this chronic disorder.45,46 In view of these, there is an urgent need for the development/identification of new compounds that inhibit the metabolic process of diabetes. Previous studies reported honey comprises about 180 chemical substances including amino acids and protein; polyphenols; vitamins such as ascorbic acid, thiamine, riboflavin, niacin, pantothenic acid, and pyridoxine; and minerals such as zinc, potassium, chlorine, calcium, sulfur, sodium, phosphorous, magnesium, iron, copper. 47 The polyphenolic fraction of honey comprises of heterogeneous chemical compounds that can be further characterized into flavonoids such as flavonols, flavones, flavanols, flavanones, anthocyanidin, chalcones, and isoflavones) and non-flavonoids such as phenolic acids.17,47 Most importantly, polyphenols of natural honey performed range of therapeutics activities for numerous human disorders,15–21 therefore this study was designed to investigate the therapeutic potential of polyphenolic fraction of honey against the glucose induced COX-2 mRNA/protein expression and PGE2 production via regulation of microRNAs in mouse pancreatic islets. To prove these, we first isolated total polyphenolic fraction from natural honey (honey TPF) using the Amberlite XAD-2 column, whereas the pancreatic islets were isolated from black C57BL/6 mice. Our data determined the role of honey TPF for the first time in suppressing the pro-inflammatory genes through regulation of miRNA in mouse pancreatic islets.
Bioinformatics has now become a powerful approach for the identification of novel microRNA target genes relevant to the disease pathogenesis, numerous miRNAs seed matched have been identified and experimentally tested successfully and now this approach has become a method of choice9,48,49 and the role of miRNAs has now well reported in numerous human disorders.
50
Here we used three different bioinformatics algorithms namely TargetScan, miRBase and miRanda for the scanning of 3’UTR of mouse COX-2 (or Ptgs2) mRNA for the determination of seed matched sequences of miRNAs. TargetScan bioinformatics algorithm detected seven conserved sites in the 3’UTR of mouse COX-2 for different microRNAs such as miR-26a-5p, miR-103-3p, miR-107-3p, miR-101-3p, miR-143-3p and miR-144-3p. Moreover, this algorithm also detected that the these miRNA seed sequences conserved in mouse 3’UTR of COX-2 mRNA are also conserved in other species such as human, chimp, rhesus, squirrel, rat, rabbit, pig, cow, cat, dog and brown bat, indicating that the seed matched sequence of these miRNAs in 3′UTR of COX-2 mRNA is highly conserved. In order to confirm these findings, we further analyzed the whole 3’UTR of COX-2 mRNA for the sequences complementary to miRNAs by another computer based bioinformatics approach miRBase and miRanda database. All three algorithms TargetScan, miRBase and miRanda databases identified sequences conserved in the 3′UTR of COX-2 are complementary to miR-26a-5p seed sequence (Supplemental data). In order to validate these computer based predictions, experimental studies were performed with all seven highly conserved miRNAs predicted in 3′UTR of mouse COX-2 mRNA using their specific taqman assays. Our experiments on purified mouse pancreatic islets showed that treatment with 25 m
After validation of inverse co-relation of COX-2 and mmu-miR-26a-5p, the efficacy of honey TPF was tested on modulation of COX-2 and mmu-miR26a-5p expressions in purified pancreatic islets or Min6 cells. To prove the therapeutic potential of honey TPF, mouse pancreatic islets were transfected with anti-miR-26a-5p or anti-miR-control and then treated with honey TPF. The data showed that transfection of islets with anti-miR-26a-5p inhibited the mmu-miR-26a-5p expression compared with non-transfected islets. Importantly, treatment of these transfected islets with honey TPF enhanced mmu-miR-26a-5p expression. These results were completely reversed, when we estimated mRNA expression of COX-2. These findings not only confirmed the negative correlation of mmu-miR26a-5p and COX-2 but also showed that honey TPF potentially inhibited COX-2 mRNA expression via upregulating the mmu-miR26a-5p expression in pancreatic islets. These results were further re-validated at the levels of COX-2 protein and PGE2 production. Transfection of islets with anti-miR-26a-5p markedly enhanced the COX-2 protein and PGE2 production and this induced COX-2 protein or PGE2 production was significantly inhibited upon honey TPF treatment. Therapeutic potential of honey TPF in the negative co-regulation of mmu-miR-26a-5p and COX-2 was confirmed in pancreatic beta cell line Min6. Treatment of honey TPF with co-transfected Min6 cells of commercially purchased luciferase reporter clone of entire 3′UTR of COX-2 mRNA and anti-miR-26a-5p, showed dose-dependent decrease in luciferase activity. This concentration-dependent reduction in luciferase activity, may indicate that honey TPF reduces luciferase activity of 3′UTR COX-2 reporter via modulation of mmu-miR-26a-5p expression. Involvement of honey TPF in the negative co-regulation of mmu-miR-26a-5p and COX-2 was further reevaluated by transfection of Min6 with anti-miR-26a-5p and then co-treated with honey TPF and glucose. The data showed that the treatment with glucose alone or transfection with anti-miR-26a-5p significantly reduced the mmu-miR-26a-5p expression in Min6 cells. Interestingly, transfection with anti-miR-26a-5p synergizes with glucose noticeably reduced mmu-miR-26a-5p expression. Whereas, treatment of anti-miR-26a-5p-transfected Min6 cells with honey TPF consistently upregulated the glucose-inhibited mmu-miR-26a-5p expression in a dose-dependent manner. In same sets of transfected Min6 cells, COX-2 mRNA was quantified and found that the Min6 cells treated with glucose alone significantly upregulated COX-2 mRNA expression. Transfection with anti-miR-26a-5p also significantly increased COX-2 expression as compared with those Min6 cells transfected with anti-miR-control, but a sharp increase of COX-2 mRNA expression was observed in anti-miR-26a-5p-transfected Min6 cells stimulated with glucose for 24 h. Importantly, honey TPF significantly inhibited glucose-induced COX-2 mRNA in a dose-dependent manner in Min6 cells transfected with anti-miR-26a-5p. To determine whether this effect of honey TPF on inhibition of COX-2 mRNA also affected the protein level, lysates of these transfected Min6 cells were prepared for COX-2 protein estimation. Western blot of COX-2 protein analysis showed that transfection of Min6 cells with anti-miR-26a-5p effectively further enhanced the glucose-induced COX-2 protein. This enhanced expression of COX-2 protein was significantly inhibited by honey TPF in a dose-dependent manner. Taken together, the data clearly indicate mmu-miR-26a-5p is a direct regulator of COX-2 expression in mouse pancreatic beta cells and honey TPF caused inhibition of COX-2 mRNA/protein expression is mediated via up-regulation of mmu-miR-26a-5p expression. We also determine whether inhibition of COX-2 expression also inhibits PGE2 production, same sets of Min6 cells culture medium were analyzed. The data showed that the transfection of Min6 cells with anti-miR-26a-5p transfection effectively further increased the glucose-induced PGE2 pancreatic beta cells. This enhanced expression of PGE2 production was consistently inhibited by honey TPF in a dose-dependent manner. These novel results clearly indicate that honey TPF inhibits COX-2 expression and PGE2 production via up-regulation of mmu-miR-26a-5p expression.
The nuclear transcription factor (NF)-κB regulates numerous aspects of immune system and is known as a pivotal mediator of proinflammatory responses.
52
Activation of NF-κB signaling events induces the expression of multiple proinflammatory genes, including those encoding cytokines, chemokines, and numerous other proinflammatory mediators including COX-2 and PGE2.53,54 Analysis of 5′-flanking sequence of COX-2 gene found two sites for NF-κB.
54
Conclusions
This is the first report that shows the purified fraction of polyphenols from natural honey inhibits glucose-induced COX-2 expression and PGE2 production via up-regulation of the expression of mmu-miR26a-5p in pancreatic islets. These findings were further verified in mouse pancreatic β-cell line Min6. Moreover, the data also determined that honey TPF inhibits glucose-induced NF-κB activity. These novel pharmacological actions of honey TPF on microRNA regulation provide new suggestions that honey TPF or honey-derived compounds may be of value for the treatment of diabetic patients where inflammation and microRNAs play an active role.
Supplemental Material
sj-pdf-1-eji-10.1177_20587392221076473 – Supplemental Material for Honey polyphenolic fraction inhibits cyclooxygenase-2 expression via upregulation of microRNA-26a-5p expression in pancreatic islets
Supplemental Material, sj-pdf-1-eji-10.1177_20587392221076473 for Honey polyphenolic fraction inhibits cyclooxygenase-2 expression via upregulation of microRNA-26a-5p expression in pancreatic islets by Syed S Ahmed, Sultan AlNohair, Waleed A Abdulmonem, Homaidan T Alhomaidan, Naila Rasheed, Mohamed SM Ismail, Manal A Albatanony and Zafar Rasheed in European Journal of Inflammation
Footnotes
Acknowledgements
The authors thank Mr Casimero A. Victoria for helping in transfection experiments.
Authors’ contributions
Syed Suhail Ahmed, Sultan AlNohair, Waleed Al Abdulmonem, Homaidan T. Alhomaidan, Manal A. Albatanony carried out experimentation and data interpretation. Naila Rasheed and Zafar Rasheed performed data interpretation and manuscript drafting. All authors have read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the Deanship of Scientific Research (Grant # med-2019-2-2-I-6736), Qassim University, KSA.
Availability of data and materials
All data and materials used in this current study are available from the corresponding author on reasonable request.
Ethical approval
The study was approved by the ‘Institutional Review Board’ Ethical Committee from the Deanship of Scientific Research, Qassim University, KSA (IRB approval #19-09-02).
Supplemental material
Supplemental material for this article is available online.
Nomenclature
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
