Abstract
Objectives
Clinical notes contain rich but unstructured information that is challenging to analyze at scale. Standardized terminologies such as Systematized Nomenclature of Medicine—Clinical Terms (SNOMED CT) support semantic interoperability by providing consistent representations of clinical concepts; however, the relationship between empirical concept co-occurrence in clinical documentation and embedding-based semantic similarity remains poorly understood. This study examines how SNOMED CT concept co-occurrence patterns relate to semantic similarity in embedding space and explores the value of jointly analyzing these signals for semantic characterization of clinical narratives and exploratory phenotyping.
Methods
We analyzed SNOMED CT–annotated clinical notes from the MIMIC-IV database. Concept co-occurrence within notes was quantified using Normalized Pointwise Mutual Information (NPMI), while semantic similarity was computed using cosine similarity between pretrained biomedical concept embeddings. We examined relationships between co-occurrence and semantic similarity across temporal stages of hospitalization, clinical documentation types, and concept frequency strata.
Results
Concept co-occurrence and embedding-based semantic similarity exhibited a weak but consistent positive association, indicating that frequently co-documented concepts are not always semantically proximate. Embeddings captured clinically meaningful associations that were infrequently documented together, highlighting latent relationships beyond documentation frequency. Temporal analysis showed more diffuse concept associations in early notes and stronger, more coherent relationships in later stages of care.
Conclusion
Co-occurrence statistics and embedding-based semantic similarity provide complementary views of clinical documentation. While co-occurrence reflects documentation practices and workflow, embeddings capture deeper conceptual relationships grounded in clinical context. Integrating these approaches enables improved understanding of clinical narratives.
Introduction
Clinical notes are among the most information-dense components of the electronic health record (EHR), capturing the full spectrum of patient care—observations, diagnoses, procedures, laboratory interpretations, and clinician reasoning. Unlike structured fields such as diagnosis codes or laboratory values, these free-text narratives provide nuanced insights into disease progression, care coordination, and clinician judgment. They represent an invaluable data source for downstream tasks such as clinical decision support, cohort identification, and outcome prediction. 1 However, the unstructured, heterogeneous, and high-volume nature of clinical text poses significant challenges for computational analysis. Clinicians often use abbreviations, idiosyncratic phrasing, and context-dependent terminology, making it difficult for automated systems to extract standardized meaning at scale. Efficient and semantically grounded methods are therefore essential to transform these unstructured narratives into actionable knowledge.
Standardized medical terminologies such as the Systematized Nomenclature of Medicine—Clinical Terms (SNOMED CT) provide a formalized vocabulary for representing clinical information. 2 SNOMED CT offers hierarchical relationships between over 350,000 active concepts encompassing diseases, procedures, findings, and body structures, enabling interoperability and consistent semantic representation across healthcare systems. By aligning clinical documentation with standardized terminologies, researchers can perform large-scale, reproducible analyses that transcend institutional or linguistic variation. Yet, the mere presence of standardization does not resolve the inherent complexity of clinical data. Extracting meaningful relationships, identifying latent patterns, and understanding the contextual semantics among concepts remain ongoing challenges, particularly in narrative notes that were not designed for computational interpretation.
Recent advances in natural language processing (NLP) have revolutionized biomedical text analysis through the development of pretrained language models and domain-adapted embeddings. Models such as ClinicalBERT, 3 BioBERT, 4 and SapBERT 5 have demonstrated exceptional capability in capturing subtle linguistic and semantic relationships within biomedical corpora. These embedding models represent concepts as dense vectors in high-dimensional space, allowing for quantification of semantic similarity and contextual relatedness. Such representations have proven effective in tasks including named entity recognition, relation extraction, and ontology alignment. However, the integration of these embedding-based methods with structured medical terminologies like SNOMED CT—particularly for large-scale, real-world EHR data—remains underexplored. Most prior work has focused on concept-level classification or entity extraction, leaving open questions about how standardized terminologies and embedding spaces jointly encode clinical semantics.
A promising and underutilized direction involves analyzing the co-occurrence of SNOMED CT concepts within clinical notes and relating these empirical patterns to semantic similarity in embedding space. Co-occurrence patterns—where two concepts repeatedly appear together in the same document—can serve as a proxy for clinical reasoning, reflecting associations between symptoms, diagnoses, procedures, and outcomes. Frequent co-occurrence of semantically similar concepts may reinforce established diagnostic or therapeutic pathways (e.g. “chest pain” and “myocardial infarction”), while frequent co-occurrence of semantically distant concepts could indicate atypical presentations, comorbidities, or gaps in documentation. Conversely, concepts that are semantically related but rarely co-occur may represent underdocumented associations or opportunities for improving note completeness. Understanding these relationships can support multiple downstream applications: improving the accuracy of EHR annotation, suggesting missing or inconsistent documentation, guiding ontology refinement, and enhancing patient phenotyping for research and population health management.
This study leverages the Medical Information Mart for Intensive Care IV (MIMIC-IV) database, 6 a large, de-identified repository of clinical data encompassing millions of notes from intensive care settings. By annotating these notes with SNOMED CT concepts, we perform a large-scale quantitative and semantic analysis of concept co-occurrence patterns. We specifically investigate how statistical co-occurrence measures (e.g. normalized pointwise mutual information, NPMI) correspond to embedding-based semantic similarity derived from pretrained biomedical language models. Our central questions include (1) Do frequently co-occurring SNOMED CT concepts also cluster semantically in embedding space? (2) Can semantic similarity be leveraged to suggest potentially missing or underdocumented concepts within notes? (3) Do embedding-based relationships align with clinically recognized associations, and can they uncover novel or atypical patient subgroups?
To address these questions, we conduct a series of complementary analyses that integrate statistical, temporal, and semantic perspectives on concept co-occurrence. Specifically, our contributions are as follows:
A large-scale quantitative analysis of SNOMED CT concept co-occurrence patterns and embedding-based semantic similarity in SNOMED-annotated MIMIC-IV clinical notes. An embedding-driven clustering analysis that reveals clinically interpretable semantic groupings of SNOMED CT concepts, including symptoms, laboratory measurements, diagnoses, and procedures.
Collectively, these contributions aim to bridge the gap between standardized clinical terminologies, modern embedding-based representations, and real-world clinical data analysis. By combining co-occurrence statistics with semantic embeddings, this study provides new insights into how clinicians document patient care, how clinical knowledge is implicitly structured within EHR narratives, and how these patterns can be leveraged to improve documentation quality, interoperability, and patient-centered analytics in healthcare.
Related work
The intersection of standardized clinical terminologies, semantic embeddings, and large-scale EHR analysis has evolved substantially over the past decade. Numerous studies have leveraged the MIMIC database in conjunction with SNOMED CT or other ontologies to explore clinical documentation, predictive modeling, and semantic relationships. Johnson et al. 7 first introduced the MIMIC-III dataset, later expanded as MIMIC-IV, 6 establishing an open and reproducible foundation for intensive care research. Since its release, MIMIC has enabled diverse studies on phenotyping, 8 risk prediction, 9 and temporal representation learning, 10 underscoring its role as a cornerstone for data-driven clinical informatics.
SNOMED CT-based analyses in clinical informatics
SNOMED CT provides a standardized and hierarchically structured terminology for representing clinical concepts, offering a basis for consistent semantic annotation and interoperability. Several researchers have integrated SNOMED CT into clinical text mining pipelines for concept extraction, data harmonization, and reasoning. For example, Sohn et al. 11 employed SNOMED CT to identify acute kidney injury patterns in MIMIC-III, using association rule mining to reveal concept pairs predictive of patient deterioration. Similarly, Yu et al. 12 combined SNOMED annotations with traditional machine learning models such as logistic regression and random forests to predict in-hospital mortality, demonstrating that concept-based representations outperform structured EHR features alone.
Beyond prediction, SNOMED CT has also been used for cohort identification, adverse event detection, and semantic data integration. Koopman et al. 13 evaluated concept mapping algorithms to standardize Australian hospital discharge summaries to SNOMED CT, revealing the challenges of polysemy and domain-specific terminology. Funk et al. 14 compared automated annotators such as MetaMap, ConceptMapper, and cTAKES, emphasizing parameter sensitivity in large-scale SNOMED-based annotation. More recent frameworks, including MedCAT, 15 have further advanced contextual SNOMED annotation by combining unsupervised learning with dictionary-based concept linking, improving recognition accuracy in real-world EHRs.
Semantic embeddings and clinical language models
Parallel to ontology-driven approaches, pretrained language models have transformed the landscape of biomedical NLP. Domain-specific embeddings such as BioBERT, 4 ClinicalBERT, 3 PubMedBERT, 16 and SapBERT 5 have achieved state-of-the-art results in tasks ranging from named entity recognition to relation extraction. SapBERT, in particular, introduced self-alignment pretraining using the Unified Medical Language System (UMLS) to align synonymic biomedical terms in embedding space, demonstrating the feasibility of ontology-informed embedding learning. Wang et al. 17 combined ClinicalBERT representations with SNOMED CT annotations in MIMIC-III, enabling patient clustering based on semantic similarity and uncovering groups associated with mortality and readmission outcomes. This work highlighted the synergy between deep contextual embeddings and standardized terminologies for semantic analysis.
Beam et al.18,19 further integrated multimodal embeddings by combining structured variables, clinical text, and SNOMED-derived concepts, uncovering latent clinical relationships through cosine similarity and hierarchical clustering. In related work, Zhang et al. 20 used contrastive learning to align structured EHR concepts and unstructured text, improving representation coherence across modalities. These studies collectively underscore the promise of embedding-based models in capturing both linguistic and clinical semantics.
Ontology-guided and knowledge-infused models
While pretrained embeddings offer rich contextual representations, purely data-driven models may overlook domain hierarchies and logical constraints encoded in ontologies. To address this limitation, ontology-guided or knowledge-infused learning approaches have emerged. OntoBERT 21 and OAG-BERT 22 incorporate ontology structures directly into transformer architectures, improving interpretability and domain alignment. In biomedical NLP, works such as Ontology-Aware Clinical Language Modeling 23 and Knowledge Graph–Enhanced BERT 24 have demonstrated that injecting ontology relationships into embeddings enhances concept disambiguation and cross-hierarchy generalization.
Within clinical informatics, several recent efforts have explicitly connected ontology-guided embeddings with retrieval-augmented generation (RAG) and reasoning. Sharma et al. 25 proposed OG-RAG, which leverages ontology-guided retrieval to improve factual grounding in clinical language generation, while Bran et al. 26 demonstrated ontology-augmented retrieval for scientific discovery. Kainer et al. 27 extended this paradigm to ontology-guided annotation, applying retrieval-augmented LLMs for SNOMED-based concept suggestion in biomedical text. These studies collectively indicate a growing trend toward integrating symbolic knowledge with neural embeddings to improve accuracy and interpretability.
Temporal and co-occurrence analyses in clinical text
Temporal modeling of clinical concepts has been another active research area. Miotto et al. 28 demonstrated that unsupervised representation learning of longitudinal EHR data can capture disease progression patterns and predict future health outcomes. Similarly, Solares et al. 29 applied temporal embeddings to EHR sequences, identifying progression trajectories for chronic conditions. In the context of concept co-occurrence, Chen et al. 30 applied association rule mining and topic modeling on SNOMED CT–annotated notes to uncover latent concept networks reflecting clinical reasoning. Co-occurrence networks have also been used for ontology enrichment: Choi et al. 31 used recurrent attention models to discover relationships between diagnosis codes, while Kamath et al. 32 examined interpretability of co-occurrence clusters in MIMIC-IV to identify comorbidity structures.
Despite this progress, relatively few studies have explicitly linked concept co-occurrence frequency with semantic similarity in embedding space. Most prior research has focused on patient-level embedding aggregation, phenotyping, or predictive modeling, rather than the fine-grained exploration of intra-note semantic relationships. The correlation between statistical co-occurrence measures—such as NPMI—and embedding-based similarity remains largely unexplored. Furthermore, the potential to leverage this relationship for identifying missing documentation, suggesting concept augmentation, or detecting atypical semantic clusters has not been systematically investigated.
Positioning of the present study
To our knowledge, no prior work has systematically examined how empirical concept co-occurrence statistics and embedding-based semantic similarity jointly characterize SNOMED CT concepts within real-world clinical documentation. This study addresses this gap by integrating SNOMED-based co-occurrence analysis with modern embedding similarity measures applied to large-scale clinical notes. By quantifying the alignment and divergence between statistical association and semantic relatedness, we provide insight into how clinical concepts are documented together and how their relationships are implicitly represented in embedding space. This integrative perspective offers a foundation for improved semantic analysis of clinical narratives and supports downstream applications such as documentation quality assessment, exploratory phenotyping, and embedding-informed clinical informatics research.
Concept co-occurrence and semantic structure in clinical notes
This work builds directly on prior analysis of SNOMED CT concept co-occurrence patterns in clinical documentation, particularly the study by Noori et al., 33 which examined statistical co-occurrence relationships among SNOMED CT concepts extracted from MIMIC-IV clinical notes. That study demonstrated that concept co-occurrence networks capture meaningful clinical structure and reflect documentation practices across note types and clinical contexts. By analyzing co-occurrence frequency and association strength, the authors showed that concept-level relationships can be leveraged to identify salient clinical themes and support downstream phenotyping and exploratory analysis.
The present study extends this prior work in several important ways. First, while the earlier analysis focused primarily on empirical co-occurrence statistics, the present article explicitly integrates embedding-based semantic similarity derived from pretrained biomedical language models. This enables a direct comparison between how often concepts are documented together and how closely related they are in semantic embedding space. Second, the present study introduces a temporal dimension, stratifying clinical notes into early and late stages of hospitalization to examine how concept associations evolve over the course of care. This temporal analysis reveals systematic differences in documentation patterns that are not captured by static co-occurrence networks alone.
Additionally, this work expands beyond descriptive co-occurrence analysis by incorporating embedding-driven clustering and concept suggestion experiments, demonstrating how semantic similarity can surface clinically meaningful relationships that are infrequently co-documented. Whereas the earlier work emphasized documentation structure and co-occurrence topology, the present study focuses on the alignment and divergence between statistical association and semantic relatedness, highlighting their complementary roles in understanding clinical narratives. Together, these contributions position the present article as a semantic and methodological extension of prior SNOMED CT co-occurrence analysis, bridging ontology-driven statistics with modern representation learning to support richer clinical interpretation and downstream applications.
Methods
Dataset: MIMIC-IV clinical notes
The MIMIC-IV database is a publicly available, de-identified dataset comprising detailed health-related information on patients admitted to intensive care units at the Beth Israel Deaconess Medical Center. A central component of MIMIC-IV is its corpus of unstructured clinical notes, stored in the
MIMIC-IV contains over 400,000 clinical notes authored between 2008 and 2019, covering more than 40,000 hospital admissions and approximately 70,000 patients. The notes span a diverse range of document types, including discharge summaries, progress notes, nursing notes, radiology reports, and emergency department documentation. Each note is timestamped and associated with metadata such as caregiver type and note category, enabling temporal and contextual stratification.
To support computational analysis of clinical text, a subset of these notes has been annotated with SNOMED CT concepts using a benchmark entity linking resource released as part of the DrivenData SNOMED CT Entity Linking Challenge. Annotations are provided at the character-span level, linking contiguous text spans to standardized SNOMED CT concept identifiers. For example, in note
In this study, we use the SNOMED CT–annotated clinical notes to examine concept co-occurrence patterns and semantic similarity relationships, with the goal of uncovering latent clinical structure embedded in real-world documentation practices.
Data preparation and preprocessing
SNOMED CT concept co-occurrences were extracted from annotated clinical notes in MIMIC-IV. Concept annotations were obtained from the DrivenData SNOMED CT Entity Linking Challenge dataset (SNOMED CT International Edition, November 2025 release), which provides precomputed, span-based entity linking annotations for a curated subset of MIMIC-IV clinical notes (version 2.2). These annotations were used directly, without additional manual curation or retraining, to ensure reproducibility and reflect realistic large-scale clinical NLP conditions.
Span-level annotations were aggregated to the note level by retaining the set of unique SNOMED CT concepts appearing within each clinical note. Note-level concept sets were constructed as described below, and co-occurrence statistics were computed from these aggregated representations. The resulting concept sets span a broad range of SNOMED CT hierarchies relevant to intensive care documentation, including diagnoses, procedures, laboratory tests, anatomical structures, and clinical findings.
Clinical notes in MIMIC-IV represent a heterogeneous mix of documentation types with differing clinical purposes and stylistic conventions. Rather than stratifying analyses by note category, we intentionally analyzed all note types together to characterize global patterns of concept co-occurrence and semantic similarity across the full course of patient care. The use of SNOMED CT provides a normalized semantic representation that mitigates surface-level variation in language and structure, allowing concept-level analyses to remain meaningful across heterogeneous documentation sources.
Preprocessing
Source datasets
The preprocessing pipeline integrated multiple data sources from MIMIC-IV v2.2 and the DrivenData SNOMED CT Entity Linking Challenge, including entity-level concept annotations, raw clinical note text and identifiers, hospital admission metadata, patient identifiers, and discharge summaries. These sources enabled linkage between annotated concepts, clinical notes, hospital encounters, and temporal context.
Annotation alignment and metadata integration
Span-level SNOMED CT annotations were merged with clinical note text using shared note identifiers, and further linked to hospital admission and patient metadata via admission and subject identifiers. This ensured that each annotated concept could be associated with a specific note, hospitalization, and patient.
Temporal feature construction
Admission and discharge timestamps were used to compute length of stay and relative note timing within each hospitalization. Notes were assigned normalized temporal positions and categorized into early or late documentation windows based on their relative timing, enabling stratified temporal analysis.
Concept normalization and deduplication
SNOMED CT concept identifiers were mapped to their fully specified names (FSNs), and parenthetical qualifiers were removed to reduce lexical variability. Within each note, duplicate mentions of the same concept were removed so that each concept was counted at most once per document.
Concept co-occurrence construction
For each note, all unordered pairs of unique SNOMED CT concepts were generated to form note-level co-occurrence sets. Co-occurrence counts were aggregated across notes separately for early and late temporal windows. Low-support concept pairs (e.g. pairs occurring only once) were optionally filtered to reduce noise in downstream analyses.
Marginal frequency and association statistics
All probabilities were estimated at the clinical note level. Span-level SNOMED CT annotations were aggregated into note-level concept sets, such that each concept was treated as present or absent within a note. Document-level marginal frequencies were computed for each concept, defined as the number of notes in which the concept appeared. These frequencies, along with joint co-occurrence counts, were used to compute association statistics as described below.
Final dataset assembly
The final analysis dataset combined co-occurrence counts, marginal frequencies, temporal labels, semantic similarity scores, and associated patient and admission identifiers. All analyses were conducted at the clinical note level rather than the patient level.
Mapping SNOMED CT concepts to clinical text
SNOMED CT concepts were mapped to clinical text using a span-based entity linking approach. Each concept mention was identified as a contiguous span of text within a clinical note, defined by character-level start and end offsets, and linked to a standardized SNOMED CT concept identifier using precomputed annotations from the DrivenData SNOMED CT Entity Linking Challenge dataset. This approach preserves an explicit alignment between unstructured clinical language and ontology concepts.
Span-level annotations were subsequently aggregated into note-level concept sets for co-occurrence analysis, such that each concept was treated as present or absent within a note. The raw clinical note text served as the input to pretrained language models for embedding generation, while SNOMED CT concepts functioned as semantic references linked to specific textual spans rather than as direct model inputs.
All probabilities were estimated at the clinical note level. Span-level SNOMED CT annotations were aggregated into note-level concept sets, such that each concept was treated as present or absent within a note. For a concept
Clinical notes were stratified into early and late hospitalization periods using admission time (
Semantic representation of SNOMED CT concepts
To analyze semantic similarity between SNOMED CT concepts, we generated vector representations using the pretrained Bio_ClinicalBERT model (emilyalsentzer/Bio_ClinicalBERT). Concept names were tokenized using the model’s native tokenizer, and contextualized token representations were extracted from the final hidden layer. Mean pooling across token embeddings (excluding special tokens) was applied to obtain a single fixed-length embedding per concept.
The model was used off-the-shelf without fine-tuning on MIMIC-IV data. All embeddings were generated in inference mode using
Strength of co-occurrence
Using the note-level probability estimates defined above, we computed NPMI for each concept pair. NPMI normalizes pointwise mutual information (PMI) to account for concept frequency and produces values bounded between
Formally, for two concepts 
NPMI values close to
Clustering analysis
Concept-level embeddings were clustered using the k-means algorithm applied to the fixed-length Bio_ClinicalBERT embeddings. Embeddings were standardized prior to clustering. The number of clusters was set to
Clusters were interpreted by examining representative and frequent concepts within each cluster, revealing clinically coherent groupings such as symptoms, laboratory measurements, cardiovascular conditions, anatomical structures, and general clinical observations. Clustering evaluation focused on semantic coherence and interpretability rather than predictive accuracy. No external clinical labels were assumed, and formal clinical validation was beyond the scope of this study.
Results
Dataset scope
The analyzed dataset consists of a curated subset of MIMIC-IV-Note v2.2 provided by the challenge organizers. In total, 204 clinical notes were analyzed, corresponding to 204 hospital admissions. Patient-level identifiers are not available in this curated subset. The notes span a de-identified time range corresponding to hospitalizations indexed between years 2112 and 2206 in the MIMIC-IV timeline.
SNOMED CT concept coverage
Across all notes, the dataset contains 51,574 span-level SNOMED CT concept mentions, corresponding to 5291 unique SNOMED CT concepts. After aggregation to the note level, the mean number of unique concepts per note is 158.6, with a median of 154.0 and an interquartile range of 123.0–191.3. Concept coverage spans a broad range of clinical domains, including laboratory measurements, physical examinations, diagnoses, procedures, anatomical structures, and clinical findings.
Concept frequency distribution
Concept usage follows a strongly right-skewed, long-tailed distribution. The most frequent concepts correspond to routine laboratory tests, vital sign measurements, and common physical examinations, while the majority of concepts appear infrequently. Figure 1 summarizes the most commonly occurring SNOMED CT concepts in the corpus and illustrates this characteristic frequency imbalance, which motivates the use of normalized association measures in downstream analysis.

Top 20 most frequently suggested Systematized Nomenclature of MedicineClinical Terms (SNOMED CT) concepts across all clinical notes. The
Co-occurrence descriptives
Using the note-level co-occurrence definition described in the Methods, we identified
Table 1 summarizes first-order descriptive statistics for the SNOMED CT–annotated MIMIC-IV clinical notes used in this study. The dataset exhibits high concept density per note and a strongly long-tailed distribution of concept co-occurrence pairs, with the majority of pairs occurring only once.
First-order descriptive statistics for the SNOMED CT–annotated MIMIC-IV clinical notes used in this study (DrivenData subset).

SNOMED CT: Systematized Nomenclature of MedicineClinical Terms; MIMIC-IV: Medical Information Mart for Intensive Care IV; IQR: interquartile range.
Figure 1 illustrates the most frequently surfaced SNOMED CT concepts produced by the embedding-based suggestion analysis across all clinical notes. For each note, a semantic representation was constructed by averaging embeddings of the SNOMED CT concepts documented in that note. The top-k nearest SNOMED CT concepts in embedding space (based on cosine similarity) were then retrieved as candidate suggestions.
The counts shown in the figure represent the number of clinical notes in which a given concept appeared among the top-k suggestions, aggregated across the corpus. Importantly, this figure does not measure predictive accuracy, recall, or agreement with a gold standard. Instead, it provides descriptive evidence of semantic plausibility, demonstrating that the suggestion mechanism tends to surface clinically meaningful concepts—such as laboratory measurements, vital signs, and common diagnoses—that are central to intensive care documentation.
Laboratory and diagnostic observations dominate, led by
Next, we examine similarity between co-occurring SNOMED CT concepts in clinical notes. Figure 2 shows that cosine similarity scores cluster around 0.75, indicating most pairs are moderately to highly related and often form coherent clinical narratives (e.g.

Histogram showing the distribution of cosine similarity scores between SNOMED CT concept pairs across all clinical notes in MIMIC-IV. Cosine similarity was computed using concept embeddings, capturing the semantic proximity between concept pairs. SNOMED CT: Systematized Nomenclature of MedicineClinical Terms; MIMIC-IV: Medical Information Mart for Intensive Care IV.
Clinical notes were classified as
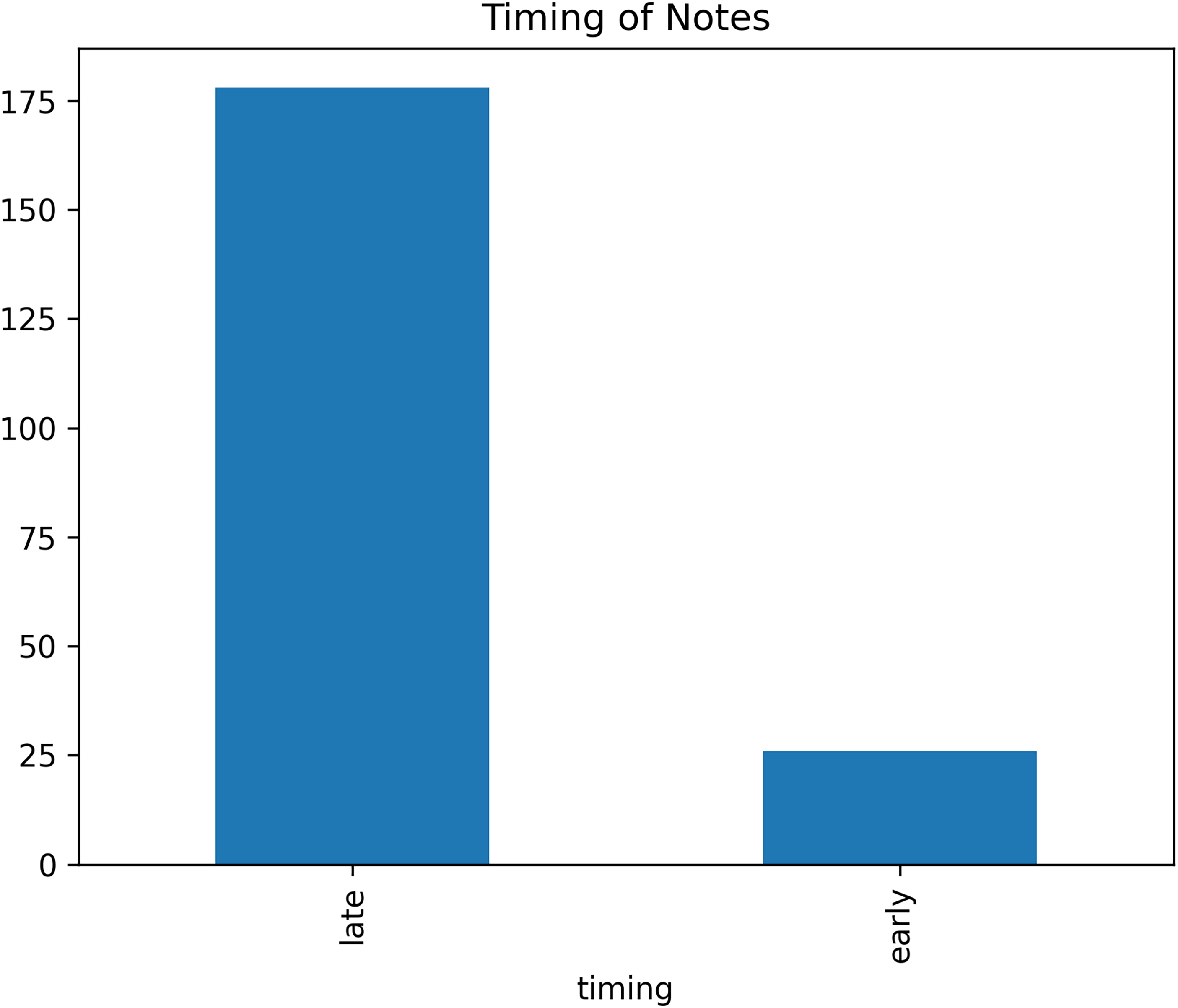
Bar chart showing the distribution of clinical notes labeled as “early” or “late” based on their timing relative to the total length of a hospital stay. Notes were labeled “early” if authored in the first 70% of the stay, and “late” otherwise.
We then computed NPMI scores for co-occurring SNOMED CT concepts, separating pairs from early versus late notes. Figure 4 shows distributions for rare pairs (fewer than five co-occurrences, blue) and frequent pairs (more than 15, orange). Rare pairs cluster near zero, suggesting chance associations or documentation noise, while frequent pairs are skewed right, reflecting strong clinical relationships such as symptom–diagnosis links. Notably, some rare pairs also show high NPMI values, pointing to underdocumented but clinically meaningful associations or niche subgroup patterns. A few extreme outliers (NPMI >0.8) represent canonical links (e.g.
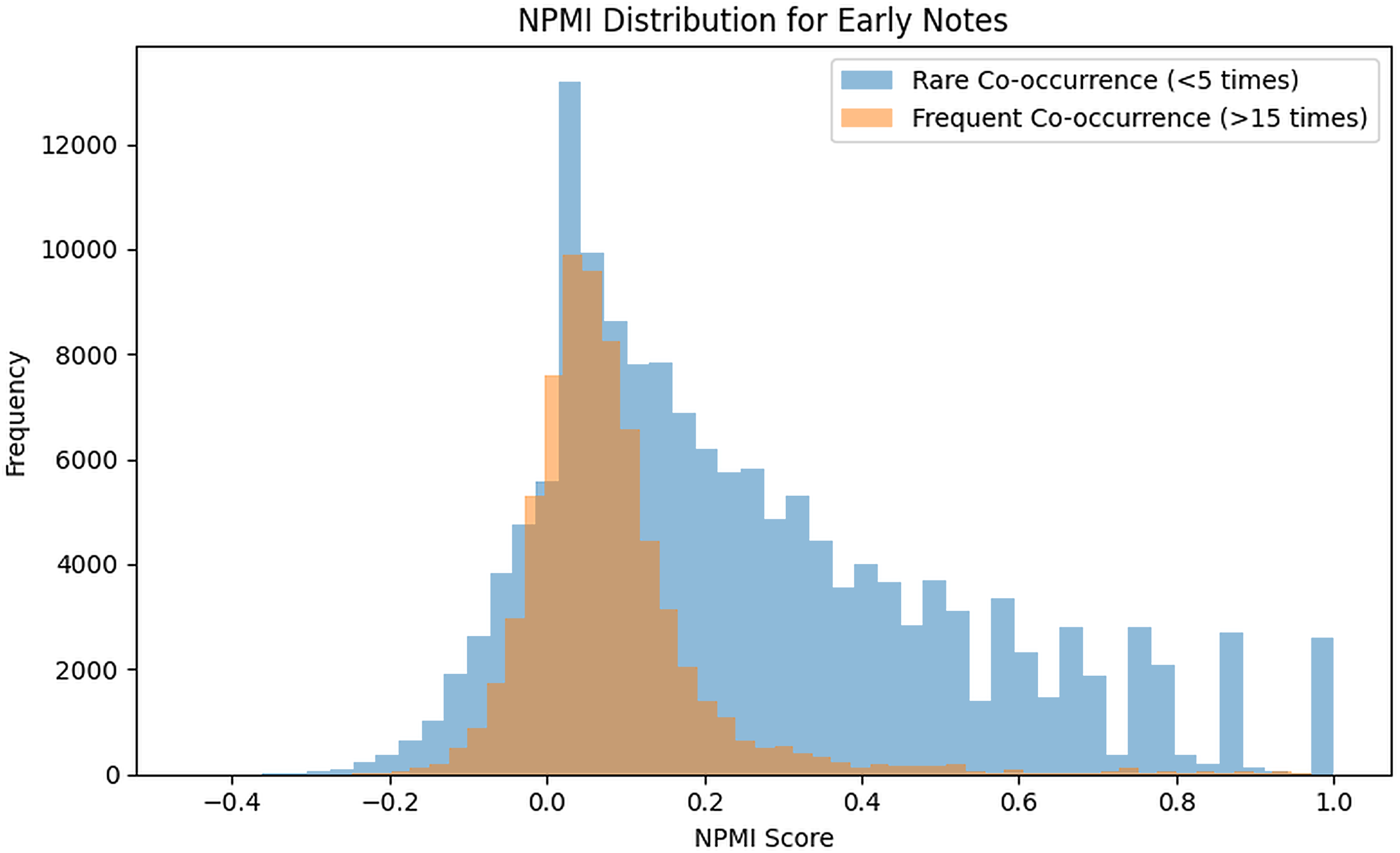
Histogram of NPMI scores for SNOMED CT concept pairs in early stage clinical notes. The distribution is stratified by co-occurrence frequency: rare co-occurrence (fewer than five notes) and frequent co-occurrence (more than 15 notes). NPMI: normalized pointwise mutual information; SNOMED CT: Systematized Nomenclature of Medicine-Clinical Terms.
We analyzed NPMI scores for concept pairs in the final 30% of hospital stays (late notes), comparing rare (blue) and frequent (orange) co-occurrences (Figure 5). Both groups peak near zero, with a sharper spike for rare pairs, indicating weak or chance associations. Frequent pairs, however, show a stronger rightward skew than in early notes, with more exceeding 0.2, reflecting stable clinical relationships common in discharge planning or chronic care. Rare pairs also display a right tail, suggesting underdocumented but meaningful associations or rare subgroup patterns. A small fraction of pairs have negative scores, pointing to mutually exclusive events or distinct care phases. Overall, late notes contain more high-frequency, high-association pairs, underscoring how concept relationships strengthen over the course of care and highlighting the utility of NPMI for temporal analysis of clinical documentation.

Histogram of NPMI scores for SNOMED CT concept pairs in late-stage clinical notes. The distribution is separated by co-occurrence frequency, showing rare co-occurrences (fewer than five notes, in blue) and frequent co-occurrences (more than 15 notes, in orange). NPMI: normalized pointwise mutual information; SNOMED CT: Systematized Nomenclature of Medicine-Clinical Terms.
Figures 6 and 7 illustrate the relationship between NPMI and cosine similarity of concepts in early versus late notes. In both cases, cosine similarity values are generally high (centered between 0.7 and 0.9), while NPMI values show a wider spread ranging from negative to positive associations.

Scatter plot illustrating the relationship between NPMI and cosine similarity for SNOMED CT concept pairs extracted from early clinical notes in MIMIC-IV. Each point represents a unique concept pair. NPMI: normalized pointwise mutual information; SNOMED CT: Systematized Nomenclature of Medicine-Clinical Terms; MIMIC-IV: Medical Information Mart for Intensive Care IV.

Scatter plot illustrating the relationship between NPMI and cosine similarity for SNOMED CT concept pairs extracted from late clinical notes in MIMIC-IV. Each point represents a unique concept pair. NPMI: normalized pointwise mutual information; SNOMED CT: Systematized Nomenclature of Medicine-Clinical Terms; MIMIC-IV: Medical Information Mart for Intensive Care IV.
Although the overall joint distributions of NPMI and cosine similarity appear visually similar across early and late notes, subtle qualitative differences are evident (Figure 8). Early notes show a broader spread of co-occurrence strength among semantically similar concepts, whereas late notes exhibit a tighter clustering around moderate positive associations. We interpret this pattern as reflecting more exploratory documentation early in hospitalization and more consolidated clinical narratives later in care.
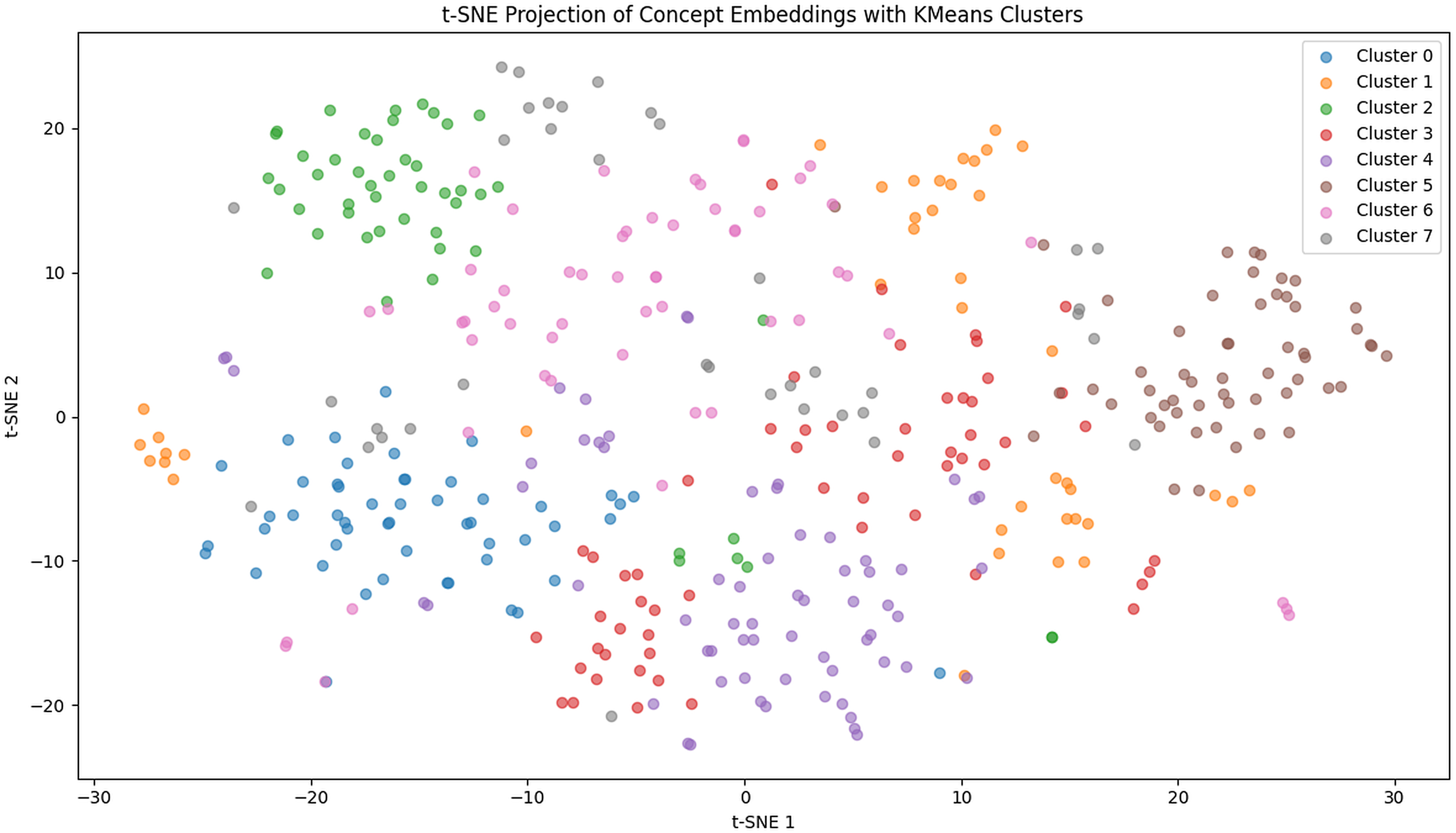
t-SNE projection of SNOMED CT concept embeddings annotated from MIMIC-IV clinical notes, color-coded by KMeans clustering (
Overall, while both note types show that high cosine similarity can occur across a wide range of NPMI values, early notes exhibit more variability, whereas late notes appear more consistent and concentrated. While the two metrics are positively associated, their differences also carry rich information that can help uncover latent clinical structures or documentation idiosyncrasies. This underscores the value of leveraging both co-occurrence statistics and embedding-based semantic measures in tandem when analyzing clinical concept relationships.
The concept clusters in Table 2 reveal clear semantic organization consistent with clinical knowledge, underscoring the effectiveness of embedding-based grouping.
A description of SNOMED concept clusters.

Overall, the clusters align with established clinical categories (e.g. cardiovascular, labs, and symptoms) while also highlighting domains with greater semantic overlap. This organization supports applications in phenotyping, ontology refinement, and decision support.
Our first research question asked whether frequently co-occurring SNOMED CT concepts also show high embedding similarity. Results indicate only a weak relationship: while some symptom–treatment pairs are both frequent and semantically close, many highly similar pairs rarely co-occur, showing that embeddings capture latent clinical associations beyond documentation frequency.
The second question explored whether semantic similarity can suggest missing concepts. By averaging embeddings of documented terms and retrieving nearby concepts, we generated contextually appropriate suggestions, many of which later appeared in follow-up notes. The third question asked if embedding-based similarity reflects clinically meaningful relationships. We found that many high-similarity pairs align with known diagnostic, procedural, or physiological links—even when absent from the SNOMED CT hierarchy—indicating that embeddings capture contextual relationships consistent with real-world practice. Finally, clustering embeddings revealed coherent clinical themes (e.g. symptoms, lab tests, anatomical structures, and diagnoses) that map to patient-level phenotypes and care patterns. This suggests embeddings can enrich patient stratification, ontology refinement, and nuanced clinical analysis. Overall, semantic similarity and co-occurrence provide complementary perspectives on documentation and patient trajectories. Embedding methods effectively identify clinically relevant relationships, support missing concept suggestion, and uncover potential anomalies or novel associations. Future work should include prospective validation and extend to broader embedding approaches and clinical contexts.
Illustrative examples of concept relationships
To provide intuition for how co-occurrence statistics and embedding-based similarity reflect real-world documentation patterns, we highlight several representative concept relationships observed in the data. These examples are intended as illustrative rather than exhaustive and do not rely on external clinical adjudication.
One common pattern involves symptom–intervention or symptom–monitoring relationships. For example, concepts such as shortness of breath frequently co-occur with oxygen administration by nasal cannula and blood oxygen saturation measurement. These pairs exhibit moderate to high NPMI values and high cosine similarity, reflecting both frequent joint documentation and strong semantic relatedness. This pattern is consistent with routine ICU documentation, where symptoms prompting respiratory support are recorded alongside monitoring and intervention terms.
In contrast, several semantically similar concept pairs show high cosine similarity but low or near-zero co-occurrence. For instance, concepts such as myocardial infarction and troponin measurement are closely related in embedding space but are not consistently documented together within the same note. This likely reflects documentation structure rather than lack of clinical relevance, as laboratory findings and diagnoses are often recorded in separate sections or different notes during the course of care.
We also observe frequent co-occurrence between semantically broader or heterogeneous concepts, such as patient’s condition stable and vital signs monitoring. These pairs show moderate NPMI but lower semantic similarity, suggesting routine co-documentation driven by templated progress notes rather than tight conceptual linkage. Such patterns highlight how co-occurrence captures documentation workflow, while embeddings encode conceptual proximity independent of note structure.
Together, these examples illustrate how co-occurrence and semantic similarity provide complementary signals: co-occurrence reflects how clinicians document information in practice, while embeddings surface latent clinical relationships that may be underdocumented or distributed across notes. These examples are presented to support qualitative interpretation of the observed patterns; formal clinical validation of specific concept relationships was beyond the scope of this study.
Discussion
This study examined how empirical co-occurrence of SNOMED CT concepts in clinical documentation relates to embedding-based semantic similarity, using large-scale ICU notes from MIMIC-IV. By integrating statistical association measures, pretrained biomedical embeddings, and temporal stratification, our analysis provides insight into how clinical knowledge is expressed, reinforced, and fragmented in real-world EHR narratives. Importantly, the results highlight that documentation frequency and semantic relatedness capture distinct, complementary dimensions of clinical meaning.
Our work is closely related to prior efforts that combine SNOMED CT annotations with embedding-based representations, particularly Wang et al., 17 who used ClinicalBERT embeddings and SNOMED concepts to construct patient-level representations for clustering and outcome prediction. That work demonstrated that semantic embeddings derived from clinical text encode clinically meaningful information and can be used effectively for downstream predictive tasks such as mortality and readmission. While methodologically similar in their use of pretrained embeddings and SNOMED CT annotations, the focus and analytical lens of the present study differ in important ways. Rather than aggregating embeddings at the patient level or evaluating predictive performance, we explicitly interrogate the relationship between two foundational signals: how often concepts are documented together (co-occurrence) and how semantically related those concepts are in embedding space. This concept-pair—level analysis allows us to examine not only where semantic similarity aligns with documentation patterns, but also where it diverges—an aspect that is largely implicit and unexplored in prior work, including Wang et al. 17
One of the central findings of this study is the consistently weak correlation between NPMI-based co-occurrence strength and embedding-based semantic similarity. Clinically, this result is both intuitive and informative. Co-occurrence in clinical notes reflects documentation practices shaped by workflow, note templates, provider roles, and temporal constraints, rather than a comprehensive encoding of all clinically related concepts. Many conceptually linked entities—such as diagnoses and their confirmatory tests, or conditions and downstream complications—may be recorded in different notes, sections, or phases of care. As a result, low co-occurrence does not imply weak clinical relevance. Embedding-based similarity, by contrast, captures latent conceptual relationships derived from broader linguistic context across the corpus. The weak correlation therefore suggests that embeddings are able to surface clinically meaningful associations that are not consistently reinforced by documentation frequency. From a clinical informatics perspective, this divergence is valuable: it highlights gaps between conceptual knowledge and narrative documentation and suggests opportunities for improving annotation completeness, decision support, and ontology refinement.
Temporal analysis further clarifies how these relationships evolve over the course of hospitalization. Early stage notes were characterized by more diffuse co-occurrence patterns and weaker association signals, even among semantically similar concepts. This reflects the exploratory nature of early clinical reasoning, where presenting symptoms, differential diagnoses, and preliminary findings are documented broadly but not yet tightly coupled. In contrast, late-stage notes exhibited stronger and more coherent co-occurrence structures, corresponding to diagnostic consolidation, treatment execution, and discharge planning. These temporal patterns suggest that late-stage documentation more faithfully reflects stabilized clinical relationships, while early stage notes may benefit most from embedding-based augmentation to surface relevant but underdocumented concepts. Treating clinical text as temporally homogeneous would obscure these clinically meaningful shifts.
Several findings that may initially appear unexpected are readily explained by real-world clinical practice. The approximately 7:1 imbalance between late and early notes is not an artifact of data processing, but rather a reflection of ICU documentation workflows. Early documentation is often limited to admission, triage, and initial assessments, whereas late-stage care involves frequent progress notes, complication tracking, multidisciplinary updates, and discharge summaries. This imbalance underscores the inherent difficulty of extracting robust early signals from sparse documentation and highlights the potential value of semantic models in compensating for limited early data.
Similarly, the presence of rare concept pairs with high NPMI values warrants careful interpretation. In some cases, these pairs represent rare but canonical clinical relationships that occur consistently within specific patient subgroups, such as uncommon complications or specialized interventions. In other cases, high NPMI may arise from templated documentation or tightly bound events that always co-occur when mentioned. Rather than being noise, these rare high-NPMI pairs may flag niche but clinically important patterns that would be missed by frequency-based filtering alone. Their identification illustrates the sensitivity of NPMI to localized association structure and reinforces the importance of examining both frequent and infrequent patterns.
Taken together, these findings suggest that co-occurrence statistics and semantic similarity should not be viewed as redundant or competing signals. Instead, they provide complementary perspectives on clinical documentation: co-occurrence reflects how clinicians write notes within workflow constraints, while embeddings capture deeper conceptual relationships grounded in language use and shared clinical context. Integrating these views enables a more nuanced understanding of clinical narratives, supports identification of underdocumented but relevant concepts, and provides a foundation for improved phenotyping, semantic quality assurance, and decision support. By explicitly examining where semantic similarity and documentation frequency align and diverge, this study extends prior embedding-based EHR analyses and offers new insight into how clinical knowledge is implicitly structured in real-world documentation.
Limitations
This study has several limitations that should be considered when interpreting the results. First, concept co-occurrence was defined at the level of entire clinical notes, such that any two SNOMED CT concepts appearing within the same note were treated as co-occurring. While this approach captures broad documentation patterns and is computationally tractable at scale, it may obscure finer-grained relationships that occur at the sentence, paragraph, or section level. Concepts documented in different sections of a note—such as past medical history versus assessment and plan—may be clinically related yet temporally or contextually separated. Future work could incorporate window-based, sentence-level, or section-aware co-occurrence definitions to refine association signals.
Second, the analysis relies on SNOMED CT annotations derived from automated concept extraction pipelines. Although SNOMED CT provides a comprehensive and standardized terminology, automated annotation is subject to errors arising from abbreviation ambiguity, negation, and context misinterpretation. Such errors may introduce noise into both co-occurrence statistics and embedding-based similarity analyses. Incorporating confidence thresholds, clinician validation, or hybrid annotation strategies may improve robustness in future studies.
Third, semantic similarity was computed using pretrained biomedical language models that were not fine-tuned specifically for SNOMED CT concept representations. While models such as ClinicalBERT and BioBERT capture rich contextual semantics, they may not fully align with SNOMED CT’s hierarchical and logical structure. Ontology-aware or graph-regularized embedding approaches could potentially improve alignment between learned representations and formal medical semantics.
Fourth, the embedding-based concept suggestion analysis was exploratory and retrospective in nature. Although many suggested concepts appeared in subsequent documentation, this does not establish clinical necessity or causal relevance. Prospective evaluation with clinician review will be required to assess whether such suggestions meaningfully improve documentation completeness, reduce omissions, or support clinical decision-making without increasing cognitive burden.
Fifth, this study was conducted using data from a single institution (MIMIC-IV), which reflects intensive care documentation practices at a specific academic medical center. Documentation styles, note structure, and clinical workflows may differ across institutions, specialties, and care settings. As a result, the generalizability of co-occurrence and semantic patterns observed here should be evaluated in other EHR datasets and healthcare environments.
Finally, while we provide illustrative examples of concept relationships to support qualitative interpretation, these examples were not formally reviewed or validated by domain experts. The study design focused on quantitative and semantic analysis at scale, and formal clinical adjudication was beyond scope. As a result, the clinical plausibility of specific high-similarity or high-association concept pairs was inferred from established medical knowledge rather than verified through structured expert evaluation. Future work should incorporate clinician review or targeted case-level validation to assess the practical relevance of embedding-based suggestions and co-occurrence patterns in real-world clinical workflows.
Despite these limitations, the study provides a large-scale, integrative analysis of statistical and semantic relationships among SNOMED CT concepts in real-world clinical text, offering a foundation for future methodological refinement and applied clinical informatics research.
Conclusion
This study presented a comprehensive semantic analysis of SNOMED CT concept co-occurrences in the MIMIC-IV clinical notes, integrating statistical association measures with embedding-based similarity to uncover latent structures in clinical documentation. By jointly examining NPMI and cosine similarity, we demonstrated that frequent co-occurrence of SNOMED CT concepts often—but not always—aligns with semantic proximity in embedding space. This partial overlap highlights the complementary nature of statistical and semantic dimensions: co-occurrence reflects documentation habits and clinical workflow, while embedding similarity captures deeper conceptual relatedness derived from contextual language use.
Our results reveal several key insights. First, while most co-occurring concept pairs exhibit moderate to high semantic similarity, a subset of frequent yet semantically distant pairs may represent atypical presentations, documentation inconsistencies, or complex multimorbidity patterns. Second, temporal analyses showed that early stage notes contain more diffuse and exploratory concept associations, whereas late-stage notes display stronger and more semantically coherent co-occurrence structures, reflecting the clinical transition from diagnostic uncertainty to treatment and discharge planning. Third, embedding-based clustering yielded distinct and clinically interpretable semantic groups—such as cardiovascular conditions, laboratory measurements, and symptom clusters—that closely correspond to real-world clinical reasoning patterns. These clusters not only validate the semantic consistency of biomedical embeddings but also offer a data-driven approach for phenotype discovery and ontology refinement.
The combined use of co-occurrence and semantic embedding analyses underscores the potential of hybrid methods for improving the interpretability and completeness of clinical documentation. Embedding-driven concept suggestion demonstrated the feasibility of identifying underdocumented yet clinically plausible concepts, paving the way for semi-automated annotation systems that enhance EHR quality and support downstream applications such as coding, cohort identification, and predictive modeling.
Future research should focus on extending this framework across multiple institutions and care settings to evaluate generalizability and robustness. Incorporating ontology-aware or graph-regularized embeddings may further align learned representations with SNOMED CT’s hierarchical semantics. Additionally, integrating multimodal data sources—such as laboratory results, imaging, and structured variables—could enable richer, context-aware modeling of clinical events.
In summary, this work advances the understanding of how semantic and statistical relationships between clinical concepts manifest in real-world documentation. By linking data-driven embeddings with standardized terminologies, it contributes to the broader goal of transforming unstructured clinical narratives into semantically grounded, interoperable knowledge representations that can meaningfully support patient care, analytics, and biomedical discovery.
Footnotes
Acknowledgements
None.
Ethical approval and consent to participate
This study used the publicly available Medical Information Mart for Intensive Care IV (MIMIC-IV) database, which contains de-identified health data. The MIMIC-IV dataset has been fully de-identified in accordance with the Health Insurance Portability and Accountability Act (HIPAA) Safe Harbor provisions. As this research involved analysis of existing, de-identified data and did not include any direct interaction with human subjects, ethical approval and informed consent were not required.
Author contributions
AN conducted the data analysis and produced the first draft of the manuscript. PM supervised AN, conceptualized the analysis, and produced the final draft of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is funded by a CAREER award (#2522386) to Manda from the Division of Biological Infrastructure at the National Science Foundation, USA.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
