Abstract
Background and Objective
Prostate cancer is one of the most common cancers in men, and early diagnosis is critical. Segmentation of cancerous regions in multiparametric MRI is a key step. Deep neural networks, such as nnU-Net, perform well, and incorporating prostate zonal information may further improve accuracy.
Methods
This study introduces four prostate cancer segmentation ensembles that integrate zonal data, compared with a baseline model, which uses zonal information as a separate input channel. Ensembles employ specific prostate zone cancer segmentation models trained with the nnU-Net method. To address variability in manual annotations, a new evaluation metric, the tolerant Dice Score Coefficient (DSC τ ), is proposed, accounting for ground truth inaccuracies.
Results
Ensemble 3 yields the best performance, with a 4.77% higher mean DSC and 6.17% higher mean DSC τ than the baseline. Although the metrics of Ensemble 4 are slightly lower, it reduces false positives by 7.79% and uses fewer models (2 vs. 3), making it more efficient. Furthermore, the application of the Conover post hoc test for unreplicated blocked data shows that there is no statistically significant difference in performance metrics between the results of two ensembles. Thus, Ensemble 4 is the preferred approach for prostate cancer segmentation. Additionally, all ensembles achieve 5.03% to 7.13% higher mean DSC τ values compared to the standard DSC, confirming the effectiveness of the new metric in handling segmentation uncertainties.
Conclusion
The experiment results indicate that the proposed Ensemble 4 is the most suitable solution for the prostate cancer segmentation task. Moreover, the results also indicate that the proposed metric, DSC τ , accounts for ground truth segmentation errors.
Keywords
Introduction
The most recent cancer statistics, as presented in the article 1 by Bray et al., indicate that among men worldwide, the incidence of prostate cancer is the second highest, and its mortality rate ranks fifth among all cancers. Considering the scale and severity of prostate cancer, an accurate segmentation or localization of cancerous regions is required to provide support for the Prostate Imaging Reporting and Data System (PI-RADS) assessment.
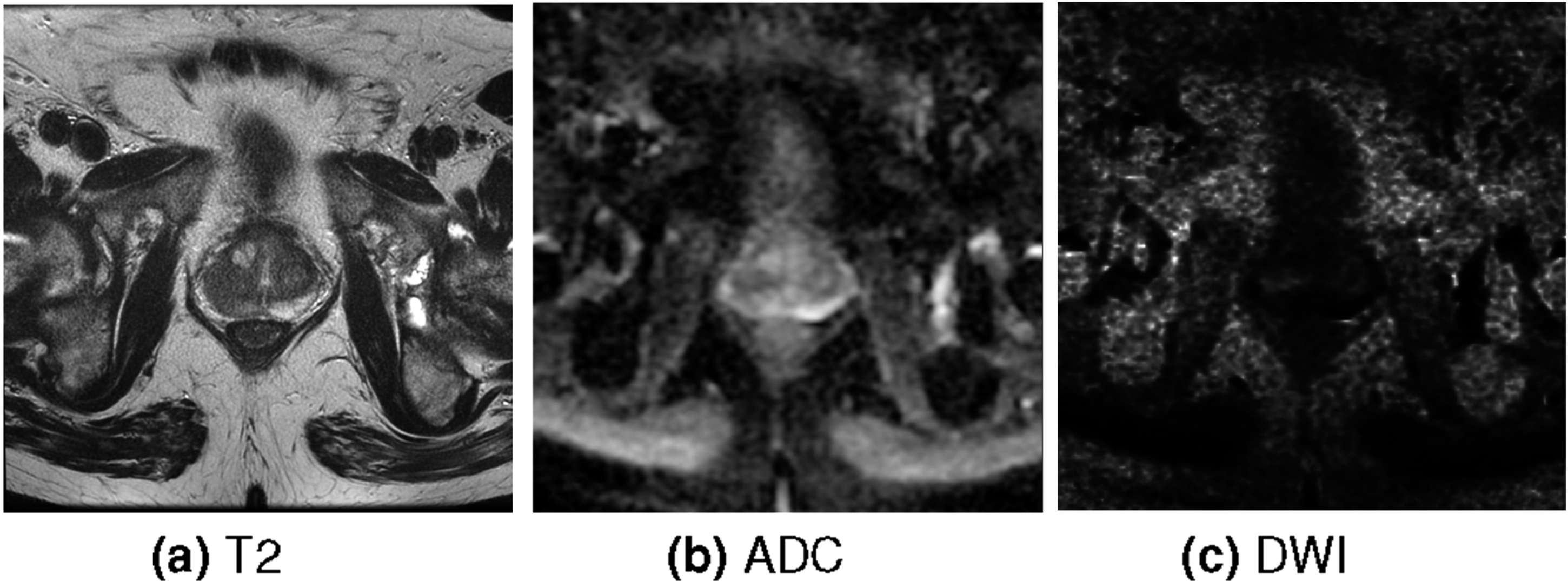
PI-RADS was introduced in the article 2 by Barentsz et al. to help radiologists evaluate mpMRI for further diagnostic assessment. It is one of the most widely used techniques for standardized interpretation in prostate cancer diagnosis. This technique employs multiparametric (mp) magnetic resonance imaging (MRI). The latest PI-RADS version is 2.1, introduced in the paper 3 by Turkbey et al. mpMRI consists of multiple sequences and the sequences used in PI-RADS v2.1 are T2-weighted (T2), Diffusion-Weighted Imaging (DWI), Apparent Diffusion Coefficient (ADC), and Dynamic Contrast-Enhanced (DCE) sequences. ADC and DWI are described in the article 4 by Baliyan et al. DWI is acquired from an MRI scanner while ADC is derived from multiple DWI scans. Moreover, the T2 sequence is described in the article 5 by Chavhan et al. Just as the DWI sequence, the T2 is acquired from an MRI scanner. All of these sequences are volumetric, anisotropic images. Lastly, the DCE sequence is described in the paper 6 by Yankeelov and Gore, which presents the methodology of dynamic contrast-enhanced MRI based on repeated volumetric imaging over time after contrast agent injection to capture contrast uptake and washout kinetics. In this study, DCE is excluded because PI-RADS v2.1 places primary diagnostic weight on T2, ADC and DWI, with DCE having secondary and limited incremental value.
The assessment within PI-RADS v2.1 employs different mpMRI imaging sequences across prostate zones 3 This is due to the fact that different imaging sequences are more effective in identifying cancerous regions within specific prostate zones. The PI-RADS v2.1 is assessed in the Transitional Zone (TZ) and the Peripheral Zone (PZ) of the prostate, as these zones account for the majority of the cancerous cases. The T2 sequence serves as the principal modality for evaluating TZ, while ADC and DWI are the primary sequences for assessing PZ. Thus, this paper proposes ensembles of deep neural network models trained on different mpMRI sequences to segment cancerous regions within specific prostate zones.
Prostate zone masks are employed for training the deep neural network models within the proposed ensembles, as well as for evaluating the performance of the ensembles. These masks are acquired by using separate models for the TZ and PZ segmentation (SM4TZPZS) workflow, which was proposed by Vaitulevičius et al. in the article. 7 This workflow employs two deep neural network models: one dedicated to the segmentation of the TZ and the other to the segmentation of the PZ. The intersection of the segmentation outputs is assigned to the TZ. The prostate zonal mask is obtained by combining the outputs of the models into a single volume, which contains TZ and PZ. The paper 7 contains the experiment results, which indicate that this workflow generalizes between different datasets better than the baseline workflow, single PZ and TZ segmentation model. Moreover, SM4TZPZS workflow has already been applied in previous work on prostate cancer localization, specifically in the study 8 by Roman et al. In addition, several post-processing steps are introduced in this study in order to correct the result of the SM4TZPZS workflow.
In total four cancerous region segmentation ensembles are proposed in this paper. The first two ensembles employ distinct deep neural network models, each trained to segment cancerous regions in the specific prostate zone. The models of the first ensemble adhere strictly to the PI-RADS v2.1 assessment guidelines, utilizing only the principal sequences specific to each zone. In contrast, the models of the second ensemble incorporate all available sequences for both prostate zones. The third ensemble integrates the baseline model together with the second ensemble. The final ensemble utilizes the baseline model and the model which employs all sequences to segment cancerous regions exclusively within the PZ. This paper presents the experiment in which the proposed ensembles are compared with the baseline model, which uses prostate zone information as a separate channel.
Moreover, all deep neural network models used in the ensembles as well as the baseline model are trained with the state-of-the-art method for medical segmentation, no-new U-Net (nnU-Net). This method was introduced by Isensee et al. 9 nnU-Net and its variations were widely employed in various medical segmentation experiments such as the ones presented in articles.7,10–14 In addition, the research presented in this paper employs the extended nnU-Net method incorporating a preprocessing step introduced by Jucevičius et al. 14 This step involves converting anisotropic volumetric MRI data into isotropic. The experiments presented by Jucevičius et al. indicate that this step improves the accuracy of the nnU-Net method.
Lastly, a new evaluation metric, the tolerant Dice Score Coefficient (DSC τ ), is proposed in this paper. The experiment results provided in the article 15 by Chen et al. indicate that prostate cancer segmentation is a very difficult task, and the results of segmentation are highly variable even among experienced radiologists. Therefore, the proposed metric accounts for segmentation errors of the ground truth that may occur as a result of the prostate cancer segmentation process.
Overall, in this paper: • Four ensembles are proposed. The proposed ensembles are compared with the baseline model, a model that uses prostate zone information as a separate channel. The results of the experiment presented in this paper show that the most suitable solution to the prostate cancer segmentation task is the ensemble comprised of the baseline model and the model that employs all sequences to segment cancerous regions exclusively within the PZ. • A novel evaluation metric is proposed, which is an extended DSC that considers the ground truth’s segmentation error. In addition to the original DSC, the results of the experiment described in this paper are evaluated using the proposed metric. The results of this experiment affirm that DSC
τ
accounts for ground truth segmentation errors, as the means obtained from the collected DSC
τ
are higher than those from the original DSC.
The rest of this paper is structured as follows: Related works section reviews related work, Methods section provides a detailed description of the ensembles, Experiment section describes the dataset used in the experiment and experiment methodology and the implementation details, Results section presents the experimental results, Discussion section discusses the results and Conclusion section concludes the paper.
Related works
This section reviews works employing deep neural networks and prostate zone information for prostate cancer segmentation tasks. The example of the ground truth of this task is displayed in Figure 1 where the green volume is the cancerous region and the transparent region is the prostate mask. The segmentation is performed using mpMRI sequences T2, ADC and DWI. The examples of these sequences are shown in Figure 2. The example of the ground truth of prostate cancer segmentation task. The green volume is the cancerous region. The transparent region is the prostate mask. The example of a single slice of each sequence.
A lot of research has already been performed on employing prostate zone information and deep neural networks for prostate cancer segmentation. One of it is published by Hosseinzadeh et al. in the papers.16,17 In this research, the authors demonstrated the advantages of incorporating probabilistic prostate zonal segmentation within a 2D computer-aided detection system. Their findings published in the article 16 indicate that including PZ and TZ segmentations leads to an average increase of 5.3% in detection sensitivity and between 0.5 and 2.0 false positives per patient. Furthermore, the findings presented in the paper 17 indicate that training models with training sets of different sizes draw the same conclusion, prostate zone masks improve the accuracy of prostate cancer segmentation.
The other research whose proposed solution uses an attention mechanism is provided in the paper. 18 The authors of this paper propose a workflow as a solution. This workflow uses a U-Net variation, Dual-Attention U-Net. Moreover, the research also contains the comparison of their proposed workflow with nnU-Net, Attention U-Net, and Dual-Attention U-Net architectures. The comparison indicates that their proposed workflow achieves the best results. Meanwhile, Dual-Attention U-Net achieves slightly better results than nnU-Net on one of their test sets and worse on the other. Attention U-Net achieves worse results than nnU-Net and Dual-Attention U-Net architectures on both test sets.
More research is provided in the article 19 by Duran et al. Their research examined the impact of employing a whole prostate mask and a single prostate zone mask for segmentation models. The use of PZ achieved 1.8% higher sensitivity and 1.4 fewer false positives per patient than the use of the whole prostate mask. However, employing a TZ mask for the segmentation model resulted in 13.9% lower sensitivity and 1.4 lower false positive per patient than employing the entire prostate mask. The authors attribute the inaccurate performance of the TZ mask to a low number of lesions in the TZ. Moreover, according to the articles 20 by Oto et al., 21 by Rosenkrantz et al. and 22 by Yu et al. cancerous regions in the TZ are difficult to distinguish from benign hyperplasia nodules. The authors of the paper 21 also indicate that the diffuse background heterogeneity and multinodularity of the TZ complicate the cancerous region segmentation of this zone. Meanwhile, the authors of the article 22 indicate that stromal asymmetry may mimic a cancerous region within TZ, whereas chronic prostatitis may produce similar appearances within the PZ. Lastly, the research presented in the paper 23 by Yu et al. indicates that cancerous regions appear less often in TZ than in PZ. Therefore, one of the ensembles proposed in this study employs a PZ mask to a greater extent than the TZ mask.
Overall, a plethora of papers contain research that uses prostate zone information in deep neural network models as a channel-wise input. Examples include the research provided by Saha et al. in the article, 18 the research published by Zheng et al. in the article 24 and the research described in the paper 25 by Mehta et al. Thus, this paper focuses on exploring other possible incorporations of prostate zone information into deep neural network models as a solution to a prostate cancer segmentation task.
Lastly, the results of prostate cancer segmentation have a high variance among even experienced radiologists, as indicated by the results of the experiment provided in the article. 15 The experiment included 4 radiologists who were tasked with segmenting prostate cancer within a single slice of the anterior zone and the PZ. Their segmentations were compared by calculating the mean DSC. The resulting mean DSC for the segmentations within PZ is 0.81, while for the segmentations within the anterior zone is 0.58. Therefore, the metric proposed in this study addresses this issue.
Methods
Compared ensembles
In this study, four ensembles are introduced. The result of the ensemble is a disjunction of the outputs of all deep neural network models used in the ensemble. As mentioned before, these models are obtained using the nnU-Net method with an additional preprocessing step proposed by Jucevičius et al.
14
The primary distinction between these models lies in the input of the model. Each model uses different sequences as input, and each used sequence is a separate channel of the input. Furthermore, a different mask is applied to the input of each model by setting voxels that do not overlap with the mask to zero. This masking strategy is an integral component of the nnU-Net framework and is therefore selected over alternative approaches, such as soft masking. Thus, the following models are used in the proposed ensembles: • TZ (T2) - this model uses only the T2 sequence as an input to segment cancerous region within TZ. • PZ (ADC, DWI) - this model uses the ADC and DWI sequences as an input to segment cancerous region within PZ. • TZ (All) - this model uses all three sequences as an input to segment cancerous region within TZ. • PZ (All) - this model uses all three sequences as an input to segment cancerous region within PZ. • prostate (All, Zones) - this model uses all three sequences to segment cancerous region within the whole prostate. The prostate masks used for the experiment are disjunctions of TZ and PZ masks. Furthermore, the state-of-the-art method to employ prostate zone information in prostate cancer segmentation tasks is to use prostate zone masks in deep neural network models as a separate input channel together with other mpMRI sequences. This method was employed in the experiments presented in papers.16,18,25 Therefore, the prostate zone masks and mpMRI sequences are used as a channel-wise input of the prostate (All, Zones) model. This model is the baseline model used in the comparison.
The first proposed and compared ensemble, denoted as Ensemble 1, is visualized in Figure 3. Green areas in Figures 3–6 represent the result of cancerous region segmentation. As mentioned in the introduction, the principal sequence for evaluating prostate cancer in TZ is T2. Meanwhile, the principal sequences for evaluating prostate cancer in PZ are ADC and DWI. Therefore, this ensemble is the disjunction of TZ (T2) model’s result and PZ (ADC, DWI) model’s result. Formally - Ensemble1 = TZ (T2) ∨ PZ (ADC, DWI). The workflow of Ensemble 1. The workflow of Ensemble 2. The workflow of Ensemble 3. The workflow of Ensemble 4.



The schematic of the second proposed ensemble, denoted as Ensemble 2, is provided in Figure 4 While each prostate zone has its own principal sequences, radiologists typically employ all three sequences to assess cancerous regions in both zones. Therefore, this ensemble is the disjunction of the results of the models TZ (All) and PZ (All). Considering that the input of the model TZ (All) is three times bigger than TZ (T2) and the input of the model PZ (All) is 1/3 times bigger than PZ (ADC, DWI), Ensemble 2 is slower than Ensemble 1 during the inference and the training processes. Therefore, if Ensemble 1 and Ensemble 2 achieve similar results, then Ensemble 1 is preferred over Ensemble 2. The equation of Ensemble 2 is Ensemble2 = TZ (All) ∨ PZ (All).
The third proposed ensemble, denoted as Ensemble 3, is provided in Figure 5. Firstly, the baseline model, prostate (All, Zones), is used to create this ensemble. Furthermore, it is more likely that Ensemble 3 will achieve better results than Ensemble 2 as its input contains more information. Therefore, this ensemble is the disjunction of the TZ (All) model’s result, PZ (All) model’s result and prostate (All, Zones) model’s result. Overall, the accurate performance of this ensemble would indicate that prostate (All, Zones) model and ensemble 2 are complementary to each other. The equation of this ensemble is Ensemble3 = TZ (All) ∨ PZ (All) ∨ prostate (All, Zones).
The schematic of the last proposed ensemble, denoted as Ensemble 4, is visualized in Figure 6. Yu et al. in the paper 23 provide a summary of the epidemiological literature. This summary contains the distribution of prostate cancer instances in prostate zones. Approximately 20% of instances appear in TZ. This means that the data is imbalanced, and prostate cancer in TZ may not have enough instances for the deep neural network model training. Moreover, the research in the articles20–22 indicates that detecting prostate cancer in TZ is generally more complicated than in PZ.
Therefore, this ensemble is the disjunction of the results of the models PZ (All) and prostate (All, Zones). Overall, if this ensemble in the experiment bears performance just as accurate as Ensemble 3, then it will indicate that the TZ (All) model does not significantly influence the result of Ensemble 3. Formally - Ensemble4 = PZ (All) ∨ prostate (All, Zones).
Performance evaluation
Accurate evaluation of prostate cancer segmentation is inherently challenging due to the substantial inter observer variability in delineating tumour boundaries, even among expert radiologists and pathologists. This variability arises from indistinct lesion margins, heterogeneous tumour appearance on mpMRI, and partial volume effects, leading to small but clinically insignificant boundary discrepancies that are nevertheless penalized heavily by conventional overlap based metrics such as the Dice Score Coefficient (DSC). Thus, in this research, a new metric, the tolerant Dice Score Coefficient, is proposed. DSC
τ
is an extended DSC
26
that considers a segmentation error of the ground truth which may arise due to the nature of prostate cancer segmentation. The original DSC is incorporated into the loss function of the nnU-net method. Moreover, DSC has been widely used in prostate MRI segmentation studies, particularly for whole gland and zonal prostate segmentation. This metric is used in experiments, such as those presented in the articles.27–29 Furthermore, DSC has also been used in lesion segmentation works to assess spatial overlap, such as.18,25,30 The equation for calculating the original DSC is provided in Equation (1). In this equation, TP is a volume of the predicted cancerous region that overlaps with the ground truth; FN is a volume of the ground truth annotation which does not overlap with the prediction, and FP is a volume of the prediction that does not overlap with the ground truth. The volumes TP, FP and, FN are calculated as numbers of voxels.
Metrics such as area under the receiver operating curve and average precision, as employed in the PI-CAI challenge, 31 and in the research published in the article, 18 are well suited for lesion level detection and case level clinical decision support. In contrast, this study focuses on voxel wise segmentation of prostate cancer lesions, where spatial overlap and boundary accuracy are of primary interest. Therefore, DSC was selected as the principal evaluation metric.
The proposed metric, DSC
τ
, relaxes the penalization of false positives and false negatives within a predefined spatial margin while preserving the definition of true positives. This design reflects the clinical reality that minor boundary deviations are often within the range of annotation uncertainty and should not be interpreted as segmentation failure. The proposed modification of DSC is Equation (2) where FN
τ
is the number of tolerant FN and FP
τ
is the tolerant FP. Tolerant FN and tolerant FP are metrics which consider segmentation errors of the ground truth. FN
τ
is calculated by dilating the predicted volume, while FP
τ
is calculated by dilating the ground truth volume. The dilation operation is described in the paper.
32
This operation is a morphological operation that expands foreground regions in a binary image by adding pixels to object boundaries. The dilation operation is performed by sliding a structuring element over the image and replacing each pixel with the maximum value within the neighbourhood defined by the structuring element. This operation is commonly used to fill small gaps, connect nearby structures, and enhance object boundaries. Both dilations are performed with an annulus-shaped kernel. The radius of the kernel is adjusted so that the result of the dilation makes the dilated regions wider by a fixed length, called tolerance. Custom metrics were also used in the experiments, described in the papers.24,33 The metric used in the paper
33
expands the prostate cancer segmentation by 5 mm. Furthermore, the authors of the paper
24
consider predicted local maxima of prostate cancer probability maps as true positives if they are located within 5 mm of any prostate cancer region. Thus, the chosen tolerance for the experiment of this paper is 5 mm. The Venn diagram which illustrates the calculation of FN
τ
and FP
τ
is presented in Figure 7. Venn diagrams of FN
τ
and FP
τ
.
Similar tolerant Dice Score Coefficient metrics have been introduced in the past. One of such metrics have been proposed in the paper. 34 However, unlike DSC τ , proposed in this study, the tolerant Dice Score Coefficient, proposed by the authors of the article, 34 contains TP calculated with tolerance. Thus, the novel metric, DSC τ , proposed in this paper, maintains the strict definition of correctly identified tumour regions, thereby preserving sensitivity to meaningful lesion detection while reducing undue penalization of ambiguous boundary regions. Another metrics similar to DSC τ are surface DSC, proposed in the article, 35 weighted DSC (WDC), proposed in the study, 36 and Added Path Length (APL), proposed in the paper. 37 These metrics differ significantly from DSC τ , introduced in this study. Surface DSC and APL assess the overlap of two surfaces rather than volumetric overlap as is done by the conventional DSC and DSC τ . Lastly, unlike the DSC τ , WDC takes into account the distance between the predicted and ground truth voxels in calculating volumetric overlap. It mitigates the impact of boundary discrepancies relative to the conventional DSC, but still penalizes them more than the DSC τ .
In this paper, the results of the experiment are evaluated with the original DSC and the proposed DSC τ .
Additionally, Friedman tests 38 are employed to evaluate statistical significance between related groups, specifically comparing ensembles and the baseline model. This non-parametric test is suitable for analysing two or more samples, particularly when the data exhibit non-normal distributions or involve small sample sizes. The Friedman test determines whether significant differences exist in the mean ranks of the groups, making it well-suited for repeated measures designs.
Finally, the lesion level metrics are reported in this study. These include the distributions of produced lesion counts for each compared ensemble and the baseline model, as well as the distributions of missed lesions, which correspond to lesion level false negatives. In addition, the distributions of spurious foci predicted by each ensemble and the baseline model are reported, representing lesion level false positives.
Experiment
Data
Statistics of echo time (TE) and repetition time (TR) for the acquired T2 and DWI sequences (milliseconds).
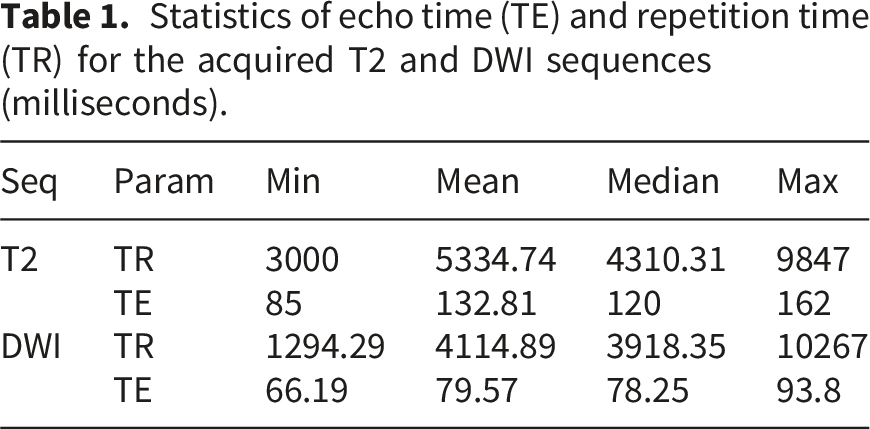
Moreover, the dataset includes prostate, cancerous region and biopsy masks. The cancerous region masks were manually delineated by a single radiologist of 10 years experience. The annotation process was completed prior to the experimental procedures. Thus, the annotator was blinded to the output of the compared ensembles and the baseline model. The biopsies were performed by the interventional radiologist with 15 years of experience. One patient had manually delineated cancerous regions that was not confirmed by a biopsy assigned a malignant diagnostic category, despite having other biopsies classified as malignant. Thus, this patient is excluded from the remaining cohort, resulting in the final dataset with the size of 119 cases. The complete flow diagram of patient selection is provided in Figure 8. The youngest patient in the final dataset is 46 years old, while the oldest one is 82. The average and median age of patients is 64 years old. On average, 15 biopsies were taken per patient, while the median number is 16. The highest number of biopsies performed on a patient is 25, while the lowest is 3. The distribution of the biopsy results by Gleason scores (GS) are reported in Table 2, where GS column indicates the Gleason Score and Category column indicates the corresponding diagnostic category. The No of patients column represents the number of patients who contain at least a single biopsy of the given GS, No of biopsies column indicates the total number of biopsies with that Gleason score and biopsies/patient represents the mean number of biopsies with the given Gleason score among patients with at least one such biopsy. Furthermore, the dataset is expanded with prostate zone masks. All sequences and masks are volumetric. Patient selection flow diagram. Gleason score distribution.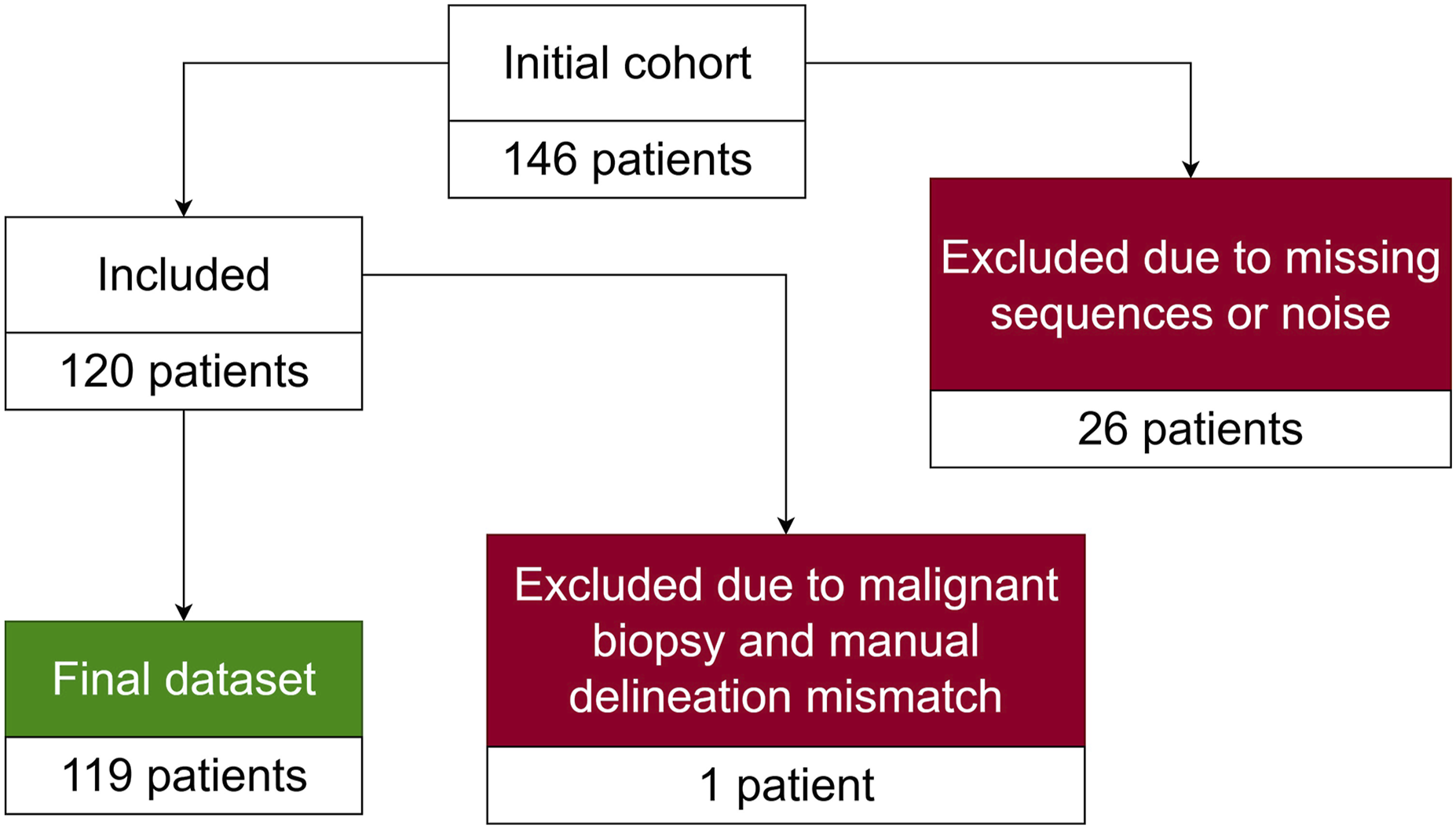

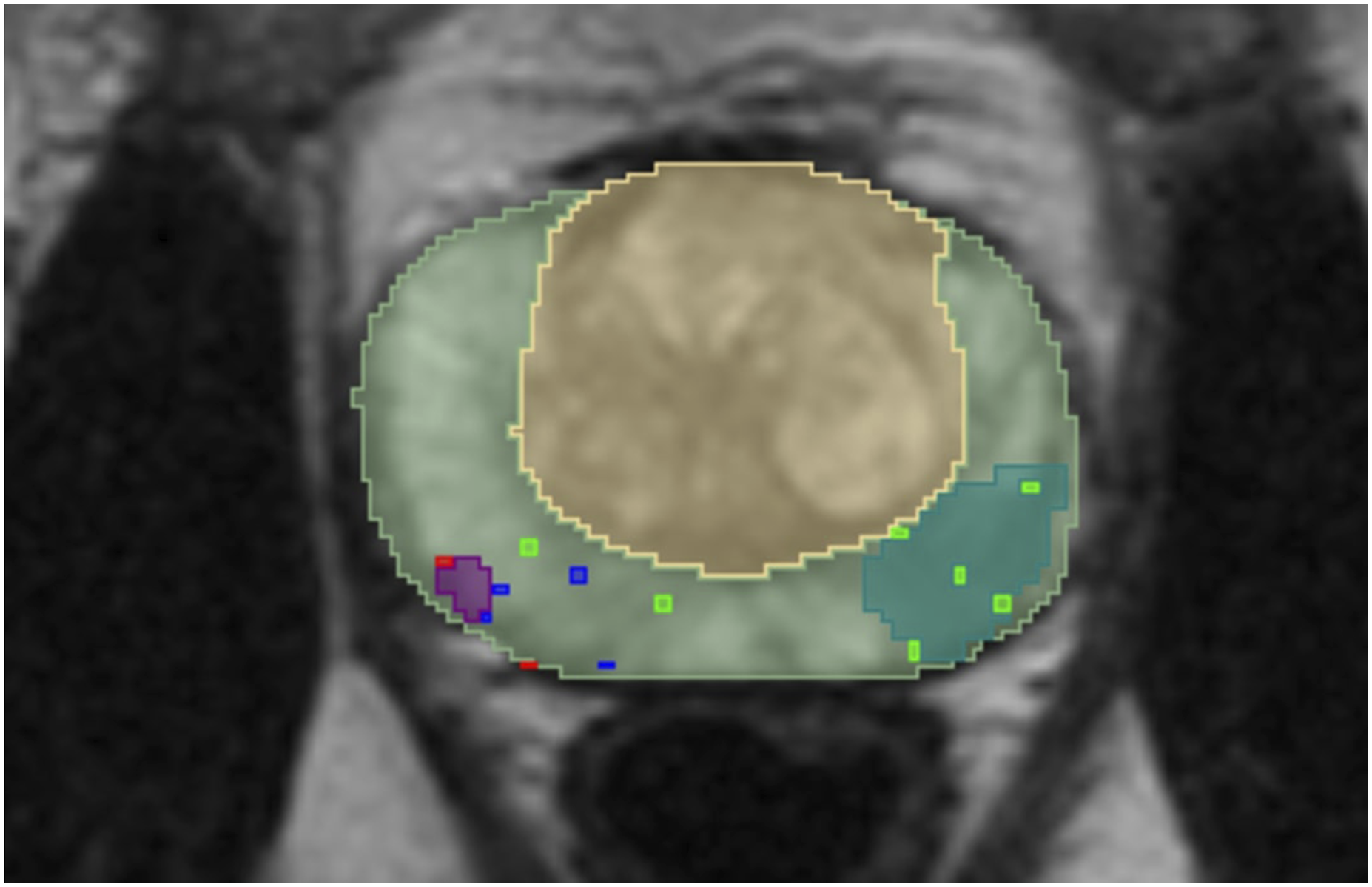
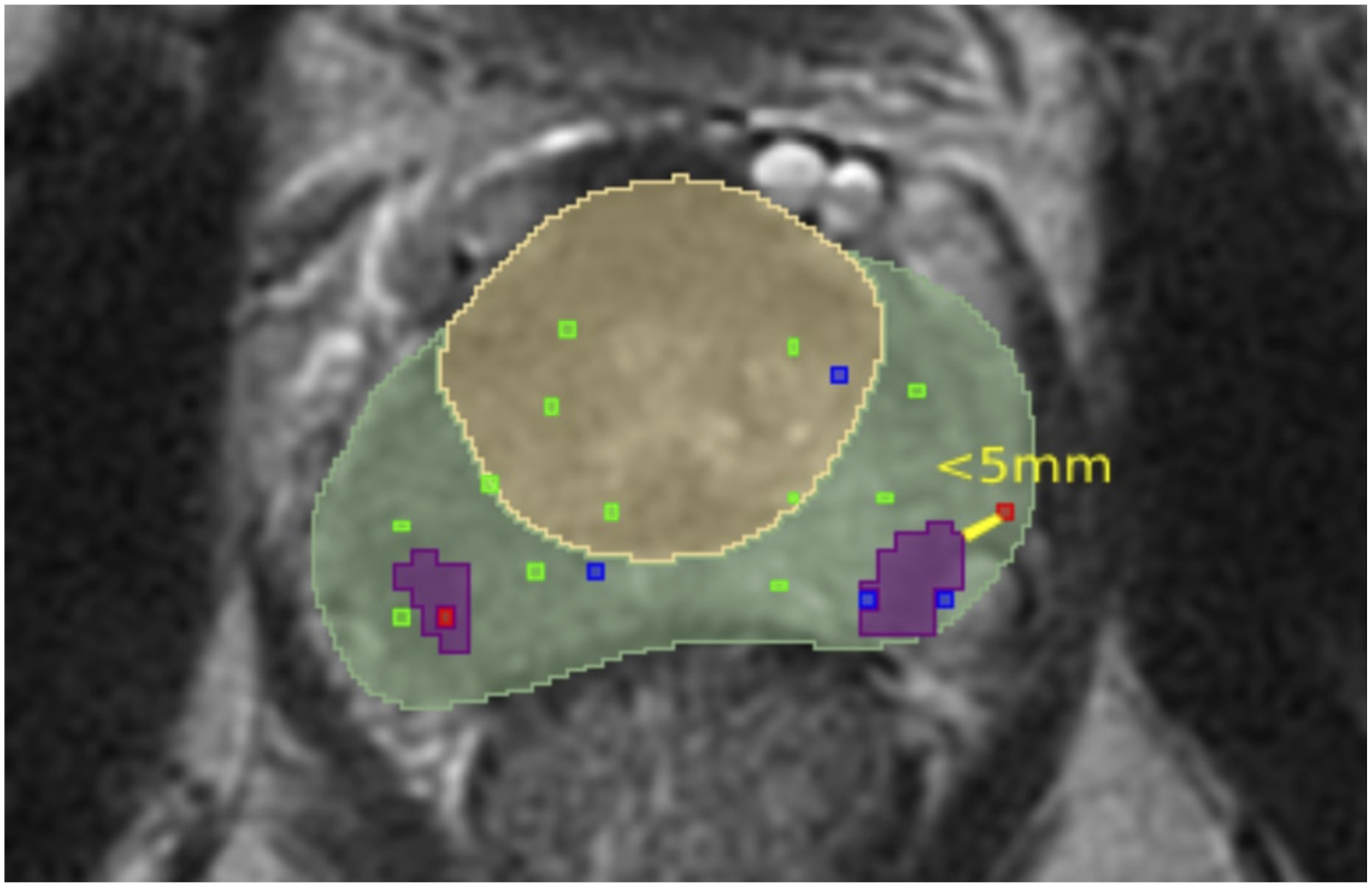
The ground truth is acquired by first calculating the connected components within manual segmentations. Each component corresponds to a single manually delineated lesion within the segmentation. The ground truth is considered manually delineated lesions confirmed by a malignant biopsy. The lesion is considered confirmed by a malignant biopsy if it satisfies one of the two conditions. The examples of those conditions are illustrated in Figures 9 and 10 where the light green area is a PZ, the yellow area is a TZ, green, blue, and red dots represent biopsies with benign, undetermined, malignant results respectively and the purple areas represent the manually delineated lesions which are considered as a ground truth while the areas coloured in the light blue/green colour are lesions which are not included in the ground truth. Those conditions are: • The manually delineated lesion intersects with a malignant biopsy mask. The example of a single slice, in which one lesion (purple area) satisfies this condition, is presented in Figure 9. • Malignant biopsy mask is located within a 5 mm radius around the manually delineated lesion. According to the articles
39
by Sahni et al. and
40
by Bhat et al. the prostatic biopsies are performed using a standard grid. The grid contains evenly spaced holes with a 5 mm distance. Hence, this spacing can introduce an error of a 5 mm radius. The example of a single slice, in which one lesion (purple area) satisfies this condition and another satisfies the first condition, is presented in Figure 10. The example of a single slice, where one manually delineated lesion intersects with a malignant biopsy mask while the other does not. The example of a single slice, in which one manually delineated lesion is closer than 5 mm from the malignant biopsy.
The distribution of cancerous lesions across prostate zones.

Lesion size statistics. The sizes are reported in mm3.

The prostate zone masks are acquired using the SM4TZPZS workflow, which was proposed by Vaitulevičius et al. in the article.
7
However, the acquired prostate zones have faults – there are gaps between the zones, the zones overlap. These faults are caused by limitations of the deep neural network models, such as prediction uncertainty at zone boundaries and partial volume effects. As a result, these errors can negatively affect the performance of compared ensembles and the baseline model. An example of such faults is presented in Figure 11 where the yellow area represents a TZ and the green area represents a PZ. Therefore, a further post-processing of the workflow result is designed in this study. The post-processed PZ is acquired using a function (3). Meanwhile, a post-processed TZ is acquired using a function (4). A single slice of unprocessed SM4TZPZS workflow result where green area represents PZ and yellow area represents TZ.
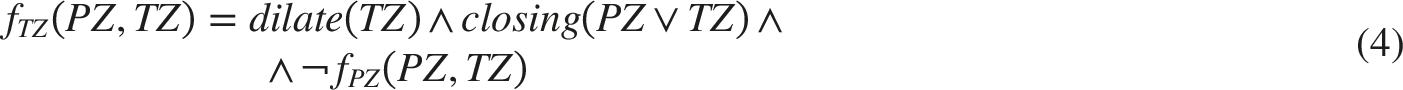

In these functions, dilate is a dilation operation with a ball kernel of 5-pixel radius. Furthermore, closing is a binary morphological closing operation, which is defined as a dilation operation followed by an erosion operation. The erosion operation was introduced in the paper 32 as well as the dilation operation. The dilation of PZ mask in the function (3) fills small gaps that occur between PZ and TZ regions. Next, the conjunction with the closing of the union of the unprocessed PZ and TZ masks ensures that the dilation only fills gaps between the zones and does not extend into unrelated areas. Finally, the conjunction with the negated TZ mask guarantees that the post-processed PZ does not overlap with the unprocessed TZ. Meanwhile, the dilation of TZ mask in the function 4 fills the remaining small gaps between the zones. The following conjunction with the closing of the union of the unprocessed zones restricts the effect of dilation to the region between the zones. The final conjunction with the negated and processed PZ ensures that the processed TZ does not overlap with the processed PZ. The ball kernel of a 10-pixel radius is used for binary morphological closing operation. The shape and size of the kernels of dilation and morphological closing operations are empirically selected to fill small gaps while preserving the anatomical shape of the prostate zones.
An example of post-processed prostate zones is provided in Figure 12 where the yellow area is a TZ, the green area represents a PZ, and the blue area is a prostate mask provided by NVI. A single slice of post-processed SM4TZPZS workflow result where green area represents PZ, yellow area represents TZ and blue area represents prostate gland.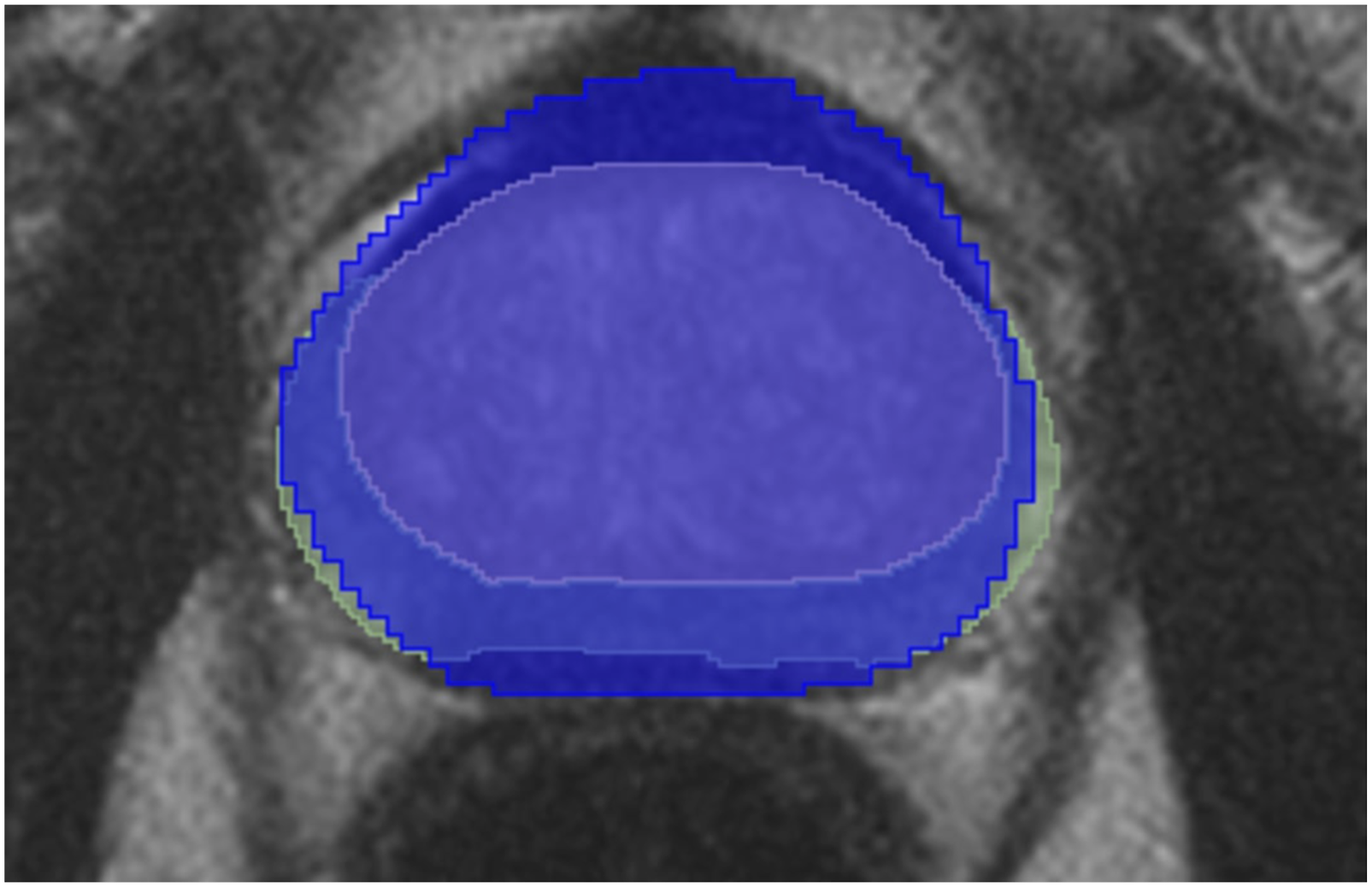
A lot of post-processed prostate zones contain regions that do not intersect the prostate mask. Therefore, all the post-processed prostate zone masks are further refined by acquiring the intersection between the prostate mask and the prostate zone mask. The refined prostate masks are used for training of the deep neural network models, while unrefined ones are used for the evaluation of ensembles and the baseline model.
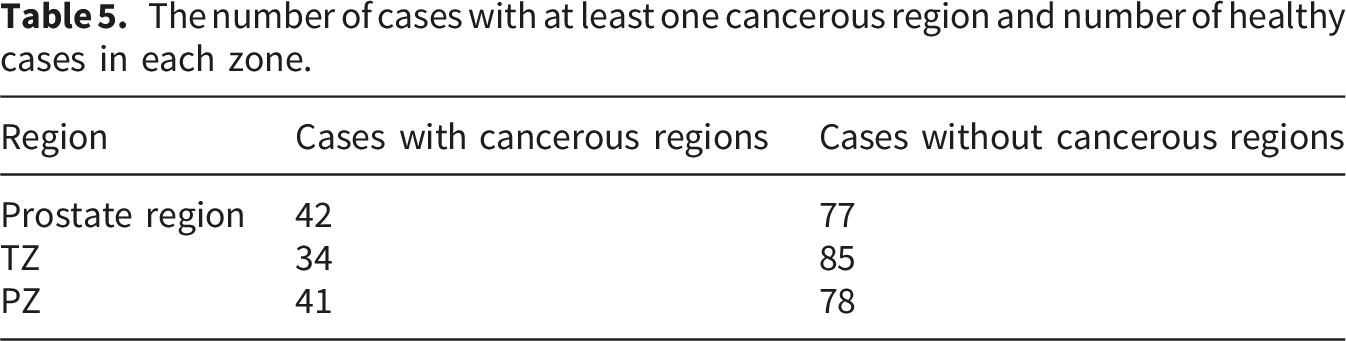
The number of cases with at least one cancerous region and number of healthy cases in each zone.

Experiment setup
As mentioned in the previous section, the dataset is imbalanced, as cases with a cancerous region are under-represented. To address this issue, the evaluation of ensembles and the baseline model consists of performing 5-fold cross-validation. Patients with cancerous regions are evenly distributed across the folds. 5-fold cross-validation is performed for each ensemble and the baseline model with the same folds. Thus, all deep neural network models are trained 5 times, each time using a different fold. Consequentially, the ensembles and the baseline model are evaluated 5 times as well, each time using a different fold. Four of the five resulting folds contain 24 patients. Therefore, for each fold models used in the ensembles are trained on 95 cases. For each fold the resulting ensembles are tested on 24 cases. The last fold consists of 23 patients. Hence, for this fold each model used in the ensembles are trained on 96 patients and each resulting ensemble is tested on 23 cases.
All deep neural network models are trained using nnU-Net method. The training data augmentation consists of random rotations, random scaling, Gaussian noise, Gaussian blur, gamma correction and mirroring. Each deep neural network model is trained using a batch size of 4 for 1000 epochs. Optimization is performed using stochastic gradient descent with Nesterov momentum (μ == 0.99) and an initial learning rate of 0.01. The learning rate is updated at the beginning of each epoch using polynomial decay schedule, defined in the equation (5). In this equation lr0 denotes the initial learning rate, e represents the current epoch, E represents total number of epochs and lr
e
corresponds to the adjusted learning rate at epoch e. Each training is performed with a different random seed. All trainings and inference experiments are conducted on the Nvidia DGX1 GPU server equipped with 8xV100 system architecture.
Overall, 4 ensembles are evaluated and compared. They are described in Methods section as well as the baseline model, prostate (All, Zones). The evaluation is performed for each patient separately by calculating the DSC and the DSC τ . Unfortunately, if the patient does not have a cancerous region, then both DSC τ and original DSC cannot be calculated. Therefore, for each ensemble and baseline model, the percentage of patients (false positive patients), for whom the result of the ensemble or the baseline model includes a cancerous region while the ground truth does not, is computed. This percentage is calculated for each fold separately as well. The dataset contains 77 patients whose ground truth does not contain a cancerous region. Hence, only these 77 patients are used for the calculation of false positive patients metric. Meanwhile, the DSC τ and the original DSC are calculated for 42 patients whose ground truth contained cancerous regions.
Results
Mean and standard deviation of prostate cancer segmentation results acquired from 5-fold cross-validation. These metrics are calculated on collected metrics DSC τ and DSC.
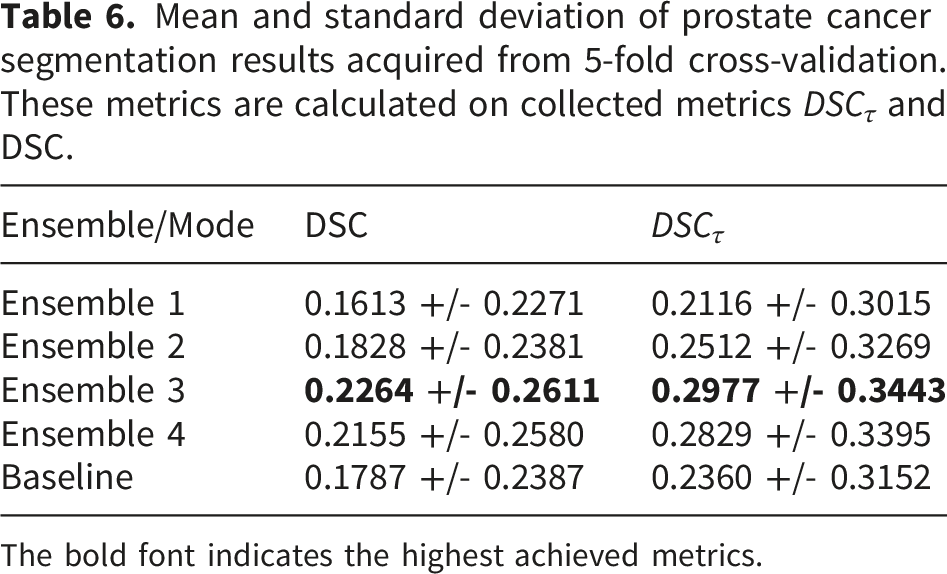
The bold font indicates the highest achieved metrics.
In addition, Friedmann test results indicate a significant difference between the compared ensembles and the baseline model. The p-value when comparing DSC τ is 0.003107, while the p-value of DSC comparison is 0.00067. Both of those values are lower than the significance level, 0.05 (confidence level 95%). Therefore, the hypothesis H0 is rejected, which indicates a statistically significant difference between the compared ensembles and the baseline model.
Conover post hoc test results. The tests are performed on collected metrics DSC τ and DSC. The metric on which the p-value is acquired is indicated in parenthesis.

The bold font indicates the p-values that confirm the statistically significant difference.
Since the p-values are below the significance level, H0 is rejected, indicating that there is a statistically significant difference between the results of the baseline model prostate (All, Zones) and Ensembles 3, 4. Meanwhile, there is no statistically significant difference between the results of the baseline model prostate (All, Zones) and Ensembles 1, 2 as p-values are higher than the significance level and therefore, H0 is not rejected. The other results of the Conover post hoc test show that there is no statistically significant difference between the results of Ensembles 3 and 4 as well as the results of Ensembles 1 and 2. However, the DSC metrics of Ensembles 1, 2 and the results of Ensembles 3, 4 statistically significantly differ while DSC τ metrics - do not.
Moreover, the DSC
τ
and DSC, calculated from 5-fold cross-validation results, are visualized as boxplots in Figure 13. The vertical axis indicates compared ensembles and the baseline model, while the vertical one indicates aggregated DSC and DSC
τ
. The orange boxplots indicate DSC
τ
while the blue ones indicate DSC. The yellow dots on boxplots represent the mean DSC and mean DSC
τ
for their respective boxplots. The boxplot indicates the same conclusions as Table 6. Boxplots of DSC (blue) and DSC
τ
(orange) prostate cancer segmentation results acquired from 5-fold cross-validation.
The percentage of patients for whom the result of the ensemble or the baseline model includes a cancerous region while the ground truth does not (false positive patients).
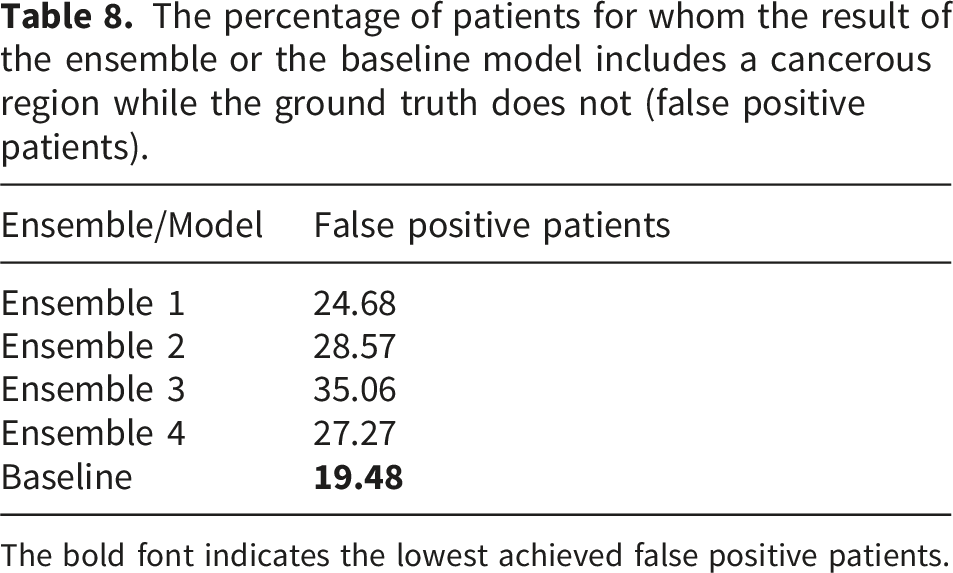
The bold font indicates the lowest achieved false positive patients.
Distributions of predicted lesions across patients. Each distribution is expressed as the number of patients for whom the ensemble or the baseline model generated the corresponding number of lesions.

Moreover, the compared ensembles tend to predict more than two lesions per patient, even though the deep neural network models were trained on a dataset in which the maximum number of lesions per patient is two. However, this overestimation affects only two to three patients per ensemble. The baseline model overestimates even less, predicting four lesions for only a single patient.
Distributions of missed lesions across patients. In total 42 patients had at least one lesion. Each distribution is expressed as the number and the percentage of patients for whom the ensemble or the baseline model missed the corresponding number of lesions.
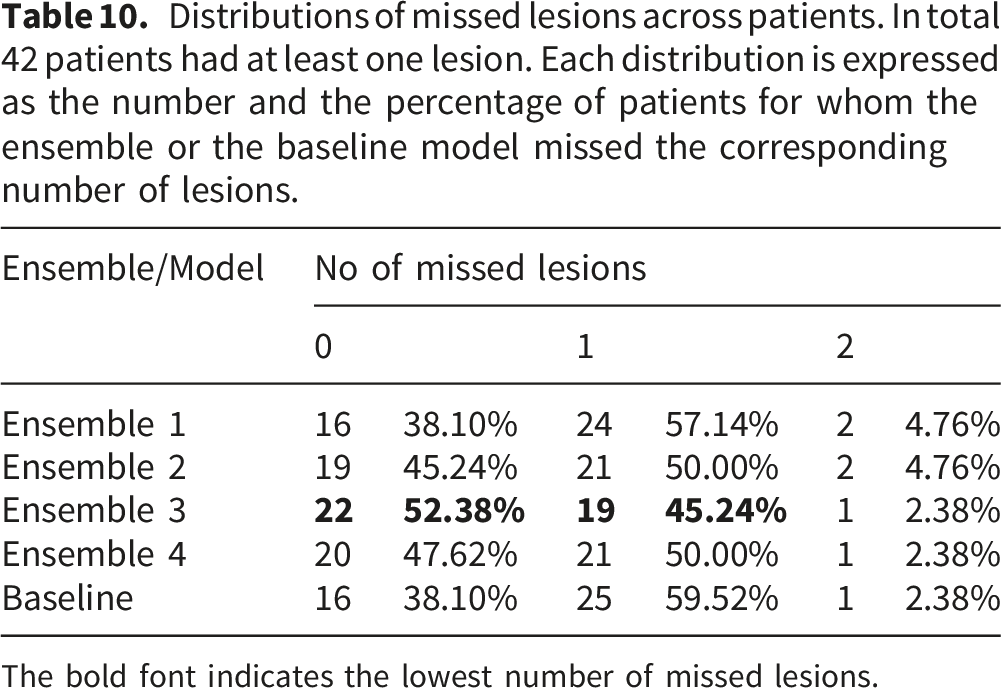
The bold font indicates the lowest number of missed lesions.
Figure 14 presents representative single slice segmentation results for each proposed ensemble alongside the baseline model, highlighting instances in which the predicted lesion masks overlap with the ground truth. All examples are derived from the same patient and correspond to the same anatomical slice. Green areas indicate the outputs of the respective ensemble or the baseline model, while, blue areas represent the ground truth. The results and the ground truth is overlayed on the T2 sequence. Furthermore, Figure 15 shows single slice examples of the cases for whom all of the proposed ensembles, as well as the baseline model, failed to localize the lesion. Blue areas represent the ground truth, and the ground truth is overlayed on the T2 sequence. Single patient, single slice examples of segmentation results (green areas) produced by each ensemble and the baseline model, highlighting cases where the predictions overlap with the ground truth lesions (blue areas). The results are overlayed on the T2 sequence. The single slice examples of patients with lesions (blue areas) that none of the ensembles as well as the baseline model localized. The results are overlayed on the T2 sequence. The examples (a) and (b) are different patients.

Two cases were observed in which the deep neural network models produced fragmented lesion predictions. In the first case, the PZ (All) model and the baseline Prostate (All, Zones) model segmented the same lesion into two separate components. Consequently, Ensemble 2 and the baseline model also produced a fragmented version of this ground truth lesion. In contrast, Ensemble 3 and Ensemble 4 did not exhibit this fragmentation, as the disjunction of their constituent models yielded a single, continuous lesion. The second instance of fragmentation was caused solely by the TZ (All) model. As a result, Ensemble 2 and Ensemble 3 produced a fragmented lesion in this case. However, this lesion was not localized at all by Ensemble 1, Ensemble 2, Ensemble 4 and the baseline model. Overall, no additional cases of lesion fragmentation were observed, allowing to conclude that these two instances are outliers and that the proposed ensembles generally do not fragment lesions.
Distributions of spurious foci across patients generated by the compared ensembles and the baseline model. Each distribution is expressed as the number of patients exhibiting each number of generated spurious foci.

The bold font indicates the lowest number of generated spurious foci.
Distributions of spurious foci across patients generated by the compared ensembles and the baseline model. Each distribution is expressed as the percentage of patients (119) exhibiting each number of generated spurious foci.

The bold font indicates the lowest percentage of generated spurious foci.

The single slice examples of generated spurious foci (green areas) by each ensemble. The results are overlayed on the T2 sequence.
Discussion
The result of the presented experiment evaluates possible improvements to the solution of the prostate cancer segmentation task. The accurate solution of this task is important as it can greatly assist radiologists in evaluating mpMRI for further diagnostic assessment of prostate cancer.
In this paper, four ensembles are proposed for the prostate cancer segmentation task. Moreover, these ensembles are compared with the baseline model, prostate (All, Zones). For the comparison, a new metric, DSC τ , is introduced, which is a modified DSC, DSC τ , and it accounts for a segmentation error in the ground truth. The comparison is performed with both the DSC τ and the original DSC metric.
All of the proposed ensembles employ models trained with nnU-net method. The baseline model is trained with nnU-net method as well. Thus, each model’s architecture is equivalent. The main difference is the prostate zone within which the model segments prostate cancer. Each model can be ran in parallel as none of them depend on the output of the other. Hence, the difference between inference time of each ensemble can be negated with sufficient computational resources. However, the ensembles differ in the number of employed models. Thus, different ensemble requires larger computational resources. The largest ensemble is Ensemble 3, which employs 3 nnU-Net models. Meanwhile, the other 3 ensembles consist of only 2 models. Therefore, it is impractical to use Ensemble 3 if at least one of the other 3 ensembles achieve similar results.
Furthermore, the compared ensembles as well as the baseline model employ prostate zone masks. Therefore, the isolated quantitative effect of zonal information cannot be directly assessed within this study. However, the consistent use of zonal masks across ensembles ensures that performance differences are not confounded by unequal access to anatomical prior, allowing a more controlled evaluation of ensemble strategies. Moreover, prostate zone masks provide an explicit anatomical prior that guides the network toward region specific representations, which is particularly relevant given the known differences in cancer prevalence and imaging characteristics between prostate zones. In addition, the positive effect on prostate cancer segmentation accuracy was indicated by research presented in prior publications.
Both collected DSC τ and DSC indicate that Ensemble 3 achieves the most accurate results in prostate cancer segmentation tasks. This ensemble achieves 4.77% higher mean DSC than the baseline model prostate (All, Zones). Furthermore, Ensemble 3 achieves 6.17% higher mean DSC τ than the baseline model prostate (All, Zones). However, Ensemble 4 is only marginally less accurate than Ensemble 3. Ensemble 4 achieves 1.09% lower mean DSC than Ensemble 3. Moreover, Ensemble 4 achieves 1.48% lower mean DSC τ and 0.84% than Ensemble 3. The result of Ensemble 3 consists of the results of the three models, while the result of Ensemble 4 is acquired from two models. Therefore, Ensemble 4 uses fewer models, and both Ensemble 3 and Ensemble 4 can be considered as the most suitable solution to the prostate cancer segmentation task. Moreover, this observation is consistent with the results of the missed lesions analysis.
Meanwhile, the baseline model prostate (All, Zones) is the least erroneous when segmenting prostate cancer among patients. This model achieves the lowest percentage of false positive patients. Moreover, this finding is further supported by the analysis of generated spurious foci. These results indicate that the use of disjunction within the ensembles leads to an accumulation of false positives. This is consistent with the observation that Ensemble 3, which combines three deep neural network models rather than two, yields substantially more false positive cases than the other ensembles. Alternative ensemble strategies, such as majority voting, could potentially reduce false positive rates, although this would likely come at the cost of decreased accuracy. Furthermore, the prostate (All, Zones) model exhibits a reduction in false positive patient cases by less than 10% compared to the next two ensembles, which achieve the lowest percentage of false positive patients, Ensemble 1 and Ensemble 4. Considering that sensitivity in medicine is more important than specificity, the difference in false positive patients between Ensemble 1, Ensemble 4 and prostate (All, Zones) model can be considered insignificant.
Moreover, Ensemble 1 and Ensemble 2 perform significantly worse than Ensemble 3 and Ensemble 4. Furthermore, the accuracy of Ensemble 4 is only marginally worse than Ensemble 3. All ensembles, except Ensemble 4, employ TZ specific prostate cancer segmentation deep neural network model. Moreover, both Ensemble 3 and Ensemble 4 use model which segments prostate cancer in the whole prostate gland. These findings indicate that TZ specific prostate cancer segmentation models do not improve overall performance. This observation is consistent with prior studies. Moreover, it aligns with anatomical considerations, including the difficulty of distinguishing cancerous regions in TZ from benign hyperplasia nodules, the diffuse background heterogeneity and multinodularity of TZ, and the effects of stromal asymmetry on cancer segmentation within TZ.
Furthermore, Ensemble 1 accuracy is worse than the other ensembles and the baseline model. The difference between Ensemble 1 and the other ensembles, as well as the baseline model, is the use of zone specific imaging sequences in prostate cancer segmentation models. These results indicate that employing all three mpMRI sequences in prostate cancer segmentation models is preferable to using zone specific sequences. However, Ensemble 1 achieves the lowest false positive patients among compared ensembles. This suggests that the use of all imaging sequences may increase the likelihood of generating more spurious foci.
Lastly, mean DSC τ is higher than the original mean DSC. Ensemble 1 achieves 5.03% higher DSC τ than the original DSC. Ensemble 2 achieves 6.84% higher DSC τ than the original DSC. Ensemble 3 achieves 7.13% higher DSC τ than the original DSC. Ensemble 4 achieves 6.74% higher DSC τ than the original DSC. The model prostate (All, Zones) achieves 5.73% higher DSC τ than the original DSC. This result indicates that the DSC τ accounts for ground truth segmentation errors. The incremental contribution of the proposed metric lies in its ability to account for clinically acceptable boundary deviations that are penalized by the conventional DSC. In prostate cancer segmentation, small spatial discrepancies are common due to inter observer variability, yet these deviations often have minimal clinical impact. The conventional DSC treats all mismatches equally, regardless of their spatial magnitude. In contrast, the proposed DSC τ introduces a tolerance margin that considers small boundary offsets as acceptable matches. This allows the metric to distinguish between clinically negligible boundary deviations and truly significant segmentation errors. Although the ranking of the evaluated methods remains similar in our experiments, the DSC τ provides a more clinically meaningful interpretation of segmentation quality by reducing the impact of minor boundary inconsistencies. This is particularly relevant for prostate cancer segmentation, where small contour variations frequently occur even among expert annotations. Therefore, the proposed metric does not aim to replace the conventional DSC but rather to complement it by providing an evaluation that better reflects clinically acceptable segmentation accuracy.
Overall, considering that Ensemble 4 is the only ensemble that achieved competitive results when comparing DSC τ , DSC and false positive patients in the experiment presented in this paper, this research concludes that Ensemble 4 is the most suitable solution to the prostate cancer segmentation task. The achieved metrics are comparable to those reported in previous studies on prostate cancer segmentation, such as the work presented in Ref. 14. However, both the baseline model and the proposed ensembles exhibited high false positive rates and low DSC, indicating that neither approach currently provides a sufficiently accurate solution for prostate cancer segmentation. These limitations are likely influenced by the relatively small dataset size, which not only restricts the learned lesion characteristics but also introduces a bias toward smaller lesions, as reflected in the lesion size distribution. Consequently, the ensembles may have been predominantly trained on features associated with smaller lesions, potentially impairing their ability to accurately segment larger ones. While the precise impact of this imbalance is difficult to assess given the limited sample size, evaluating the proposed ensembles on substantially larger and more representative datasets is a critical direction for future work. Such evaluation could help mitigate false positives, clarify the effect of lesion size distribution, and better assess robustness across a wider range of lesion sizes. Moreover, if low DSC values and high false positive rates persist on larger datasets, the proposed ensembles may still be valuable for prostate cancer localization, in which case alternative performance metrics, such as the area under the receiver operating characteristic curve or average precision, may be more appropriate than DSC, as DSC penalizes small boundary discrepancies. Furthermore, prior studies on prostate cancer segmentation that utilized larger datasets, such as,18,19 reported notably higher performance metrics. Lastly, it should be emphasized that the primary objective of this study is the empirical comparison of the proposed ensembles with the baseline model, rather than the development of a clinically deployable prostate cancer segmentation method. The findings highlight the most feasible solution among the compared approaches for the prostate cancer segmentation task. Nevertheless, further validation on emerging large scale publicly available datasets is necessary.
The dataset was collected from 11 different institutions, which supports the generalizability of the findings across institutional settings. In addition, the data were acquired using a wide range of mpMRI scanners and imaging parameters, further enhancing the applicability of the results to heterogeneous clinical environments. However, the substantial variability in acquisition protocols may have negatively affected the accuracy of the evaluated ensemble. This limitation could be mitigated in future work by training and evaluating the models on a larger dataset.
The results obtained in the experiment of this paper indicate one further possible future research direction on the prostate cancer segmentation task, the incorporation of another mpMRI sequence, Dynamic Contrast Enhancement, into the result of the ensembles described in this paper.
Conclusion
This study demonstrates that the proposed ensembles incorporating prostate zone masks can significantly improve the accuracy of prostate cancer segmentation. In contrast, the use of zone specific mpMRI sequences did not yield additional performance gains. The highest segmentation accuracy was achieved by the ensemble combining two deep neural network models: one trained to segment cancerous regions across the entire prostate and another focused specifically on the peripheral zone. Despite these promising results, the experiments were conducted on a dataset of limited size, which negatively affect the accuracy of the compared ensembles. Consequently, further validation on larger datasets is required to confirm the robustness of the proposed approach. Finally, the results indicate that the proposed tolerant Dice similarity coefficient effectively accounts for high inter observer variability, highlighting its suitability as an evaluation metric for prostate cancer segmentation tasks.
Footnotes
Acknowledgements
The authors are thankful for the HPC resources provided by the ITOAC of Vilnius University.
ORCID iDs
Ethical considerations
The dataset is provided by the Lithuanian National Cancer Institute under the terms of the bioethical agreement (number 2020/5-1229-714).
Author contributions
Authors contributed equally to this work. Aleksas Vaitulevičius was responsible for software development, validation, formal analysis, investigation, writing the original draft and editing after the review and visualization. Jolita Bernatavičienė was responsible for conceptualization, validation and supervision. Ieva Naruševičiūtė was responsible for data curation, annotations, resources and clinical expertise. Jurgita Markevičiutė was responsible for validation and statistical expertise. Mantas Trakymas was responsible for data curation, resources, funding and clinical expertise. Povilas Treigys was responsible for conceptualization, methodology, project administration, funding acquisition, supervision and validation.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Publication/Research is funded by the Research Council of Lithuania under the Programme “University Excellence Initiatives” of the Ministry of Education, Science and Sports of the Republic of Lithuania (Measure No. 12-001-01-01-01 “Improving the Research and Study Environment”). Project No.: S-A-UEI-23-11.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data are not publicly available due to restrictions of the bioethical agreement (number 2020/5-1229-714).
Guarantor
Povilas Treigys serves as the guarantor of this study. He foresaw the study processes, including the study design, data collection, analysis, and the accuracy of the reported results. He affirms that all aspects of the research adhere to ethical and scientific standards and that the conclusions drawn are a true reflection of the study findings.


