Abstract
Background
Although tumor response evaluated with radiological imaging is frequently used as a primary endpoint in clinical trials, it is difficult to obtain precise results because of inter- and intra-observer differences.
Purpose
To evaluate usefulness of a cloud-based local-read paradigm implementing software solutions that standardize imaging evaluations among international investigator sites for clinical trials of lung cancer.
Material and Methods
Two studies were performed: KUMO I and KUMO I Extension. KUMO I was a pilot study aiming at demonstrating the feasibility of cloud implementation and identifying issues regarding variability of evaluations among sites. Chest CT scans at three time-points from baseline to progression, from 10 patients with lung cancer who were treated with EGFR tyrosine kinase inhibitors, were evaluated independently by two oncologists (Japan) and one radiologist (France), through a cloud-based software solution. The KUMO I Extension was performed based on the results of KUMO I.
Results
KUMO I showed discordance rates of 40% for target lesion selection, 70% for overall response at the first time-point, and 60% for overall response at the second time-point. Since the main reason for the discordance was differences in the selection of target lesions, KUMO I Extension added a cloud-based quality control service to achieve a consensus on the selection of target lesions, resulting in an improved rate of agreement of response evaluations.
Conclusion
The study shows the feasibility of imaging evaluations at investigator sites, based on cloud services for clinical studies involving multiple international sites. This system offers a step forward in standardizing evaluations of images among widely dispersed sites.
Introduction
*Equal contributors.
Widely used standards for measuring the response of tumor are the Response Evaluation Criteria In Solid Tumor (RECIST) and the World Health Organization (WHO) criteria. The former is one-dimensional (1D) and the latter is two-dimensional (2D) (9,10). RECIST was proposed and established in 2000 and an updated version was published in 2009 (11). It is based on the rationale that maximum diameters are linearly related to cell kill compared to bi-dimensional evaluation, and on evidence showing the agreement between 1D and 2D evaluations (10). RECIST has been applied in many clinical trials, and although quite useful, it does have some problems. Considerable intra- and inter-observer variability has been noted, especially in tumors with complex shapes or located in poorly-contrasted regions (12–15). To compensate for this variability, regulatory authorities recommend an ICR with several readers to mitigate potential bias resulting from variance among investigator sites (2,5–8).
The software solution used in this study was Lesion Management Solutions (LMS) developed by MEDIAN Technologies (16,17). LMS is an image analysis software application for evaluating CT images. It allows lesion identification, quantification, and comparison of successive CT scans of the same patient. The comparison is achieved by synchronous navigation between two scans and automatic pairing of the lesions. Using these tools, we made possible a cloud-based local-read paradigm for imaging evaluations which consists of implementing a cloud software solution and a quality control at investigator sites. Readers working at distant locations were able to reliably perform radiological evaluations from the same cloud system. The objective is to standardize the review of images at investigator sites in the frame of a clinical study. The purpose of this study was to investigate usefulness of a cloud implementation of this system in terms of evaluation of tumor response of lung cancers according to the RECIST criteria and the inter-observer agreement among sites.
Material and Methods
Study design and patient inclusion
This was a retrospective study of CT images of lung cancer patients at Saga University Hospital (Japan), Nice University Hospital (France), and other facilities. KUMO I, the first part of the study, was intended to demonstrate the feasibility of cloud implementation and to suggest technical improvements, as well as to identify issues regarding variability of evaluations among sites. The workflow of the KUMO I study is shown in Fig. 1. The Japanese investigator sites, the French independent reviewer site, the data manager (MEDIAN Technologies, Valbonne, France, www.mediantechnologies.com), and the data center (Canon IT Solutions, Tokyo, Japan, www.canon-its.co.jp) are all connected to the Canon cloud infrastructure service SOLTAGE (Canon IT Solutions, Tokyo, Japan) through a virtual private network (VPN). SOLTAGE provides storage, computing, and application services. The investigator sites provide the medical images and perform the analysis and interpretation of the medical images on Web-based MEDIAN Technologies Imaging Service. The analysis and interpretation results are centrally stored and available to the independent reviewer to perform its analysis and interpretation. This workflow is under the supervision of the data manager and data center that also have access to central information and progress status of the study.
KUMO I and KUMO I Extension Study work flow. Scan data were imported into the web client and anonymized by the data center, Canon IT Solutions, Japan. The data center, Canon, and the data managers (MEDIAN) stored and processed the images, and the image database was sent to each reader, in Japan and France. The study compared evaluations between readers and analyzed the reasons for discordances. Readers with different medical training and education, working at distant locations were able to reliably perform radiological evaluations from the same cloud system. The cloud quality control service detected non-conformance in applying RECIST 1.1 and had the readers change their evaluations, resolving the discrepancies. Based on KUMO I, KUMO I Extension was designed to improve the system. KUMO I Extension included additional quality controls to arrange the consensus of lesion selection (*). LMS, lesion management solutions; PACS, picture archiving and communication system, VPN; virtual private network.
CT images of 10 lung cancer patients were acquired at three time-points (baseline, best response, and progression) in the course of treatment. Patients who were treated with EGFR tyrosine kinase inhibitors (EGFR-TKI) were randomly selected, since obvious changes in image were observed when EGFR-TKI was administered to lung cancer patients with
Imaging technique
All images were taken with multi-detector CT scanners (LightSpeed VCT®, GE Healthcare Japan, Tokyo, Japan; SOMATOM Definition®, SIEMENS, Munich, Germany) at Saga University Hospital, or selected in MEDIAN images database. Slice thickness of the scans was 5 mm for KUMO 1 and 1–2.5 mm for KUMO 1 Extension. Tube voltage was 120 kV, and field of view (FOV) was 300–500 mm. All images were properly anonymized and copied to a virtual server at the data center operated by Canon IT Solutions, Inc. The images were processed by a cloud-based prototype of Lesion Management Solutions (LMS) (MEDIAN Technologies, Valbonne, France). LMS is at the core of MEDIAN’s Clinical Trial Imaging Services, which include image and workflow management, and image processing specifically designed for multi-site oncology clinical trials (Fig. 2). The image processing component of LMS offers software for detection, segmentation, and quantification of thoracic lesions (Fig. 2a). The segmentation process, which is based on a three-dimensional (3D) region-growing algorithm, begins with a simple point-and-click on the lesion of interest. Readers can make manual adjustments to the contour of the lesion as necessary. After segmentation is complete, the longest axial diameter, short axis, and volume of each lesion are extracted automatically. In the follow-up evaluation, two scans from two time-points are displayed side by side while automatic registration points to the volume of interest in the newer scan (Fig. 2b). A reader points to the corresponding lesions in the newer scan, which are then analyzed in the same manner as at the baseline evaluation. Change in size and volume between time-points are then calculated and reported (Fig. 2c). LMS graphically displays the evolution of the tumor burden based on both diameter and volume of lesions. Finally all of the review data are used to compute the response evaluation and to categorize the response as complete response, partial response, stable disease, or progressive disease.
The LMS system consists of three steps: (a) auto-segmentation and quantification; (b) follow-up segmentation; and (c) response evaluation. Just one click on a lesion leads to automatic segmentation, and quantification of longest diameter, short axis, and tumor volume. To compare images from two time-points, the system automatically registers the images to match the position of the lesion. Evaluation is also automatic, based on RECIST criteria and volumetry analysis.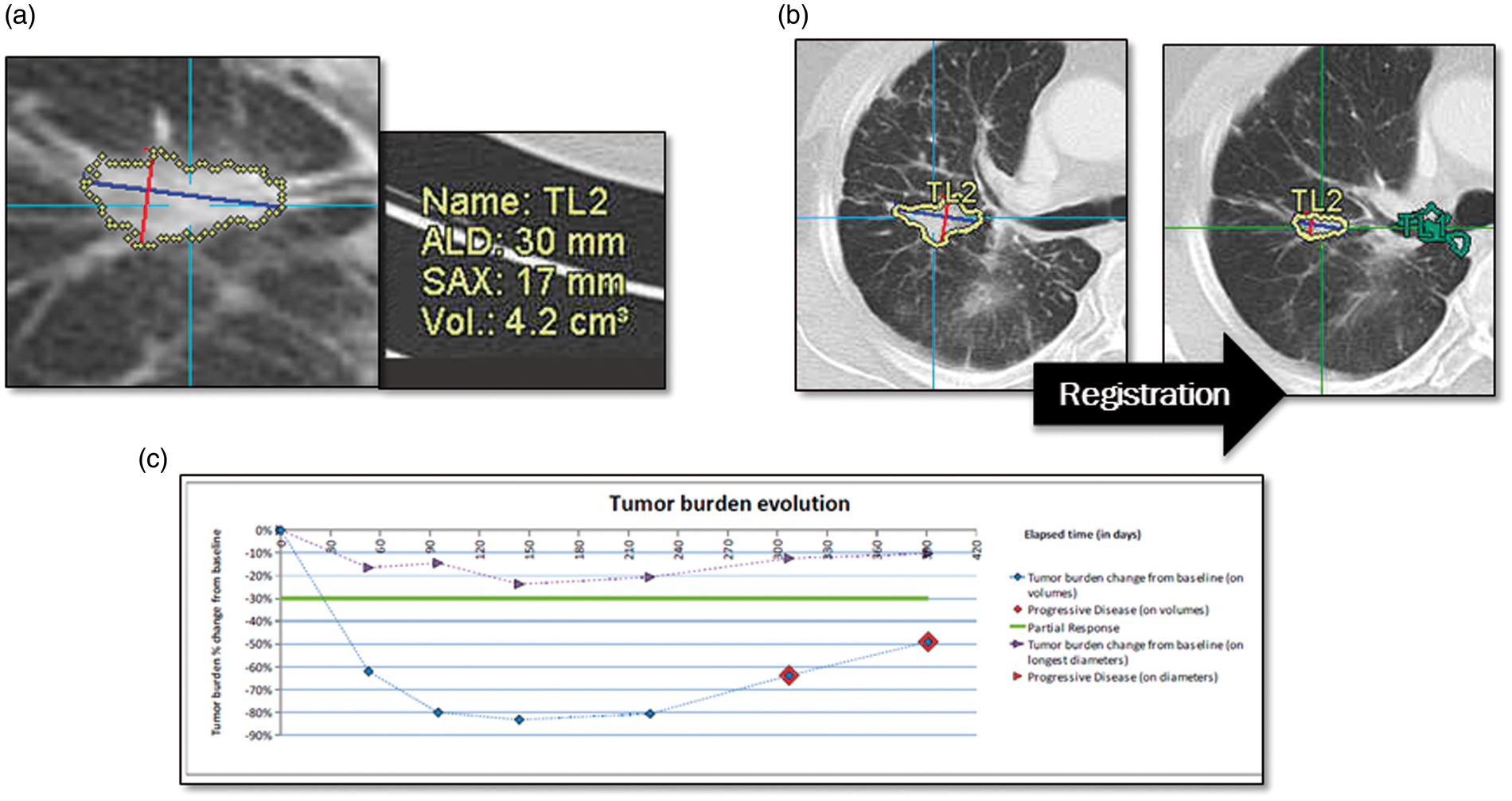
Statistical analysis
Kappa analysis was performed to evaluate inter-observer agreements between the reviewer and each investigator using IBM SPSS Statistics 19 (SPSS, Inc., IBM Company, Tokyo, Japan). The strength of agreement indicated with kappa values has been reported as: <0, poor; 0–0.20, slight; 0.21–0.40, fair; 0.41–0.60, moderate; 0.61–0.80, substantial; and 0.81–1.00, almost perfect (18).
Results
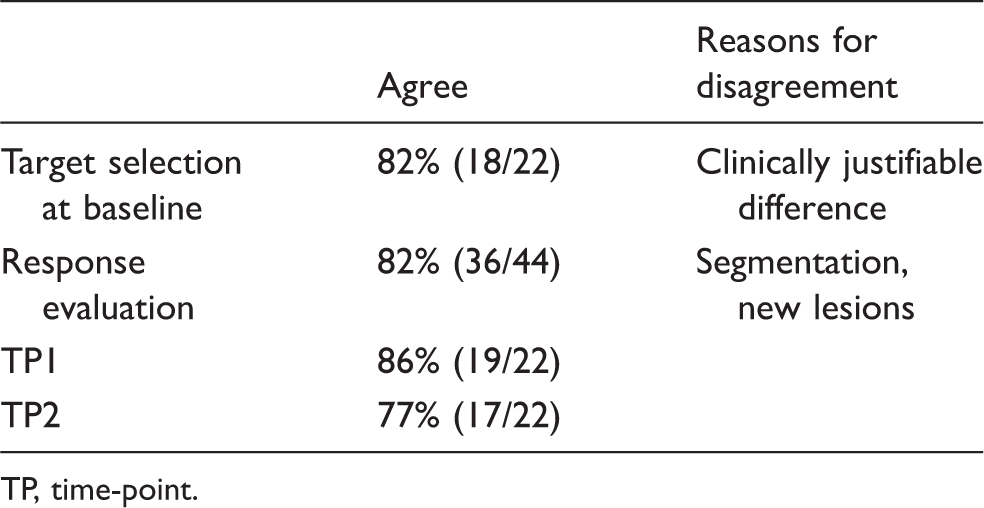
Inter-reader agreement between two investigators and a reviewer (KUMO I Study).

TP, time-point.
Improvement of inter-reader agreement was observed after reaching consensus between a reviewer and investigator. Based on the results of KUMO I study, KUMO I Extension added a cloud-based quality control service to achieve a consensus on the selection of target lesions (Fig. 1). The KUMO I Extension, raised the agreement rate of response evaluations to 82% from 65% in KUMO I (Table 2). Kappa coefficient value of inter-observer agreement for response evaluations between the reviewer and the investigators was 0.724 ± 0.17 (95% CI). The cloud software solution gives the possibility for a given clinical study that all investigator sites work with the same tools, on the same database. Thanks to the data centralization provided by this cloud configuration, an ongoing quality control is made possible. A quality control performed by a clinician regarding the choice of target lesions can be implemented: in case of disagreement between the reviewer and the investigator site, the site has to explain and/or revise his choice. In spite of this system, target selection at baseline did not completely agree even after adjustment because of clinically justifiable differences between the reviewer and the investigators as shown in Fig. 3. The investigators selected a primary lesion as the target lesion even though segmentation was difficult because of the complicated shape of the lesion and its location adjacent to the pleura (Fig. 3a). The reviewer selected a metastatic lesion, however, because evaluation of the lesion is easily reproducible (Fig. 3b). Differences in lesion segmentation between the reviewer and the investigators are shown in Table 3. Among total lesions which were evaluated in all images in KUMO I Extension Study, the frequency of indicated differences in axial diameter between a reviewer and each investigator was investigated. The lesions adjacent to the mediastinum and pleura showed bigger differences in axial diameter (Fig. 4) compared to those in the lung field. Lesions in the lung field had enough contrast on imaging so they could be segmented by automated tools, while other lesions, including lesions in the lymph nodes, could not.
Discordance of lesion selection between a reviewer (a) and investigators (b). The reviewer selected a metastatic lung lesion, and the investigator selected a primary lesion adjacent to the pleura. Comparison of segmentations performed manually between a reviewer (a) and two investigators (b, c). Inter-reader agreement between two investigators and a reviewer (KUMO I Extension Study). TP, time-point. Difference in lesion segmentation between two investigators and a reviewer (KUMO I Extension Study).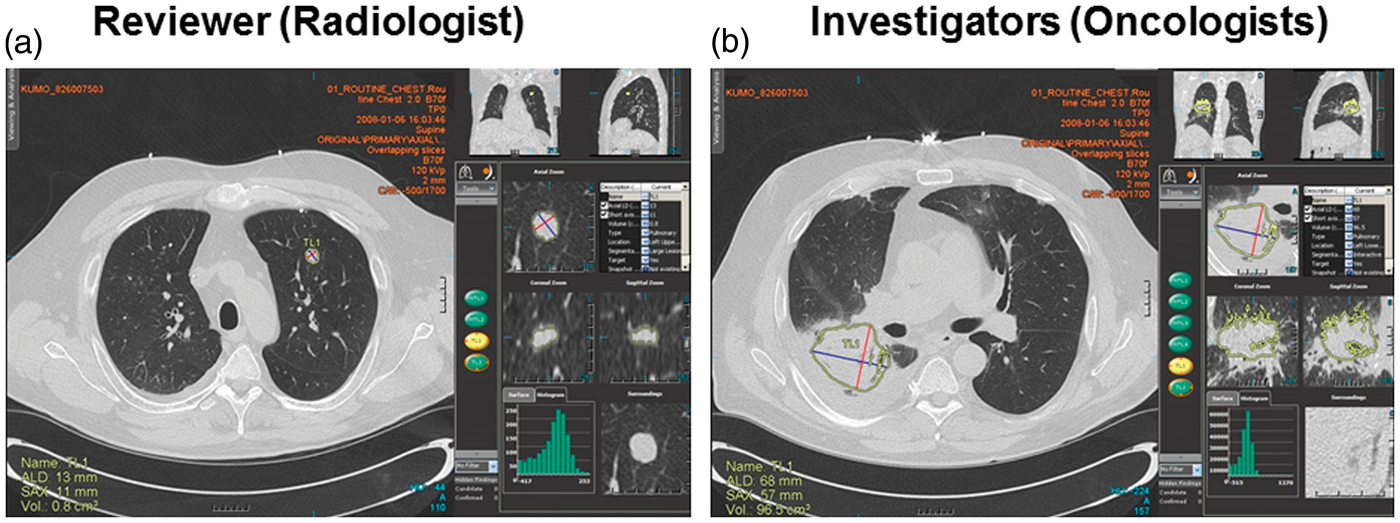


Discussion
Building on the results of KUMO I, modification by the cloud quality control service could lead to improved concordance between readers in the KUMO I Extension study. Ongoing monitoring of evaluations through specialized services to reduce variability among sites was made possible by centralized data management.
As for cost-effectiveness, the cost required to implement solutions depends on the number of sites in usual central review systems. However, a cloud-based solution developed in this study contains centralized data processing system, and readers just need a minimum set of computers connecting to the Internet. Therefore, the total cost to set up such a cloud-based service is much lower than using locally installed software from both direct and indirect perspectives including maintenance. To secure the participants’ privacy, all necessary de-identification processes are done before putting images to the LMS system. Personal information is not shared among readers. We also chose VPN connection and a highly secure data center.
Some limitations of the study are the small sample size and the lack of evaluation of lymph nodes. These studies pave the way for further investigations such as the improvement of the automated segmentation tools to better address the lesions adjacent to the pleura and mediastinum as well as the lymph nodes, since discordance in response evaluation occurred mainly when segmentation was performed manually. In addition, it would also be interesting to take advantage of this implementation to investigate the limitations of the RECIST criteria: volumetric measurement of tumor size has been reported to be reproducible and accurate compared to 1D or 2D measurements (19). LMS provides automatic evaluation of tumor volume even when the tumor contour is complicated, as with caveating lesions. With such modification of the system, a prospective clinical study involving several hospitals around the world should be performed to confirm its feasibility.
In conclusion, the cloud-based local-read automatic imaging analysis system could become an integral component of global clinical trials for solid tumors after some modifications.
Declaration of conflicting interests
Colette Charbonnieris an employee of MEDIAN Technologies, France. Junta Yamamichi and Hideaki Mizobe are employees of Global Healthcare IT Project, Canon Inc., Japan.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
